ISSN: 1449-2288International Journal of Biological Sciences
Int J Biol Sci 2008; 4(4):202-207. doi:10.7150/ijbs.4.202 This issue Cite
Research Paper
Generation and Validation of a Mouse Line with a Floxed SRC-3/AIB1 Allele for Conditional Knockout
1. Department of Molecular and Cellular Biology, Baylor College of Medicine, Houston, TX, USA.
2. Institute of Biosciences and Technology, Texas A&M University Health Science Center, Houston, TX, USA
# Z. Liu and L. Liao contributed equally to this work.
Abstract
The steroid receptor coactivator-3 (SRC-3), also known as AIB1, ACTR, p/CIP and NCOA3, is a transcriptional coactivator for nuclear receptors and certain other transcription factors. SRC-3 is widely expressed and plays important physiological functions and pathogenic roles in breast and prostate cancers. SRC-3 knockout (SRC-3-/-) mice display genetic background-dependent embryonic lethality and multiple local and systemic abnormalities. Since both the partial lethality and the systemic effects caused by global SRC-3 knockout interfere with downstream investigation of tissue-specific function of SRC-3, we have generated floxed SRC-3 (SRC-3f/f) mice with conditional alleles carrying loxP sites in introns 10 and 12 by a gene-targeting strategy. The two SRC-3f/f mouse lines (A and B) are indistinguishable from wild type mice. To test if deletion of the floxed exons 11 and 12 for SRC-3 nuclear receptor interaction domains and disruption of its downstream sequence for transcriptional activation domains would inactivate SRC-3 function, SRC-3f/f mice were crossbred with EIIa-Cre mice to generate SRC-3d/d mice with germ line deletion of the floxed SRC-3 gene. Both lines of SRC-3d/d mice exhibited growth retardation and low IGF-I levels, which was similar to that observed in SRC-3-/- mice. The line A SRC-3d/d mice showed normal viability, while line B SRC-3d/d mice showed partial lethality similar to SRC-3-/- mice, probably due to variable distributions of genetic background during breeding. These results demonstrate that the floxed SRC-3 mouse lines have been successfully established. These mice will be useful for investigating the cell type- and developmental stage-specific functions of SRC-3.
Keywords: SRC-3, AIB1 Allele, knockout mice
Introduction
The steroid receptor coactivator-3 (SRC-3), also known as AIB1, ACTR, p/CIP and NCOA3, is a member of the p160 SRC coactivator family that also contains SRC-1 (NCOA1) and SRC-2 (TIF2, GRIP1, NCOA2) (1). SRC-3 interacts with ligand-activated nuclear receptors such as receptors for estrogen, androgen and progesterone and certain other transcription factors such as E2F1, PEA3 and AP-1 (1-5). SRC-3 enhances gene transcription by recruiting histone remodeling enzymes including CBP (cAMP response element-binding protein binding protein), p300 and p/CAF (p300 and CBP associated protein) histone acetyltransferases and the coactivator-associated methyltransferase 1 (CARM-1) to the target gene promoter (reviewed in (1)). SRC-3 has been designated as a proto-oncogene based on the following: SRC-3 is overexpressed in subgroups of human breast and prostate cancers (6-8); SRC-3 is required for breast and prostate cancer cell proliferation and survival in culture (8-10); SRC-3-deficient mice are resistant to oncogene- and carcinogen-induced mammary gland and prostate carcinogenesis and metastasis (11-14); and ectopic overexpression of SRC-3 in mouse mammary epithelial cells is sufficient to induce mammary tumorigenesis (15).
SRC-3 is expressed in brain, vascular endothelial and smooth muscle cells, intestinal smooth muscle cells, mammary gland epithelium, oocyte, prostate stromal and basal cells, mammary gland and prostate tumor cells, Sertoli cells in the testis and liver cells (16, 17). SRC-3 knockout (SRC-3-/-) mice have been previously generated by conventional gene-targeting strategies to explore the in vivo function of SRC-3 (16, 18). Global inactivation of the SRC-3 gene in mice causes partial embryonic lethality depending on strain genetic backgrounds ((16) and unpublished data) and growth retardation accompanied with low IGF-I and IGF-binding protein 3 (IGFBP-3) levels, delays pubertal development, reduces female reproductive function and mammary gland growth accompanied with low estrogen levels (16, 18, 19), decreases estrogen-dependent vascular protection (20), inhibits adipogenesis (21) and increases inflammatory responses (22). These findings indicate that SRC-3 is widely expressed, and it regulates physiological functions through both its cell autonomous cellular actions and its systemic effects on hormones such as IGF-I and estrogen.
Although SRC-3-/- mice have provided valuable information regarding fundamental genetic functions of SRC-3, the diverse and systemic effects of global SRC-3 deficiency have hindered our understanding of tissue- and cell type-specific and developmental stage-specific functions of SRC-3. In order to circumvent these limitations, we have generated and validated mouse lines carrying a floxed SRC-3 allele that is appropriate for Cre-mediated tissue-specific inactivation of the SRC-3 gene.
Materials and Methods
DNA cloning, embryonic stem (ES) cell culture and gene targeting
The mouse SRC-3 genomic DNA was obtained from phage clones containing large fragments of the SRC-3 genomic DNA from the 129SvEv mouse genome (16, 23). A 5.65-kb DNA fragment for the 5' targeting arm was taken from the SRC-3 genomic DNA in a phage clone by Apa I and Sca I co-digestion and subcloned into the Pac I/Asc I sites of the pFRT-LoxP plasmid. A 1.9-kb SRC-3 DNA fragment to be floxed was taken from the same clone by Sca I and Mfe I co-digestion and subcloned into the Not I and Kpn I sites of the same plasmid. A 5-kb SRC-3 DNA fragment for the 3' targeting arm was taken from another SRC-3 genomic DNA clone by Mfe I and Bmg BI co-digestion and subcloned into the Xho I and Bam HI sites of the same plasmid for finishing construction of the targeting vector (Fig. 1). The vector was linearized by Xho I (inserted after the 5' arm) digestion and transfected into the TC-1 129SvEv mouse ES cells (24) by electroporation. Transfected ES cells were cultured in selection medium with G418 (Geneticin) and FIAU (1-(2-deoxy-2-fluoro-beta-D-arabinofuranosyl)-5-iodouracil) and survived clones were isolated for Southern blot analysis as described previously (16, 25). The 5' probe DNA was taken from the SRC-3 genomic DNA clone by Sal I and Apa I digestion, which was located upstream at the 5' targeting arm (Fig. 1). The 3' probe DNA was taken from the SRC-3 genomic DNA clone by Bmg BI and Kpn I digestion, which was located downstream at the 3' targeting arm (Fig. 1). Southern blot analysis also was preformed with a neo probe to exclude clones with any random integrations of the targeting vector DNA.
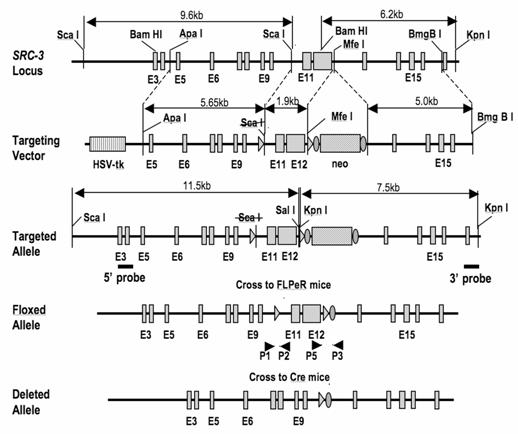
The gene targeting strategy for generation and validation of the floxed SRC-3 mice. A part of the mouse SRC-3 gene structure is shown with indicated exons, restriction enzyme sites used in construction of the targeting vector and Southern blot analysis. The targeting vector structure is shown with indicated regions and lengths of its 3' and 5' targeting arms. The loxP sites are indicated by triangles. The Frt sites are indicated by ovals. The targeted allele is drawn with indicated 5' and 3' probe positions. After DNA is digested with Sca I and Sal I, the 5' probe should detect a 9.6-kb band for WT allele and a 11.5-kb band for the targeted allele as indicated. After digested with Bam HI and Kpn I, the 3' probe should detect a 6.2-kb band for the WT allele and a 7.5-kb band for the targeted allele as indicated. The floxed allele can be obtained after excision of the Frt-flanked neo cassette through crossing to FLPeR transgenic mice. The P1, P2, P5 and P3 primers for PCR-based genotype analyses are indicated (arrowheads). The structure of the deleted allele obtained after Cre-mediated excision of the floxed region is also sketched.
Microinjection and mouse breeding
The correctly targeted ES clones were microinjected into C57/BL6 morulas to generate chemeric founder mice. The male chimeric mice were kept for breeding with female C57BL/6 mice to test germ line transmission of the targeted SRC-3 locus. Next, the male chimeric mice with proven germ line transmission ability were bred with the 129SvEv FLPeR mice (Stock #003946, Jackson Lab) (26) to remove the Frt-flanked neo expression cassette (Fig. 1). Finally, the FLPeR allele was excluded in incoming generations and the floxed SRC-3 allele was excised in germ line by breeding with the C57BL/6 EIIa-Cre transgenic mice (Stock #003724, Jackson Lab) (27).
Mouse genotype analysis by PCR
The locations of PCR primers used for genotyping WT and manipulated SRC-3 alleles are indicated in Fig. 1. These primers include SRC3-P1 (5'-tgacatctggtggtgtgacc), SRC3-P2 (5'-tagcccctccttttcctgtt), SRC3-P3 (5'-gaaacctcaaggttatccttcaatt), and SRC3-P5 (5'-tgctctgcttagatacctgctg). Mouse genomic DNA samples were prepared from tail tips by following standard protocols. PCR reactions were performed using genomic DNA as templates and SRC-3 genotype-specific primer pairs with the following program: 94oC, 3 minutes; 94oC, 30 seconds, 57oC 40 seconds, 72oC, 1 minute for 35 cycles. The methods for genotyping EIIa-Cre and FLPeR mice were described previously (26, 27).
Body weight and IGF-1 measurements
Male mice were weighed at 4, 8 and 12 weeks of age. Blood samples were collected from 12-week-old mice and serum samples were prepared and frozen at -80°C until analysis. Serum IGF-1 concentrations were measured using an ELISA kit (DSL-10-29200, Diagnostic Systems Lab.) by following the manufacturer's instruction. Results are presented as mean ± standard deviation. Statistical analysis was performed using unpaired Student's t test with p<0.05 to be considered as significant difference.
Results
Generation of ES cells with targeted SRC-3 allele by homologous recombination
The mouse SRC-3 gene spans about 79 kb in length and contains 23 exons, and it encodes a 160-kD protein with 1403 amino acid residues (23). SRC-3 protein contains an N-terminal basic helix-loop-helix/PAS (Per/ARNT/Sim) domain, a central nuclear receptor interaction domain with multiple LXXLL (L, leucine, X, any amino acid residue) motifs, and a C-terminal transcriptional activation domain (1, 28). To obtain a floxed SRC-3 allele appropriate for Cre-mediated conditional inactivation of the SRC-3 gene, we designed and constructed a targeting vector that could knock in one loxP site into intron 10 and another loxP site into intron 12 through homologous recombination in mouse ES cells. The targeting vector contains a 5.65-kb 5' targeting arm, a floxed 1.9-kb DNA fragment spanning exons 11 and 12 and intron 11, a neo expression cassette for positive selection, a 5-kb 3' targeting arm and a tk expression cassette for negative selection (Fig. 1). The floxed exons 11 and 12 encode the nuclear receptor interaction domain of SRC-3 (23, 28). In addition, the deletion of exons 11 and 12 would also cause a coding-frame shift between exons 10 and 13 and result in disruption of the C-terminal polypeptide sequence of SRC-3 (Fig. 1).
After the TC-1 mouse ES cells were electroporated with the targeting vector DNA and cultured in selection medium with G418 and FIAU, 288 survived clones were isolated and DNA samples were prepared from these clones for Southern blot screening. The 3' probe detected 10 clones with a 7.5-kb targeted allele band in addition to a 6.2-kb wild type (WT) allele band after DNA samples were digested with Bam H1 and Kpn I (Fig. 1 and Fig. 2B). Four of these positive clones also showed correct recombination at the 5' location, exhibiting a 11.5-kb targeted allele band in addition to a 9.6-kb WT allele band on Southern blot using the 5' probe after DNA was digested with Sca I and Sal I (Fig. 1 and Fig. 2A). A neomycin probe was also used to confirm the sole homologous integration locus of the targeting vector at the SRC-3 gene in the targeted ES cells (Fig. 2C). These results demonstrate that multiple targeted ES clones with the floxed SRC-3 allele had been successfully generated. Since this targeted allele contains an Frt-flanked neo cassette, this allele is designated as SRC-3f-neo.
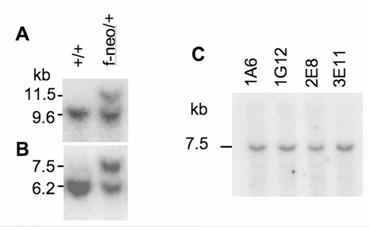
Identification of targeted ES clones by Southern blot analysis. A & B. Southern blot analyses of ES clones using the 5' and the 3' probes. DNA was digested with Sca I and Sal I in panel A and with Bam HI and Kpn I in panel B. The genotypes of WT (+/+) and targeted (f-neo) ES clones are indicated. C. Southern blot analysis of ES clones using the neo probe. The correctly targeted clones (1A6, 1G12, 2E8 & 3E11) contain only one neo integration site within the 7.5-kb band for the targeted SRC-3 allele. DNA was digested as that in panel B.
Generation of SRC-3f/f mice
Correctly targeted ES clones were microinjected into C57BL/6 blastocysts to generate chimeric mice. Male chimeras were kept for testing germ line transmission of the SRC-3f-neo allele by crossing with C57BL/6 female mice and by examining the coat color of their offspring. Agouti coat indicates a germ line transmission from the 129SvEv ES cell, while black coat indicates a C57BL/6 blastocyst donor origin. A total of 7 chimeric founder mice were identified to have germ line transmission capability. Two of these founder mice (lines A and B) were crossed with female FLPeR mice (26) to remove the Frt-flanked neo cassette in their pups with an SRC-3f-neo/+/FLPeR+ genotype. In these pups, the flippase recombinase (FLP) was ubiquitously expressed and its expression resulted in excision of the Frt-flanked neo cassette in the SRC-3f-neo allele. As shown in Fig. 3, PCR analyses detected WT and targeted alleles in FLPeR-negative SRC-3f-neo/+ mice (Fig. 3A) and WT and floxed alleles in FLPeR-positive and neo-negative SRC-3f/+ mice (Fig. 3B). Subsequently, SRC-3f/+ breeding pairs were used to generate WT and SRC-3f/f mice that were negative of both neo cassette and FLPeR. Because the normal function of the SRC-3 gene is required for somatic growth and normal IGF-I levels, we measured body growth rates and serum IGF-I levels and found no differences between WT and SRC-3f/f littermates with a pure 129SvEv strain background (Fig. 4, A & B). These results indicate that the SRC-3f allele has normal function identical to the SRC-3+ allele.
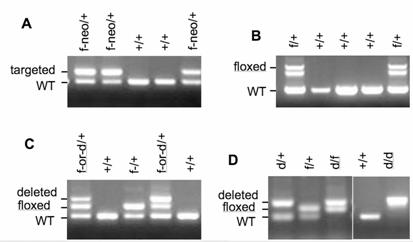
Genotype analyses of mice with WT (+), targeted (f-neo), floxed (f) or deleted (d) SRC-3 alleles. A. Genotype analysis of WT and SRC-3f-neo/+ mice. Both SRC-3+ and SRC-3f-neo alleles were detected by PCR using SRC3-P1 and SRC3-P2 primers. B. Genotype analysis of SRC-3f/+ mice after the neo cassette is excised by crossing to FLPeR mice. Both SRC-3+ and SRC-3f alleles were detected by PCR using SRC3-P3/SRC3-P5 primers. Due to both loxP and Frt palindromic sequences in the floxed allele, the product from PCR using primers SRC3-P3 and SRC3-P5 produces two bands for the SRC-3f allele and one band for the SRC-3+ allele. C. Genotype analyses of mice derived from SRC-3f/+ and EIIa-Cre breeding pairs. Both SRC-3+ and SRC-3f were detected by PCR using SRC3-P1 and SRC3-P2 primers as shown in panel A. The SRC-3d allele was detected by PCR using SRC3-P1 and SRC3-P3 primers. Both SRC-3d and SRC-3f alleles were detected in SRC-3f/+/EIIa-Cre+ mice due to partial excision of the floxed SRC-3 allele. D. Genotype analyses of mice derived from SRC-3f/+/EIIa-Cre+ (SRC-3f-or-d/+) breeding pairs. PCR was performed as in panel C. From this breeding, mice with germ line deletion of the floxed SRC-3 (SRC-3d/d) were generated and identified.
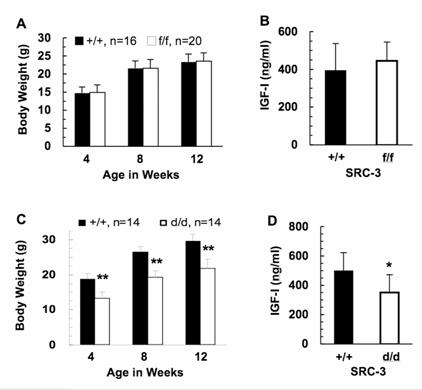
Measurements of body weight and serum IGF-I levels. A & B. SRC-3f/f and WT mice showed identical somatic growth patterns and similar serum IGF-I levels. The body weights of male WT (n=16) and SRC-3f/f (n=20) mice with a 129SvEv strain background were measured at ages indicated. Blood samples were collected from 12-week-old male WT (n=11) and SRC-3f/f (n=11) mice for IGF-I measurement. C & D. SRC-3d/d mice exhibit growth retardation and low IGF-I levels. The body weights (Panel C) and serum IGF-I concentrations (Panel D) of male WT and SRC-3d/d mice with a mixed strain background of C57BL/6 and 129SvEv were measured as that in Panels A & B. **, p < 0.005 and *, p < 0.05 by unpaired t-test.
Germ-line deletion of the floxed SRC-3 allele recapitulates an SRC-3 knockout phenotype
To test whether excision of the floxed region of the SRC-3f allele would completely inactivate the gene function, we crossed SRC-3f/f mice with a 129SvEv strain background (both A and B lines) with EIIa-Cre transgenic mice with a C57BL/6 strain background (27) and generated SRC-3f/+/EIIa-Cre+ mice with a mixed genetic background. In these mice, the expressed Cre resulted in a partial deletion of the SRC-3f allele and produced mosaic mice with both the deleted SRC-3 (SRC-3d) and the remaining SRC-3f alleles in addition to the WT SRC-3 allele in the same mouse as determined by PCR genotyping using tail-tip DNA samples (Fig. 3C). Subsequently, SRC-3d/+ and SRC-3d/d mice were generated by breeding SRC-3f/+/EIIa-Cre+ mice and their progenies (Fig. 3D). For the A line, 25 of WT, 57 of SRC-3d/+ and 25 of SRC-3d/d mice were obtained at weaning stage from the SRC-3d/+ breeding pairs, which was consistent with expected Mendelian ratios. For the B line, 22 of WT, 33 of SRC-3d/+ and 10 of SRC-3d/d mice were obtained at weaning stage from the SRC-3d/+ breeding pairs, which was consistent with the partial lethal phenotype observed in SRC-3-/- mice. The differences between line A and line B SRC-3d/d mice in viability may be caused by variable distributions of strain background during the breeding process.
For both lines A and B, SRC-3d/+ mice were slightly smaller than age-matched WT mice (Data not shown). However, both male and female SRC-3d/d mice were significantly smaller than WT mice at all ages examined, including 4, 8 and 12 weeks (Fig. 4C for line A, and data for line B not shown). When measured at 12 weeks of age, SRC-3d/d mice also had a remarkable reduction in serum IGF-I levels compared with age-matched WT mice (Fig. 4D for line A, and data for line B not shown). The growth defect and low levels of IGF-I phenotypes observed in SRC-3d/d mice closely resemble the SRC-3 knockout mice (16). These results demonstrate that excision of the floxed region (exons 11 and 12) of the SRC-3f allele results in a loss-of-function mutant of the SRC-3 allele.
Discussion
Since SRC-3 is identified as a transcriptional coactivator, many studies have been performed to explore its normal functions in development, metabolism, reproduction and cardiovascular system, and its pathogenetic roles in hormonally promoted breast and prostate cancers (11-13, 15, 16, 18-20, 29). Because no germ line mutation has been found to link SRC-3 to a human disease, the knowledge regarding most of the SRC-3 in vivo functions is gained from SRC-3 knockout and transgenic mice. Although these mouse models provide valuable information, their limitations of systemic effects caused by global SRC-3 deficiency have made it difficult to assign certain phenotypes and their molecular basis to a specific cell type or a specific developmental stage. In order to use the Cre-loxP system to inactivate SRC-3 in either a cell type-specific or a developmental stage-specific way, we have generated SRC-3f/f mice using a gene targeting strategy in this study. SRC-3f/f mice exhibit normal somatic growth and normal IGF-I levels and are indistinguishable from WT mice, indicating that insertion of the loxP sites has no effect on the SRC-3 gene function.
In SRC-3f/f mice, the paired loxP sites are located in introns 10 and 12. The floxed exons 11 and 12 encode the essential nuclear receptor interaction domain and the deletion of these two exons also disrupts the downstream coding frame. As expected, in our testing experiments deletion of the floxed SRC-3 alleles in SRC-3d/d mice caused postnatal growth retardation and low levels of circulating IGF-I, which are two of the major defects observed in SRC-3 knockout mice (1, 16, 18). These results demonstrate that Cre-mediated excision of the floxed SRC-3 allele disrupt the function of the SRC-3 gene. Intriguingly, the two lines of SRC-3d/d mice exhibit different viability. The line A SRC-3d/d mice have a normal viability while the line B SRC-3d/d mice show a partial lethality. This discrepancy may be caused by uneven distributions of 129SvEv and C57BL/6 strain backgrounds in these two lines of mice during the breeding process. In agreement with this explanation, our recent studies have revealed that the lethal degrees of SRC-3-/- mice are dependent on strain background. Lesser, medium and severe degrees of lethality are found in SRC-3-/- mice with a 129SvEv, a mixed 129SvEv/C57BL/6 and a C57BL/6 strain background, respectively ((16) and unpublished results).
Taken together, we have generated and validated the SRC-3f/f mouse line appropriate for conditional inactivation of the SRC-3 gene after crossing with a Cre mouse line of interest. The SRC-3f/f mouse line will be useful for investigating the cell type- and developmental stage-specific functions of SRC-3 in mice.
Acknowledgements
We thank Dr. Chuxia Deng for ES cells, Dr. James Martin for the pFRT-loxP plasmid, and Ms. Jenny Xia and Dr. Robert B. York for critical reading of the manuscript. This work is partially funded by DK58242, CA112403 and CA119689 grants from National Institutes of Health.
Conflict of Interest
The authors have declared that no conflict of interest exists.
References
1. Xu J, Li Q. Review of the in vivo functions of the p160 steroid receptor coactivator family. Mol Endocrinol. 2003;17(9):1681-92
2. Mussi P, Yu C, O'Malley BW, Xu J. Stimulation of steroid receptor coactivator-3 (SRC-3) gene overexpression by a positive regulatory loop of E2F1 and SRC-3. Mol Endocrinol. 2006;20(12):3105-19
3. Louie MC, Zou JX, Rabinovich A, Chen HW. ACTR/AIB1 functions as an E2F1 coactivator to promote breast cancer cell proliferation and antiestrogen resistance. Mol Cell Biol. 2004;24(12):5157-71
4. Goel A, Janknecht R. Concerted activation of ETS protein ER81 by p160 coactivators, the acetyltransferase p300 and the receptor tyrosine kinase HER2/Neu. J Biol Chem. 2004;279(15):14909-16
5. Yan J, Yu CT, Ozen M, Ittmann M, Tsai SY, Tsai MJ. Steroid receptor coactivator-3 and activator protein-1 coordinately regulate the transcription of components of the insulin-like growth factor/AKT signaling pathway. Cancer Res. 2006;66(22):11039-46
6. Anzick SL, Kononen J, Walker RL. et al. AIB1, a steroid receptor coactivator amplified in breast and ovarian cancer. Science. 1997;277(5328):965-8
7. Osborne CK, Bardou V, Hopp TA. et al. Role of the estrogen receptor coactivator AIB1 (SRC-3) and HER-2/neu in tamoxifen resistance in breast cancer. J Natl Cancer Inst. 2003;95(5):353-61
8. Zhou HJ, Yan J, Luo W. et al. SRC-3 is required for prostate cancer cell proliferation and survival. Cancer Res. 2005;65(17):7976-83
9. Planas-Silva MD, Shang Y, Donaher JL, Brown M, Weinberg RA. AIB1 enhances estrogen-dependent induction of cyclin D1 expression. Cancer Res. 2001;61(10):3858-62
10. List HJ, Lauritsen KJ, Reiter R, Powers C, Wellstein A, Riegel AT. Ribozyme targeting demonstrates that the nuclear receptor coactivator AIB1 is a rate-limiting factor for estrogen-dependent growth of human MCF-7 breast cancer cells. J Biol Chem. 2001;276(26):23763-8
11. Kuang SQ, Liao L, Wang S, Medina D, O'Malley BW, Xu J. Mice lacking the amplified in breast cancer 1/steroid receptor coactivator-3 are resistant to chemical carcinogen-induced mammary tumorigenesis. Cancer Res. 2005;65(17):7993-8002
12. Kuang SQ, Liao L, Zhang H, Lee AV, O'Malley BW, Xu J. AIB1/SRC-3 deficiency affects insulin-like growth factor I signaling pathway and suppresses v-Ha-ras-induced breast cancer initiation and progression in mice. Cancer Res. 2004;64(5):1875-85
13. Chung AC, Zhou S, Liao L, Tien JC, Greenberg NM, Xu J. Genetic ablation of the amplified-in-breast cancer 1 inhibits spontaneous prostate cancer progression in mice. Cancer Res. 2007;67(12):5965-75
14. Fereshteh MP, Tilli MT, Kim SE. et al. The nuclear receptor coactivator amplified in breast cancer-1 is required for Neu (ErbB2/HER2) activation, signaling, and mammary tumorigenesis in mice. Cancer Res. 2008;68(10):3697-706
15. Torres-Arzayus MI, De Mora JF, Yuan J. et al. High tumor incidence and activation of the PI3K/AKT pathway in transgenic mice define AIB1 as an oncogene. Cancer Cell. 2004;6(3):263-74
16. Xu J, Liao L, Ning G, Yoshida-Komiya H, Deng C, O'Malley BW. The steroid receptor coactivator SRC-3 (p/CIP/RAC3/AIB1/ACTR/TRAM-1) is required for normal growth, puberty, female reproductive function, and mammary gland development. Proc Natl Acad Sci U S A. 2000;97(12):6379-84
17. Mark M, Yoshida-Komiya H, Gehin M. et al. Partially redundant functions of SRC-1 and TIF2 in postnatal survival and male reproduction. Proc Natl Acad Sci U S A. 2004;101(13):4453-8
18. Wang Z, Rose DW, Hermanson O. et al. Regulation of somatic growth by the p160 coactivator p/CIP. Proc Natl Acad Sci U S A. 2000;97(25):13549-54
19. Liao L, Chen X, Wang S, Parlow AF, Xu J. Steroid receptor coactivator 3 maintains circulating insulin-like growth factor I (IGF-I) by controlling IGF-binding protein 3 expression. Mol Cell Biol. 2008;28(7):2460-9
20. Yuan Y, Liao L, Tulis DA, Xu J. Steroid receptor coactivator-3 is required for inhibition of neointima formation by estrogen. Circulation. 2002;105(22):2653-9
21. Louet JF, Coste A, Amazit L. et al. Oncogenic steroid receptor coactivator-3 is a key regulator of the white adipogenic program. Proc Natl Acad Sci U S A. 2006;103(47):17868-73
22. Yu C, York B, Wang S, Feng Q, Xu J, O'Malley BW. An essential function of the SRC-3 coactivator in suppression of cytokine mRNA translation and inflammatory response. Mol Cell. 2007;25(5):765-78
23. Ning G, Jurecic V, Baldini A, Xu J. Structure and chromosomal locations of mouse steroid receptor coactivator gene family. In Vitro Cell Dev Biol Anim. 1999;35(8):481-6
24. Deng C, Wynshaw-Boris A, Zhou F, Kuo A, Leder P. Fibroblast growth factor receptor 3 is a negative regulator of bone growth. Cell. 1996;84(6):911-21
25. Kuang SQ, Liao L, Zhang H. et al. Deletion of the cancer-amplified coactivator AIB3 results in defective placentation and embryonic lethality. J Biol Chem. 2002;277(47):45356-60
26. Farley FW, Soriano P, Steffen LS, Dymecki SM. Widespread recombinase expression using FLPeR (flipper) mice. Genesis. 2000;28(3-4):106-10
27. Lakso M, Pichel JG, Gorman JR. et al. Efficient in vivo manipulation of mouse genomic sequences at the zygote stage. Proc Natl Acad Sci U S A. 1996;93(12):5860-5
28. Liao L, Kuang SQ, Yuan Y, Gonzalez SM, O'Malley BW, Xu J. Molecular structure and biological function of the cancer-amplified nuclear receptor coactivator SRC-3/AIB1. J Steroid Biochem Mol Biol. 2002;83(1-5):3-14
29. Wang Z, Qi C, Krones A. et al. Critical roles of the p160 transcriptional coactivators p/CIP and SRC-1 in energy balance. Cell Metab. 2006;3(2):111-22
Author contact
![]() Correspondence to: Jianming Xu, Ph.D., Department of Molecular and Cellular Biology, Baylor College of Medicine. One Baylor Plaza, Houston, TX 77030. E-mail: jxutmc.edu
Correspondence to: Jianming Xu, Ph.D., Department of Molecular and Cellular Biology, Baylor College of Medicine. One Baylor Plaza, Houston, TX 77030. E-mail: jxutmc.edu
Received 2008-7-11
Accepted 2008-7-21
Published 2008-7-23
