Impact Factor ISSN: 1449-2288
Int J Biol Sci 2010; 6(7):769-772. doi:10.7150/ijbs.6.769 This issue Cite
Letter To Editor
The Glypican 3-Hosted Murine Mir717 Gene: Sequence Conservation, Seed Region Polymorphisms and Putative Targets
1. Department of Animal Science, University of Ljubljana, Groblje 3, SI- 1230 Domzale, Slovenia
2. National Institute of Chemistry, Hajdrihova 19, SI-1001 Ljubljana, Slovenia
3. Department of Animal Sciences, Washington State University, Pullman, Washington 99164-6351, USA
*Authors contributed equally to the work.
Received 2010-10-20; Accepted 2010-11-25; Published 2010-12-2
Abstract
Mir717 (mmu-mir-717) was first reported in mouse and resides in the intron 3 of glypican 3 (Gpc3) gene. Our present study revealed that this microRNA (miRNA) gene is conserved among 26 mammalian species and harbors polymorphic sites within the mature seed region in mice. Our finding represents a rare four layer genomic overlap consisting of growth associated quantitative trait locus (QTL), body mass associated Gpc3 gene, highly conserved miRNA gene and mature miRNA seed single nucleotide polymorphism (SNP) identified in the lean mouse strain 129/Sv. Additionally, genes potentially targeted by Mir717 include 91 genes associated with obesity and related phenotypes in mammals. Our analysis provides a basis for further experiments to causally connect the identified SNP and Mir717 gene itself to obesity regulation. Furthermore, our bioinformatics analysis now enables functional annotation of Mir717 orthologs in other species, thus determining the effect of its target genes on fat-related traits.
Keywords: Obesity, glypican 3 (Gpc3) gene, microRNA (miRNA), Mir717 (mmu-miR-717), comparative genomics.
MicroRNA (miRNA) Mir717 (mmu-mir-717) was first reported in mouse, which is located in the intron 3 of the glypican 3 (Gpc3) host gene on X chromosome (http://www.mirbase.org/). The human miR-717 is also expressed, and plays important regulatory roles in renal osmoregulation [1]. It has been shown that intronic miRNAs could be coordinately expressed with their host gene mRNA, implying that they might share a common transcriptional mechanism, however, anti-correlative miRNA/host gene expression have also been demonstrated [2-4]. Glypican 3 is a member of the glypican-related integral membrane proteoglycan family. The protein binds to and inhibits the dipeptidyl peptidase activity of CD26, thus inducing apoptosis in certain cell types. In human, mutations in GPC3 gene cause the Simpson-Golabi-Behmel overgrowth syndrome [5], whereas Gpc3 knockout mice show increased body mass, renal dysplasia, and perinatal mortality [6]. Additionally, a QTL for growth was also mapped to the region containing Gpc3 [7]. Therefore, the objectives of our present study were to 1) elucidate evolutionary conservation of Mir717 among the mammalian species, 2) identify genetic variants of the Mir717 in mice, and 3) identify potential target genes of Mir717, particularly those that are associated with obesity and related conditions [8].
According to the miRBase 13.0 (http://microrna.sanger.ac.uk/) and Ensembl (http://www.ensembl.org/index.html) databases, the Mir717 resides within the intron 3 of the Gpc3 gene (Figure 1A). Orthologs of mouse Mir717 were retrieved from the Ensembl genome browser using “alignments / multispecies view / eutherian mammals” option. Multiple species sequence alignment was performed using the MultAlin program (http://bioinfo.genotoul.fr/multalin). Multiple alignment of known (mouse) and predicted Mir717 ortholog sequences showed a high level of conservation among 26 eutherian mammalian species (Figure 1B). So far, the Mir717 gene has been found to be expressed in the mouse and human only. Due to the high sequence conservation this region could now be analyzed by miRNA prediction algorithms and experimentally to functionally annotate the corresponding orthologous miRNA gene in other species. The mouse mature Mir717 differs from the corresponding rat and human sequences in only one and two nucleotides, respectively (Figure 1B). Seven species had 100% mature sequence similarity including human (Homo sapiens): chimpanzee (Pan troglodytes), orangutan (Pongo pygmaeus), macaque (Macaca mulatta), Philippine tarsier (Tarsius syrichta), cat (Felis catus) and European rabbit (Oryctolagus cuniculus) (Figure 1B).
Gene organization, comparative genomics, and genetic variability of the Mir717 gene. A: genomic organization of the Gpc3 with Mir717 in its intron 3. B: Mir717 alignment among 26 mammalian species, mature miRNA region is marked with the square. C: Mir717 genetic variability; location of the premature SNP rs30373504 (light blue arrow) and seed SNP rs30372501 (green arrow) polymorphic between the low fat and high fat mouse strains. Mature Mir717 is underlined.
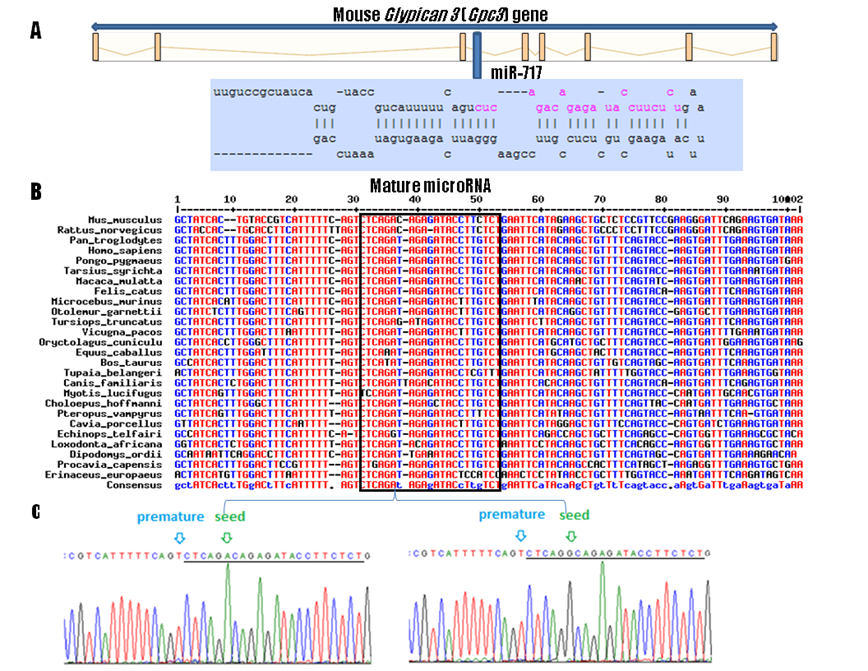
Genetic variability of Mir717 in mouse was evaluated by sequencing PCR products (primers Mir717-F: CCAAATCACCACCTTTGTCC and Mir717-R: AGGAAGCTTGGAGGCAGATT). PCR was carried out in a total volume of 10 μl including ~50 ng of DNA, 1 x PCR buffer, 1 mM MgCl2, 200 µM dNTPs, 0.5 U Taq DNA polymerase (Applied Biosystems, Foster City, CA, USA) and 5 pmol of each primer using Applied Biosystems Gene Amp PCR System 9700 and conditions: 95oC for 10 min, 32 cycles of 94oC for 30 sec, 59oC for 30 sec and 72oC for 30 sec, followed by a further 5‑min extension at 72oC. PCR products were treated with ExoI and SAP enzymes (Fermentas) to remove primers before using a BigDye terminator v3.1 cycle sequencing kit (Applied Biosystems). Capillary electrophoresis was performed on an ABI3130xl genetic analyzer. Sequencing of the 475 bp region flanking the Mir717 was performed with 2-3 mice per inbred strain: a DNA panel contained the “high growth” mouse mutant (genetic background of C57BL/6J) [9], DBA/2J from Harlan (Italy) and DNA from embryonic stem cell line HM1 isolated from 129/Sv strain [10]. Analysis revealed an A>G substitution SNP within the mature Mir717 among these three strains at position 49775654 on Chromosome X (Ensembl release 60). The allele A was shared by high fat strains DBA/2J and C57BL/6J. Interestingly, the allele A is also present in 25 out of 26 compared mammalian species (Figure 1B), suggesting an important functional role of this site. However, the 129/Sv strain contained the allele G at the SNP site. As this line exhibits lower values for all obesity related traits (blood lipids, total body fat, weights of individual fat depots etc.; http://phenome.jax.org/) this SNP allele G could therefore be potentially associated with leanness in mice. However, to causally confirm this association, segregation analyses should be conducted between the lean line 129/Sv and other high fat strains sharing the SNP allele of C57BL/6J and DBA/2J.
Bioinformatic analysis of Mir717 was performed using the Ensembl, MGI (http://www.informatics.jax.org/) and Patrocles (http://www.patrocles.org/) and revealed two SNPs in the Mir717. The pre-miRNA SNP (rs30373504) was monomorphic in our tested mouse population, but the polymorphic A>G SNP resided in the mature miRNA (rs30372501). This mature miRNA SNP (miR-SNP) is located within the “seed region”; defined as six or seven nucleotides between the nucleotides 2-7 or 2-8 of the miRNA 5' end that are responsible for mRNA binding [11] (Figure 1C).
The term miR-SNP refers to the variation that occurs in the miRNA gene sequence, whereas miR-TS-SNP refers to the SNP that occurs in the miRNA target site (TS) [11]. MiR-TS-SNPs have been associated with many diseases, including tumor susceptibility [12]. Because one miRNA can have multiple targets, miR-SNPs would be expected to exhibit more profound and broader biological effects than miR-TS-SNPs [11]. The causal effect of the seed miR-SNPs on phenotype variability has been shown recently - two groups discovered that miR-96 seed SNP was responsible for hearing loss in human and mouse [13,14]. Our results further imply the importance of the seed SNP by linking our identified SNP to fat deposition traits.
Additionally, we performed the search for the putative obesity associated Mir717 targets. Obesity genes were collected from the Human obesity gene map [15] (http://obesitygene.pbrc.edu), Rat genome database (RGD) (http://rgd.mcw.edu/), Mouse genome informatics (MGI) (http://www.informatics.jax.org) and Pubmed (http://www.ncbi.nlm.nih.gov/). The search for putative Mir717 targets was performed using DIANA - microT v3.0 target prediction tool (http://diana.cslab.ece.ntua.gr/), TargetScan, Release 5.1 software (http://www.targetscan.org/), MirGen (http://diana.pcbi.upenn.edu/cgi-bin/miRGen/v3/Targets.cgi), miRDB (http://mirdb.org/miRDB/), and PITA (http://genie.weizmann.ac.il/pubs/mir07/index.html). We compiled a total of 2610 genes potentially targeted by Mir717. The analysis revealed only eight targets discovered by all five tools and they are 2010002N04Rik, Camta1, Coq4, Dedd, Dixdc1, Spry1, Syp, and Tyro3. From the database searches of Human obesity gene map, RGD (rat), MGI (mouse) and Pubmed (cattle) 679 fat deposition related genes were collected. Comparison of Mir717 target genes to the obesity gene list revealed that 91 target genes were described previously as obesity related.
Several miRNAs have been shown to play an essential role in adipogenesis, such as miR-27 [16], miR-21 [17], miR-27b [18], and miR-335 [19]. The SNP identified in our study is located in the seed region of the miRNA that has to be complementary to the mRNA in order to recognize the target. MiR-SNP could impact on the catalogue of miRNA targets, not only by disrupting the interaction of the mutant Mir717 with some target genes but also by creating illegitimate targets that are not targeted by the wild type Mir717. Our finding represents a four layer genomic overlap: growth associated QTL, body mass associated Gpc3 gene, highly conserved miRNA gene and seed miRNA SNP which is a relatively rare event. The analyzed Mir717 SNP should now be used in future crosses between the low fat strain 129/Sv and the high fat strains DBA/2J and C57BL/6J to test for direct associations of this marker with obesity/leanness related traits. Furthermore, our bioinformatics analysis also provides a basis for functional annotation of Mir717 orthologs in other species as well as identification of posttranscriptional regulation of fatness-related targets.
Acknowledgements
This work was supported by the Slovenian Research Agency (ARRS) through the Research programme Comparative genomics and genome biodiversity (P4-0220) and project Syntol. This activity was also funded, in part, with an Emerging Research Issues Internal Competitive Grant from the Washington State University, College of Agricultural, Human, and Natural Resource Sciences, Agricultural Research Center to Z.J.
Conflict of Interests
The authors have declared that no conflict of interest exists.
References
1. Huang W, Liu H, Wang T. et al. Tonicity-responsive microRNAs contribute to the maximal induction of osmoregulatory transcription factor OREBP in response to high-NaCl hypertonicity. Nucleic Acids Res. 2010 [Epub ahead of print]
2. Baskerville S, Bartel DP. Microarray profiling of microRNAs reveals frequent coexpression with neighboring miRNAs and host genes. RNA. 2005;11:241-7
3. Li SC, Tang P, Lin WC. Intronic microRNA: discovery and biological implications. DNA Cell Biol. 2007;26:195-207
4. Polster BJ, Westaway SK, Nguyen TM. et al. Discordant expression of miR-103/7 and pantothenate kinase host genes in mouse. Mol Genet Metab. 2010;101:292-5
5. Pilia G, Hughes-Benzie RM, MacKenzie A. et al. Mutations in GPC3, a glypican gene, cause the Simpson-Golabi-Behmel overgrowth syndrome. Nat Genet. 1996;12:241-7
6. Cano-Gauci DF, Song HH, Yang HL. et al. Glypican-3-deficient mice exhibit developmental overgrowth and some of the abnormalities typical of Simpson-Golabi-Behmel syndrome. J Cell Biol. 1999;146:255-64
7. Oliver F, Christians JK, Liu X. et al. Regulatory variation at glypican-3 underlies a major growth QTL in mice. PLoS Biol. 2005;3:872-7
8. Rozman D, McWhir J, Horvat S. Defects in cholesterol synthesis genes in mouse and in humans: lessons for drug development and safer treatments. Drug Metab Rev. in press
9. Horvat S, Medrano JF. Lack of Socs2 expression causes the high-growth phenotype in mice. Genomics. 2001;72:209-12
10. Magin TM, McWhir J, Melton DW. A new mouse embryonic stem-cell line with good germ line contribution and gene targeting frequency. Nucleic Acids Res. 1992;20:3795-6
11. Sun G, Yan J, Noltner K. et al. SNPs in human miRNA genes affect biogenesis and function. RNA. 2009;15:1640-51
12. Nicoloso MS, Sun H, Spizzo R. et al. Single-nucleotide polymorphisms inside microRNA target sites influence tumor susceptibility. Cancer Res. 2010;70:2789-98
13. Mencía A, Modamio-Høybjør S, Redshaw N. et al. Mutations in the seed region of human miR-96 are responsible for nonsyndromic progressive hearing loss. Nat Genet. 2009;41:609-13
14. Lewis MA, Quint E, Glazier AM. et al. An ENU-induced mutation of miR-96 associated with progressive hearing loss in mice. Nat Genet. 2009;41:614-8
15. Rankinen T, Zuberi A, Chagnon YC. et al. The human obesity gene map: the 2005 update. Obesity (Silver Spring). 2006;14:529-44
16. Lin Q, Gao Z, Alarcon RM. et al. A role of miR-27 in the regulation of adipogenesis. FEBS J. 2009;276:2348-58
17. Kim YJ, Hwang SJ, Bae YC. et al. MiR-21 regulates adipogenic differentiation through the modulation of TGF-beta signaling in mesenchymal stem cells derived from human adipose tissue. Stem Cells. 2009;27:3093-102
18. Karbiener M, Fischer C, Nowitsch S. et al. microRNA miR-27b impairs human adipocyte differentiation and targets PPARgamma. Biochem Biophys Res Commun. 2009;390:247-51
19. Nakanishi N, Nakagawa Y, Tokushige N. et al. The up-regulation of microRNA-335 is associated with lipid metabolism in liver and white adipose tissue of genetically obese mice. Biochem Biophys Res Commun. 2009;385:492-6
Author contact
Corresponding author: Dr. Tanja Kunej, Department of Animal Science, Biotechnical Faculty, University of Ljubljana, Domzale, Slovenia. Email: tanja.kunejuni-lj.si. Dr. Zhihua Jiang, Department of Animal Sciences, Washington State University, Pullman, WA 99164-6351. Email: jiangzedu

 Global reach, higher impact
Global reach, higher impact