ISSN: 1449-2288International Journal of Biological Sciences
Int J Biol Sci 2015; 11(8):961-969. doi:10.7150/ijbs.11943 This issue Cite
Research Paper
Exosomal Transfer of Vasorin Expressed in Hepatocellular Carcinoma Cells Promotes Migration of Human Umbilical Vein Endothelial Cells
Beijing Institute of Basic Medical Sciences, Beijing 100850, China
Abstract
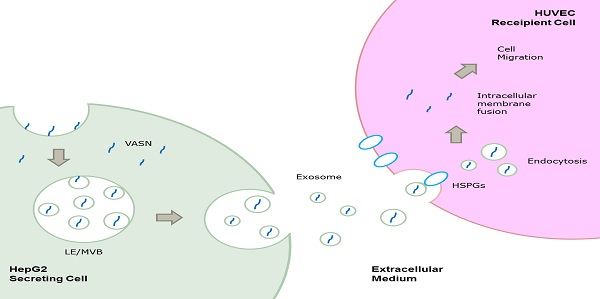
Vasorin (VASN) is a type I transmembrane protein that plays important roles in tumor development and vasculogenesis. In this paper, we showed that VASN could be a key mediator of communication between tumor cells and endothelial cells. We confirmed for the first time that HepG2-derived VASN can be transferred to human umbilical vein endothelial cells (HUVECs) via receptor mediated endocytosis of exosomes, at least in part through HSPGs. The HepG2-derived VASN containing exosomes promote migration of recipient HUVECs cells. Our results identify a novel pathway by which a functional protein expressed in tumor cells affects the biological fate of endothelial cells via exosomes.
Keywords: Vasorin, exosomes, endocytosis, angiogenesis, tumor metastasis
Introduction
Hepatic carcinoma is a common cancer with poor prognosis. China accounts for about 50% of the incidence of liver cancer in the world[1]. We recently identified a serum biomarker of hepatocarcinoma, vasorin (VASN), screened by subtractive-EMSA-SELEX to be highly expressed in sera and tissues samples of hepatocarcinoma[2]. The VASN was further confirmed to promote cell proliferation and migration[2].
VASN, also known as SLIT-like 2, shares structural similarities with large secreted proteins of the SLIT family. It is a transmembrane glycoprotein[3]. The extracellular domain of VASN consists of tandem leucine-rich repeats, an epidermal growth factor (EGF)-like motif, and a fibronectin type III-like motif; this domain is cleaved by tumor necrosis factor-alpha converting enzyme (TACE). The cleaved ectodomain directly binds to transforming growth factor-β (TGF-β) and inhibits TGF-β signaling[4]. The truncated protein is subsequently cleaved by the intramembrane aspartyl protease γ-secretase to release an intracellular domain of approximately 15 kDa[5].
VASN was originally identified as a TGF-β inhibitor that plays a role in neointima formation after vascular injury and partly regulates response to pathological stimuli in blood vessel walls[3, 6, 7]. Slit-like 2, the zebrafish homolog of VASN, is a developmental protein that involve in zebrafish central neural and vascular morphogenesis[6]. Consistent with this expression pattern in zebrafish, VASN expression is found to increase gradually during embryonic development in mice[7].
Recent reports show that VASN is also highly expressed in some cancer cell lines and facilitate the progression of tumor[3, 4, 8, 9]. However, the mechanisms underlying VASN in cancer have hitherto not been elucidated. Because angiogenesis is essential for tumor progression and VASN is an indispensable protein of the vascular system in addition to its role in tumor progression, we hypothesize that VASN might be a mediator between tumor progression and angiogenesis besides it promotes cancerous cell proliferation and migration.
In the present study, we found that VASN could be released and transferred from HepG2 cells to human umbilical vein endothelial cells (HUVECs) via exosomes through HSPGs mediated endocytosis pathway and could promote the migration of HUVECs. Our results helped identify a novel pathway by which a functional protein expressed in tumor cells affects the biological fate of endothelial cells via exosomes.
Methods
Cell culture
Human hepatocellular carcinoma cells HepG2 (ATCC# HB-8065), human embryonic hepatic cells HL-7702 (L02)[10], human cervical carcinoma cell HeLa(ATCC#CCL-2), human leukemic monocyte lymphoma cell U937(ATCC# CRL-1593.2), and HUVECs (ATCC# CRL-1730) were cultured in Dulbecco's Modified Eagle Medium (DMEM; Gibco, USA) containing 10% fetal bovine serum (FBS; Hyclone, USA), 100 units/mL penicillin, and 100 μg/mL streptomycin. The cells were maintained in cell culture flask (Thermo Fisher Scitific, USA) in a humidified chamber at 37°C with 5% CO2.
For antidody blocking procedure, cells (80,000/well) were seeded in a 24-well plate 24h before the antibodies were added to culture. 50 μg/ml mouse monoclonal anti-VASN antibody, nonspecific mouse IgG, or no treatment (mock group) were added to the cell culture and further incubated at 37°C. 24 hours later, cells were washed with 1 mL of PBS and harvested for further experiments.
Quantitative reverse transcriptase-polymerase chain reaction
RNA was isolated using the TRIzol reagent (Invitrogen, USA) according to the manufacturer's instructions, and 1 μl total RNA were performed reverse transcription using with the Moloney murine leukemia virus reverse transcriptase (Promega, USA). For quantitative real-time PCR analysis, we used the Mx3000p Real-Time PCR system (Stratagene, USA) and SYBR Green Realtime PCR Master Mix (Toyobo, Japan). Primers were synthesized by Sangon (China). All measurements were performed in triplicate and the relative VASN mRNA levels were analysed using the Mx3000p software (version 4.1). After normalization with β-actin mRNA, the VASN mRNA level in each group was expressed as fold change relative to the expression level in HepG2 cells.
The following primers were used:
β-actin: forward 5′-GGCATCGTGATGGACTCCG-3′,
reverse 5′-GCTGGAAGGTGGACAGCGA-3′;
VASN: forward 5′-GAGAGCCACGTCACACTGG-3′,
reverse 5′-CAAAGTCGGCGTAGTCAAGC-3′.
Exosome purification
Exosomes were prepared from HepG2 cell supernatants using the ExoQuick-TC™ Exosome Precipitation Solution (System Biosciences, USA) as described by the manufacturer. First, cells were cultured fusion to 60% in 75cm2 cell culture flask in DMEM. Conditioned medium was collected after 24 h of serum-free DMEM incubation and centrifuged at 3000 g for 15 min to precipitate cells and cell debris. Supernatants (50 ml) were ultrafiltered into 1 ml concentrate via Amicon Ultra-15 centrifugal filter(Millipore, Germany). Concentrate was then transferred to a new tube. One mL of the ExoQuick-TC solution was diluted with 5 mL of the supernatant. The resuspended pellet was refrigerated overnight (for at least 12 h) and centrifuged at 1500 g for 30 min on the next day. Finally, the exosomes pellets were resuspended in phosphate-buffered saline (PBS) and stored at -70°C until further use. The density and protein concentration of purified exosomes were determined using the BCA Protein Assay Kit (TianGen, China).
Transmission electron microscopy (TEM)
The exosomes pellets were resuspended in PBS and layered onto carbon-coated electron microscopy grids (Electron Microscopy Sciences, USA). After incubation for 5 min at room temperature, grids were negatively stained with 3% tungstophosphoric acid hydrate for 2 min. The grids were then examined and photographed with a Hitachi H7650 electron microscope (Hitachi, Japan).
DNA and siRNA transfection
Cells at 70% confluency were transfected with vectors or siRNAs. All transfections were performed using Lipofectamine 2000 (Invitrogen) according to the manufacturer's protocol. Plasmids pCMV-myc and pCMV-VASN-myc were purchased from Sino Biological (China). VASN siRNA and scrambled RNA as negative control (NC) were synthesized by GenePharma (China). The siRNA sequences were as follows:
VASN siRNA: sense 5′-CCAACAGGCUGCAUGAAAU-3′ and antisense 5′-GGUUGUCCGACGUACUUUA-3′.
Negative control: sense 5′-UUCUCCGAACGUGUCACGU-3′ and antisense 5′-ACGUGACACGUUCGGAGAA-3′.
Western blotting
Mouse monoclonal anti-VASN antibody was purchased from Abnova, USA. Rabbit polyclonal anti-CD63 antibody was purchased from Abcam, USA. Rabbit anti-HSPG2 antibody was purchased from ProteinTech, China. GAPDH was used as an internal control, and anti-GAPDH antibody was purchased from CoWin Biotech, China. Proteins were prepared by lysing exosomes in radioimmunoprecipitation assay (RIPA) buffer (Applygen, China), separated by sodium dodecyl sulfate-polyacrylamide gel electrophoresis (SDS-PAGE), and transferred via 100 voltage to polyvinylidene difluoride (PVDF) membranes (Millipore). After blocked with 5% milk, the membranes were incubated with primary antibodies overnight at 4°C, followed by incubation with horseradish peroxidase-conjugated secondary antibodies for 1 h. Immunoreactivity was detected via electrochemiluminescence (ECL) by using the SuperSignal® West Pico Chemiluminescent Substrate (Thermo Fisher Scientific).
Immunofluorescence microscopy
HUVECs (5000 cells) were planted on glass cover slip, fixed with methanol, washed with PBS, and blocked with 2% normal rabbit serum. The coverslips were then incubated with primary antibodies overnight and the corresponding FITC-conjugated secondary antibodies at room temperature for 1 h. After antibodies incubation, cells were washed twice with PBS and stained with 4′,6-diamidino-2-phenylindole (DAPI) for 10 minutes. The cells were then viewed under a LSM 510 META confocal microscope (Zeiss, Germany).
Cell proliferation assay
Cell proliferation was determined by the 3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide (MTT) assay. HUVECs were seeded in 96-well culture plates (Thermo Fisher Scientific) at a concentration of 4 × 103 cells per well. After incubation under different conditions for various periods of time (0h, 24h, 48h, 72h), 5 μL of 10 mg/mL MTT was added to each well. The plates were further incubated for 4 h at 37°C and 1M NaOH solution supplemented with 1% SDS were added to each well for at least 12 h. Absorbance was measured at 490 nm with a microplate spectrophotometer (Bio-Tek, USA).
Cell migration assay
Cell migration was determined by the wound-healing assay. HUVECs (1×105 cells per well) were grown to 80% confluence in a 24-well plate. Wounds were made by scraping a conventional pipette tip across the cell monolayer. Cell migration was induced by DMEM supplemented with 10% FBS. The wound images were captured by microscopy (Olympus, Japan) immediately after wounding and after 24 h. The wound width was measured using the ImageJ software, and the percentage of wound healed was calculated using the following formula:
Wounded area filled (%) = 100% - (width after 24h/width at the beginning) × 100%.
Statistical analysis
At least three independent experiments were performed for each assay. Data are shown as means ± SEM. Statistical differences were analyzed by unpaired Student t test, and P-values less than 0.05 were considered statistically significant.
Results
VASN is highly expressed in cancerous HepG2 cells and lower expressed in normal HUVECs cells
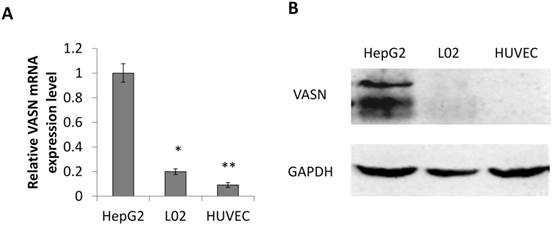
Consistent with our previous result, the expression level of VASN in human hepatocellular carcinoma HepG2 cells is higher than in human embryonic hepatic L02 cells[2]. We further confirmed that human umbilical vein endothelial cell line HUVECs expressed even lower VASN both at mRNA and protein levels (Fig. 1A-B).
VASN is released in the exosomes from HepG2 cells
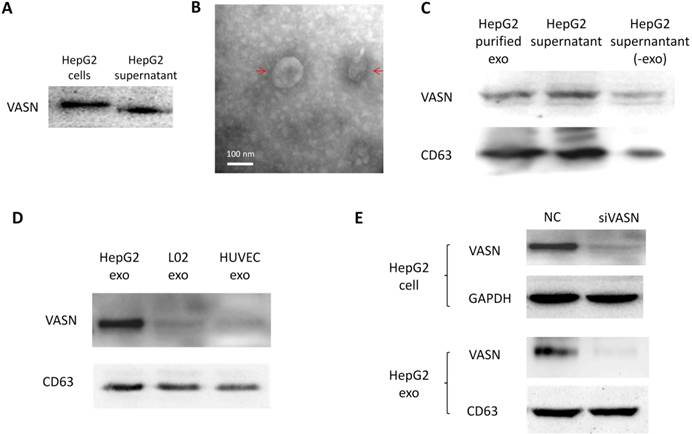
Cancer cells may communicate with endothelial cells by secreting free vascular endothelial growth factors such as VEGF[11], or by releasing membrane vesicles such as “microvesicles” and “exosomes” to transfer functional molecules including “oncoproteins” into recipient cells[12, 13]. We showed that VASN was detectable in HepG2 supernatant (Fig. 2A). The shift rate of VASN in supernatant is fast than that in whole cell extract because the former is cleaved by TACE and lack of intracellular domain. We then purified exosomes from the supernatant of HepG2 cells and confirmed them by TEM (Fig. 2B). VASN expression in exosomes was confirmed by western blotting. VASN was detected both in isolated exosomes and supernatant, but its expression was low in exosomes-depleted supernatant (Fig. 2C). CD63, an exosomal marker protein, was also detecteded. Consistent with the intracellular expression levels of VASN in these cell lines, VASN expression was found to be higher in HepG2-derived exosomes than in L02-derived or HUVECs-derived exosomes (Fig. 2D). When HepG2 cells were treated with VASN siRNA, the expression levels of VASN in exosomes derived from these cells were decreased (Fig. 2E), indicating exosomal protein levels correlate with intracellular VASN expression levels.
VASN secreted from HepG2 cells is transferred to HUVECs via exosomes
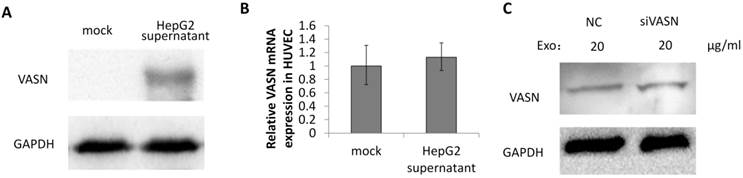
To explore whether VASN is a mediator between tumor progression and angiogenesis, the secreted VASN in HepG2 supernatant was added to culture medium of vascular cell line HUVECs. VASN was up-regulated in whole cell extracts of supernatant-treated HUVECs (Fig. 3A). VASN mRNA levels were unchanged in these cells (Fig. 3B). Furthermore, transfection of VASN siRNA into the co-cultured HUVECs could not prevent the increase in VASN protein levels (Fig. 3C). These results indicate that the source of increased VASN protein was extracellular, i.e., from the supernatant of HepG2 cells.
VASN expression in various cell lines. (A) Real-time PCR analysis of VASN mRNA level in HepG2, L02, and HUVECs cell lines. VASN mRNA level was normalized to that of β-actin as an internal control. Values are represented as means of three independent experiments. (B) VASN expression in lysates of HepG2, L02, and HUVECs cells measured by Western Blot. GAPDH was used as a loading control. *P< 0.05 and **P< 0.01 were considered as statistically significant.
VASN protein secretion and localization. (A) Western blot analysis of VASN protein in cell extracts and supernatant of HepG2 cells. (B) Electron micrograph of exosomes isolated from supernatants of HepG2 cells. Bar represents 100 nm. (C) VASN expression in exosomes, supernatant, and exosomes-depleted supernatants. The exosomal marker protein CD63 was also detected. (D) Western blot analysis of VASN protein in exosomes derived from HepG2, L02, and HUVECs. CD63 was used as a loading control. (E) Western blot analysis of VASN protein in cell lysates (top panel) and exosomes (bottom panel) from HepG2 cells transfected with NC or VASN siRNA (siVASN). GAPDH and CD63 were used as loading controls for cell lysates and exosomes, respectively.
VASN was transferred from HepG2 supernatant to HUVECs. (A) HUVECs were incubated with or without HepG2 supernatant and the cell lysates of HUVECs were subjected to western blotting using the anti-VASN antibody. GAPDH was used as a loading control. (B) Real-time PCR analysis of mRNA from HUVECs after co-cultured with or without HepG2 supernatants. VASN mRNA was normalized by using β-actin as an internal control. Values are represented as means of three independent experiments. (C) HUVECs were transfected with scrambled siRNA (NC) or VASN siRNA, and incubated with HepG2 supernatants, cell lysates were extracted and subjected to western blot analysis using the anti-VASN antibody. GAPDH was used as a loading control.
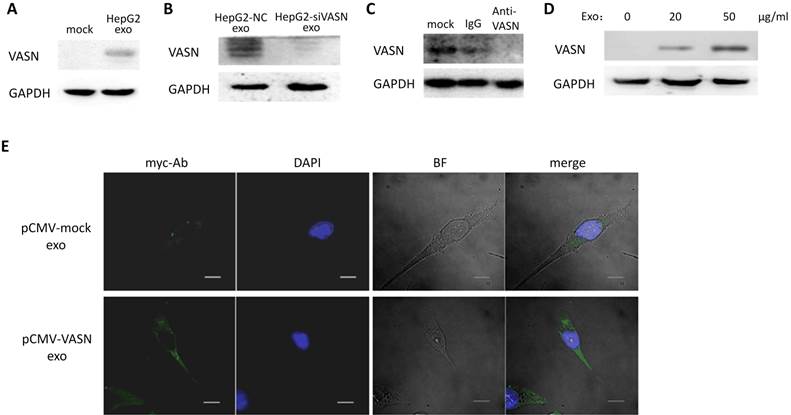
To determine whether VASN could be transferred between two different cell lines via exosomes, we isolated HepG2-derived exosomes and incubated them with HUVECs for 24 h. Result showed that the protein levels of VASN in whole cell extracts of HUVECs were increased (Fig. 4A). Pre-silencing VASN expression in HepG2 with siRNA could block the VASN elevation in HepG2-derived exosomes treated HUVECs cells maybe because of lower VASN in exosomes (Fig. 4B). Similar results were obtained when mouse monoclonal antibody against VASN was added into the co-culture system of HepG2 derived exosomes and HUVECs (Fig. 4C). The transfer of VASN into HUVECs cells by HepG2-derived exosomes showed a dose-dependent manner (Fig. 4D). The exogenous VASN with myc tag was transiently expressed in HepG2 cells, and the internalization of exosomal myc-VASN into HUVECs cells was visualized by immunofluorescence with anti-myc antibody and fluorescence labeled secondary antibody. (Fig. 4E). All the above implies an exosomes specific intercellular transfer of VASN from HepG2 to HUVECs.
The uptake of VASN from HepG2 derived exosomes by HUVECs through HSPGs mediated endocytosis
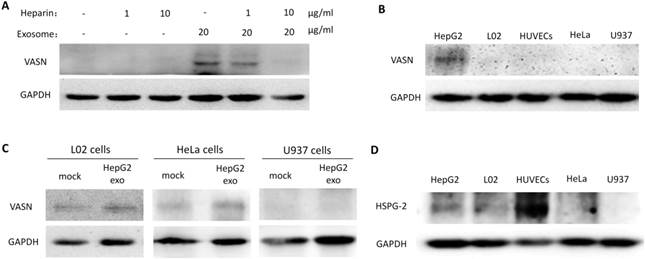
Cells appear to take up exosomes by a variety of mechanisms including endocytosis[14], macropinocytosis[15], phagocytosis[16], and lipid raft_mediated internalization[17]. Among which, the heparin sulphate (HS) proteoglycans (HSPGs) are recently reported to play an important role in the cell surface adsorption and internalization of exosomes[18]. We use heparin (HS mimetic), a competitive inhibitor of cell surface receptors dependent on HSPGs coreceptors to block the HSPGs pathway. After culture the HepG2-derived exosomes with HUVECs in presence of heparin, the increased VASN in recipient cells were decreased in a heparin dose-dependent manner (Fig. 5A). This implies HSPGs involves in the transfer of exosomes VASN into HUVECs.
We isolated HepG2-derived exosomes and incubated them with other VASN low expressing cell lines including HeLa, U937 and LO2 (Fig. 5B) for 24 h. None of the U937, LO2 and HeLa cell lines showed significant changes in VASN abundance (Fig. 5C). Because we showed that HSPGs plays important role in the VASN transfer via exosomes previously, we suspected that the cell type specificity might be due to the different expression, structure or functional epitopes of HSPGs in tested cell lines. We detected the expression level of one member of the HSPGs, HSPG2 in the tested cells. Results showed that HSPG2 is high expressed only in HUVECs (Fig. 5D) while other cells have low expressed HSPG2. Taken together, our data demonstrated that exosomes could act as carriers of VASN protein from HepG2 cells to HUVECs.
Effect of exosomal VASN on proliferation and migration of HUVECs
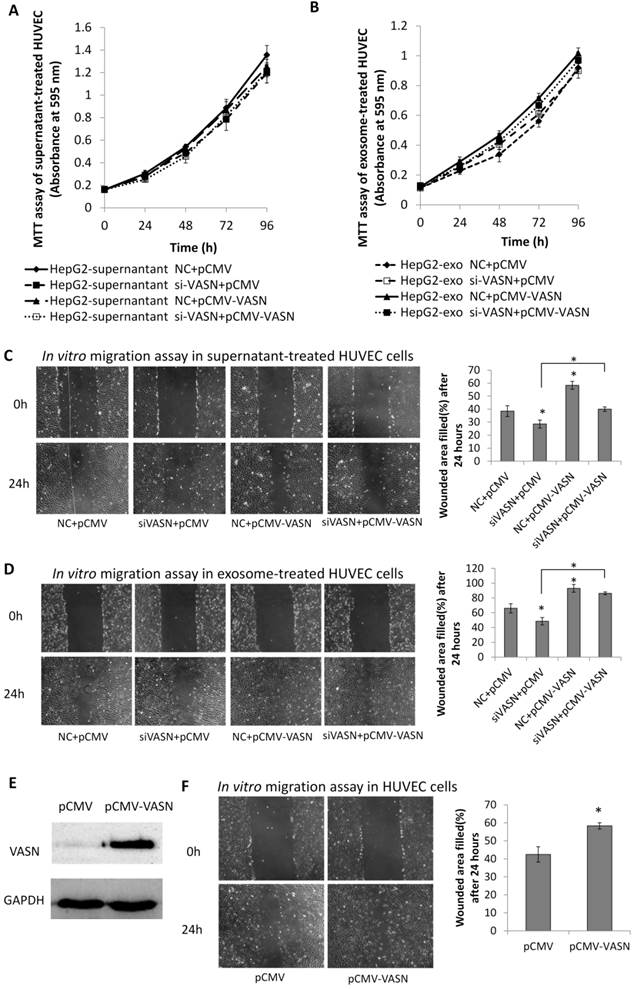
To analyze the biological effects of intercellular transfer of VASN from HepG2 to HUVECs, we evaluated the proliferation and migration of HUVECs treated with HepG2 supernatant and HepG2-derived exosomes. Neither HepG2 supernatant nor HepG2-derived exosomes significantly affected the proliferation of HUVECs (Fig. 6A-B). However, significant effects were observed on HUVECs migration. HUVECs showed decreased migration after treated with supernatants from HepG2 cells expressing VASN siRNA compared with control (Fig. 6C). Stimulation with purified exosomes showed similar effects on migration as HepG2 supernatants (Fig. 6D). In agreement with these findings, intracellular over-expression of VASN promoted HUVECs cell migration (Fig. 6E-F), indicating that VASN transported by HepG2-derived exosomes can effectively accelerate HUVECs migration.
Discussion
Tumors are complex tissues comprising various cell types, such as endothelial cells, mesenchymal cells, cancerous cells, etc[19]. Cell-cell communication between cancer cells and their microenvironment is crucial to tumor progression[20]. As a rich vascular tumor of hepatocarcinoma, the communication between cancer cells and endothelial cells is very important to cancer progression[21].
Transfer of VASN protein from HepG2 to HUVECs via exosomes. (A) Western blot shows increased levels of VASN in HUVECs pretreated with HepG2-derived exosomes. GAPDH was used as a loading control. (B) Western blot analysis of VASN protein in HUVECs treated with exosomes derived from HepG2 cells transfected with NC or VASN siRNAs. GAPDH was used as a loading control. (C) HUVECs were incubated with 20 μg/mL HepG2-derived exosomes in the absence (mock) or in the presence of normal mouse IgG, mouse monoclonal anti-VASN antibody for 24hr in, and then analyzed for VASN levels in the whole cell lysates of incubated HUVECs. GAPDH was used as a loading control. (D) Concentration-dependent uptake of HepG2-derived exosomal VASN in HUVECs. GAPDH was used as a loading control. (E) The exogenous myc tagged VASN expressed in exosomes of HepG2 cells was transferred into HUVECs cells as analyzed by immunofluorescence with anti-myc antibody and fluorescence labeled secondary antibody. HUVECs were incubated with exosomes derived from HepG2 cells transfected with plasmids expressing myc-labeled VASN and analyzed by immunofluorescence under confocal microscopy. DAPI stains showed the nuclear areas.
Exosomal VASN was transferred into HUVECs through HSPGs pathways. (A) HUVECs were co-cultured with HepG2-derived exosomes in presence of different dose of heparin (0, 1 μg/ml, 5 μg /ml, 10 μg /ml) for 24hr. The whole cell lysates of HUVECs were subjected to western blotting using the anti-VASN antibody. GAPDH was used as a loading control. (B) Western blot analysis of VASN express level in HepG2, L02, HUVECs, HeLa, U937 cells; (C) Exosomal VASN cannot be transferred to HeLa, U937 and L02; (D) Western blot analysis of HSPG2 express level in HepG2, L02, HUVECs, HeLa, U937 cells.
Effects of exosomal VASN on cellular processes in HUVECs. (A) Supernatant VASN has no effect on recipient HUVECs proliferation. (B) Exosomal VASN has no effect on recipient HUVECs proliferation. HepG2 cells were transfected with NC or VASN siRNAs and pCMV or pCMV-VASN plasmids. HepG2 supernatants (A) or purified exosomes (B) were collected and incubated with HUVECs for 24 h. MTT assay was performed to determine the effect of these treatments on HUVECs proliferation. (C) Supernatant VASN enhanced the migration of recipient HUVECs. (D) Exosomal VASN enhanced the migration of recipient HUVECs. Wound-healing assay was performed on HUVECs treated with (C) supernatants or (D) exosomes isolated from HepG2 cells transfected with NC or VASN siRNAs and pCMV or pCMV-VASN plasmids. (E) Western blot analysis of VASN protein in HUVECs transfected with pCMV or pCMV-VASN plasmids. (F) HUVECs transiently overexpressed VASN showed increased migration by wound-healing assay. Values are represented as means of three independent experiments. *P < 0.05 was considered statistically significant.
Cells communicate with each other using diverse processes, such as membrane-to-membrane contact[22], ligand-receptor binding[23], or release of soluble mediators[24]. Recent studies suggest that exosomes are small vesicles that are shed from one type of cell and transduce signals to another cell, thereby acting as important mediators in cell-cell communication[25]. Exosomes are 30-100 nm in diameter. These vesicles are formed through fusion of multivesicular bodies (MVBs) with the plasma membrane[26]. Most cells, including lymphocytes[27], dendritic cells[28], reticulocytes[29], epithelial cells[30], mesenchymal stem cells [31]and tumor cells[32], are capable of releasing exosomes. The exact functions of exosomes are still unknown. There are reports regarding exosomes produced by tumor cells have effects on target cells, including stimulation of the target by surface-expressed ligands, receptor transfer between the tumor cell and the target, and horizontal transfer of functional biological molecules into the target et al[33]. Whether or not extracellular vesicles(EVs) uptake is a cell type specific process is controversial. Some believe that exosomes uptake is a specific process that occurs when cell and exosomes share the right combination of ligand and receptor, while others have negative opinions. However, there are certainly examples of cell type specific internalization of exosomes[34, 35].
Here, we identified VASN, as an abundantly expressed protein in cancerous HepG2 cells, could be secreted into cell culture supernatantsand mainly existed in the exosomes. More importantly, VASN was detectable in recipient HUVECs delivered by HepG2-derived exosomes when the two cell lines were co-cultured, and exosomal cell-cell transfer of VASN is dependent on HSPGs mediated internalization of exosomes. Finally, the exosomal divery of VASN correlated with migratory changes in the recipient cells. Furthermore, the data obtained in our study shown that the transportation of VASN are not significant in L02, HeLa and U937 cells, indicating that the exosomal transfer of VASN might be cell type specific. One of HSPGs member, HSPG2 may partially responsible for this cell type specificity. Nevertheless, the detailed molecular interplay mediating this process needs elucidation. It will be interesting to explore the mechanism by which VASN affects recipient cell migration. Further investigation is also required to determine whether VASN inhibition of TGF-β signaling mediates it effect on recipient cell migration.
In conclusion, we identified a novel pathway regulating tumor metastasis via the transmembrane protein VASN. We believe that VASN-positive exosomes provide a metastatic signal from primary tumor cells to the surrounding cells. Exosomes act as a carrier of this signal by mediating cell-cell communication in the progression of cancer, such as hepatocellular carcinoma, and exosomes-mediated transfer of metastatic factors could be a means for cells to induce cell migration in paratumor tissues. In the present study, we discovered that a functional protein highly expressed in a transformed cell could affect the biological fate of an endothelial cell via exosomes-mediated transfer Detailed investigation of this mechanism could help elucidate a novel pathway in the progression of tumor metastasis.
Acknowledgements
We thank the staff of Beijing Institute of Basic Medical Sciences for their assistance.
This work was supported by grants from the Major State Basic Research Development Program of China (973 Program) [grant numbers 2011CB811300, 2010CB912801] and the Natural Science Foundation of China [grant numbers 31100569, 30971630, 31370794].
Competing Interests
The authors have declared that no competing interest exists.
References
1. Cornella H, Alsinet C, Sayols S, Zhang Z, Hao K, Cabellos L. et al. Unique genomic profile of fibrolamellar hepatocellular carcinoma. Gastroenterology. 2015;148:806-18.e10
2. Li S, Li H, Yang X, Wang W, Huang A, Li J. et al. Vasorin is a potential serum biomarker and drug target of hepatocarcinoma screened by subtractive-EMSA-SELEX to clinic patient serum. Oncotarget. 2015;6:10045-59
3. Ikeda Y, Imai Y, Kumagai H, Nosaka T, Morikawa Y, Hisaoka T. et al. Vasorin, a transforming growth factor beta-binding protein expressed in vascular smooth muscle cells, modulates the arterial response to injury in vivo. Proc Natl Acad Sci U S A. 2004;101:10732-7
4. Malapeira J, Esselens C, Bech-Serra JJ, Canals F, Arribas J. ADAM17 (TACE) regulates TGFbeta signaling through the cleavage of vasorin. Oncogene. 2011;30:1912-22
5. Hemming ML, Elias JE, Gygi SP, Selkoe DJ. Proteomic profiling of gamma-secretase substrates and mapping of substrate requirements. PLoS Biol. 2008;6:e257
6. Chen L, Yao JH, Zhang SH, Wang L, Song HD, Xue JL. Slit-like 2, a novel zebrafish slit homologue that might involve in zebrafish central neural and vascular morphogenesis. Biochem Biophys Res Commun. 2005;336:364-71
7. Krautzberger AM, Kosiol B, Scholze M, Schrewe H. Expression of vasorin (Vasn) during embryonic development of the mouse. Gene Expr Patterns. 2012;12:167-71
8. Choksi S, Lin Y, Pobezinskaya Y, Chen L, Park C, Morgan M. et al. A HIF-1 target, ATIA, protects cells from apoptosis by modulating the mitochondrial thioredoxin, TRX2. Mol Cell. 2011;42:597-609
9. Caccia D, Zanetti DL, Micciche F, De Bortoli M, Carniti C, Mondellini P. et al. Secretome compartment is a valuable source of biomarkers for cancer-relevant pathways. J Proteome Res. 2011;10:4196-207
10. Yang Y, Yang L, You QD, Nie FF, Gu HY, Zhao L. et al. Differential apoptotic induction of gambogic acid, a novel anticancer natural product, on hepatoma cells and normal hepatocytes. Cancer Lett. 2007;256:259-66
11. Zhu HF, Wan D, Luo Y, Zhou JL, Chen L, Xu XY. Catalpol increases brain angiogenesis and up-regulates VEGF and EPO in the rat after permanent middle cerebral artery occlusion. Int J Biol Sci. 2010;6:443-53
12. Cho JA, Park H, Lim EH, Lee KW. Exosomes from breast cancer cells can convert adipose tissue-derived mesenchymal stem cells into myofibroblast-like cells. Int J Oncol. 2012;40:130-8
13. Skog J, Wurdinger T, van Rijn S, Meijer DH, Gainche L, Sena-Esteves M. et al. Glioblastoma microvesicles transport RNA and proteins that promote tumour growth and provide diagnostic biomarkers. Nat Cell Biol. 2008;10:1470-6
14. Tian T, Zhu YL, Zhou YY, Liang GF, Wang YY, Hu FH. et al. Exosome uptake through clathrin-mediated endocytosis and macropinocytosis and mediating miR-21 delivery. J Biol Chem. 2014;289:22258-67
15. Fitzner D, Schnaars M, van Rossum D, Krishnamoorthy G, Dibaj P, Bakhti M. et al. Selective transfer of exosomes from oligodendrocytes to microglia by macropinocytosis. J Cell Sci. 2011;124:447-58
16. Feng D, Zhao WL, Ye YY, Bai XC, Liu RQ, Chang LF. et al. Cellular internalization of exosomes occurs through phagocytosis. Traffic. 2010;11:675-87
17. Mulcahy LA, Pink RC, Carter DR. Routes and mechanisms of extracellular vesicle uptake. J Extracell Vesicles. 2014:3
18. Christianson HC, Svensson KJ, van Kuppevelt TH, Li JP, Belting M. Cancer cell exosomes depend on cell-surface heparan sulfate proteoglycans for their internalization and functional activity. Proc Natl Acad Sci U S A. 2013;110:17380-5
19. Wilson J, Balkwill F. The role of cytokines in the epithelial cancer microenvironment. Semin Cancer Biol. 2002;12:113-20
20. Hui EE, Bhatia SN. Micromechanical control of cell-cell interactions. Proc Natl Acad Sci U S A. 2007;104:5722-6
21. Heindryckx F, Gerwins P. Targeting the tumor stroma in hepatocellular carcinoma. World J Hepatol. 2015;7:165-76
22. Mou Y, Wang Y, Li J, Lu S, Duan C, Du Z. et al. Immunohistochemical characterization and functional identification of mammary gland telocytes in the self-assembly of reconstituted breast cancer tissue in vitro. J Cell Mol Med. 2013;17:65-75
23. Uchida N, Lee JS, Horst RJ, Lai HH, Kajita R, Kakimoto T. et al. Regulation of inflorescence architecture by intertissue layer ligand-receptor communication between endodermis and phloem. Proc Natl Acad Sci U S A. 2012;109:6337-42
24. Fan Y, Potdar AA, Gong Y, Eswarappa SM, Donnola S, Lathia JD. et al. Profilin-1 phosphorylation directs angiocrine expression and glioblastoma progression through HIF-1alpha accumulation. Nat Cell Biol. 2014;16:445-56
25. Kourembanas S. Exosomes: vehicles of intercellular signaling, biomarkers, and vectors of cell therapy. Annu Rev Physiol. 2015;77:13-27
26. Pan BT, Teng K, Wu C, Adam M, Johnstone RM. Electron microscopic evidence for externalization of the transferrin receptor in vesicular form in sheep reticulocytes. J Cell Biol. 1985;101:942-8
27. McLellan AD. Exosome release by primary B cells. Crit Rev Immunol. 2009;29:203-17
28. Yang X, Meng S, Jiang H, Chen T, Wu W. Exosomes derived from interleukin-10-treated dendritic cells can inhibit trinitrobenzene sulfonic acid-induced rat colitis. Scand J Gastroenterol. 2010;45:1168-77
29. Blanc L, Barres C, Bette-Bobillo P, Vidal M. Reticulocyte-secreted exosomes bind natural IgM antibodies: involvement of a ROS-activatable endosomal phospholipase iPLA2. Blood. 2007;110:3407-16
30. Walker JD, Maier CL, Pober JS. Cytomegalovirus-infected human endothelial cells can stimulate allogeneic CD4+ memory T cells by releasing antigenic exosomes. J Immunol. 2009;182:1548-59
31. Vallabhaneni KC, Penfornis P, Dhule S, Guillonneau F, Adams KV, Mo YY. et al. Extracellular vesicles from bone marrow mesenchymal stem/stromal cells transport tumor regulatory microRNA, proteins, and metabolites. Oncotarget. 2015;6:4953-67
32. Lundholm M, Schroder M, Nagaeva O, Baranov V, Widmark A, Mincheva-Nilsson L. et al. Prostate tumor-derived exosomes down-regulate NKG2D expression on natural killer cells and CD8+ T cells: mechanism of immune evasion. PLoS One. 2014;9:e108925
33. Camussi G, Deregibus MC, Bruno S, Cantaluppi V, Biancone L. Exosomes/microvesicles as a mechanism of cell-to-cell communication. Kidney Int. 2010;78:838-48
34. Rana S, Yue S, Stadel D, Zoller M. Toward tailored exosomes: the exosomal tetraspanin web contributes to target cell selection. Int J Biochem Cell Biol. 2012;44:1574-84
35. Zech D, Rana S, Buchler MW, Zoller M. Tumor-exosomes and leukocyte activation: an ambivalent crosstalk. Cell Commun Signal. 2012;10:37
Author contact
![]() Corresponding authors: Ningsheng Shao: E-mail: shaonshac.cn, Tel: +86 10 66932311; Jie Li: E-mail: jannbiocom, Tel: +86 10 66932313;
Corresponding authors: Ningsheng Shao: E-mail: shaonshac.cn, Tel: +86 10 66932311; Jie Li: E-mail: jannbiocom, Tel: +86 10 66932313;
Received 2015-2-20
Accepted 2015-5-18
Published 2015-6-11
