ISSN: 1449-2288International Journal of Biological Sciences
Int J Biol Sci 2015; 11(9):1088-1099. doi:10.7150/ijbs.10603 This issue Cite
Research Paper
The Small Molecule R-(-)-β-O-Methylsynephrine Binds to Nucleoporin 153 kDa and Inhibits Angiogenesis
1. Department of Biotechnology, Translational Research Center for Protein Function Control, College of Life Science & Biotechnology, Yonsei University, Seoul 120-749, Republic of Korea
2. Eskitis Institute, Griffith University, Brisbane QLD 4111, Australia
3. Faculty of Health Sciences, University of Macau, Av. Universidade, Taipa, Macau SAR, China
4. Department of Ophthalmology, Seoul National University College of Medicine, Seoul 110-799, Republic of Korea
5. Department of Microbiology and Immunology, Seoul National University College of Medicine, Seoul 110-799, Republic of Korea
6. Department of Internal Medicine, Yonsei University College of Medicine, Seoul 120-752, Republic of Korea
Abstract
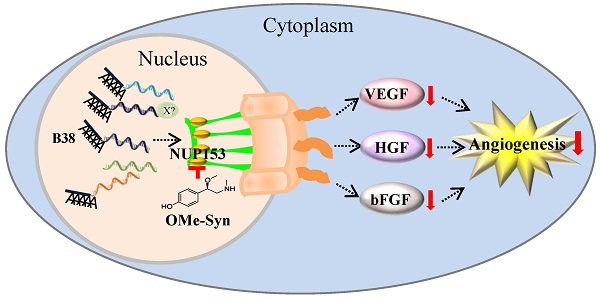
R-(-)-β-O-methylsynephrine (OMe-Syn) is a naturally occurring small molecule that was identified in a previous screen as an inhibitor of angiogenesis. In this study, we conducted two animal model experiments to investigate the in vivo antiangiogenic activity of OMe-Syn. OMe-Syn significantly inhibited angiogenesis in a transgenic zebrafish model as well as in a mouse retinopathy model. To elucidate the underlying mechanisms responsible for the antiangiogenic activity of OMe-Syn, we used phage display cloning to isolate potential OMe-Syn binding proteins from human cDNA libraries and identified nucleoporin 153 kDa (NUP153) as a primary binding partner of OMe-Syn. OMe-Syn competitively inhibited mRNA binding to the RNA-binding domain of NUP153. Furthermore, depletion of NUP153 in human cells or zebrafish embryos led to an inhibition of angiogenesis, in a manner similar to that seen in response to OMe-Syn treatment. These data suggest that OMe-Syn is a promising candidate for the development of a novel antiangiogenic agent and that inhibition of NUP153 is possibly responsible for the antiangiogenic activity of OMe-Syn.
Keywords: Angiogenesis, Natural product, Small molecule, NUP153, Target identification
Introduction
Angiogenesis is a physiological process through which new blood vessels form from existing vasculature. It is an essential process in embryonic development and wound healing and is precisely regulated by a number of endogenous regulatory factors [1]. However, aberrant regulation of angiogenesis is involved in several diseases, including cancer, retinopathy, and macular degeneration [2, 3]. Thus, regulation of pathological angiogenesis by pharmacological agents such as small molecules or monoclonal antibodies has become a promising strategy to treat angiogenesis-related diseases [4, 5].
We previously conducted an unbiased drug screen for angiogenesis inhibitors from a small natural product fragment library with improved aqueous solubility and membrane permeability profiles [6]. A natural product, R-(-)-β-O-methylsynephrine (OMe-Syn) isolated from plants from the Rutaceae family was one of the most potent hits from the screen and was subsequently demonstrated to inhibit vascular endothelial growth factor (VEGF)-induced angiogenesis in vitro and in the chorioallantoic membrane (CAM) in chick embryos [6]. The simple and unique chemical structure and good membrane permeability of OMe-Syn highlight it as a potentially useful candidate in the development of novel antiangiogenic agents. However, a more thorough in vivo demonstration of the antiangiogenic activity of OMe-Syn is required for further preclinical and clinical development of this compound. Moreover, the mechanisms through which OMe-Syn suppresses angiogenesis are not yet understood. Therefore, in this study, we employed two animal models of angiogenesis to investigate the antiangiogenic activity of OMe-Syn in vivo: a transgenic zebrafish model and a mouse oxygen-induced retinopathy model. In addition, we synthesized a biotinylated affinity probe of OMe-Syn to identify potential cellular binding partners. Using a T7 phage display of human cDNA libraries, we identified nucleoporin 153 kDa (NUP153) as one of the potential binding partners of OMe-Syn.
NUP153 is a component of the nuclear pore complex (NPC), which serves as a gated channel to regulate macromolecular transport between the cytoplasm and the nucleus [7, 8]. Misregulation of this gated channel, including modification of the cargo, changes in nuclear transport machinery, or reformation of the NPC itself, may influence cellular function and subsequently lead to tumorigenesis [9]. Among the nucleoporins, NUP153 has been highlighted as a key player in the NPC. NUP153 plays an important role in the export of RNAs and proteins from the nucleus to the cytoplasm [10]. A recent report showed that reduced expression of NUP153 resulted in a delay in mitosis and cell cycle progression without affecting global nucleocytoplasmic transport in cancer cells [11]. Notably, depletion of NUP153 in human breast carcinoma cells affected both cell migration and the cell cycle [12]. These studies indicate that NUP153 may have previously unrecognized roles in cell proliferation and cancer progression in addition to its basic function in nucleocytoplasmic transport.
In this study, we show that OMe-Syn effectively inhibits angiogenesis in both transgenic zebrafish and a mouse retinopathy model. We further report that OMe-Syn binds to NUP153, a key component of the NPC, and inhibits the binding of RNA to NUP153. Depletion of NUP153 in endothelial cells and in zebrafish resulted in the inhibition of angiogenesis in a manner similar to that seen with OMe-Syn treatment; this observation indicates that NUP153 may be a possible molecular target of OMe-Syn.
Materials and Methods
Zebrafish angiogenesis model
Zebrafish were maintained on a 14-hour light/dark cycle at 28 °C. All zebrafish experiments were done according to the National Institute of Health guidelines. Embryos were collected in Ringer's solution according to standard protocols [13]. For the drug treatment, DMSO stock solution of OMe-Syn (10 mM in DMSO) was diluted in Ringer's solution to the appropriate concentration. The final concentration of DMSO did not exceed 1% of the total volume of media. Embryos at the one-cell stage were added to the solutions. For the morpholino (MO) knockdown experiments, zebrafish embryos at the one-cell stage were added to Ringer's solution containing MOs (Gene Tools; Philomath, OR, USA) designed to block translation (5ʹ-TTTTCCCTCCTCCTGTCGCCGCCAT-3ʹ) or splicing (5ʹ-ACATTGAGACATGTTCAGACCTGAT-3ʹ) of NUP153. Tricaine was diluted in Ringer's solution (0.6%, w/v) to anesthetize the embryos for imaging. Zebrafish images were taken with a Fluoview FV1000 confocal microscope (Olympus; Center Valley, PA, USA).
Mouse oxygen-induced retinopathy model
C57BL/6J mice were purchased from Central Laboratory Animal Inc. (Seoul, Republic of Korea). Protocols for animal care, use, and treatment were in agreement with the “ARVO Statement for the Use injected intravitreously once with 20 μM OMe-Syn in 1 μL PBS on P14, when of Animals in Ophthalmic and Vision Research.” Mice were kept at room temperature (~23°C) under a standard 12-hour light/dark cycle. For the induction of mouse oxygen-induced retinopathy (OIR), we followed a procedure described by Smith et al. [14] with some modifications [15]. At postnatal day (P) 7, pups were exposed to hyperoxia (75 ± 0.5% O2) for 5 days (P7-P12) and then returned to normoxia (room air) for 5 days. To assess the antiangiogenic activity of OMe-Syn, the pups were retinal neovascularization begins after hyperoxia treatment. Retinal neovascularization was evaluated at P17, when it peaks after hyperoxia treatment. Ten animals were used in each group.
Assessment of retinal neovascularization by fluorescein angiography
Deeply anesthetized OIR mice were intracardially injected with a fluorescein-conjugated dextran (molecular weight = 500,000; Sigma-Aldrich; St. Louis, MO, USA) in PBS. After 1 hour of perfusion, the eyes were enucleated and fixed in 4% paraformaldehyde in 0.1 M phosphate buffer for 1 hour. The retinas were dissected, flat-mounted and viewed by fluorescence microscopy (Eclipse 80i; Nikon Instruments Inc. Tokyo, Japan) at 40× magnification.
Quantitative assessment of retinal neovascularization
After being fixed for 24 hours, the eyes were embedded in paraffin after fixation for another 24 hours. Sagittal sections (5 μm) were cut at 30 μm intervals through the cornea parallel to the optic nerve. The paraffin sections were stained with hematoxylin and eosin. To quantify retinal neovascularization, vascular lumens on the vitreal side of the inner limiting membrane were counted in 10 sections from each eye in a blinded manner and plotted using the Graphpad prism software (GraphPad Software, San Diego, CA, USA).
Cell culture and proliferation assay
Human umbilical vein endothelial cells (HUVECs, 7-11 passages) were grown in endothelial cell growth medium-2 (EGM-2) using the EGM-2 Bullet Kit (Lonza; Allendale, NJ, USA). HeLa (cervical carcinoma) cells were grown in Dulbecco's modified Eagle's medium containing 10% FBS. The cells were maintained at 37 °C in a humidified 5% CO2 incubator. A cell proliferation colorimetric assay was conducted using 3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide, and cytotoxicity was examined using a trypan blue exclusion assay.
Biotinylation of OMe-Syn
OMe-Syn was purified from a member of the Rutaceae family as described previously [6] and biotinylated as follows. Biotin-N-hydroxysuccinimide (biotin-NHS; 20 mg; Thermo Scientific; Waltham, MA, USA) was added to a solution of OMe-Syn (10 mg) in methanol (3 mL). The reaction was adjusted to pH 8 with sodium hydroxide (0.2 M), stirred for 5 hours at room temperature, and monitored by LC-MS. Upon completion of the reaction, the reaction solvent was evaporated. The crude product was subjected to chromatography on a Betasil C18 column (5 μm, 150 mm×21.2 mm) in a gradient of 100% water to 100% methanol for 50 min at a flow rate of 10 mL/min to produced biotinylated OMe-Syn (biotinyl-OMe-Syn; retention time, 23 min; 90% yield. NMR spectra were recorded at 30 °C on a Varian Inova 600 MHz spectrometer. High-resolution mass measurement was acquired on a Bruker Daltonics Apex III 4.7e Fourier transform mass spectrometer (Bruker Daltonics; Billerica, MA, USA), fitted with an Apollo API source. A Betasil C18 column (5 μm, 150 mm×21.2mm) and a Phenomenex Luna column (C18, 3 μm, 2.0 mm×150 mm) were used for semi preparative HPLC and LC-MS analysis, respectively. All solvents used for the reactions and chromatography were OmniSolv HPLC grade, and the water used was filtered using Millipore Milli-QPF (Billerica, MA, USA): 1HNMR (600 MHz, DMSO-d6, δH), 7.12 (d, 2H, J=8), 6.77 (d, 2H, J=8), 6.42 (s, 1H), 6.36 (s, 1H), 4.27 (m, 1H), 4.24 (t, 1H, J=7), 4.14 (m, 1H), 3.56 (dd, 1H, J=8.2,4.6), 3.39 (m, 2H), 3.10 (s, 3H), 2.85 (m, 2H), 2.81 (s, 3H), 2.22 (m, 2H), 1.61 (m, 2H), 1.45 (m, 2H), 1.30 (m, 2H); 13CNMR (600 MHz, DMSO-d6, δC), 172.3, 162.8, 157.2, 129.8, 127.9 (2C), 115.3 (2C), 80.9, 61.1, 59.2, 56.1, 55.5, 53.6, 40.0, 36.7, 33.8, 32.4, 28.3, 24.8; and (+) HRESIMS m/z 408.1957 [M + H]+ (calculated for C20H30N3O4S1, 408.1951, Δ 1.47 ppm).
Phage display cloning
Biotinyl-OMe-Syn was immobilized onto a streptavidin-coated plate (Thermo Scientific), and phage display was performed using T7 phages encoding human cDNA libraries from colon tumors, lung tumors, and heart tissue (Millipore) as described previously [16]. Specific OMe-Syn-binding phages were isolated and analyzed by DNA sequencing. The sequence homologies of the obtained sequences were compared with sequences in the GenBank database by using the BLAST program.
Cloning and purification of the RNA-binding domain of NUP153
A PCR-amplified N-terminal (amino acids [aa] 235-326) fragment of NUP153 cDNA (NM_005124) was cloned into the EcoRI/SalI site of the pET-28a expression vector (Millipore). The plasmid was transformed into RosettaTM (DE3) Escherichia coli, and the RNA-binding domain (RBD) of NUP153 was purified using Ni-NTA agarose beads (Qiagen, Valencia, CA, USA). Bead-bound protein was eluted with elution buffer containing a high concentration of imidazole, according to the manufacturer's instructions.
Competitive gel electrophoretic mobility shift assay
The DNA template used to synthesize the RNA probe was prepared by hybridizing the following DNA oligonucleotides: 5ʹ-TAATACGACTCACTATAG-3ʹ and 5ʹ-GGCGGCCGCTCTTGAACTAGTGGATCCCCCGGGCTGCACTATAGTGAGTCGTATTA-3ʹ. The resulting template was used for in vitro transcription with the T7MEGAscript Kit (Life Technologies; Grand Island, NY, USA) and [α-32P]UTP to produce an internally labeled version of B38, a 38-nucleotide RNA that is derived from pBluescript-SK(+) and has been shown to bind to the NUP153-RBD [17]. Gel-purified radioisotope-labeled B38 (20 fmol) was incubated for 1 hour at 4 °C with the NUP153-RBD (2 μM), which had been purified by Ni-affinity chromatography as described previously [17], in a total volume of 10 μL of binding buffer containing 25 mM HEPES (pH 7.5), 100 mM KCl, 0.5% Triton X-100, 0.5 mg/mL bovine albumin, and 7.5% glycerol. For the competitive electrophoretic mobility shift assay (EMSA), increasing amounts (0.04, 0.4, and 1.0 mM) of NUP153 inhibitors were preincubated with the NUP153-RBD for 1 hour at 4 °C prior to incubation with the labeled B38. Samples were electrophoresed at 4 °C on a pre-run 20% native acrylamide gel. Phosphorimage analysis and quantification was performed using a Fuji BAS-2500 phosphorimager. The experiment was conducted three times independently.
RNA interference of NUP153
The StealthTM RNA Interference Kit (Life Technologies) was used to synthesize small interfering RNA (siRNA) specific to human NUP153. For knockdown of NUP153 mRNA, HUVECs were transfected with either control siRNA or various concentrations of NUP153 siRNA by using the LipofectamineTM 2000 transfection reagent (Life Technologies) according to the manufacturer's instructions. Knockdown efficiency of endogenous NUP153 was determined by Western blot analysis.
Western blot analysis
The cell lysates were separated by 10% SDS-PAGE and then transferred to PVDF membranes (Millipore) using standard electroblotting procedures. The blots were then blocked and incubated overnight at 4 °C with anti-NUP153 (Abcam, Cambridge, MA, USA), anti-VEGF (Abcam), anti-basic fibroblast growth factor (Abcam), anti-hepatocyte growth factor (Abcam), and anti-tubulin (Millipore) primary antibodies. Immunolabeling was detected using an enhanced chemiluminescence kit (GE Healthcare, Pittsburgh, PA, USA) according to the manufacturer's instructions.
Preparation of conditioned medium (CM) from HeLa cell culture
HeLa cells (7×104 cells/well) were seeded in a 12-well plate. Cells were then transfected with a scrambled siRNA or siRNA specific for human NUP153 for 24 hours. Conditioned medium (CM) from each siRNA treated HeLa cells was collected and concentrated using the Amicon ultra-0.5 centrifugal filter unit (Millipore). The CMs from each culture condition of HeLa cells were then used for HUVEC experiments in serum-depleted media.
Capillary tube formation assay
A 48-well plate was coated with Matrigel (10 mg/mL), which was allowed to polymerize for 1 hour at 37 °C. HUVECs (6×104 cells) were then placed on the surface of the Matrigel. The CM from NUP153-depleted HeLa cells was added the cells and the incubation was continued for 4-16 hours at 37°C. The HUVEC tube formation was observed under an Olympus IX71 microscope and photographed at 100× magnification with a DP70 camera (Olympus).
Chemoinvasion assay
To examine the invasiveness of HUVECs in vitro, we used a Transwell chamber (Corning Costar; Tewksbury, MA, USA) system according to the manufacturer's instruction. Briefly, the lower side of the filter was coated with gelatin (10 μL, 1 mg/mL), and the upper side was coated with Matrigel (10 μL, 3 mg/mL). The CM from NUP153-depleted HeLa cells was added to the lower part of the chamber, and HUVECs (7×105 cells) were placed in the upper part. Cells were then incubated at 37 °C for 18 hours, and were fixed with 70% methanol and stained with hematoxylin and eosin. Cell invasiveness was measured by counting the number of cells in the lower side of the filter using a microscope at 100× magnification.
Statistical analysis
Results are expressed as mean ± standard error (SE). Student's t-test was used to determine statistical significance between two groups. To compare the mean of the test group with a hypothetical value (control), one sample t-test was used. A p-value that was <0.05 was considered statistically significant.
Results
OMe-Syn inhibits intersegmental vessel angiogenesis in fli1a:EGFP transgenic zebrafish embryos
We previously reported that OMe-Syn inhibited VEGF-induced angiogenesis in vitro and in a CAM angiogenesis model [6]. In the present study, the antiangiogenic activity of OMe-Syn was investigated using fli1a:EGFP transgenic zebrafish, a well-established model for visualizing angiogenesis in zebrafish. Prior to assessing angiogenesis, we tested several concentrations of OMe-Syn to find an optimal drug dosage for the treatment in zebrafish. Since treatment of zebrafish with OMe-Syn at higher than 10 μM showed a sign of toxicity and morphological change, we used 5 μM as a maximal safe dose of OMe-Syn for subsequent zebrafish experiments (data not shown). Zebrafish embryos (n=30 per group) at the one-cell stage were incubated in medium containing DMSO or OMe-Syn (2.5 and 5 μM). Angiogenesis was assessed by counting the number of complete intersegmental vessels (ISV) at 30 hours post fertilization (hpf). Treatment of the zebrafish embryos with OMe-Syn significantly and dose-dependently inhibited ISV formation (Fig. 1A-C). The compound at this concentration range was not significantly toxic to zebrafish embryos, as assessed by the viability of the embryos (Fig. 1D). These data demonstrate that OMe-Syn is a potent inhibitor of angiogenesis in a zebrafish model.
OMe-Syn inhibits retinal neovascularization in mice
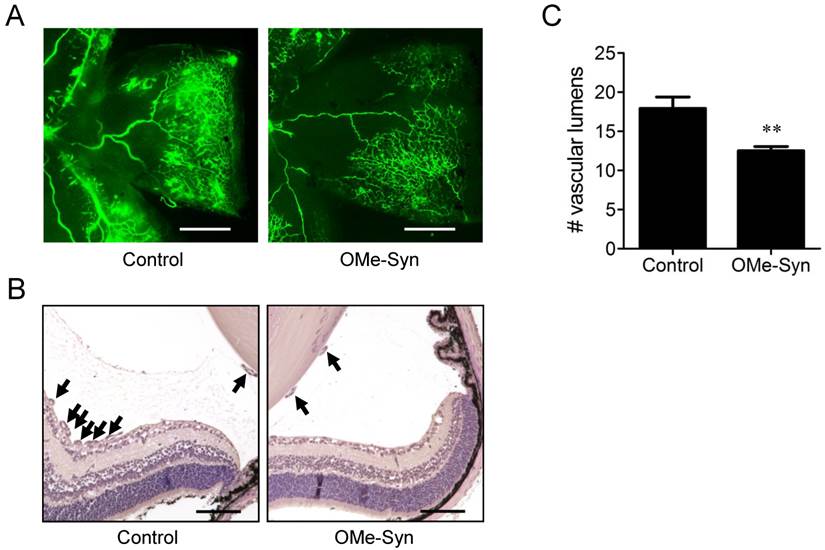
To further investigate the in vivo antiangiogenic activity of OMe-Syn, we employed a mouse model of OIR [14]. This is a well-established animal model for stimulating retinal neovascularization with high reproducibility and is used to assess the activity of antiangiogenic agents [18]. OMe-Syn was intravitreously injected once into OIR mice (10 mice per group) on P14, and the inhibitory effect of the compound was examined at P17. To qualitatively evaluate the effect of OMe-Syn on retinal neovascularization, retinal flat mounts were evaluated with fluorescein-conjugated dextran. Neovascular tufts of intravitreal neovascularization and vascular leakage were observed at the border of vascular and avascular retina in control OIR mice (Fig. 2A and B). In OMe-Syn-treated OIR mice, however, the neovascular tufts and vascular tortuosity were decreased. To quantitatively evaluate the effect of OMe-Syn on retinal neovascularization, vascular lumens between the posterior surface of the lens and the anterior surface of the inner limiting membrane were counted in a blinded manner. We found that the average number of vascular lumens in OMe-Syn-treated OIR mice (12.4 ± 0.9) was significantly lower than that in control OIR mice (17.9 ± 2.7), suggesting the possibility of OMe-Syn as a new candidate treatment for angiogenesis-related diseases (Fig. 2C).
The effect of R-(-)-β-O-methylsynephrine on angiogenesis in fli1a:EGFP transgenic zebrafish embryos. Zebrafish embryos (n=30 per group) at the one-cell stage were used for the compound treatment. (A) Light microscopic images of whole embryos at 30 hours post fertilization (hpf) are shown. (B) Fluorescence images show dose-dependent suppression of intersegmental vessel (ISV) formation (indicated by arrows) by R-(-)-β-O-methylsynephrine (OMe-Syn). (C) Complete ISVs were counted at 30 hpf and quantified using Graphpad Prism software; **p<0.01 versus control (Student's t-test). (D) The viability of zebrafish embryos at 30 hpf upon treatment with OMe-Syn is shown.
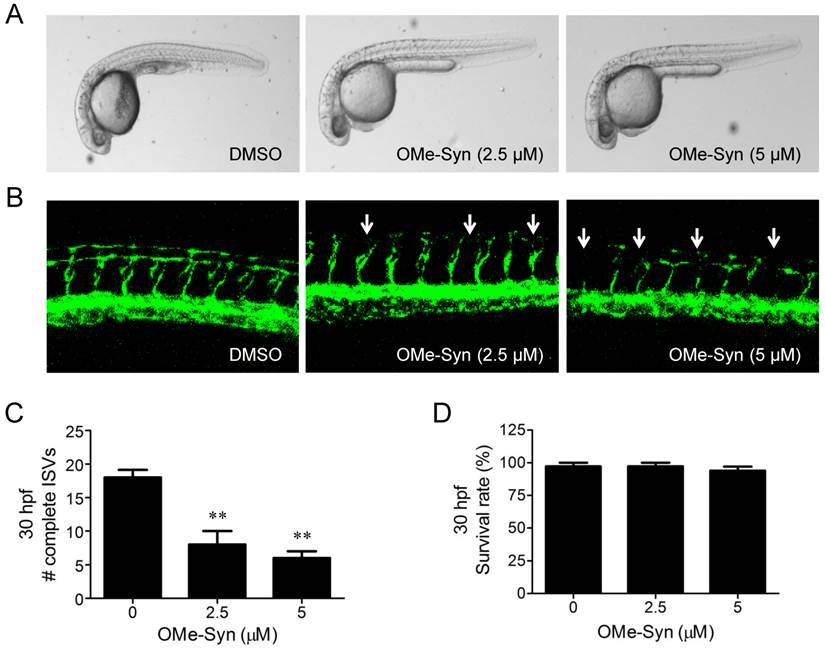
The effect of R-(-)-β-O-methylsynephrine on retinal angiogenesis in mice with oxygen-induced retinopathy. The compound was intravitreously injected once into the mice (10 mice per group) on P14, and the inhibitory effect of the compound was examined at P17. (A) Retinal vasculatures in control and R-(-)-β-O-methylsynephrine (OMe-Syn; 20 μM)-treated mice with oxygen-induced retinopathy (OIR) were evaluated with fluorescein angiography. Neovascular tufts of intravitreous region were observed at the border of vascular and avascular retina. Scale bar: 500 μm. (B, C) Vascular lumens were counted to quantify retinal neovascularization in OIR mice. Cross-sections were prepared from postnatal day 17 control and OMe-Syn (20 μM)-treated mice for hematoxylin and eosin staining. Arrows indicate the vascular lumens of new vessels growing into the vitreous region. The number of vascular lumens was counted in randomly selected images under 200× magnification. Scale bar: 100 μm; **p<0.01 versus control (Student's t-test).
OMe-Syn binds to the RNA-binding domain of NUP153
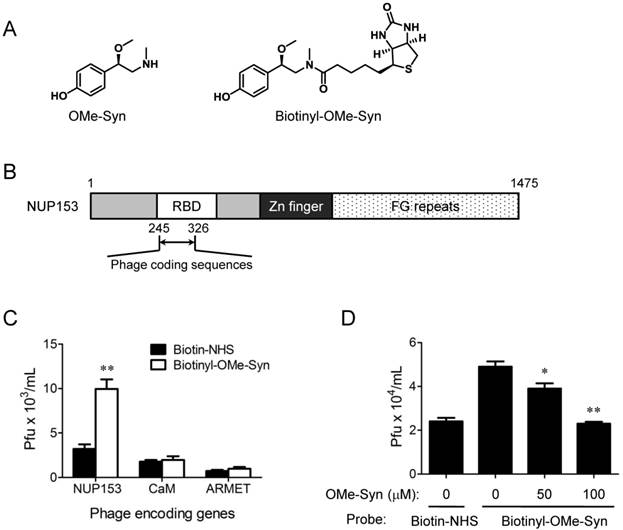
OMe-Syn has a simple but unique phenethylamine core structure (Fig. 3A) that is distinct from other known angiogenesis inhibitors, implying that it may have a unique mechanism of action in endothelial cells. To explore possible mechanisms of action for OMe-Syn, we attempted to identify OMe-Syn binding proteins from human cDNA libraries using phage display cloning, which is a well-established affinity-based method for the identification of small molecule target proteins [16, 19, 20]. We first synthesized biotinyl-OMe-Syn by coupling NHS-biotin to the amino group of OMe-Syn (Fig. 3A). To test whether biotinyl-OMe-Syn retains its original biological activity, we examined the compound in endothelial cell proliferation assays. Biotinyl-OMe-Syn inhibited the proliferation of HUVECs in a dose-dependent manner similar to that shown by OMe-Syn (Supplementary Fig. S1), suggesting that this compound is a suitable molecular probe for identifying OMe-Syn binding proteins. Biotinyl-OMe-Syn was then immobilized on a streptavidin-coated plate, and T7 phages encoding human cDNAs (a combined pool of cDNAs from colon tumor, lung tumor, and heart tissues) were incubated on the immobilized OMe-Syn to isolate proteins that bind OMe-Syn. These T7 phages typically display 50-200 amino acid protein fragments on the surface of the capsid, which provides the secondary structure of the protein for binding to the small molecule. Phage particles specifically bound to biotinyl-OMe-Syn were enriched through four rounds of biopanning and the resulting phage plaques were randomly selected for sequencing analysis of their cDNA inserts. Total 18 phage clones were selected for sequencing and the results were summarized in the Supplementary Table 1. Among the 18 phage clones sequenced, two of them had an open reading frame (ORF) that encodes 82 amino acids identical to the N-terminal region (245-326aa) of human NUP153. Interestingly, the coding region is the RNA-binding domain (RBD) within the N-terminal region of NUP153 (Fig. 3B). Based on the sequencing data, only three genes including NUP153 (2 clones), calmodulin (2 clones) and mesencephalic astrocyte-derived neurotrophic factor precursor (ARMET, 7 clones) were properly inserted into the phage plasmid (Supplementary Table 1). The other two genes including Sjogren's syndrome/scleroderma autoantigen 1 (1 clone) and putative p150 (2 clones) were frame-shifted in the phage plasmid and therefore were not studied further. The sequences of the remaining 4 clones had no significant sequence similarity found from the database, thus being removed from further analysis (data not shown). The NUP153, calmodulin and ARMET phage clones were individually subjected to the biotinyl-OMe-Syn binding assays and only the NUP153 phages showed a reproducible binding to the immobilized biotinyl-OMe-Syn (Fig. 3C). In a phage competition assay, pre-incubation of NUP153-RBD phages with OMe-Syn blocked the binding of the phages to the immobilized biotinyl-OMe-Syn in a dose-dependent manner (Fig. 3D), further demonstrating the specific binding of OMe-Syn to NUP153-RBD. These data suggested that NUP153 is a possible binding protein of OMe-Syn in mammalian cells.
OMe-Syn inhibits RNA binding to NUP153
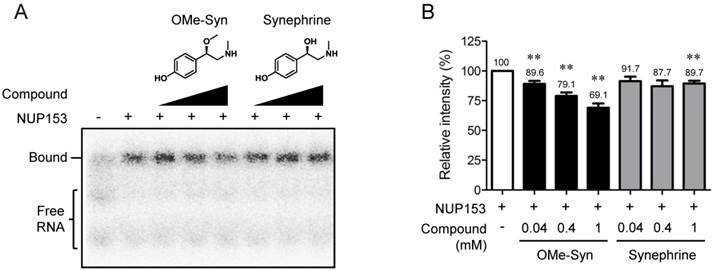
NUP153 plays an important role in nuclear export of several classes of RNA [10]. The RBD was identified within N-terminal region of NUP153 (amino acids 250-400 in human NUP153), which is responsible for binding to RNA [21]. To examine effect of OMe-Syn on RNA binding to NUP153-RBD, we expressed a recombinant NUP153-RBD protein and performed a competitive EMSA with the radioisotope-labeled B38 RNA. The ability of the radioisotope-labeled RNA probe to bind to the RBD of NUP153 was characterized in detail in a previous study [17]. As a result, OMe-Syn dose-dependently inhibited the binding of B38 RNA to NUP153-RBD, whereas synephrine, a biologically inactive analogue of OMe-Syn did not significantly block the binding except at the highest concentration (Fig. 4A and B). These data demonstrated that OMe-Syn blocked RNA association to NUP153 in vitro.
Phage display cloning to identify R-(-)-β-O-methylsynephrine binding proteins expressed by human cDNA libraries. (A) Chemical structures of R-(-)-β-O-methylsynephrine (OMe-Syn) and its biotinylated analog are shown. (B) Domain structure of nucleoporin 153 kDa (NUP153) and the phage coding region within the NUP153 RNA-binding domain (RBD) are shown. (C) The binding of NUP153-encoding phages to biotinylated OMe-Syn (biotinyl-OMe-Syn) is shown. Other candidates, including calmodulin-encoding phages (CaM) or mesencephalic astrocyte-derived neurotrophic factor precursor-encoding phages (ARMET) do not show significant binding to biotinyl-OMe-Syn. Data represent mean ± standard error from four independent experiments; **p<0.01 versus N-hydroxysuccinimide biotin-NHS (Student's t-test). (D) A phage competition assay using NUP153-encoding phages is shown. NUP153-encoding phages were preincubated with DMSO or OMe-Syn (50 or 100 μM), and the resulting mixture was used for a binding assay on immobilized biotinyl-OMe-Syn. Data represent mean ± standard error from four independent experiments; *p<0.05 and **p<0.01 versus control (Student's t-test).
The effect of R-(-)-β-O-methylsynephrine on the association of RNA with the RNA-binding domain of nucleoporin 153 kDa. (A) A competitive electrophoretic mobility shift assay was performed using a radiolabeled B38 RNA and the purified recombinant RNA-binding domain (RBD) of nucleoporin 153 kDa (NUP153). Various concentrations of R-(-)-β-O-methylsynephrine (OMe-Syn) and its inactive analog synephrine were tested for the competition of binding between B38 RNA and the NUP153-RBD. The representative image from three independent experiments is shown. (B) The relative intensity of the signals from the RNA-protein complexes was analyzed by phosphorimager. Data represent mean ± standard error from three independent experiments; **p<0.01 versus no compound control (One sample t-test).
NUP153 depletion inhibits angiogenesis in vitro
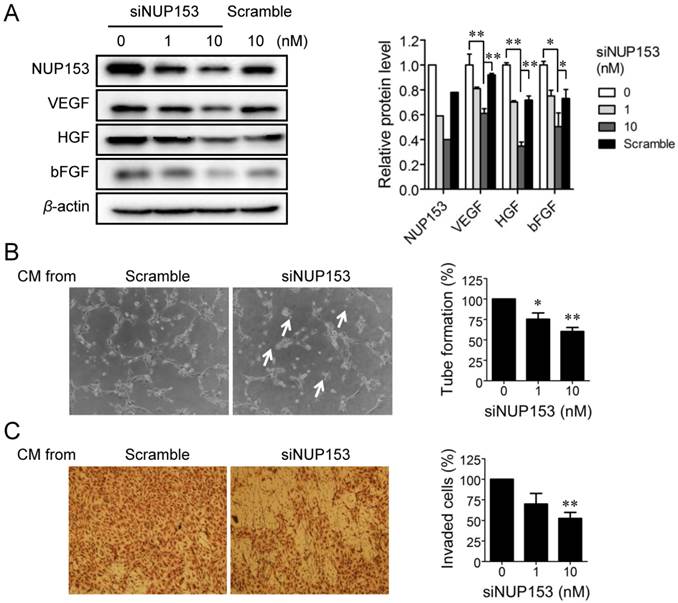
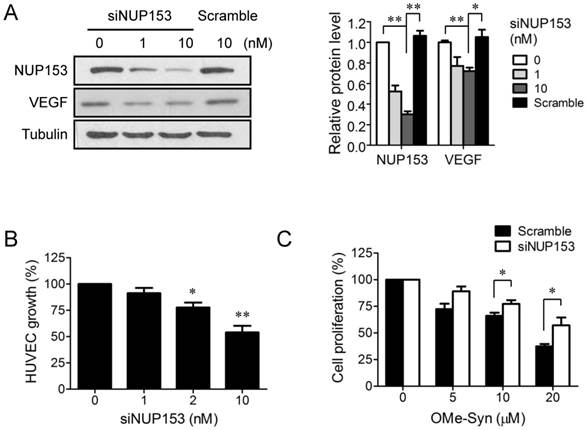
Although NUP153 has been known to be involved in cancer cell mitosis and cancer progression, its functional association with endothelial cell proliferation and angiogenesis has not been reported yet. To test whether inhibition of NUP153 function is responsible for the antiangiogenic activity of OMe-Syn, we knocked down NUP153 in human cancer and endothelial cells and then assessed their angiogenesis phenotypes. HeLa cells were treated with siRNA against NUP153, and then the protein levels of angiogenic growth factors including VEGF, basic fibroblast growth factor (bFGF), and hepatocyte growth factor (HGF) were analyzed. Surprisingly, NUP153 depletion in HeLa cells suppressed the expression of VEGF, bFGF, and HGF (Fig. 5A) in a manner similar to that seen with OMe-Syn treatment (Supplementary Fig. S2). Next, CM from HeLa cells in which NUP153 had been depleted was used to treat HUVECs, and then in vitro angiogenesis was assessed through tube formation and chemoinvasion assays. The CM from HeLa cells that had been treated with scrambled siRNA was used as a control. Compared to the control CM, the CM from NUP153-depleted HeLa cells resulted in a dose-dependent decrease in tube formation and chemoinvasion in HUVECs (Fig. 5B and C). The CM from NUP153-depleted HeLa cell culture also led to a decrease in HUVEC proliferation compared to the CM from scrambled siRNA-treated HeLa cells (Supplementary Fig. S3). Next, HUVECs were treated with siRNA for NUP153 and examined its effect on cell proliferation. Depletion of NUP153 in HUVECs reduced the level of VEGF and suppressed HUVEC proliferation (Fig. 6A and B). To determine whether the cell growth inhibition by OMe-Syn was dependent on the inhibition of NUP153, the effect of OMe-Syn on the proliferation of NUP153-depleted HUVECs was examined. Treatment with OMe-Syn resulted in weaker growth inhibition in NUP153-depleted HUVECs than in wild type HUVECs; this result indicates that the inhibition of HUVEC proliferation by OMe-Syn was at least in part through the inhibition of NUP153 (Fig. 6C).
NUP153 depletion inhibits angiogenesis in zebrafish
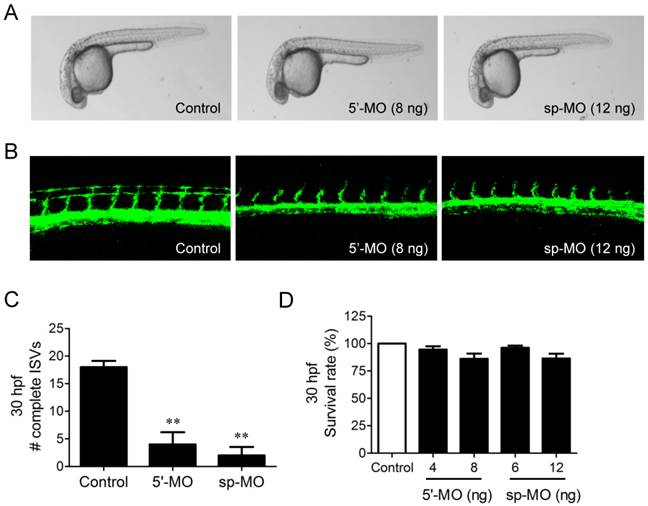
We further examined the possible involvement of NUP153 in angiogenesis in a zebrafish angiogenesis model. We used two different MOs specific to NUP153 to inhibit NUP153 protein expression in zebrafish, as follows. NUP153-specific MOs that block NUP153 splicing (n=31) or translation (n=28) were injected into fli1a:EGFP transgenic embryos at the one-cell stage. The blockade of NUP153 splicing in zebrafish was monitored by RT-PCR (Supplementary Fig. S4). Angiogenesis was assessed by counting the number of completely formed ISVs in zebrafish embryos. Treatment of the zebrafish embryos with either type of MO significantly inhibited ISV formation in the embryos at 30 hpf (Fig. 7A-C). The inhibitory effect of NUP153-specific MOs on zebrafish angiogenesis was much stronger than that of OMe-Syn (Fig. 1B and C), suggesting that the MOs acted more specifically on NUP153 than the small molecule OMe-Syn. There were subtle decreases in the viability of zebrafish embryos upon treatment with higher concentrations of NUP153-specific MOs, although the decreases were not statistically significant (Fig. 7D); this observation is presumably because developmental angiogenesis is more strongly inhibited by MOs than by OMe-Syn. These data demonstrate that inhibition of NUP153 was sufficient to block angiogenesis in vivo.
Discussion
Natural products are useful sources for drug development owing to their structural diversity and low toxicity. Of the wide variety of natural products, we were particularly interested in groups of compounds with favorable physicochemical properties for drug development, such as high water solubility and absorption rate. Therefore, we assembled a small natural product fragment library under following criteria: MW<250, clogP≤4, hydrogen bond donor≤4, hydrogen bond acceptor≤5, percentage polar surface<45, and rotational bond ≤6 [6]. From this library, we have previously identified OMe-Syn as a novel antiangiogenic natural product. Here, we investigated the in vivo efficacy of the OMe-Syn using two well-established animal models of angiogenesis: zebrafish and mouse OIR. OMe-Syn significantly inhibited angiogenesis in both animal models without apparent systemic toxicity. These results demonstrated the in vivo therapeutic efficacy of OMe-Syn. The zebrafish model is used to study angiogenesis and vasculogenesis during embryo development, hence enabling the assessment of physiological angiogenesis. In contrast, the OIR mouse model is used to study the pathological angiogenesis occurring mainly during the development of retinopathy of prematurity [22]. Interestingly, OMe-Syn inhibited angiogenesis in both the developmental model and the pathological model. These results suggest that this compound inhibits fundamental step(s) in angiogenesis. Here, we have shown that OMe-Syn decreases expression levels of VEGF, bFGF, and HGF, which are angiogenic growth factors responsible for activating receptors on endothelial cells. This could partially explain how OMe-Syn inhibited both types of angiogenesis. However, the underlying mechanisms that lead to a decrease in angiogenic growth factors by OMe-Syn in cancer or in endothelial cells remained to be elucidated.
The effect of nucleoporin 153 kDa knockdown via small interfering RNA on angiogenesis in cancer cells in vitro. (A) HeLa cells were treated with small interfering RNA against nucleoporin 153 kDa (siNUP153), and then the protein levels of nucleoporin 153 kDa (NUP153), vascular endothelial growth factor (VEGF), basic fibroblast growth factor (bFGF), and hepatocyte growth factor (HGF) were analyzed by Western blot. Scrambled small interfering RNA (Scramble) was used as the negative control. Quantitation of each protein level is shown in the bar graph. The level of β-actin protein expression was used for normalization; *p<0.05 and **p<0.01 between 0 vs 10 nM siNUP153 or 10 nM siNUP vs scrambled siRNA (Student's t-test). (B, C) Human umbilical vein endothelial cells (HUVECs) were incubated with the conditioned media (CM) from HeLa cells treated with either siNUP153 or Scramble, and then tube formation (B) and invasion (C) assays were performed. HUVEC tube formation and invasion were quantified and plotted using Graphpad Prism (right panels). Arrows indicate broken tube networks following NUP153 knockdown. Data represent mean ± standard error from four independent experiments; *p<0.05 and **p<0.01 versus control (One sample t-test).
The effect of nucleoporin 153 kDa knockdown via small interfering RNA on cell proliferation in endothelial cells. (A) Human umbilical vein endothelial cells (HUVECs) were treated with small interfering RNA against nucleoporin 153 kDa (siNUP153), and then the protein levels of nucleoporin 153 kDa (NUP153) and vascular endothelial growth factor (VEGF) were analyzed by Western blot. Scrambled small interfering RNA (Scramble) was used as the negative control. Quantitation of each protein level is shown in the bar graph. The level of tubulin was used for normalization; *p<0.05 and **p<0.01 between 0 vs 10 nM siNUP153 or 10 nM siNUP vs scrambled siRNA (One sample t-test for NUP153 and Student's t-test for VEGF). (B) HUVECs were treated with various concentrations of Scramble or siNUP153for 48 hours, and then cell proliferation was assessed using 3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide; *p<0.05 and **p<0.01 versus control (One sample t-test). (C) HUVECs were treated with 10 nM Scramble or siNUP153 for 24 hours. The cells were then treated with various concentrations of R-(-)-β-O-methylsynephrine (OMe-Syn) for 72 hours, and then cell proliferation was assessed. The values were normalized to the no-drug control from each siRNA treatment; *p<0.05 between the indicated two groups (Student's t-test).
The effect of morpholino-based knockdown of nucleoporin 153 kDa (NUP153) in zebrafish on in vivo angiogenesis. (A) Bright-field microscope images of whole embryos are shown at 30 hours post fertilization (hpf). Two types of morpholinos (MO) were used to knockdown NUP153 in zebrafish: 5ʹ-MO, which is a translational inhibitor of NUP153, and sp-MO, which is a splicing inhibitor of NUP153. (B) Zebrafish intersegmental (ISV) formation at 30 hpf was observed under a confocal microscope. (C) Complete ISVs were counted at 30 hpf and quantified using Graphpad Prism software; **p<0.01 versus control (Student's t-test). (D) Viability of zebrafish embryos at 30 hpf upon treatment with the MOs.
The phage display cloning approach has been successfully employed for identification of small molecule binding partners [16, 19, 20]. It is particularly useful for identifying small molecules with low binding affinities [16, 20]. As this approach utilizes the “enrichment” of binders, it also makes it possible to identify binding partners that are expressed at a very low level. Using phage display cloning with biotinyl-OMe-Syn as the affinity probe, we found that the NUP153-RBD is a binding partner of OMe-Syn. Specific binding of OMe-Syn to the NUP153-RBD was verified through binding competition assays. To test if the binding of OMe-Syn to NUP153 affected the function of NUP153, we used purified recombinant NUP153-RBD to conduct an isotope-labeled RNA binding assay, which is a well-established system for studying the RNA transport function of NUP153 [17]. OMe-Syn, but not its inactive analog, competitively inhibited the binding of B38 RNA to the recombinant NUP153-RBD, thus demonstrating the inhibitory effect of OMe-Syn on the association of RNA with NUP153. Although we showed that OMe-Syn binds to NUP153-RBD and inhibits its RNA-binding capacity, it is unclear whether NUP153 is the primary binding protein responsible for the biological activity of OMe-Syn. In this study, we were unable to show the binding of OMe-Syn to the endogenous full-length NUP153 in whole cell lysates. However, this assay was quite challenging to perform because OMe-Syn has a low affinity (micromolar concentration range) for NUP153-RBD and does not have covalent modifier functional groups that could be used to successfully pull down the full-length protein. Therefore, the development of a photo affinity linker conjugated OMe-Syn will greatly help in verifying whether NUP153 binds to endogenous full-length NUP153.
To test whether the inhibition of NUP153 function by OMe-Syn is sufficient to cause antiangiogenic activity, NUP153 was depleted using siRNA in cultured cells or MOs in zebrafish embryos, and the resulting angiogenesis phenotypes were observed. The depletion of NUP153 in cancer or endothelial cells caused a decrease in the levels of angiogenic growth factors, including VEGF, bFGF, and HGF, and inhibited angiogenesis in CM-based assays. The depletion of NUP153 in zebrafish embryos also resulted in a striking degree of angiogenesis inhibition. These in vitro and in vivo results demonstrated that inhibition of NUP153 was sufficient to suppress angiogenesis. To determine whether there was a causal relationship between the antiangiogenic activity of OMe-Syn and inhibition of NUP153, NUP153-depleted cells were treated with OMe-Syn. Compared to wild type endothelial cells, NUP153-depleted cells showed a significant degree of resistance to OMe-Syn treatment with respect to cell proliferation. These data strongly suggest that OMe-Syn inhibits angiogenesis, at least in part, through the inhibition of NUP153.
Although we have provided substantial evidence to show that OMe-Syn binds to the NUP153-RBD and that depletion of NUP153 inhibits angiogenesis, it is unclear how the regulation of global RNA transport could specifically affect angiogenesis. This is a critical question that will be addressed in subsequent studies. Previously, in an unbiased genome-wide RNAi screen, NUP153 was identified as a regulator of mitosis [23]. Depletion of NUP153 caused a delay in mitosis and cell cycle progression without affecting global nucleocytoplasmic transport [11]. It is interesting to speculate that NUP153 may play an important role in endothelial cell mitosis and cell cycle progression independently of its roles in global nucleocytoplasmic transport. On the other hand, the fact that either OMe-Syn treatment or NUP153 depletion caused a decrease in the levels of angiogenic growth factors indicates an involvement of gene expression regulation in response to NUP153 inhibition. NUP153 is located on the surface of the basket structure of nuclear pore and hence is one of the first pore components that mRNA cargo binds during export [10]. Because mRNA export is one of essential steps in gene expression in eukaryotic cells, the inhibition of NUP153RBD could largely impact the expression of certain genes, especially those genes whose messages or products have a short half-life. The half-lives of growth factors such as VEGF and bFGF are known to be relatively short (several tens of minutes) [24-26]. Further studies on endothelial cell mitosis and global nucleocytoplasmic transport upon treatment with OMe-Syn or depletion of NUP153 will help to decipher the underlying mechanisms in greater detail.
In conclusion, the present study shows that OMe-Syn has promise as a novel antiangiogenic agent and suggests that NUP153 is a possible molecular target mediating the antiangiogenic activity of OMe-Syn. With its simple structure and potent bioavailability, OMe-Syn is an excellent lead compound for the development of antiangiogenic agents; thus, further investigation into its optimization with respect toNUP153 binding is warranted.
Supplementary Material
Supplementary Fig.S1, S2, S3 and S4; Supplementary Table 1.
Acknowledgements
This work was supported by grants from the National Research Foundation of Korea funded by the Korea government (MSIP; 2009-0092964, 2009-0083522, 2010-0017984, 2012M3A9D1054520), the Center for Food and Drug Materials of Agriculture Science &Technology Development (PJ0079772012), the Rural Development Administration, the National R&D Program, the Ministry of Health &Welfare (0620360-1), the Brain Korea 21Project of the Republic of Korea, and the Queensland Smart State Innovation Project Fund-National and International Research Alliances and Partnerships Program.
Competing Interests
The authors have declared that no competing interest exists.
References
1. Ashton N. Retinal angiogenesis in the human embryo. Br Med Bull. 1970;26:103-6
2. Folkman J. Tumor angiogenesis: therapeutic implications. N Engl J Med. 1971;285:1182-6
3. Ng EW, Adamis AP. Targeting angiogenesis, the underlying disorder in neovascular age-related macular degeneration. Can J Ophthalmol. 2005;40:352-68
4. Ellis LM. Bevacizumab. Nat Rev Drug Discov. 2005(Suppl):S8-9
5. Llovet JM, Ricci S, Mazzaferro V, Hilgard P, Gane E, Blanc JF. et al. Sorafenib in advanced hepatocellular carcinoma. N Engl J Med. 2008;359:378-90
6. Kim NH, Pham NB, Quinn RJ, Kwon HJ. R-(-)-beta-O-methylsynephrine, a natural product, inhibits VEGF-induced angiogenesis in vitro and in vivo. Biochem Biophys Res Commun. 2010;399:20-3
7. Ahn NG. PORE-ing over ERK substrates. Nat Struct Mol Biol. 2009;16:1004-5
8. Vasu SK, Forbes DJ. Nuclear pores and nuclear assembly. Curr Opin Cell Biol. 2001;13:363-75
9. Kau TR, Way JC, Silver PA. Nuclear transport and cancer: from mechanism to intervention. Nat Rev Cancer. 2004;4:106-17
10. Ullman KS, Shah S, Powers MA, Forbes DJ. The nucleoporin nup153 plays a critical role in multiple types of nuclear export. Mol Biol Cell. 1999;10:649-64
11. Mackay DR, Elgort SW, Ullman KS. The Nucleoporin Nup153 Has Separable Roles in Both Early Mitotic Progression and the Resolution of Mitosis. Molecular Biology of the Cell. 2009;20:1652-60
12. Zhou LX, Pante N. The nucleoporin Nup153 maintains nuclear envelope architecture and is required for cell migration in tumor cells. Febs Letters. 2010;584:3013-20
13. Westerfield M. The zebrafish book: a guide for the laboratory use of zebrafish (Brachydanio rerio). Eugene, OR: M. Westerfield. 1993
14. Smith LEH, Wesolowski E, Mclellan A, Kostyk SK, Damato R, Sullivan R. et al. Oxygen-Induced Retinopathy in the Mouse. Investigative Ophthalmology & Visual Science. 1994;35:101-11
15. Zhang D, Kaufman PL, Gao G, Saunders RA, Ma JX. Intravitreal injection of plasminogen kringle 5, an endogenous angiogenic inhibitor, arrests retinal neovascularization in rats. Diabetologia. 2001;44:757-65
16. Shim JS, Lee J, Park HJ, Park SJ, Kwon HJ. A new curcumin derivative, HBC, interferes with the cell cycle progression of colon cancer cells via antagonization of the Ca2+/calmodulin function. Chemistry & Biology. 2004;11:1455-63
17. Ball JR, Dimaano C, Bilak A, Kurchan E, Zundel MT, Ullman KS. Sequence preference in RNA recognition by the nucleoporin Nup153. Journal of Biological Chemistry. 2007;282:8734-40
18. Park SW, Kim JH, Kim KE, Jeong MH, Park H, Park B. et al. Beta-lapachone inhibits pathological retinal neovascularization in oxygen-induced retinopathy via regulation of HIF-1a. Journal of Cellular and Molecular Medicine. 2014;18:875-84
19. Piggott AM, Karuso P. Rapid identification of a protein binding partner for the marine natural product kahalalide F by using reverse chemical proteomics. Chembiochem. 2008;9:524-30
20. Jung HJ, Shim JS, Lee J, Song YM, Park KC, Choi SH. et al. Terpestacin Inhibits Tumor Angiogenesis by Targeting UQCRB of Mitochondrial Complex III and Suppressing Hypoxia-induced Reactive Oxygen Species Production and Cellular Oxygen Sensing. Journal of Biological Chemistry. 2010;285:11584-95
21. Dimaano C, Ball JR, Prunuske AJ, Ullman KS. RNA association defines a functionally conserved domain in the nuclear pore protein Nup153. Journal of Biological Chemistry. 2001;276:45349-57
22. Sears JE, Hoppe G, Ebrahem Q, Anand-Apte B. Prolyl hydroxylase inhibition during hyperoxia prevents oxygen-induced retinopathy. Proceedings of the National Academy of Sciences of the United States of America. 2008;105:19898-903
23. Rines DR, Gomez-Ferreria MA, Zhou Y, DeJesus P, Grob S, Batalov S. et al. Whole genome functional analysis identifies novel components required for mitotic spindle integrity in human cells. Genome Biology. 2008:9
24. Dibbens JA, Miller DL, Damert A, Risau W, Vadas MA, Goodall GJ. Hypoxic regulation of vascular endothelial growth factor mRNA stability requires the cooperation of multiple RNA elements. Mol Biol Cell. 1999;10:907-19
25. Eppler SM, Combs DL, Henry TD, Lopez JJ, Ellis SG, Yi JH. et al. A target-mediated model to describe the pharmacokinetics and hemodynamic effects of recombinant human vascular endothelial growth factor in humans. Clin Pharmacol Ther. 2002;72:20-32
26. Murphy PR, Guo JZ, Friesen HG. Messenger RNA stabilization accounts for elevated basic fibroblast growth factor transcript levels in a human astrocytoma cell line. Mol Endocrinol. 1990;4:196-200
Author contact
![]() Corresponding author: Ho Jeong Kwon, Department of Biotechnology, Translational Research Center for Protein Function Control, College of Life Science & Biotechnology, Yonsei University, Seoul 120-749, Republic of Korea. Email: kwonhjac.kr; Fax: +82 2 362 7265; Tel: +82 2 2123 5883
Corresponding author: Ho Jeong Kwon, Department of Biotechnology, Translational Research Center for Protein Function Control, College of Life Science & Biotechnology, Yonsei University, Seoul 120-749, Republic of Korea. Email: kwonhjac.kr; Fax: +82 2 362 7265; Tel: +82 2 2123 5883
Received 2014-9-22
Accepted 2015-1-21
Published 2015-7-16
