ISSN: 1449-2288International Journal of Biological Sciences
Int J Biol Sci 2016; 12(1):100-108. doi:10.7150/ijbs.13498 This issue Cite
Review
Genome Wide Sampling Sequencing for SNP Genotyping: Methods, Challenges and Future Development
1. Department of Animal Sciences, Washington State University, Pullman, WA 99164-7620, USA;
2. Key Laboratory of Agricultural Animal Genetics, Breeding and Reproduction of Ministry of Education and The Cooperative Innovation Center for Sustainable Pig Production, Huazhong Agricultural University, Wuhan, China;
3. Department of Pediatrics, Human and Molecular Genetics Center and Children's Research Institute, Medical College of Wisconsin, Milwaukee, WI 53226, USA;
4. Department of Integrative Physiology and Neuroscience, Washington State University College of Veterinary Medicine, Pullman, WA 99164-7620, USA.
Abstract
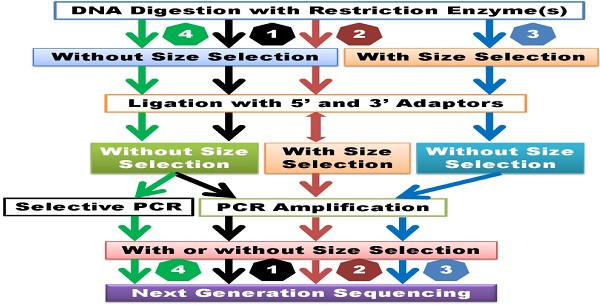
Genetic polymorphisms, particularly single nucleotide polymorphisms (SNPs), have been widely used to advance quantitative, functional and evolutionary genomics. Ideally, all genetic variants among individuals should be discovered when next generation sequencing (NGS) technologies and platforms are used for whole genome sequencing or resequencing. In order to improve the cost-effectiveness of the process, however, the research community has mainly focused on developing genome-wide sampling sequencing (GWSS) methods, a collection of reduced genome complexity sequencing, reduced genome representation sequencing and selective genome target sequencing. Here we review the major steps involved in library preparation, the types of adapters used for ligation and the primers designed for amplification of ligated products for sequencing. Unfortunately, currently available GWSS methods have their drawbacks, such as inconsistency in the number of reads per sample library, the number of sites/targets per individual, and the number of reads per site/target, all of which result in missing data. Suggestions are proposed here to improve library construction, genotype calling accuracy, genome-wide marker density and read mapping rate. In brief, optimized GWSS library preparation should generate a unique set of target sites with dense distribution along chromosomes and even coverage per site across all individuals.
Keywords: restriction site associated DNA sequencing, genotyping by sequencing, reduced representation library sequencing, complexity reduction of polymorphism sequencing, dominant and co-dominant markers, genome wide association study, human, animals and plants
