10
Impact Factor
ISSN: 1449-2288
Int J Biol Sci 2005; 1(2):67-79. doi:10.7150/ijbs.1.67 This issue Cite
Review
The Multiple Personalities of the Regulatory Subunit of Protein Kinase CK2: CK2 Dependent and CK2 Independent Roles Reveal a Secret Identity for CK2β
Department of Biochemistry, Siebens-Drake Research Institute, University of Western Ontario, London, Ontario, Canada, N6A 5C1
Received 2004-10-1; Accepted 2005-2-1; Published 2005-4-1
Abstract
Protein kinase CK2 (formerly casein kinase II), an enzyme that participates in a wide variety of cellular processes, has traditionally been classified as a stable tetrameric complex consisting of two catalytic CK2α or CK2α' subunits and two regulatory CK2β subunits. While consideration of CK2 as a tetrameric complex remains relevant, significant evidence has emerged to challenge the view that its individual subunits exist exclusively within these complexes. This review will summarize biochemical and genetic evidence indicating that the regulatory CK2β subunit exists and performs functions independently of CK2 tetramers. For example, unbalanced expression of catalytic and regulatory CK2 subunits has been observed in a variety of tissues and tumors. Furthermore, localization studies including live cell imaging have demonstrated that while the catalytic and regulatory subunits of CK2 exhibit extensive co-localization, independent mobility of the individual CK2 subunits can also be observed within cells. Identification of proteins that interact with CK2β in the absence of catalytic CK2 subunits reinforces the notion that CK2β has functions distinct from CK2 and begins to offer insights into these CK2-independent functions. In this respect, the discovery that CK2β can interact with and modulate the activity of a number of other serine/threonine protein kinases including A-Raf, c-Mos and Chk1 is particularly striking. This review will discuss the interactions between CK2β and these protein kinases with special emphasis on the properties of CK2β that mediate these interactions and on the implications of these interactions in yielding new prospects for elucidation of the cellular functions of CK2β.
Keywords: Protein kinase CK2, CK2β, protein kinase, CK2-independent interactions, cyclin
1. Introduction
Despite the fact that protein kinase CK2 (formerly known as casein kinase II) was discovered approximately 50 years ago it remains a perplexing biological molecule [1] . It is now abundantly clear that it is a promiscuous enzyme as a diverse and somewhat bewildering array of more than 300 potential substrates have been identified [2] . CK2 is a serine/threonine kinase that participates in a wide variety of cellular processes including cell differentiation, proliferation and survival [3, 4] . Genetic studies in organisms such as yeast and slime mould have revealed that CK2 is essential for viability [5, 6] . Recent studies of molecular clock machinery in Drosophila and Arabidopsis have also provided evidence for the involvement of CK2 in circadian oscillator function [7, 8, 9, 10] . Collectively, these studies demonstrate that CK2 participates in the regulation of processes that are fundamental to many aspects of life. Accordingly, it is not surprising that perturbations in the expression of CK2 are associated with human disease.
There is an increasing body of evidence indicating that CK2 is involved in protein kinase networks controlling cell cycle progression and cellular responses to stress that are associated with various cancers. In this respect, CK2 is involved in pathways that respond to a variety of stresses including ultraviolet light, anisomycin, heat shock, tumor necrosis factor α and arsenite [11, 12, 13, 14] . Investigations in yeast and mammalian cells have revealed requirements for CK2 at various stages of the cell cycle including G1 phase and the G1/S and G2/M transitions [15, 16, 17, 18] . Furthermore, abnormally high levels of CK2 have been observed in various types of cancer (breast, prostate, lung, kidney, head and neck) and in transformed cells [19, 20, 21, 22, 23] . In a related vein, CK2 co-operates with several oncogenes including c-Myc, Tal-1 and Ha-Ras which subsequently leads to transformation [24, 25, 26, 27, 28] . A direct link has been established between CK2 and tumorigenesis in transgenic mice, where in T-cells and mammary glands targeted expression of CK2 leads to lymphomagenesis and mammary tumors [24, 23] . Based on this involvement in transformation and tumorigenesis, CK2 has recently attracted attention as a potential therapeutic target [29, 30] . This participation in cancer also emphasizes the importance of understanding comprehensively how CK2 works as an enzyme and how it functions to regulate specific biological events.
2. CK2: General Features and Subunit Composition
2.1 The Catalytic CK2 Subunits
CK2 is ubiquitously expressed in eukaryotic cells and exhibits extensive sequence and functional conservation across species. As compared to the catalytic subunits of many other protein kinases, one notable feature of the catalytic subunits of CK2 is that they possess constitutive activity. In this respect, the catalytic subunits of CK2 exhibit enzymatic activity when expressed as individual recombinant proteins in bacteria [31, 32] . Traditionally, the physiological significance of this observation was questioned since the catalytic subunits of CK2 appeared to be accompanied by stoichiometrically equivalent quantities of the regulatory CK2β subunit when CK2 was purified from most sources including yeast, flies and mammalian tissues [33, 34, 35, 36] . Consequently, CK2 has typically been viewed as a tetrameric complex consisting of two catalytic subunits (38-42kDa in mammals) and two regulatory subunits (27kDa in mammals) [2] .
In humans, two different forms of its catalytic subunits (designated CK2α or CK2α'), which are encoded by distinct genes, were initially characterized [34] . With the exception of their unrelated C-terminal domains, these two isoforms are very similar to one another exhibiting approximately 90% identity within their catalytic domains. Recently a third isoform (designated CK2α'') that is almost completely identical to CK2α with respect to the predicted amino acid sequence of its catalytic domain was also identified [37] . In fact, the only significant distinguishing feature between CK2α and CK2α'' lies in their completely distinct C-terminal domains. While it is clear that the different CK2 isoforms are closely related and exhibit considerable functional overlap, there is also evidence for functional specialization of the individual CK2 isoforms in yeast, mice and mammals [3] . Since this issue has been addressed in detail elsewhere it will not be further discussed.
2.2 The Regulatory CK2 Subunit
In contrast to the catalytic isoforms of CK2, only one known form of the regulatory subunit (designated CK2β) has been identified in mammals [38] . CK2β does not display extensive homology with other protein kinase regulatory subunits, but is remarkably conserved among species [39, 38] . In fact, the amino acid sequence of CK2β is completely identical between birds and mammals [40] . Studies conducted by several different groups using a variety of approaches including X-ray crystallography have determined that a dimer of the CK2β subunits forms the core of the CK2 tetramer (Figure 1A) [41, 42, 43, 44, 45, 46] . CK2β has several notable features that have been characterized by an extensive body of work (summarized in Figure 1). As a prelude to a detailed consideration of emerging CK2-independent functions of CK2β, the following discussion will summarize a number of its notable features.
2.3 CK2β: Phosphorylation sites
A large proportion of CK2β has been shown to be phosphorylated at an autophosphorylation site consisting of serines 2, 3 and possibly 4 at its N-terminus [47, 48] . Based on kinetic measurements, it was previously concluded that autophosphorylation occurs by an intramolecular mechanism [49] . However, this view was challenged by the determination of the high-resolution structure of tetrameric CK2, which revealed that the N-terminus of CK2β was located more than 40Å away from the active site of either of the catalytic subunits [50] . Thus, an intermolecular reaction mediated by the formation of higher order CK2 structures is hypothesized [3] . Although the precise function of this autophosphorylation remains unknown, studies conducted by Zhang et al. suggest that phosphorylation of these sites enhances CK2β stability [51] . CK2β is also phosphorylated at S209 near its C-terminus, a residue which is phosphorylated in a cell-cycle dependent manner by p34cdc2 in vitro and in mammalian cells [47, 52, 53] . The functional significance of this latter phosphorylation site remains unknown.
2.4 CK2β: Putative Degradation Motifs
As the regulatory subunit of a protein kinase that has functions associated with cell cycle progression, CK2β is reminiscent of cyclins that are the regulatory subunits of cyclin-dependent kinases. In a related respect, it is particularly intriguing that CK2β has motifs that have been previously characterized as motifs that regulate cyclin degradation. For example, CK2 has a sequence resembling that of the nine amino acid motif called the destruction box that plays a key role in the specific degradation of cyclin B at the end of mitosis [54, 55] . This motif, which was first recognized in CK2β by Allende and Allende, contains three highly conserved residues conforming to the general destruction box consensus RXXLXXXXN/D [38, 55] . Interestingly, this motif is located on a surface exposed α-helix (Figure 1) where it would be available for recognition by the cellular degradation machinery. Several proteins including mitotic cyclins, Aurora A and B, and Nek2A also contain one or more KEN boxes, a signal which has also been shown to play a role in mediating cell cycle dependent protein degradation [56, 57, 58, 59] . This degradation motif is characterized by the minimal consensus sequence KEN, but is often followed shortly by either an N or D residue and often preceded by another N or D reside (N/DKENX0-4N/D). Notably, CK2β contains a similar sequence, namely DKFNLTGLN between amino acids 32 and 40. With the exception of a single mutation within its destruction box that did not exert any apparent affect, the functional relevance of the putative destruction and KEN boxes of CK2β have not yet been characterized [51] . Nevertheless, given the intriguing parallels between CK2β and cyclins, more detailed investigation of the functional significance of these motifs is warranted.
CK2β: The regulatory subunit of CK2 A. High-resolution crystal structure of the CK2 holoenzyme. The catalytic subunits are represented as an alpha carbon trace (grey) while the CK2β dimer forming the core of the enzyme is represented by blue ribbons. Important motifs are coloured as indicated in the schematic diagram (B). CK2β monomers are distinguished by different shades of the appropriate colour. (prepared using Swiss PDB Viewer; 1JWH [116, 117, 50] ) B. Schematic illustration of CK2β depicting important regions. The phosphorylation sites are represented by black spheres; while the KEN box (orange) and D box (yellow) represent putative degradation motifs. The acidic loop is involved in modulation of catalytic subunit activity by mediating polyamine binding. Cysteines 109, 114, 137 and 140 of the zinc-finger region mediate CK2β dimer formation and the positive regulatory region mediates binding between CK2β and the catalytic subunits. (adapted from [3] )
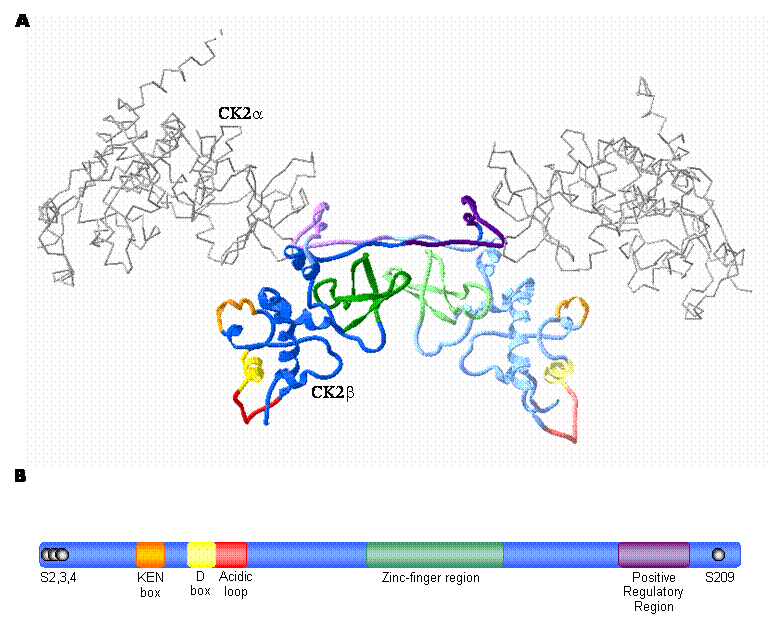
2.5 CK2β: CK2 Subunit Interactions
Beginning in the 1990's, a number of groups examined the mechanisms responsible for CK2 subunit interactions [41, 42, 43, 44, 45, 46] . In accordance with these studies, the elucidation of the structure of CK2β by X-ray crystallography revealed the importance of the zinc-finger in CK2β dimerization [44] . The zinc-finger region is characterized by four cysteine residues (109, 114, 137 and 140) which mediate the interaction allowing the CK2β dimer to form the core of the CK2 holoenzyme [44, 46] . As expected, disruption of the zinc-finger by mutation of Cys 109 and 114 resulted in a loss of CK2β dimer formation [46] . Intriguingly, it was observed that the zinc-finger mutants also failed to interact with the CK2 catalytic subunits both in vitro and in vivo [46] . Studies examining the interactions between the catalytic and regulatory subunits during CK2 assembly revealed that CK2β dimerization precedes catalytic subunit binding and in fact was a prerequisite for CK2 tetramer formation [45] . Each catalytic subunit of the CK2 tetramer is associated with one CK2β molecule at its positive regulatory domain delineated by residues 181-203 in the CK2β C-terminus [43, 44, 60] (Figure 1). This region, which was confirmed by the high-resolution structure of tetrameric CK2, is responsible for the ability of CK2β to enhance and stabilize CK2 activity [43, 44, 60, 50] .
One additional noteworthy sequence within CK2β is a surface exposed acidic loop encoded by residues 55-64 [61, 62] . Given the presence of a basic cluster between residues 74-80 in CK2α it had been tempting to propose that the acidic region may represent an auto-inhibitory sequence analogous to that seen in other kinases [50, 3] . However, crystal structure data indicates that this is not likely to be the case since these regions are separated by more than 30Å [50] . This acidic region has also been identified as the site on CK2 that binds polyamines which are known to stimulate CK2 activity in vitro [63, 64] . Mutation of residues in the acidic loop to remove negative charge raises the basal enzyme activity and abolishes polyamine stimulation [65, 63, 64] . Consequently, although the precise mechanistic details remain to be defined, it does appear that the acidic loop may have a role in CK2 regulation.
3. A Traditional View of CK2
In general, the CK2 tetramer exhibits constitutive activity that can be readily detected in most cell or tissue extracts in the absence of any stimulation. Although the catalytic subunits of CK2 are active in the absence of CK2β, the traditional prevailing view in the field was that the regulatory CK2β subunit plays a key role in CK2 regulation through a variety of distinct actions. In this respect, CK2β was shown to enhance the stability of the catalytic subunits and to modulate their substrate selectivity. For example, CK2β is capable of stimulating CK2 activity towards certain substrates such as topoisomerase II or p53, while inhibiting its activity toward other substrates like calmodulin [66] . As noted previously, CK2β also harbours a binding site for polyamines that stimulate CK2 holoenzyme activity, but not free catalytic subunit activity [67, 64] . Collectively, these observations reinforced the view that CK2β is a central component of tetrameric CK2 complexes that plays a vital role in its regulation.
4. Challenges to the Traditional View of CK2
It is important to emphasize that the traditional view of CK2 as a stable tetrameric enzyme arose primarily through biochemical studies of CK2 and its enzymatic characteristics. The stable nature of CK2 tetramers (dissociation constant Kd = 5.4nM), as well as their spontaneous formation from individual recombinant subunits in vitro supports the traditional view of CK2 [68, 69, 70] . Consequently, it was particularly striking when X-ray crystallography revealed that the CK2α/CK2β interface was relatively small (832Å2) and flexible compared to the average interface size of 1722Å2 for permanent protein subunit interactions [71, 50] . This result clearly raises the spectre that CK2 tetramers are subject to disassembly and re-assembly. As discussed in the following sections, the emergence of new technologies has led to additional perspectives that further challenge the traditional view of CK2 as a stable tetrameric complex.
4.1 Expression and Assembly of CK2
It is clear that CK2 is involved in cell cycle progression and while it is known that all CK2 subunits are expressed throughout the cell cycle, relatively little is known about how either CK2α or CK2β expression is regulated [72] . Characterization of both the CK2α and CK2β genes, as well as adjacent upstream sequences led to identification of a pair of highly conserved Ets1 response elements [73, 74] . While these observations offer an attractive mechanism for the coordinated expression of CK2 subunits destined to form tetrameric complexes, these results are not entirely consistent with earlier studies that had shown that CK2β protein was synthesized in excess of the catalytic subunit in exponentially growing tissue culture cells [75] . In a similar respect, several reports have revealed evidence for the unbalanced expression of the catalytic and regulatory subunits in a variety of different tissues [20, 68, 76] . For example, a systematic analysis in different mouse organs showed that all of the CK2 subunits were expressed at the highest level in brain and testes [76] . However, in comparison to the levels of CK2α, the level of CK2β in testis was significantly higher than in the brain indicating an asymmetric distribution of the different CK2 subunits in these tissues. The existence of free CK2β in both brain and testes was confirmed by immunoprecipitation [76] . The intriguing demonstration that aberrantly high levels of CK2β relative to CK2α have also been observed in tumors, highlights the importance of understanding the dynamic role of CK2β both within the context of the CK2 holoenzyme and as an independent protein [77] .
4.2 Localization of CK2 Subunits
Although initial immunofluorescence localization studies emphasized the predominant nuclear localization of CK2, it is evident that CK2 can be found at other sites within cells. These initial studies also emphasized the extensive degree of overlap that was observed between the independent CK2 subunits; an interpretation consistent with the prevailing view of CK2 as a stable, tetrameric complex [78] . Notably, these authors did observe modest populations of CK2β that did not co-localize with either CK2α or CK2α' [78] . In retrospect, this latter observation could have received more attention for raising the prospect of CK2-independent functions of CK2β.
Subsequent localization studies confirmed the conclusion that the catalytic and regulatory subunits of CK2 are not exclusively co-localized. For example, in a Chironomus tentans epithelial cell line CK2α and CK2β were readily detected in holoenzyme-free pools [79] . While the majority of both subunits were localized to nuclear fractions, a major proportion of the CK2α subunits were tightly bound to other nuclear components whereas CK2β was only loosely associated with other nuclear components [79] . In addition, using CK2 subunit-specific antibodies, immunofluorescence studies demonstrated in mammalian cells that all three subunits of CK2 were localized to the smooth endoplasmic reticulum and the Golgi complex [80] whereas, only the CK2α and CK2α' subunits could be detected in the rough endoplasmic reticulum. By examining the localization of individual CK2 subunits in fixed cells and in extracts, these studies clearly reveal the asymmetric distribution of the individual CK2 subunits.
As noted previously, the three-dimensional structure of tetrameric CK2 that was revealed by X-ray crystallography is not entirely consistent with its existence as stable complexes. However, classical strategies for evaluating the localization of CK2 in cells were limited because they could only offer static views of CK2 in fixed cells. To overcome these limitations, elegant live cell imaging strategies were performed using fluorescent variants of individual CK2 subunits [81, 82] . In addition to confirming the predominantly nuclear and moderately cytoplasmic localization of both of the CK2 subunits, these studies demonstrated that each of the CK2 subunits was independently imported into the nucleus. Further examination of NIH 3T3 cells stably expressing either GFP-α or GFP-β using the FRAP (fluorescence recovery after photobleaching) technique revealed that the CK2 subunits were highly mobile within the nucleus as they relocated to the photobleached area within 10 seconds [81] . The independent movement of the catalytic and regulatory subunits within the nucleus was also observed on a relatively short time-scale [82] . Collectively, these observations suggest that the CK2 tetramer is not a stable complex but more closely resembles a dynamic, transient heterocomplex [50, 82] . Furthermore, the finding that the individual CK2 subunits exhibit distinct patterns of localization provides additional support for the prospect that they perform distinct tetramer-dependent and tetramer-independent functions.
4.3 Genetic Manipulation of CK2
Budding yeast strains harbouring disruptions of either of the two catalytic isoforms of CK2 are viable indicating that any one of the tasks performed by CK2 may be dispensable [6] . However, deletion of both of the catalytic isoforms of CK2 is synthetic lethal indicating that some CK2 activity is required for viability [18, 6, 2] . In contrast to the requirement for at least one catalytic CK2 isoform, budding yeast are viable in the absence of regulatory CK2 subunits, an observation that suggests that the constitutive activity of the catalytic CK2 subunits is sufficient to maintain survival. Disruption of the gene encoding the catalytic isoform CK2α' has also been performed in mice while the gene encoding CK2α has not yet been disrupted. Homozygous CK2α'-/- mice are viable, but the males are infertile due to a defect in spermatogenesis [83] . This result suggests that, with the exception of apparently unique functions in sperm progenitors, CK2α can compensate for the loss of CK2α' in the maintenance of survival.
In contrast to the situation in yeast where disruption of regulatory CK2 subunits does not compromise viability, disruption of CK2β in mice is embryonic lethal [84] . Mouse embryos homozygous for the CK2β null allele (CK2β-/-) showed severe growth retardation and were resorbed around E7.5 [84] . Together with the finding that the development of CK2β-/- blastocysts was impaired in vitro, these results suggest that embryos lacking CK2β cease to develop after implantation. Furthermore, the inability to derive homozygous CK2β knockout embryonic stem cells using classical and conditional knockout strategies strongly suggests that a loss of CK2β leads to immediate cell death [84] . These findings suggest that CK2β is required for the appropriate modulation of CK2 activity since it would appear that the constitutive activity of the catalytic CK2 subunits is not sufficient to maintain viability. In addition, this finding could also reflect requirements for CK2β to perform functions that are independent of CK2.
5. CK2 Independent Functions of CK2β
Although traditionally viewed exclusively as a component of tetrameric CK2 complexes, several independent lines of investigation that were highlighted in the preceding sections demonstrated that CK2β does not exist exclusively within stable CK2 complexes and raises the prospect that CK2β could have CK2-independent functions. In fact, there are a number of additional studies that further reinforce the notion that CK2β does indeed have functions that are independent of its role as the regulatory subunit of CK2. For example, CK2β over-expression studies conducted in the fission yeast S. pombe revealed severe growth defects and a multiseptated phenotype whereas over-expression of CK2α had no effect [85] . Genetic studies in S. cerevisiae have also linked CK2β to checkpoint regulation. In this regard, Toczyski et al (1997) demonstrated that CK2β was required for adaptation, a process where yeast recover from G2 arrest to resume proliferation despite continued DNA damage [86] . The participation of CK2β in DNA damage responses does not appear to be restricted to yeast since human Xeroderma pigmentosum cells gained UV resistance when transfected with CK2β cDNA [87] . Since no defects in CK2 activity are apparent in xeroderma pigmentosum cells, it was hypothesized that the UV resistance conferred by over-expression of CK2β arose through a CK2-independent function [87] . Although these observations are intriguing they only begin to characterize the many potential independent functions of CK2β.
6. Interaction Partners of CK2β
In many cases, detailed examination of the interaction partners of a protein can lead to a better understanding of not only the functional processes in which it is involved but also the regulatory mechanisms controlling its expression, function and degradation. Accordingly, it is anticipated that considerable insight into the regulation and functions of CK2β, including those functions that may be independent of CK2, will be illuminated by a thorough investigation of its interaction partners. In this respect, over the last decade a plethora of CK2β-specific interaction partners have been identified through studies that have been performed both in vitro and in vivo [88, 89, 90, 91, 92, 93, 94, 95, 96] . While many of these interactions have not been further investigated, some of these proteins have undergone more extensive validation allowing for their classification as either CK2 dependent or CK2 independent binding partners of CK2β. A summary of interaction partners from mammalian cells (or Xenopus in the case of c-Mos) that can be distinguished on the basis of whether they interact with CK2-independent forms of CK2β or CK2-associated CK2β is presented in Table 1.
Although beyond the scope of the present article, it is also noteworthy that a considerable amount of new information has begun to emerge from systematic studies performed on a genome-wide scale in model organisms such as yeast [97, 98] . Information from these studies is continually being compiled in databases such as BIND or PreBIND allowing for in silico determination of protein-protein interaction networks [99, 100] . Indeed, such databases identify numerous interaction partners of the CK2 subunits both within the context of the CK2 holoenzyme and as independent proteins. For example, two independent studies, one employing a tandom-affinity purification (TAP)/mass spectrometry approach and the other using an immunoprecipitation/mass spectrometry approach, showed that individual CK2 subunits were differentially localized to a number of multi-protein complexes [97, 98] . Each of these studies demonstrate incomplete overlap between the catalytic and regulatory subunits of CK2 as all four CK2 subunits (yeast contain 2 forms of both catalytic and regulatory subunit) were detected in some multi-protein complexes while other complexes only contained one, two or three of the CK2 subunits [97, 98] . Although these studies are limited to the yeast proteome and await independent confirmation, the emergence of CK2 subunit specific interactions in a model system such as yeast will undoubtedly be instructive of the situation in mammalian systems. Furthermore, it is evident that while several CK2β specific interaction partners offering many new insights into its CK2-independent functions have already been characterized in mammalian cells, it can be anticipated that our current understanding remains far from complete.
CK2β specific interacting proteins
| Interaction partner | Function | Detected | References |
| CK2 dependent binding partners of CK2β | |||
| p90Rsk | S/T protein kinase | in vitro | 92, 93 |
| PKCζ | S/T protein kinase, mediate NF-κB activation | in vitro | 88, 93 |
| Topoisomerase II | DNA remodelling, essential during mitosis and meiosis | in vitro | 92, 101, 103 |
| p53 | Tumor supressor gene product | in vitro/in vivo | 92, 102, 115 |
| p27KIP1 | CDK inhibitor, cell cycle progression | in vitro | 107 |
| p21WAF1/CIP1 | CDK inhibitor, cell cycle progression | in vitro/in vivo | 92, 105 |
| Cdc25B | Phophatase, CDK activator, cell cycle progression | in vitro/in vivo | 106 |
| CD5 | Cell surface receptor, thymocytes, T-cells, some B1a B-cells | in vitro | 92 |
| FGF-2 | Fibroblast growth factor 2, cell proliferation | in vitro | 92, 108 |
| Nopp 140 | Nucleolar protein, potential chaperone for nucleolar transport | in vitro | 92, 104 |
| L5 | Ribosomal protein | in vitro/in vivo | 92, 93 |
| L41 | Ribosomal protein | in vitro | 92 |
| HHV-6 IE2 | Human herpesvirus 6 immediate-early 2 protein, gene promoter transactivator | in vitro/in vivo | 96 |
| CK2 independent binding partners of CK2β | |||
| c-Mos | S/T protein kinase, MAPK activation, cell cycle progression | in vitro/in vivo | 89, 92, 95 |
| Chk1 | S/T protein kinase, regulator of DNA damage induced G2 arrest, checkpoint control | in vitro/in vivo | 94 |
| A-Raf | S/T protein kinase, mitogenic signalling, cell proliferation | in vitro/in vivo | 90, 92, 93 |
6.1 CK2 Dependent Binding Partners of CK2β
As outlined in table 1, a number of CK2-dependent binding partners of CK2β, that is, proteins that bind to tetrameric CK2 through binding sites on CK2β, have been identified. Although detailed examination of each of these interactions is beyond the scope of this article, consideration of a few of the binding partners provides insights into the general roles of CK2β in regulating the cellular functions of CK2. From this perspective, one of the main functions of CK2β in the context of CK2 appears to be that of a substrate docking protein. Topoisomerase II and p53 are likely bona fide physiological CK2 substrates that are dependent on interactions with CK2β in order to be phosphorylated by CK2 [101, 102] . In each case, a specific N-terminal region of CK2β (residues 51-110 for topoisomerase II and residues 72-149 for p53) was identified as the binding site [102, 103] . A number of other CK2-dependent CK2β binding partners including Cdc25B, p27KIP1, Nopp140 and p21WAF1/CIP1 have also been shown to bind to the extreme N-terminal region of CK2β [104, 105, 106, 107] . As more potential CK2 substrates are characterized, it will be interesting to note whether groups of structurally similar proteins that bind to the same N-terminal binding site emerge.
A second function of CK2β in the context of CK2 appears to involve transmission of regulatory signals provided by other proteins in a manner that could be analogous to that seen with polyamines. FGF-2 exemplifies this as binding of FGF-2 to CK2β stimulates CK2 activity toward nucleolin in vitro [108] . In this situation, the existence of numerous potential binding sites in the N-terminus of CK2β may represent a mechanism where binding of a regulatory protein to a given site will elicit a particular effect on CK2 activity. Collectively, these studies highlight complementary mechanisms by which CK2β modulates the ability of CK2 to phosphorylate specific cellular targets.
6.2 CK2 Independent Binding Partners of CK2β
In addition to the CK2-dependent binding partners of CK2β, a number of proteins that bind to CK2β in the absence of catalytic CK2 subunits have been identified. Intriguingly, these proteins include the A-Raf, c-Mos and Chk1 serine/threonine protein kinases [89, 90, 94] . As noted in Table 1, CK2β interacts with two other serine/threonine protein kinases, namely p90Rsk and PKCζ [88, 91, 93] . Since these latter two protein kinases, represent proteins that interact with CK2β within tetrameric CK2 complexes, they will not be further discussed. Instead, the remaining discussion will focus on the interaction between CK2β and those serine/threonine protein kinases that interact with it in the absence of catalytic CK2 subunits.
6.2.1 A-Raf
A-Raf along with B-Raf and c-Raf-1 comprise a family of three cytoplasmic serine/threonine protein kinases that contain three conserved regions, CR1 (Ras-binding and cysteine rich domains), CR2 (serine/threonine-rich domain) and CR3 (catalytic domain) [109] . Raf kinases are essential in MAPK (mitogen activated protein kinase) pathways as they relay the signal from the receptor/Ras complex to the cytosolic kinase cascade by phosphorylating and activating the appropriate MAPK kinase (MKK or MEK) [109] . CK2β was identified as an interaction partner of A-Raf in two independent yeast two hybrid screens [110, 90] . Subsequent examination of this interaction in vitro revealed the specificity of this association since CK2β was not able to interact with either B-Raf or c-Raf-1. The CK2β binding site on A-Raf was initially mapped to residues 255-569, which includes sequences that can be aligned with the CK2β-interaction region of CK2α (Figure 2). A potential regulatory role of CK2β was suggested by co-expression of CK2β and A-Raf in insect cells which resulted in a 10-fold enhancement of A-Raf kinase activity toward MEK [90] . Although the possibility of complex formation between CK2α, β and A-Raf was not excluded in this study it was shown that CK2α was not able to induce a similar activation. In fact inclusion of CK2α in the assay abolished the activation observed in the presence of CK2β, suggesting that CK2α was competing with A-Raf for binding to CK2β. As discussed below, the CK2β binding site on A-Raf is similar to the CK2β binding sites of other serine/threonine protein kinases (Figure 2) [50] .
6.2.2 c-Mos
Upon stimulation with progesterone, Xenopus oocytes undergo meiotic maturation, a process which requires the activity of c-Mos, a germ cell-specific serine/threonine protein kinase [89] . c-Mos initiates signalling via the MAPK pathway as it also phosphorylates MAPK kinase (MKK or MEK) which in turn phosphorylates and activates MAPK. Interaction between CK2β and c-Mos was originally identified using a yeast two hybrid screen of a Xenopus oocyte cDNA library [89] . Interactions between CK2β and c-Mos were subsequently confirmed in several other systems including in vitro binding assays, co-transfection studies in human 293T cells and in Xenopus oocytes overexpressing CK2β. Using a series of deletion mutants, the c-Mos binding domain was mapped to the C-terminal 55 amino acids of CK2β, a region which also mediates interaction with CK2α (or α').
In stark contrast with its effect on A-Raf, CK2β inhibited the activity of v-Mos, the constituitively active counterpart of c-Mos, by 40% in vitro [89, 90] . CK2β also interfered with the Mos-mediated MAPK activation and progesterone-induced oocyte maturation while a mutant of CK2β lacking its C-terminal 55 amino acids and therefore not able to bind c-Mos lacked these inhibitory abilities. Direct in vivo evidence further indicated that CK2β inhibits the ability of c-Mos to induce mitotic arrest in rapidly dividing embryonic cells. Based on their studies, Chen and colleagues proposed that CK2 binds and inhibits c-Mos during its initial synthesis [89] . Inhibition is alleviated when c-Mos overcomes a certain threshold that is set by the amount of free CK2β available [89] . Free c-Mos is then able to activate the MAPK pathway. This mechanism is consistent with the observation that CK2β is synthesized in excess of the CK2 catalytic subunits and that newly synthesized CK2β is slowly incorporated into the holoenzyme [75] . This model is also supported by recent studies that identified the CK2β binding region in the N-terminus of c-Mos, between amino acids 52 and 115 [95] . As is the case with A-Raf, this region closely resembles the CK2β binding region of catalytic CK2 subunits (Figure 2).
Sequence alignment of serine/threonine protein kinases containing a putative CK2β binding domain: Comparison of the CK2β binding region of the CK2 catalytic subunits (α and α') with the putative CK2β binding regions identified in other serine/threonine protein kinases, namely Chk1 (aa1-87), c-Mos (aa52-115) and A-Raf (aa323-373). Basic residues – black, acidic residues – blue, small or hydrophobic residues – red, remaining residues - green. Invariant residues are highlighted in yellow while other highly conserved residues are marked by an asterisk (*).

6.2.3 Chk1
Studies conducted in S. pombe led to the identification of Chk1 as a protein serine/threonine kinase that is essential for DNA damage-induced G2 arrest [111] . In mammalian systems, Chk1 is not only required for activation of the G2 checkpoint but is also required for the viability of embryonic stem cells [112, 113] . Recently, Guerra and colleagues (2003) employed co-expression and immunoprecipitation studies to demonstrate interactions between Chk1 and CK2β in mammalian cells [94] . Furthermore, complexes between Chk1 and CK2β excluded the CK2α subunit while complexes between the CK2α and CK2β excluded Chk1 [94] . As was the case for A-Raf and c-Mos as well as CK2α and CK2α', the C-terminal region of CK2β was required to bind to Chk1. Like A-Raf, Chk1 activity is enhanced upon interaction with CK2β [90, 94] . Although the mechanistic basis for this activation remains to be determined, it may relate to the observation that the activity of the kinase domain of Chk1 (1-265aa) is 20-fold more active than full length Chk1 [94] . In this respect, the C-terminal region of Chk1 may exert an autoinhibitory effect on kinase activity that is relieved upon CK2β binding. As yet, the CK2β-binding region of Chk1 has not been experimentally determined. However, as noted by Guerra and colleagues, comparison of the high-resolution three-dimensional structures of CK2α and Chk1 suggests that the highly conserved N-terminal lobe of these proteins may contain a CK2β binding domain [114, 50, 94] . Support for this notion comes from the alignment of the amino acid sequences of the CK2β binding regions of CK2α, A-Raf and c-Mos with Chk1 which reveals extensive similarity with the sequence corresponding to the N-terminal lobe of Chk1 (Figure 2).
Although speculative at the present time, one intriguing connection between CK2β and Chk1 is that both of these proteins have been linked to adaptation checkpoints in yeast. Of particular significance is the observation that the effects of CK2β that are required for this adaptation are independent of the catalytic CK2 subunits [86] . Since this result suggests that the effects of CK2β are mediated through interactions with other cellular factors, it is tantalizing to speculate that these factors may include other proteins involved in adaptation, possibly even Chk1.
6.3 A Common CK2β Binding Domain in CK2β-Interacting Protein Kinases?
It is particularly striking that three of the known CK2-independent CK2β interaction partners identified to date are serine/threonine protein kinases containing sequences reminiscent of the CK2β binding region present in the CK2 catalytic subunits (Figure 2) [89, 90, 94, 95] . There are a number of residues within this region, originally noted by Niefind and colleagues including L41, G46, G48, V53, V65 and K68 (numbering based on CK2α sequence) that are absolutely conserved within each of the kinases [50] .
While the conservation of some of these residues, notably G46, G48, V53 and K68 is not entirely surprising because they are highly conserved throughout the serine/threonine protein kinase family, the invariance of the other residues is intriguing. In addition to the invariant residues, a significant proportion of the other residues in this region are also very highly conserved (Figure 2). Examination of the crystal structure of CK2 (Figure 3) indicates that several residues within this region may be important in maintenance of the secondary and tertiary structure of the enzyme. However, residues such as L41 together with an adjacent hydrophobic residue (V42 in CK2α) are located at the CK2α-CK2β interface. While rigorous testing is still required, the location of these residues suggests that they may be important in mediating the interaction (Figure 3C).
Overall, the alignment shown in Figure 2 agrees with the suggestion that CK2β is able to bind a portion of the N-terminal lobe of a set of protein kinases to act as either a positive or negative regulator of activity [94, 95] . Evidence supporting this suggestion is derived from an investigation where residues 52-115 of c-Mos, representing its CK2β binding domain, were fused to Xenopus cyclin A2 and tested for ability to act as an independent CK2β binding domain. Indeed, this domain was able to confer specific CK2β binding to the cyclin A2 protein suggesting that this region does represent a portable CK2β binding domain [95] . Therefore, although the precise determinants responsible for specific interactions with CK2β remain to be delineated, it is evident that the CK2β binding does not exclusively reside within the catalytic subunits of CK2. Accordingly, it can be envisaged that other protein kinases, and possibly other proteins, harbouring the appropriate structural elements remain to be identified as CK2β-interacting proteins. In this respect, it is intriguing that recent genetic screens performed on a global scale in yeast revealed synthetic lethality between Cdc7, a protein kinase involved in regulation of DNA replication [99] . While direct interactions with CK2β have not yet been examined, this observation is consistent with the prediction that other CK2β-regulated protein kinases remain to be identified.
6.4 Implications: CK2β as a Promiscuous Kinase Regulatory Subunit
As noted earlier, a major role for CK2β within CK2 complexes appears to be substrate docking or recruitment where it brings the substrate protein and the catalytic subunit into close enough proximity to facilitate the phosphorylation reaction [115, 101, 102, 108, 104, 91, 103, 105, 106, 107] . Based on these examples, a similar role for CK2β in the context of A-Raf, c-Mos or Chk1 can be envisaged. In this respect, it will be interesting to determine whether CK2β will have the capacity to modulate the phosphorylation of specific proteins by a number of distinct protein kinases. A second function of CK2β within CK2 complexes appears to be its ability to translate interactions with regulatory factors such as FGF-2 and polyamines into changes in CK2 activity. Accordingly, it is tempting to speculate that CK2β may be able to transmit regulatory signals to A-Raf, c-Mos or Chk1 in an analogous manner. In addition to these possibilities, it remains to be seen whether two distinct kinases can be simultaneously bound to the CK2β dimer, allowing CK2β to act as a scaffold to co-ordinate two distinct signal transduction pathways. Although these suggestions remain speculative, it is evident that CK2β offers intriguing new prospects for understanding the regulation and functions of protein kinases that are distinct from CK2.
Structural comparison of the CK2β binding regions of CK2α and Chk1. A. Crystal structure depicting the interaction between CK2α (grey and green) and CK2β (blue). The CK2β binding region of CK2α is represented by the green ribbon. CK2β is shown for context. (1JWH [50] ) B. Crystal structure of Chk1 (1IA8 [114] ). The putative CK2β binding region of Chk1 is represented as a red ribbon. Residues that are absolutely conserved between the two CK2β binding regions are shown in black. C. Alignment of the high resolution crystal structures of the CK2β binding regions of tetrameric CK2α (green), CK2α alone (purple; 1PJK [118] ) and Chk1 (red). A backbone representation of the CK2β binding domain of each protein is shown. The amino acid side chains of invariant residues are shown in black. In particular, L11 in Chk1 is analogous to L41 in CK2α and may be important in mediating binding to CK2β.
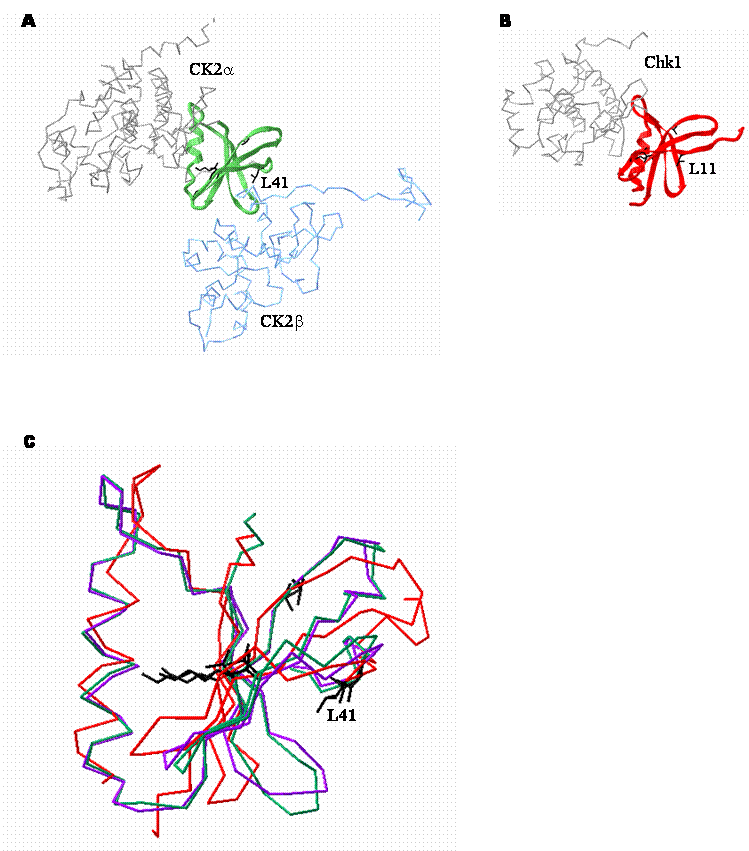
7. Perspectives and Other Considerations
Although traditionally considered to be a stable tetrameric enzyme, there is now an abundance of evidence that challenges this static view of CK2. In this respect, information derived from diverse experimental strategies ranging from X-ray crystallography, to gene knockout studies and live cell imaging experiments suggest that the individual subunits of CK2 do indeed exist outside stable tetrameric complexes. In fact, it now appears certain that CK2β, a remarkably conserved protein that has classically been known only for its role as the regulatory subunit of CK2, has cellular functions that are independent of CK2. These CK2-independent functions of CK2β reinforce the importance of delineating its regulation, both within and external to, CK2 complexes. In this respect, it will be instructive to determine the precise mechanisms that govern cellular levels of CK2β and to determine how its interactions with different partners are regulated.
It is striking that, in addition to the catalytic subunits of CK2, three different protein kinases, namely A-Raf, c-Mos and Chk1 have been identified as proteins that interact with CK2β. While the effects of CK2β on each of these protein kinases appear to differ, it is apparent that CK2β has the potential to modulate each of their kinase activities in some manner. Since many details regarding the physiological significance of these findings remain to be elucidated, it is premature to consider the formal establishment of a new classification of CK2β-regulated kinases. Nevertheless, it is intriguing that the ability of CK2β to regulate different protein kinases is analogous to that observed with cyclins, a noteworthy observation given the presence of sequences within CK2β that resemble two distinct types of destruction signals that are present in members of the cyclin family.
Overall, while many questions remain, the identification of CK2-independent partners for CK2β has dramatically affected our view of how this remarkably conserved protein is involved in cellular regulation. Furthermore, we can certainly anticipate that the identification of new partners for CK2β will accompany the emergence of new systematic strategies for the evaluation of protein-protein interactions and determination of protein complex composition. Based on past performance, we can thus expect more surprises regarding the biological functions of CK2β, both within, and external to, tetrameric CK2 complexes.
Acknowledgements
We wish to thank past and present members of the Litchfield laboratory for contributions to our work. Research on CK2 has been supported by grants from the National Cancer Institute of Canada with funds from the Canadian Cancer Society and the Terry Fox Foundation raised through the Terry Fox Run, the Canadian Institutes of Health Research and the Premier's Research Excellence Awards Program of the Province of Ontario. Ashley Bibby was supported by a Postgraduate Scholarship from the Natural Sciences and Engineering Research Council of Canada. We apologize for any lack of citations of important advances or publications because of space limitations.
Conflict of Interest
The authors have declared that no conflict of interest exists.
References
1. Burnett G, Kennedy E.P. The enzymatic phosphorylation of proteins. J Biol Chem. 1954 ;211(2):969-980
2. Pinna L.A. Protein kinase CK2: a challenge to canons. J Cell Sci. 2002 ;115(20):3873-3878
3. Litchfield DW. Protein kinase CK2: structure, regulation and role in cellular decisions of life and death. Biochem J. 2003 ;369(1):1-15
4. Filhol O. et al. Protein kinase CK2: a new view of an old molecular complex. EMBO Rep. 2004 ;5(4):351-355
5. Kikkawa U. et al. Molecular cloning of casein kinase II alpha subunit from Dictyostelium discoideum and its expression in the life cycle. Mol Cell Biol. 1992 ;12(12):5711-5723
6. Glover CV. On the physiological role of casein kinase II in Saccharomyces cerevisiae. Prog Nucleic Acid Res Mol Biol. 1998 ;59:95-133
7. Lin J.M. et al. A role for casein kinase 2 alpha in the Drosophila circadian clock. Nature. 2002 ;420(6917):816-820
8. Akten B. et al. A role for CK2 in the Drosophila circadian oscillator. Nat Neurosci. 2003 ;6(3):251-257
9. Blau J. A new role for an old kinase: CK2 and the circadian clock. Nat Neurosci. 2003 ;6(3):208-210
10. Daniel X. et al. CK2 phosphorylation of CCA1 is necessary for its circadian oscillator function in Arabidopsis. Proc Natl Acad Sci U S A. 2004 ;101(9):3292-3297
11. Gerber D.A. et al. Heat-induced relocalization of protein kinase CK2. Implication of CK2 in the context of cellular stress. J Biol Chem. 2000 ;275(31):23919-23926
12. Sayed M. et al. Stress-induced activation of protein kinase CK2 by direct interaction with p38 mitogen-activated protein kinase. J Biol Chem. 2000 ;275(22):16569-16573
13. Ghavidel A, Schultz MC. TATA binding protein-associated CK2 transduces DNA damage signals to the RNA polymerase III transcriptional machinery. Cell. 2001 ;106(5):575-584
14. Keller D.M. et al. A DNA damage-induced p53 serine 392 kinase complex contains CK2, hSpt16, and SSRP1. Mol Cell. 2001 ;7(2):283-292
15. Pepperkok R. et al. Cell growth stimulation by EGF: inhibition through antisense-oligodeoxynucleotides demonstrates important role of casein kinase II. Exp Cell Res. 1991 ;197(2):245-253
16. Lorenz P. et al. Cell biological studies with monoclonal and polyclonal antibodies against human casein kinase II subunit beta demonstrate participation of the kinase in mitogenic signaling. J Biol Chem. 1993 ;268(4):2733-2739
17. Pepperkok R. et al. Casein kinase II is required for transition of G0/G1, early G1, and G1/S phases of the cell cycle. J Biol Chem. 1994 ;269(9):6986-6991
18. Hanna D.E. et al. Casein kinase II is required for cell cycle progression during G1 and G2/M in Saccharomyces cerevisiae. J Biol Chem. 1995 ;270(43):25905-25914
19. Daya-Makin M. et al. Activation of a tumour-associated protein kinase (p40TAK) and casein kinase II in human squamous cell carcinomas and adencarcinomas of the lung. Cancer Research. 1994 ;54:2262-2268
20. Stalter G. et al. Asymmetric expression of protein kinase CK2 subunits in human kidney tumours. Biochem Biophys Res Commun. 1994 ;202:141-147
21. Yenice S. et al. Nuclear casein kinase 2 (CK-2) activity in human normal, benign hyperplastic and cancerous prostate. Prostate. 1994 ;24:11-16
22. Faust R.A. et al. Elevated protein kinase CK2 activity in chromatin of head and neck tumours: association with malignant transformation. Cancer Letters. 1996 ;101:31-35
23. Landesman-Bollag E. et al. Protein kinase CK2 in mammary gland tumorigenesis. Oncogene. 2001 ;20(25):3247-3257
24. Seldin DC, Leder P. Casein kinase II alpha transgene-induced murine lymphoma: relation to theileriosis in cattle. Science. 1995 ;267(5199):894-897
25. Kelliher M.A. et al. Tal-1 induces T cell acute lymphoblastic leukemia accelerated by casein kinase II alpha. Embo J. 1996 ;15(19):5160-5166
26. Landesman-Bollag E. et al. p53 deficiency and misexpression of protein kinase CK2alpha collaborate in the development of thymic lymphomas in mice. Oncogene. 1998 ;16(23):2965-2974
27. Orlandini M. et al. Protein kinase CK2alpha' is induced by serum as a delayed early gene and cooperates with Ha-ras in fibroblast transformation. J Biol Chem. 1998 ;273(33):21291-21297
28. Channavajhala P, Seldin DC. Functional interaction of protein kinase CK2 and c-Myc in lymphomagenesis. Oncogene. 2002 ;21(34):5280-5288
29. Ravi R, Bedi A. Sensitization of tumor cells to Apo2 ligand/TRAIL-induced apoptosis by inhibition of casein kinase II. Cancer Res. 2002 ;62(15):4180-4185
30. Romieu-Mourez R. et al. Protein kinase CK2 promotes aberrant activation of nuclear factor-kappaB, transformed phenotype, and survival of breast cancer cells. Cancer Res. 2002 ;62(22):6770-6778
31. Hu E, Rubin CS. Expression of wild-type and mutated forms of the catalytic (alpha) subunit of Caenorhabditis elegans casein kinase II in Escherichia coli. J Biol Chem. 1990 ;265(33):20609-20615
32. Grankowski N. et al. Isolation and characterization of recombinant human casein kinase II subunits alpha and beta from bacteria. Eur J Biochem. 1991 ;198(1):25-30
33. Heller-Harrison R.A. et al. Cloning and characterization of a cDNA encoding the beta subunit of human casein kinase II. Biochemistry. 1989 ;28(23):9053-9058
34. Lozeman F.J. et al. Isolation and characterization of human cDNA clones encoding the alpha and the alpha' subunits of casein kinase II. Biochemistry. 1990 ;29(36):8436-8447
35. Jauch E. et al. In vivo functional analysis of Drosophila protein kinase casein kinase 2 (CK2) beta-subunit. Gene. 2002 ;298(1):29-39
36. Zelada A. et al. cDNA cloning, biochemical and phylogenetic characterization of beta- and beta'-subunits of Candida albicans protein kinase CK2. Yeast. 2003 ;20(6):471-478
37. Shi X. et al. A novel casein kinase 2 alpha-subunit regulates membrane protein traffic in the human hepatoma cell line HuH-7. J Biol Chem. 2001 ;276(3):2075-2082
38. Allende JE, Allende C.C. Protein kinases: Protein kinase CK2: an enzyme with multiple substrates and a puzzling regulation. Faseb J. 1995 ;9(5):313-323
39. Litchfield DW, Luscher B. Casein kinase II in signal transduction and cell cycle regulation. Mol Cell Biochem. 1993 ;127:187-200
40. Maridor G. et al. Casein kinase II. cDNA sequences, developmental expression, and tissue distribution of mRNAs for alpha, alpha', and beta subunits of the chicken enzyme. J Biol Chem. 1991 ;266(4):2362-2368
41. Gietz R.D. et al. Interactions between the subunits of casein kinase II. J Biol Chem. 1995 ;270(22):13017-13021
42. Boldyreff B. et al. Structure of protein kinase CK2: dimerization of the human beta-subunit. FEBS Lett. 1996 ;379(2):153-156
43. Marin O. et al. Physical dissection of the structural elements responsible for regulatory properties and intersubunit interactions of protein kinase CK2 beta-subunit. Biochemistry. 1997 ;36(23):7192-7198
44. Chantalat L. et al. Crystal structure of the human protein kinase CK2 regulatory subunit reveals its zinc finger-mediated dimerization. Embo J. 1999 ;18(11):2930-2940
45. Graham KC, Litchfield DW. The regulatory beta subunit of protein kinase CK2 mediates formation of tetrameric CK2 complexes. J Biol Chem. 2000 ;275(7):5003-5010
46. Canton D.A. et al. Assembly of protein kinase CK2: investigation of complex formation between catalytic and regulatory subunits using a zinc-finger-deficient mutant of CK2beta. Biochem J. 2001 ;358(Pt 1):87-94
47. Litchfield DW. et al. Phosphorylation of the beta subunit of casein kinase II in human A431 cells. Identification of the autophosphorylation site and a site phosphorylated by p34cdc2. J Biol Chem. 1991 ;266(30):20380-20389
48. Boldyreff B. et al. Ser2 is the autophosphorylation site in the beta subunit from bicistronically expressed human casein kinase-2 and from native rat liver casein kinase-2 beta. Eur J Biochem. 1993 ;218(2):515-521
49. Sarno S. et al. Cooperative modulation of protein kinase CK2 by separate domains of its regulatory beta-subunit. Biochemistry. 1999 ;39:2324-2329
50. Niefind K. et al. Crystal structure of human protein kinase CK2: insights into basic properties of the CK2 holoenzyme. Embo J. 2001 ;20(19):5320-5331
51. Zhang C. et al. Phosphorylation regulates the stability of the regulatory CK2beta subunit. Oncogene. 2002 ;21(23):3754-3764
52. Litchfield D.W. et al. The protein kinase from mitotic human cells that phosphorylates Ser-209 on the casein kinase II beta-subunit is p34cdc2. Biochim Biophys Acta. 1995 ;1269(1):69-78
53. Meggio F. et al. Phosphorylation and activation of protein kinase CK2 by p34cdc2 are independent events. Eur J Biochem. 1995 ;230(3):1025-1031
54. Glotzer M. et al. Cyclin is degraded by the ubiquitin pathway. Nature. 1991 ;349(6305):132-138
55. King RW. et al. Mutagenic analysis of the destruction signal of mitotic cyclins and structural characterization of ubiquitinated intermediates. Mol Biol Cell. 1996 ;7(9):1343-1357
56. Bastians H. et al. Cell cycle-regulated proteolysis of mitotic target proteins. Mol Biol Cell. 1999 ;10(11):3927-3941
57. Hames R.S. et al. APC/C-mediated destruction of the centrosomal kinase Nek2A occurs in early mitosis and depends upon a cyclin A-type D-box. Embo J. 2001 ;20(24):7117-7127
58. Littlepage L.E. and Ruderman, J.V. Identification of a new APC/C recognition domain, the A box, which is required for the Cdh1-dependent destruction of the kinase Aurora-A during mitotic exit. Genes Dev. 2002 ;16(17):2274-2285
59. Littlepage L.E. et al. Identification of phosphorylated residues that affect the activity of the mitotic kinase Aurora-A. Proc Natl Acad Sci U S A. 2002 ;99(24):15440-15445
60. Sarno S. et al. A multifunctional network of basic residues confers unique properties to protein kinase CK2. Mol Cell Biochem. 1999 ;191(1-2):13-19
61. Li D. et al. The physical association of casein kinase 2 with nucleolin. J Biol Chem. 1996 ;271(26):15662-15668
62. Litchfield D.W. et al. Analysis of interactions between the subunits of protein kinase CK2. Biochem Cell Biol. 1996 ;74(4):541-547
63. Leroy D. et al. Direct identification of a polyamine binding domain on the regulatory subunit of the protein kinase casein kinase 2 by photoaffinity labeling. J Biol Chem. 1995 ;270(29):17400-17406
64. Leroy D. et al. Binding of polyamines to an autonomous domain of the regulatory subunit of protein kinase CK2 induces a conformational change in the holoenzyme. A proposed role for the kinase stimulation. J Biol Chem. 1997 ;272(33):20820-20827
65. Meggio F. et al. Casein kinase 2 down-regulation and activation by polybasic peptides are mediated by acidic residues in the 55-64 region of the beta-subunit. A study with calmodulin as phosphorylatable substrate. Biochemistry. 1994 ;33(14):4336-4342
66. Marin O. et al. Structural features underlying the unusual mode of calmodulin phosphorylation by protein kinase CK2: A study with synthetic calmodulin fragments. Biochem Biophys Res Commun. 1999 ;256(2):442-446
67. Hathaway GM, Traugh JA. Kinetics of activation of casein kinase II by polyamines and reversal of 2,3-bisphosphoglycerate inhibition. J Biol Chem. 1984 ;259(11):7011-7015
68. Pinna LA, Meggio F. Protein kinase CK2 ("casein kinase-2") and its implication in cell division and proliferation. Prog Cell Cycle Res. 1997 ;3:77-97
69. Battistutta R. et al. The crystal structure of the complex of Zea mays alpha subunit with a fragment of human beta subunit provides the clue to the architecture of protein kinase CK2 holoenzyme. Eur J Biochem. 2000 ;267(16):5184-5190
70. Martel V. et al. Dynamic localization/association of protein kinase CK2 subunits in living cells: a role in its cellular regulation? Ann N Y Acad Sci. 2002 ;973:272-277
71. Jones S, Thornton J.M. Principles of protein-protein interactions. Proc Natl Acad Sci U S A. 1996 ;93(1):13-20
72. Bosc DG. et al. Expression and regulation of protein kinase CK2 during the cell cycle. Mol Cell Biochem. 1999 ;191(1-2):213-222
73. Krehan A. et al. Ets1 is a common element in directing transcription of the alpha and beta genes of human protein kinase CK2. Eur J Biochem. 2001 ;268(11):3243-3252
74. Pyerin W, Ackermann K. Transcriptional coordination of the genes encoding catalytic (CK2alpha) and regulatory (CK2beta) subunits of human protein kinase CK2. Mol Cell Biochem. 2001 ;227(1-2):45-57
75. Luscher B, Litchfield DW. Biosynthesis of casein kinase II in lymphoid cell lines. Eur J Biochem. 1994 ;220(2):521-526
76. Guerra B. et al. Protein kinase CK2: evidence for a protein kinase CK2beta subunit fraction, devoid of the catalytic CK2alpha subunit, in mouse brain and testicles. FEBS Lett. 1999 ;462(3):353-357
77. Vilk G. et al. Inducible expression of the regulatory protein kinase CK2beta subunit: incorporation into complexes with catalytic CK2 subunits and re-examination of the effects of CK2beta on cell proliferation. J Cell Biochem. 2001 ;84(1):84-99
78. Krek W. et al. Casein kinase II is a predominantly nuclear enzyme. J Cell Biol. 1992 ;116(1):43-55
79. Stigare J. et al. A majority of casein kinase II alpha subunit is tightly bound to intranuclear components but not to the beta subunit. Mol Cell Biochem. 1993 ;129(1):77-85
80. Faust M. et al. Localization of individual subunits of protein kinase CK2 to the endoplasmic reticulum and to the Golgi apparatus. Mol Cell Biochem. 2001 ;227(1-2):73-80
81. Martel V. et al. Visualization and molecular analysis of nuclear import of protein kinase CK2 subunits in living cells. Mol Cell Biochem. 2001 ;227(1-2):81-90
82. Filhol O. et al. Live-cell fluorescence imaging reveals the dynamics of protein kinase CK2 individual subunits. Mol Cell Biol. 2003 ;23(3):975-987
83. Xu X. et al. Globozoospermia in mice lacking the casein kinase II alpha' catalytic subunit. Nat Genet. 1999 ;23(1):118-121
84. Buchou T. et al. Disruption of the regulatory beta subunit of protein kinase CK2 in mice leads to a cell-autonomous defect and early embryonic lethality. Mol Cell Biol. 2003 ;23(3):908-915
85. Roussou I, Draetta G. The Schizosaccharomyces pombe casein kinase II alpha and beta subunits: evolutionary conservation and positive role of the beta subunit. Mol Cell Biol. 1994 ;14(1):576-586
86. Toczyski D.P. et al. CDC5 and CKII control adaptation to the yeast DNA damage checkpoint. Cell. 1997 ;90(6):1097-1106
87. Teitz T. et al. Expression of the cDNA for the beta subunit of human casein kinase II confers partial UV resistance on xeroderma pigmentosum cells. Mutat Res. 1990 ;236(1):85-97
88. Diaz-Meco M.T. et al. zeta PKC induces phosphorylation and inactivation of I kappa B-alpha in vitro. Embo J. 1994 ;13(12):2842-2848
89. Chen M. et al. The casein kinase II beta subunit binds to Mos and inhibits Mos activity. Mol Cell Biol. 1997 ;17(4):1904-1912
90. Hagemann C. et al. The regulatory subunit of protein kinase CK2 is a specific A-Raf activator. FEBS Lett. 1997 ;403(2):200-202
91. Grein S. et al. Searching interaction partners of protein kinase CK2beta subunit by two-hybrid screening. Mol Cell Biochem. 1999 ;191(1-2):105-109
92. Guerra B, Issinger OG. Protein kinase CK2 and its role in cellular proliferation, development and pathology. Electrophoresis. 1999 ;20(2):391-408
93. Kusk M. et al. Interactions of protein kinase CK2beta subunit within the holoenzyme and with other proteins. Mol Cell Biochem. 1999 ;191(1-2):51-58
94. Guerra B. et al. Modulation of human checkpoint kinase Chk1 by the regulatory beta-subunit of protein kinase CK2. Oncogene. 2003 ;22(32):4933-4942
95. Lieberman SL, Ruderman JV. CK2 beta, which inhibits Mos function, binds to a discrete domain in the N-terminus of Mos. Dev Biol. 2004 ;268(2):271-279
96. Shimada K. et al. Human herpesvirus 6 immediate-early 2 protein interacts with heterogeneous ribonucleoprotein K and casein kinase 2. Microbiol Immunol. 2004 ;48(3):205-210
97. Gavin A.C. et al. Functional organization of the yeast proteome by systematic analysis of protein complexes. Nature. 2002 ;415(6868):141-147
98. Ho Y. et al. Systematic identification of protein complexes in Saccharomyces cerevisiae by mass spectrometry. Nature. 2002 ;415(6868):180-183
99. Bader G.D. et al. BIND: the Biomolecular Interaction Network Database. Nucleic Acids Res. 2003 ;31(1):248-250
100. Donaldson I. et al. PreBIND and Textomy--mining the biomedical literature for protein-protein interactions using a support vector machine. BMC Bioinformatics. 2003 ;4(1):11-
101. Bojanowski K. et al. DNA topoisomerase II and casein kinase II associate in a molecular complex that is catalytically active. J Biol Chem. 1993 ;268(30):22920-22926
102. Appel K. et al. Mapping of the interaction sites of the growth suppressor protein p53 with the regulatory beta-subunit of protein kinase CK2. Oncogene. 1995 ;11(10):1971-1978
103. Leroy D. et al. Mutations in the C-terminal domain of topoisomerase II affect meiotic function and interaction with the casein kinase 2 beta subunit. Mol Cell Biochem. 1999 ;191(1-2):85-95
104. Li D. et al. Specific interaction between casein kinase 2 and the nucleolar protein Nopp140. J Biol Chem. 1997 ;272(6):3773-3779
105. Romero-Oliva F, Allende JE. Protein p21(WAF1/CIP1) is phosphorylated by protein kinase CK2 in vitro and interacts with the amino terminal end of the CK2 beta subunit. J Cell Biochem. 2001 ;81(3):445-452
106. Theis-Febvre N. et al. Protein kinase CK2 regulates CDC25B phosphatase activity. Oncogene. 2003 ;22(2):220-232
107. Tapia J.C. et al. Cell cycle regulatory protein p27KIP1 is a substrate and interacts with the protein kinase CK2. J Cell Biochem. 2004 ;91(5):865-879
108. Bonnet H. et al. Fibroblast growth factor-2 binds to the regulatory beta subunit of CK2 and directly stimulates CK2 activity toward nucleolin. J Biol Chem. 1996 ;271(40):24781-24787
109. Chong H. et al. Mechanisms of regulating the Raf kinase family. Cell Signal. 2003 ;15(5):463-469
110. Boldyreff B, Issinger OG. A-Raf kinase is a new interacting partner of protein kinase CK2 beta subunit. FEBS Lett. 1997 ;403(2):197-199
111. Walworth N. et al. Fission yeast chk1 protein kinase links the rad checkpoint pathway to cdc2. Nature. 1993 ;363(6427):368-371
112. al-Khodairy F. et al. Identification and characterization of new elements involved in checkpoint and feedback controls in fission yeast. Mol Biol Cell. 1994 ;5(2):147-160
113. Takai H. et al. Aberrant cell cycle checkpoint function and early embryonic death in Chk1(-/-) mice. Genes Dev. 2000 ;14(12):1439-1447
114. Chen P. et al. The 1.7 A crystal structure of human cell cycle checkpoint kinase Chk1: implications for Chk1 regulation. Cell. 2000 ;100(6):681-692
115. Filhol O. et al. Casein kinase II and the tumor suppressor protein P53 associate in a molecular complex that is negatively regulated upon P53 phosphorylation. J Biol Chem. 1992 ;267(29):20577-20583
116. Guex N, Peitsch MC. SWISS-MODEL and the Swiss-PdbViewer: an environment for comparative protein modeling. Electrophoresis. 1997 ;18(15):2714-2723
117. Berman H.M. et al. The Protein Data Bank. Nucleic Acids Research. 2000 ;28:235-242
118. Ermakova I. et al. Crystal structure of a C-terminal deletion mutant of human protein kinase CK2 catalytic subunit. J Mol Biol. 2003 ;330(5):925-934
Author biography
Ashley Bibby is pursuing graduate studies in David Litchfield's laboratory at The University of Western Ontario.
David Litchfield obtained Ph.D. under the supervision of Eric Ball at the University of Western Ontario followed by postdoctoral training in the laboratory of Edwin G. Krebs at the University of Washington. Litchfield established an independent research program at the Manitoba Cancer Treatment and Research Foundation before moving to the University of Western Ontario where he is currently a Professor in the Departments of Biochemistry and Oncology.
![]() Corresponding address:
Corresponding address:
David W. Litchfield, Department of Biochemistry, Siebens-Drake Research Institute, University of Western Ontario, London, Ontario, Canada, N6A 5C1. Telephone: 519-661-4186 Fax: 519-661-3175 Email: litchfica

 Global reach, higher impact
Global reach, higher impact