Impact Factor ISSN: 1449-2288
Int J Biol Sci 2006; 2(3):93-94. doi:10.7150/ijbs.2.93 This issue Cite
Editorial
Amphioxus: a peaceful anchovy fillet to illuminate Chordate Evolution (II)
Departament de Genètica, Facultat de Biologia, Universitat de Barcelona, Spain
Received 2006-5-19; Accepted 2006-5-22; Published 2006-5-22
Abstract
The genome of the amphioxus is on the horizon. With Linda Holland and Jeremy Gibson-Brown at the forefront, with all the amphioxus community behind, and with the Joint Genome Institute, the amphioxus genome will see the light this year, 2006. Hope that it will reflect the “prototypical” preduplicative genome of vertebrates. It may answer definitively what the human genome did not: Are we vertebrates octaploid? Will it shed light on the novelties that helped non-chordates to be chordates? And more, will amphioxus, with a simpler genome, be developed to a senior “experimental model system”, allowing the testing of molecular functions in a non-duplicated genome background and allowing genetic modification to “recapitulate” evolution? Thanks to an outstanding collaboration between labs, the laboratory culture of amphioxus is underway after years of hard work in the field. 2007 looks promising for amphioxus research.
Keywords: amphioxus, Evo-Devo, vertebrates, chordates
Introduction
The year 2006 may be remembered in history as “the year of the amphioxus”. First, the genome will, after years of waiting, see the light. Amphioxus and non-amphioxus specialists are heavy annotating and analysing the first draft. The efforts for sequencing the amphioxus genome started in USA and Europe in 1999. European agencies failed to fund (or we European researchers failed to convince the agencies of the importance of the amphioxus genome). Fortunately, at the other side of the Atlantic, Linda Holland (Scripps Institution, University of California) and Jeremy Gibson-Brown (Washington University) did not fail. They collaborated on a white paper to the Joint Genome Institute of the Department of Energy [1] that was supported by the relatively few labs concentrated on amphioxus research. [Ricard Albalat (Spain), Marianne Bronner-Fraser (USA), Hector Escrivà (France), Jordi Garcia-Fernàndez (Spain), Kaoru Kubokawa (Japan), Vincent Laudet (France), Thruston Lacalli (Canada), Leo Pezzement (USA), Georgia Panopoulou (Germany), Mario Pestarino (Italy), Pierre Pontarotti (France), T.V. Ventakatesh (USA), Hiroshi Wada (Japan), Ulrich Welsch (Germany), Kinya Yasui (Japan), and Shi-Cui Zhang (China)]. A number of outstanding scientists outside the amphioxus community also gave support to the proposal (the list is available in ref [1]). That the amphioxus genome was crucial to the understanding of the origin of the human genome maybe was not clear to funding agencies but it was for the Human Genome Consortium, as in the landmark Human Genome paper in 2000 [2]: “Our results so far are insufficient to settle whether two rounds of WGD occurred around 500 Myr ago [...]. Another approach to determining whether a widespread duplication occurred at a particular time in vertebrate evolution would be to sequence the genomes of organisms whose lineages diverged from vertebrates at appropriate times, such as amphioxus). Now we are there. Led by Dan Rokhsar (JGI), and also thanks to the EST collections made by the team of Nori Satoh (Kyoto University) in collaboration with Jr-Kai Yu and Linda Holland (University of California) we are about to know it: are we, human, octaploid? And which were the gene novelties (if any) that characterised chordate evolution? Coming soon, very soon...
And second, the big “drama” with amphioxus: how they “do it”. To be a model, a genetic, developmental and functional one, amphioxus “has” to allow researchers to use its eggs and sperm! And they do it not very often, the American species Branchiostoma floridae once every two weeks in the summer, and not much more often the Eastern species B. belchei. Still, pioneer experiments of gene knockdown, reporter constructs, and cell lineage have been performed [3-5]. As with the genome, there is a big new, again not thanks to the European funding agencies, but to the effort of European Research Groups. An Amphioxus Consortium headed by Hector Escrivá (Banyuls sur mer) and composed by members of the labs in Barcelona, Lyon, Paris and Genova have been, during three years, trying to understand the reproductive behaviour of the European amphioxus, B. lanceolatum (Fig. 1). And, finally, they succeeded! Now, nearly every day, we are able to obtain eggs and sperm on demand during the breeding season (April-July) [6]. This is possible not only in place, at the Laboratoire Arago, the marine station at Banyuls sur Mer. but also in our laboratory in Barcelona. As far as we know, this is the only “dry” lab (not a marine station) in where amphioxus eggs and sperm are obtained in a controlled manner, a small but big step towards the dream: developing amphioxus as an experimental model system (Fig. 2).
Amphioxus in the lab: Banyuls (left), Barcelona (center) and egg injection system (right).

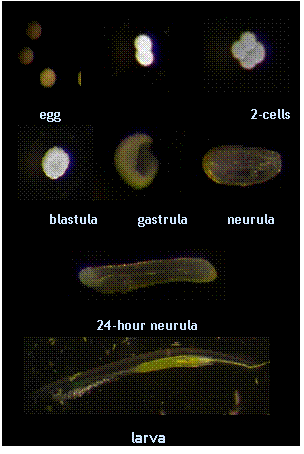
Developmental series of the amphioxus Branchiostoma floridae. Photograph courtesy of Hector Escrivà.
In this set of special issues, most of the amphioxus researchers that have fought for the genome and the embryos are giving their particular data or views in their particular area of research. All are focused on understanding the mechanisms behind evolution and the origin of vertebrates. This second part focuses particularly in the nervous system, one of the major steps forwards to the complexity of vertebrates.
Homologies, innovations, brain, neural crest, colinearity, ..., interesting, promising and provocative hypothesis arise from the study of our lovely little lancelet.
Genome by the corner, embryos by the hand, things are getting better, every day.
Acknowledgements
The author is very grateful to all the groups that are participating in this special issue, for their effort and friendship. Author's lab research is funded by grants BFU2005-00252 (Spanish Ministry of Education and Science), by the “Distinció” of the Generalitat de Catalunya, and by the European Community's Human Potential Programme HPRN-CT-2002-00263/BMC 'Neurogenome'.
Conflict of interest
The author has declared that no conflict of interest exists.
References
1. Gibson-Brown JJ, Osoegawa K, McPherson JD, Waterston RH, de Jong PJ, Rokhar DS, Holland LZ. A proposal to sequence the amphioxus genome submitted to the Joint Genome Institute of the US department of Energy. J Exp Zool (Mol Dev Evol). 2003 ;300B:5-22
2. International Human Genome Sequence Consortium. Initial sequencing and analysis of the human genome. Nature. 2001 ;409:860-921
3. Yu JK, Holland ND, Holland LZ. Tissue-specific expression of FoxD reporter constructs in amphioxus embryos. Dev Biol. 2004 ;274:452-461
4. Schubert M, Yu JK, Holland ND, Escriva H, Laudet V, Holland LZ. Retinoic acid signalling acts via Hox1 to establish the posterior limit of the pharynx in the chordate amphioxus. Development. 2005 ;132:61-73
5. Benito-Gutierrez E, Nake C, Llovera M, Comella JX, Garcia-Fernandez J. The single AmphiTrk receptor highlights increased complexity of neurotrophin signalling in vertebrates and suggests an early role in developing sensory neuroepidermal cells. Development. 2005 ;132:2191-2202
6. Fuentes M, Schubert M, Dalfo D, Candiani S, Benito E, Gardenyes J, Godoy L, Moret F, Illas M, Patten I, Permanyer J, Oliveri D, Boeuf G, Falcon J, Pestarino M, Fernandez JG, Albalat R, Laudet V, Vernier P, Escriva H. Preliminary observations on the spawning conditions of the European amphioxus (Branchiostoma lanceolatum) in captivity. J Exp Zool (Mol Dev Evol). 2004 ;302B:384-391
Author contact
Corresponding address:

 Global reach, higher impact
Global reach, higher impact