10
Impact Factor
ISSN: 1449-2288
Int J Biol Sci 2006; 2(3):133-141. doi:10.7150/ijbs.2.133 This issue Cite
Review
Genome duplications of early vertebrates as a possible chronicle of the evolutionary history of the neural crest
1. Graduate School of Life and Environmental Sciences, University of Tsukuba, Tsukuba 305-8572, Japan
2. Faculty of Integrated Arts and Sciences, University of Tokushima, 1-1 Minami-Josanjima, Tokushima 770-8502, Japan
Received 2006-2-1; Accepted 2006-5-22; Published 2006-5-23
Abstract
It is now accepted that ancestral vertebrates underwent two rounds of genome duplication. Here we test the possible utility of these genome duplication events as a reference time for the evolutionary history of vertebrates, by tracing the molecular evolutionary history of the genes involved in vertebrate neural crest development. For most transcription factors that are involved in neural crest specification, more than two paralogs are involved in that process. These were likely involved in the specification of the neural crest before the genome duplications occurred in ancestral vertebrates, although FoxD3 may have acquired that role after the genome duplications. By contrast, the epithelial-mesenchymal transition of neural crest cells is controlled by genes that evolved after the genome duplications, such as cadherin6, cadherin7, cadherin11, and rhoB. This suggests that primitive neural crest cells control their delamination by using a small or distinct set of cell adhesion molecules. Alternatively, these observations suggest that delamination of the neural crest evolved after the genome duplications. In that case, the neural crest might have evolved in sequential steps; the specification of the neural crest occurred before the genome duplications, and the neural crest acquired a new cell migration property after the genome duplications.
Keywords: vertebrates, genome duplication, neural crest, evolutionary sequence
1. Introduction
Several important novel structures evolved in ancestral vertebrates, including a complicated nervous system equipped with integrated sensory systems, bones, and acquired immune systems. Importantly, these novel structures probably did not evolve all at once, but via sequential steps. Therefore, to understand the early evolutionary history of the vertebrates, we need to reveal the evolutionary sequence of the emergence of these novel structures. This information will have rich implications concerning how the bodies of ancestral vertebrates became more elaborate, and what kinds of adaptation promoted the evolution of the body plan.
To infer the evolutionary sequence of vertebrate characteristics, fossil evidence is the richest source of information. In addition, the genome is another possible chronicle of the early history of vertebrates. It is now widely accepted that vertebrates experienced two rounds of whole genome duplication, which can be traced from conserved genomic segments in chromosomes [1, 2]. Traces of these genome duplications are also evidenced in the observations that most invertebrate genes have two to four counterparts in vertebrate genomes.
In this article, we test the possible relationship of these genome duplications to the evolution of the neural crest. Since Gans and Northcutt (1983) [3] proposed the new head theory, the origin and evolution of the neural crest has been one of the main issues in vertebrate evolutionary biology. The neural crest is a population of cells that differentiates at the boundary between the neural tube and the surface ectoderm. The neural crest has two peculiar cell characters: migration and pluripotency. Protochordates represented by ascidians and amphioxus, which are the closest relatives of vertebrates, lack comparable cells, such as migratory cells or multipotent cells, at the boundary between the neural tube and the surface ectoderm (but see Jeffery et al. (2004) [4], which will be discussed below). Therefore, the neural crest was once regarded as a new cell type that emerged at the origin of the vertebrates. In addition, the neural crest gives rise to several structures that are characteristic of vertebrates, such as cranial sensory organs and the craniofacial skeleton. Therefore, Gans and Northcutt (1983) [3] emphasized that the evolution of the neural crest together with the placode was a key evolutionary event in vertebrate evolution.
Recent analyses of morphogenetic genes in protochordates have led to a revision of this idea. The expression of amphioxus Dll and ascidian Pax3/7 homologs in the dorsal midline epidermis abutting the neural tube has led some authors to postulate that the differentiation of the dorsal midline epidermis and dorsal part of the neural tube of protochordates is regulated by machinery comparable to that of the vertebrate neural crest [5, 6]. However, the neural crest of vertebrates is quite distinct from its protochordate counterpart in terms of pluripotency and migration. Although cells in the amphioxus dorsal midline epidermis migrate, they do so as a cell sheet and do not delaminate from each other. Ascidian dorsal midline epidermal cells do not migrate. Therefore, the evolution of the neural crest may be regarded not as the birth of a new cell population, but as the acquisition of new cell characters, such as migration-delamination and pluripotency [7]. To gain insight into how these new cell characters evolved, the genes whose vertebrate homologs are involved in the delamination of the neural crest were analyzed.
Here, we attempt to reconstruct the evolutionary history of the neural crest by examining the molecular evolution of the genes involved in neural crest development, focusing on whether these genes were involved in neural crest development before or after the genome duplications. When more than two vertebrate paralogs are involved in a certain process of development, it is likely that these genes were involved in the developmental process before the genome duplications occurred. Conversely, when only one of the duplicated genes is involved in a certain developmental process, the gene may have acquired its function after the genome duplications occurred. Nevertheless, there are some instances in which the primitive function of a gene has been retained by only one of the duplicated genes, such as for the three hedgehog genes of vertebrates (Sonic [Shh], Desert [Dhh], and Indian [Ihh] hedgehog). Amphioxus possesses a single ancestral type of the gene (hh) that is expressed in the midline structure endoderm, the notochord, and the floorplate [8]. This primitive function of hh has been inherited by Shh only, and not by Ihh or Dhh. Therefore, when only one of the duplicated genes is involved in a certain developmental process, the function of the gene in protochordates must be considered. If only one of the duplicated genes possesses a certain role, which is not shared by protochordate homologs, it is more likely that the role was acquired after the genome duplications.
Since vertebrates experienced two rounds of genome duplication, they should ideally possess four genes corresponding to each protochordate gene. However, the duplicated genes were redundant, and some of them were lost secondarily. When only two paralogs are retained in vertebrates, it is not easy to deduce which round of duplication gave rise to the two genes. Moreover, some genes have experienced gene duplications that occurred independently of the genome duplications. For example, it is accepted that teleost fishes experienced a third genome duplication. To avoid the confusion that arises from additional gene duplications, phylogenetic analyses are mainly conducted using avian and mammalian genes.
Here, we analyzed when genes acquired their functions in neural crest development, and based on this information, we deduced the evolutionary history of the neural crest. We found that many, but not all, of the transcription factors were involved in neural crest development before the genome duplications. In contrast, only some of the paralogs of the cell adhesion molecules and RhoB that are involved in neural crest delamination are involved in neural crest development.
2. Transcription factors involved in the neural crest
Several transcription factors are known to be involved in neural crest development [9]. For most of these transcription factors, more than two paralogs are involved in neural crest specification. Both Pax3 and Pax7 are expressed at the neural plate border, i.e., at the lateral edge of the neural plate and the neural crest [10, 11]. Protochordates have single counterpart genes that are also expressed in the neural plate border [6, 12]. Therefore, it is likely that Pax3/7 acquired a role in neural crest specification before the genome duplications. Similarly, of the three mouse Msx genes, Msx1 and Msx2 are expressed in the neural plate border [13-17], and their protochordate counterpart also marks the neural plate border [18, 19]. Four Zic genes are expressed in the neural plate border of vertebrates [20]. Molecular phylogenetic studies have indicated that two ascidian Zic genes [macho and ZicL(N)] emerged from an ancestral gene independent of the duplication that gave rise to the vertebrate Zic genes [21, 22]. Although Gostling and Shimeld (2003) [23] did not resolve the molecular evolutionary status of amphioxus Zic, their tree is consistent with the idea that genome duplications in early vertebrates gave rise to multiple vertebrate Zic genes. Two ascidian Zic genes are expressed in the notochord, muscle, and neural tube, while a restricted expression pattern along the dorsoventral axis of the neural tube was not observed [21, 22, 24]. Amphioxus Zic is also expressed in the neural tube and somatic mesoderm, and later expression in the neural tube is restricted to the dorsal part [23].
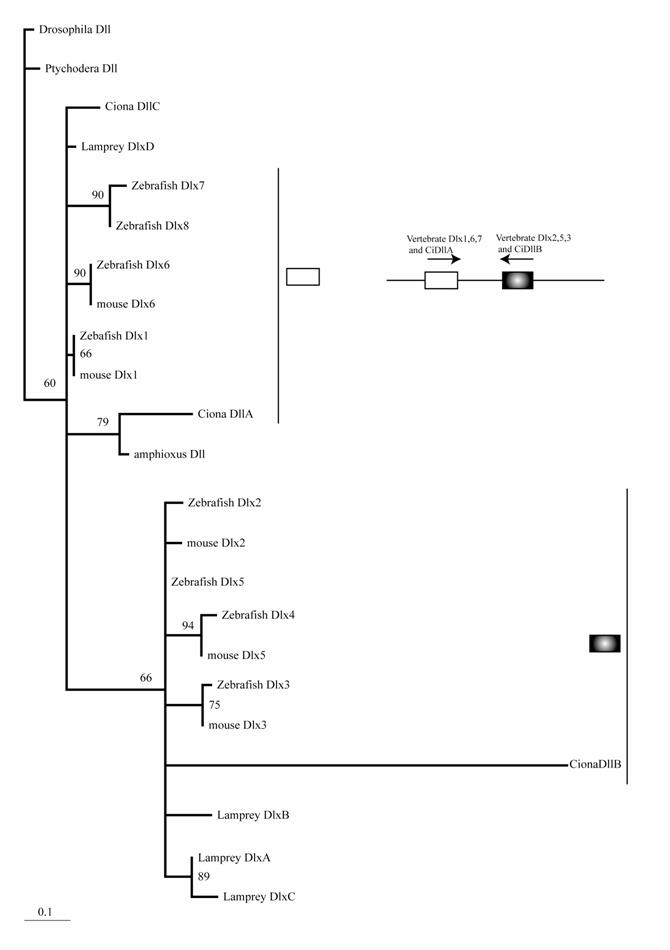
The mouse possesses six Dlx genes, all of which are expressed in the neural crest or its derivatives [25]. Although molecular phylogenetic analysis did not resolve the phylogeny of chordate Dlx genes, it is consistent with the idea that multiple Dlx genes have evolved through the genome duplications of ancestral vertebrates (Fig. 1). Stronger support for this conclusion comes from the genomic organization. The six vertebrate Dlx genes are located on chromosomes as three pairs, and the three pairs are linked with the Hox clusters of vertebrates [26]. Duplication of the Hox cluster of vertebrates is the best-known example of the genome duplications. Therefore, an ancestral Dlx gene linked to the Hox cluster duplicated in tandem. Subsequently, the genome duplications gave rise to four pairs of Dlx. The one linked to the HoxC cluster may have been lost. Ciona has three Dlx genes. Interestingly, two of these (DllA and DllB) are linked on a chromosome in the tail-to-tail direction as vertebrate homologs [27]. Our phylogenetic analysis weakly support the close relationship of Ciona DllA to vertebrate Dlx1, 6, and 7, while DllB is more closely related to Dlx2 , 3, and 5 of vertebrates. This suggests that the tandem duplication of Dlx genes predates the divergence of ascidians and vertebrates; i.e., the common ancestors of ascidians and vertebrates possessed Dlx genes linked in tandem, although the high substitution rate of Ciona Dlx genes makes the tree topology less convincing (Fig. 1). One of the Ciona genes, DlxC, and amphioxus Dlx are expressed in the neural plate border [5, 27, 28].
Molecular phylogenetic tree of Dlx genes. Molecular phylogenetic tree of Dlx genes. The tree is constructed by the quartet maximum likelihood method using TreePuzzle [54] based on amino acid sequences of the homeodomain. Vertebrate genes can be classified into two groups, and each protochordate gene shows phylogenetic affinity to one of them. This suggests that the tandem duplication of Dlx genes predates the divergence of the chordate groups.
In the chicken and mouse, the expression of snail and slug in the neural crest have been swapped; snail, but not slug, is expressed in the mouse pre-migratory neural crest, while slug, but not snail, is expressed in the chicken pre-migratory neural crest [29]. This swap can be explained easily if the common ancestors of vertebrates had acquired snail/slug expression before the gene duplication occurred. Indeed, in Xenopus, both snail and slug are expressed in the neural crest [30]. The molecular phylogeny supports the evolution of snail and slug in ancestral vertebrates via gene duplication, and amphioxus and ascidians possess the ancestral type of gene [31]. Both amphioxus and ascidian snail are expressed in the neural tube, although the expression profiles differ [31-32]. While amphioxus snail marks the edge of the neural plate that forms the dorsal part of the neural tube [31], the ascidian genes are expressed predominantly in the lateral cells of the neural tube and not in the dorsal part [33,32].
Three vertebrate paralogs of SoxE (Sox8, 9, and 10) are expressed in the neural crest. Both ascidian and amphioxus possess a single ancestral type of SoxE gene [9, 21] (HW, unpublished data). Therefore, although SoxE expression has not been examined in either ascidians or amphioxus, SoxE was likely involved in neural crest specification before the genome duplications.
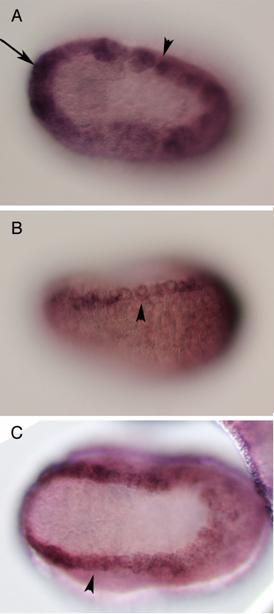
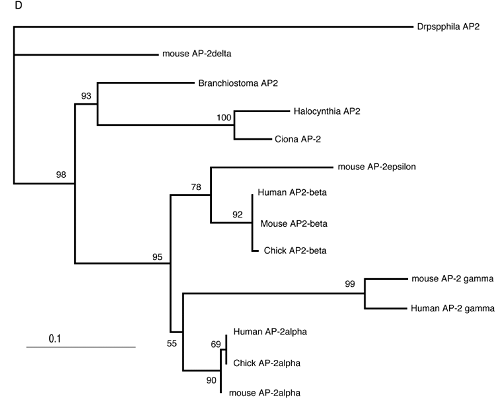
Vertebrates possess five AP-2 genes; three of them, AP-2a, b, and g, are expressed in the neural crest [34]. Molecular phylogenetic studies support the evolution of four vertebrate genes in the ancestral vertebrates, although we could not resolve the phylogenetic position of AP-2d [35] (Fig. 2D). Amphioxus AP-2 is expressed uniformly in epidermal cells [35]. Ciona has two AP-2 genes, and one of them is not expressed in early embryogenesis, while the other one (AP-2-like2) is expressed in epidermal cells [28]. Interestingly, careful examination has revealed that the expression of AP-2-like2 is up-regulated in the dorsal midline epidermis (Fig. 2A, B). Similar up-regulation is also observed in Halocynthia AP-2 (Fig. 2C). This expression pattern is reminiscent of Xenopus AP-2a, which is expressed rather broadly in epidermal cells, and their expression is subsequently up-regulated in the neural crest cells [36].
Characterization of the ascidian homologs of AP-2. (A-C) The expression of CiAP-2 (A, B) and HrAP-2 (C) at neurula stage. Embryos in A and C are viewed from dorsal side, while B is the lateral view (anterior to the left). Upregulation in the dorsal midline epidermis of AP-2 is observed in both Ciona and Halocynthia (arrowheads). Note that epidermal cells abutting the anterior edge of the neural plate also show upregulation of the AP-2 (arrow). (D) Molecular phylogenetic tree of AP-2 genes. AP-2 from Halocynthia roretzi (HrAP-2) was isolated from a cDNA library of the gastrula stage (Acc. No.: XXXX). The tree is constructed by quartet maximum likelihood method using Treepuzzle [54] based on the conserved amino acid sequences of the helix-span-helix motif.
Vertebrates have five paralogs of the D group Fox genes, of which FoxD1 and D2 are relatively closely related [37]. Ascidian and amphioxus possess the ancestral type of FoxD gene [38, 39]. Molecular phylogenetic studies support the hypothesis that the genome duplications gave rise to FoxD1-2, D3, D4, and D5 [38]. In contrast to the genes mentioned above, only one of the paralogs of FoxD, FoxD3, is involved in neural crest specification [40, 41]. Ascidian and amphioxus genes are expressed in the endoderm, notochord, and neural tube, and no restricted expression pattern was observed along the dorsoventral axis of the neural tube [38, 39]. Therefore, FoxD3 genes might have become involved in neural crest specification after the genome duplication occurred, although the possibility cannot be excluded that FoxD became involved in neural crest specification in ancestral vertebrates before the genome duplications, and that function was inherited by one of the duplicated genes, FoxD3.
3. Neural crest effectors involved in delamination
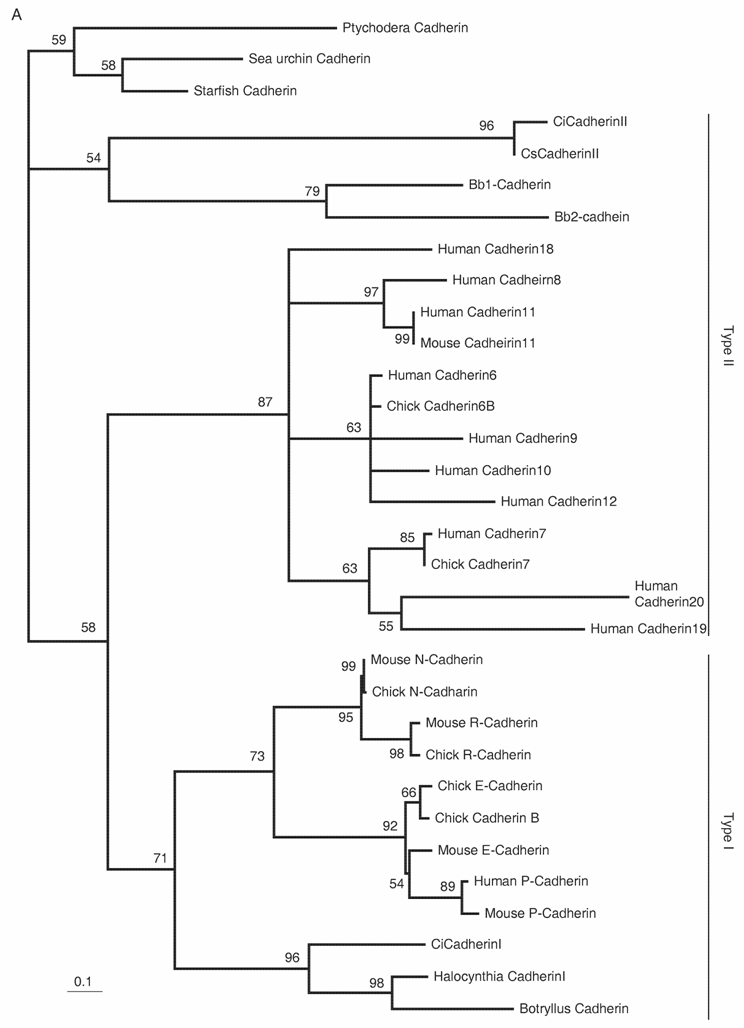
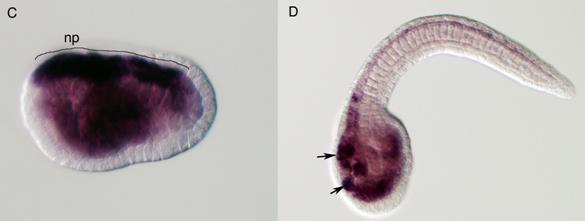
Delamination of the neural crest depends on the regulated expression of cell adhesion molecules, i.e., cadherins. N-cadherin and cadherin6 are expressed in the pre-migratory neural crest, while others, such as cadherin7 and cadherin11, are expressed in post-migratory neural crest cells [42,43]. The ectopic expression of N-cadherin or cadherin7 in the neural tube prevents neural crest delamination without affecting differentiation of the neural crest [44]. Molecular phylogenetic analyses of cadherin genes have shown that vertebrate cadherin genes can be classified into two families: type I cadherins include E‑, N-, P-, and R-cadherins, while type II includes other classic cadherins, such as cadherin6, 7, and 11 [45] (Fig. 3A). Ascidians have two cadherins corresponding to type I and II cadherins of vertebrates [46]. Type I cadherin is expressed ubiquitously both in Ciona and Halocynthia [47] (data not shown), whereas type II cadherin (CiCadherinII) is predominantly expressed in the neural tube [48] (Fig. 3C). Subsequently, CiCadherinII marks subpopulations of cells in the brain (Fig. 3D). This expression pattern of CiCadherinII suggests that in addition to its role in neurulation, type II cadherin played a role in the formation of neuromeres in the brain of the ancestral chordate. Amphioxus possesses two cadherins, whose molecular structures are odd and show similarity to those of non-chordate invertebrates [49, 50]. We could not tell when the gene duplication that gave rise to type I and type II cadherin occurred in the present molecular phylogenetic analysis [50]; however, multiple copies of type I and type II genes obviously emerged in ancestral vertebrates (Fig. 3A). Therefore, the evolution of the molecular machinery using these cadherins for neural crest delamination must have occurred after the genome duplications.
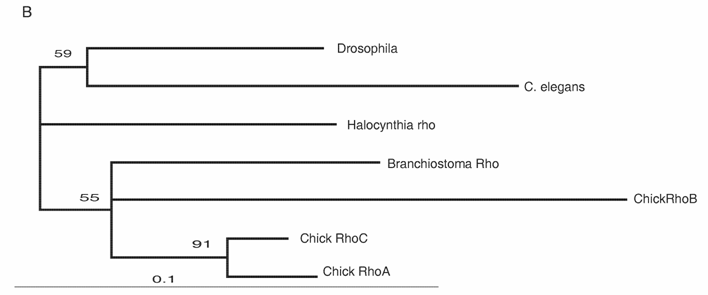
The Rho family of small GTPases is involved in several aspects of regulating cell motility. RhoB is responsible for regulating the delamination of the neural crest [51]. Molecular evolutionary analyses have presented evidence that vertebrates acquired three Rho genes that were derived from a single ancestral chordate gene [52] (Fig. 3B). Halocynthia has a Rho gene that is expressed ubiquitously (data not shown). Therefore, RhoB acquired specific neural crest expression after the genome duplications.
In contrast to the transcription factors involved in neural crest specification, the molecular evolutionary histories of cadherins and Rho genes suggest that the genome duplications of vertebrates were essential for the innovation of neural crest cells as migrating cells. It is not certain whether the gene duplications of rho GTPases were essential for the evolution of neural crest delamination, i.e., whether the neural crest requires a specific type of rho GTPase for its control of delamination. The ancestral vertebrate could possibly control the delamination of neural crest cells via the posttranscriptional regulation of the activity of a single rho gene. In the case of cadherins, however, without genome duplication, the ancestral vertebrates could not possess specific repertoires of cadherin molecules for the pre- and post-migratory stages. Interestingly, FoxD3, which was recruited in neural crest specification possibly after the genome duplications, is involved in regulating cadherin7 expression [53]. Assuming that the ancestral vertebrates acquired a migratory neural crest before the genome duplications, the delamination of the neural crest must have been regulated using a small set of adhesion molecules. This is not impossible because invertebrates, such as Drosophila and sea urchins, regulate several aspects of epithelial-mesenchymal transitions using a small set of cell adhesion molecules. However, vertebrates are far richer in mesenchymal cells than are any other chordates. This may result because vertebrates possess more repertoires of cell adhesion molecules, which arose via the genome duplications, and the neural crest may be one of the new cell types that acquired mesenchymal cell properties by using the rich repertoires of cell adhesion molecules. In this sense, it is interesting to determine how the delamination of the lamprey neural crest is regulated. It is now accepted that the genome duplications occurred before the divergence of gnathostomes. However, it remains to be elucidated whether extant agnathans, such as lampreys and hagfish, experienced the second round of gene duplication [2]. If the ancestral vertebrate possessed true migrating neural crest cells before the genome duplication, they must have been controlled by mechanisms distinct from that in present gnathostomes, which utilize multiple sets of cadherins.
Jeffery et al. (2004) found that one colonial species of ascidian (Ecteinascidia turbinate) possesses migrating cells from the neural tube, and that they differentiate into pigment cells, just like vertebrate neural crest cells [4]. As mentioned above, however, the delamination of these cells is unlikely to be regulated by switching several paralogs of cadherin genes. The control of delamination in the ascidian neural crest-like cells is an interesting issue. The mechanism regulating those cells may represent the ancestral machinery of the genetic control of neural crest delamination before the genome duplications. Conversely, because the neural crest-like cells begin migration well after neural tube closure, the cells might have acquired migration independently from the vertebrate neural crest.
Molecular evolution of cadherin and Rho genes. (A, B) Molecular phylogenetic trees of cadherin (A) and rho (B) genes, constructed by quartet maximum likelihood method using TreePuzzle 5.0 [54]. The amino acid sequences of the C-terminus cytoplasmic domain were used for the cadherin gene analysis, while entire amino acid sequences were used for the rho genes. Nucleotide sequences of rho genes from Branchiostoma belcheri (BbRho) and Halocynthia roretzi (HrRho) by following Acc. Nos. (YYYY for BbRho and ZZZZ for HrRho). (C, D) The expression of CiCadherinII in the neurula (C: lateral view) and tailbud stage embryo (D). CiCadherinII is expressed uniformly in the neural plate of neurula embryo (C), while in tailbud stage, it shows more restricted expression in the neural tube (D), suggesting its role in neuromere formation of ascidian CNS.
4. Evolutionary sequence of the vertebrate neural crest
By referring to the molecular evolutionary histories of the genes involved in neural crest cell differentiation and delamination, we reconstructed the hypothetical evolutionary steps of neural crest evolution. Most of the transcription factors involved in the neural crest specification have ancestral functions in the neural plate border or neural tube of protochordates. These transcription factors likely acquired roles in neural plate border and neural crest specification before the genome duplications, except for FoxD3, which might have been co-opted for neural crest specification after the genome duplications. By contrast, the epithelial-mesenchymal transition of neural crest cells is controlled by genes that evolved after the genome duplications. This suggests that delamination of the neural crest evolved after the genome duplications, although the possibility that primitive neural crest cells control their delamination via a distinct mechanism using a small set of cell adhesion molecules cannot be dismissed. The key factor to determine whether the primitive neural crest delaminated prior to the genome duplications may be the lamprey neural crest. The lamprey has true migratory neural crest cells, but it may not have experienced both rounds of genome duplication.
Acknowledgements
We thank Nori Satoh for providing the Ciona gene collection, which greatly facilitated this study, and Hiroki Oda for useful discussions on cadherin evolution. We also thank Nina Sugino and Kyosuke Kawahata for technical assistance of Ciona in situs. Part of the work presented here is supported by Kato Memorial Bioscience Foundation, and by Grants-in-Aid from the ministry of Education, Science, Sports and Culture, Japan (Nos. 12026219, 13045020, 14034228) to HW.
Conflict of interest
The authors have declared that no conflict of interest exists.
References
1. Ohno S. Evolution by gene duplication. New York: Springer-Verlag. 1970
2. Furlong RF, Holland PWH. Were vertebrate octoploid. Phil Trans R Soc Lond B. 2002 ;357:531-544
3. Gans C, Northcutt RG. Neural crest and the origin of vertebrates: a new head. Science. 1983 ;220:268-274
4. Jeffery WR, Strickier AG, Yamamoto Y. Migratory neural crest-like cells form body pigmentation in a urochordate embryo. Nature. 2004 ;431:696-699
5. Holland ND, Panganiban G, Henyey EL, Holland LZ. Sequence and developmental expression of AmphiDll, an amphioxus Distal-less gene transcribed in the ectoderm, epidermis and nervous system: insights into evolution of craniate forebrain and neural crest. Development. 1996 ;122:2911-2920
6. Wada H, Holland PWH, Sato S, Yamamoto H, Satoh N. Neural tube is partially dorsalized by overexpression of HrPax-37: the ascidian homologue of Pax-3 and Pax-7. Dev Biol. 1997 ;187:240-252
7. Wada H. Origin and evolution of the neural crest: a hypothetical reconstruction of its evolutionary history. Dev Growth Differ. 2001 ;43:509-520
8. Shimeld SM. The evolution of the hedgehog gene family in chordates: insights from amphioxus hedgehog. Dev Genes Evol. 1999 ;209:40-47
9. Meulemans D, Bronner-Fraser M. Gene-regulatory interactions in neural crest evolution and development. Dev Cell. 2004 ;7:291-299
10. Liem KFJ, Tremml G, Roelink H, Jessel TM. Dorsal differentiation of neural plate cells induced by BMP-mediated sugnals from epidermal ectoderm. Cell. 1995 ;82:969-979
11. Mansouri A, Stoykova A, Torres M, Gruss P. Dysgenesis of cephalic neural crest derivatives in Pax7-/- mutant mice. Development. 1996 ;122:831-838
12. Holland LZ, Schubert M, Kozmik Z, Holland ND. AmphiPax3/7, an amphioxus paired box gene: insights into chordate myogenesis, neurogenesis, and the possible evolutionary precursor of definitive vertebrate neural crest. Evol Dev. 1999 ;1:153-165
13. Hill RE. et al. A new family of mouse homeo box-containing genes: molecular structure, chromosomal location, and developmental expression of Hox-7.1. Genes Gev. 1989 ;3:26-37
14. Robert B, Sassoon D, Jacq B, Gehring W, Buckingham M. Hox-7, a mouse homeobox gene with a novel pattern of expression during embryogenesis. EMBO J. 1989 ;8:91-100
15. MacKenzie A, Ferguson MW, Sharpe PT. Hox-7 expression during murine craniofacial development. Development. 1991 ;113:601-611
16. MacKenzie A, Ferguson MW, Sharpe PT. Expression patterns of the homeobox gene, Hox-8, in the mouse embryo suggest a role in specifying tooth initiation and shape. Development. 1992 ;115:403-420
17. Shimeld SM, MaKay IJ, Sharp PT. Th emurine homeobox gene Msx-3 hows highly restricted expression in the developing neural tube. Mech Dev. 1996 ;55:201-210
18. Sharman AC, Shimeld SM, Holland PWH. An amphioxus Msx gene expressed predominantly in the dorsal neural tube. Dev Genes Evol. 1999 ;209:260-263
19. Aniello F. et al. Identification and developmental expression of Ci-msxb: a novel homologue of Drosophila msh gene in Ciona intestinalis. Mech Dev. 1999 ;88:123-126
20. Nakata K, Koyabu Y, Aruga J, Mikoshiba K. A novel member of Xenopus Zic family, Zic5, mediates neural crest development. Mech Dev. 2000 ;99:83-91
21. Imai KS, Hino K, Yagi K, Satoh N, Satou Y. Gene expression profiles of transcription factors and signaling molecules in the Ciona intestinalis embryo. Development. 2004 ;131:4047-4058
22. Wada S, Saiga H. HrzicN, a new Zic family gene of ascidians, plays essential roles in the neural tube and notochord development. Development. 2002 ;129:5597-5608
23. Gostling NJ, Shimeld SM. Protochordate Zic genes define primitive somite compartments and highlight molecular changes underlying neural crest evolution. Evol Dev. 2003 ;5:136-144
24. Satou Y. et al. macho-1-related genes in Ciona embryos. Dev Genes Evol. 2002 ;212:87-92
25. Panganiban G, Rubenstein JLR. Developmental functions of the Distal-less/Dlx homeobox genes. Development. 2002 ;129:4371-4386
26. Stock DW. et al. The evolution of the vertebrate Dlx gene family. Proc Natl Acad Sci USA. 1996 ;93:10858-10863
27. Caracciolo A, Gregorio AD, Aniello F, DiLauro R, Branno M. Identification and developmental expression of three Distal-less homeobox containing genes in the ascidian Ciona intestinalis. Mech Dev. 2000 ;99:173-176
28. Imai KS, Hino K, Yagi K, Satoh N, Satou Y. Gene expression profiles of transcription factors and signaling molecules in the Ciona intestinalis embryo. Development. 2004 ;131:4047-4058
29. Sefton M, Sanchez S, Nieto MA. Conserved and divergent roles for members of the Snail family of transcription factors in the chick and mouse embryo. Development. 1998 ;125:3111-3121
30. Linker C, Bronner-Fraser M, Mayor R. Relationship between Gene Expression Domains of Xsnail, Xslug, and Xtwist and Cell Movement in the Prospective Neural Crest of Xenopus. Dev Biol. 2000 ;224:215-225
31. Langeland J, Tomsa JM, Jackman WR, Kimmel CB. An amphioxus snail gene: Expression in paraxial mesoderm and neural plate suggests a conserved role in patterning the chordate embryo. Dev Genes Evol. 1998 ;208:569-577
32. Wada S, Saiga H. Cloning and embryonic expression of Hrsna, a snail family gene of the ascidian Halocynthia roretzi: implication in the origins of mechanisms for mesoderm specification and body axis formation in chordates. Dev Growth Differ. 1999 ;41:9-18
33. Corbo JC, Erives A, DiGregorio A, Chang A, Levine M. Dorsoventral patterning of the vertebrate neural tube is conserved in a protochordate. Development. 1997 ;124:2335-2344
34. Hilger-Eversheim K, Moser M, Schorle H, Buettner R. Regulatory roles of AP-2 transcription factors in vertebrate development, apoptosis and cell-cycle control. Gene. 2000 ;260:1-12
35. Meulemans D, Bronner-Fraser M. Amphioxus and lamprey AP-2 genes: implications for neural crest evolution and migration pattern. Development. 2002 ;129:4953-4962
36. Luo T, Lee YH, Saint-Jeannet JP, Sargent TD. Induction of neural crest in Xenopus by transcription factor AP2a. Proc Natl Acad Sci USA. 2003 ;100:532-537
37. Kaestner KH, Knöchel W, Martínez DE. Unified nomenclature for the winged helix/forkhead transcription factors. Genes Dev. 2000 ;14:142-146
38. Imai KS, Satoh N, Satou Y. An essential role of FoxD gene in notochord induction in Ciona embryos. Development. 2002 ;129:3441-3453
39. Yu J, Holland ND, Holland LZ. Tissue-specific expression of FoxD reporter constructs in amphioxus embryos. Dev Biol. 2004 ;274:452-461
40. Kos R, Reedy MV, Johnson RL, Erickson CA. The winged-helix transcription factor FoxD3 is important for establishing the neural crest lineage and repressing melanogenesis in avian embryos. Development. 2001 ;128:1467-1479
41. Sasai N, Mizuseki K, Sasai Y. Requirement of FoxD3-class signaling for neural crest determination in Xenopus. Development. 2001 ;128:2525-2536
42. Nakagawa S, Takeichi M. Neural crest cell-cell adhesion controlled by sequential and subpopulation-specific expression of novel cadherins. Development. 1995 ;121:1321-1332
43. Borchers A, David R, Wedlich D. Xenopus cadherin-11 restrains cranial neural crest migration and influences neural crest specification. Development. 2001 ;128:3049-3060
44. Nakagawa S, Takeichi M. Neural crest emigration from the neural tube depends on regulated cadherin expression. Development. 1998 ;125:2963-2971
45. Nollet F, Kools P, van Roy F. Phylogenetic analysis of the cadherin superfamily allows identification of six major subfamilies besides several solitary members. J Mol Biol. 2000 ;299:551-572
46. Sasakura Y. et al. A genomewide survey of developmentally relevant genes in Ciona intestinalis X. Genes for cell junctions and extracellular matrix. Dev Genes Evol. 2003 ;213:303-313
47. Satou Y. et al. A cDNA resource from the basal chordate Ciona intestinalis. Genesis. 2002 ;33:153-154
48. Imai KS. Isolation and characterization of b-catenin downstream genes in early embryos of the ascidian Ciona savignyi. Differentiation. 2003 ;71:346-360
49. Oda H. et al. A novel amphioxus cadherin that localizes to epithelial adherens junctions has an unusual domain organization with implication for chordate phylogeny. Evol Dev. 2002 ;4:426-434
50. Oda H, Akiyama-Oda Y, Zhang S. Two classic cadherin-related molecules with no cadherin extracellular repeats in the cephalochordate amphioxus: distinct adhesive specificities and possible involvement in the development of multicell-layered structures. J Cell Sci. 2003 ;117:2757-2767
51. Liu J, Jessel TM. A role for rhoB in delamination of neural crest cells from the dorsal neural tube. Development. 1998 ;125:5055-5067
52. Hotta K, Takahashi H, Ueno N, Gojobori T. A genome-wide survey of the genes for planar polarity signaling or convergent extension-related genes in Ciona intestinalis and phylogenetic comparisons of evolutionary conserved signaling components. Gene. 2003 ;317:165-185
53. Dottori M, Gross MK, Labosky P, Goulding M. The winged-helix transcription factor Foxd3 suppresses interneuron differentiation and promotes neural crest cell fate. Development. 2001 ;128:4127-4138
54. Strimmer K, von Haeseler A. Qualtet puzzling: a quartet maximum likelihood method for reconstructing tree topologies. Mol Biol Evol. 1996 ;13:964-969
Author contact
![]() Corresponding address:
Corresponding address:

 Global reach, higher impact
Global reach, higher impact