Impact Factor
ISSN: 1449-2288
Int J Biol Sci 2006; 2(4):216-226. doi:10.7150/ijbs.2.216 This issue Cite
Research Paper
Homeostatic restitution of cell membranes. Nuclear membrane lipid biogenesis and transport of protein from cytosol to intranuclear spaces.
Research Center C-873, University of Medicine and Dentistry of New Jersey, New Jersey Dental School, Newark, NJ 07103, USA
Received 2006-6-27; Accepted 2006-8-29; Published 2006-8-30
Abstract
Our studies on homeostatic restitution of cellular and subcellular membranes showed that vesicular intracellular transport is engaged in systematic and coordinated replacement of lipids and proteins in the membranes of the secretory, non-dividing epithelial cells (Slomiany et al., J. Physiol. Pharmacol. 2004; 55: 837-860). In this report, we present evidence on the homeostatic restitution of lipids in the biomembranes that constitute nuclear envelopes. We investigated nuclear membranes lipid synthesis by employing purified intact nuclei (IN), the outer nuclear membrane (ONM), the inner nuclear membrane (INM) and the cell cytosol (CC). In contrast to Endoplasmic Reticulum (ER) which in the presence of CC generates new biomembrane that forms ER vesicles transporting ER products to Golgi, the IN, ONM and INM are not producing transport vesicles. Instead, the newly synthesized lipids remain in the nuclear membranes. The membranes (INM, ONM) of IN incubated with CC become enriched with newly synthesized phosphatidylcholine (PC), phosphatidylinositol (PI), phosphatidylinositol phosphates (PIPs) and phosphatidic acid (PA). The incubation of separated ONM and INM with CC also enriched the membranes with IN specific lipids identified above. Moreover, the incubation of IN or its membranes with CC afforded retention of numerous CC proteins on the nuclear membrane. Here, we concentrated on 30kDa CC protein that displayed affinity to nuclear membrane PIP2. The 30kDa CC protein bound to PIP2 of IN, INM, and ONM. With IN, initially the PIP2-30kDa CC protein complex was detected on ONM, after 30-120 min of incubation, was found on INM and in nuclear contents. At the same time when the 30 kDa protein was released from INM and found in nuclear contents, the PIP2 of INM and ONM became undetectable, while the lipid extract from the membrane displaced from IN contained labeled PI only. Since ONM is an uninterrupted continuum of ER and INM, we speculate that the synthesis of the lipids in the ER, in the region adjacent to nucleus, is defining nuclear outer and inner biomembrane composition, is responsible for transport of the cytosolic protein into the nucleus and, replenishment of ER membrane used for vesicular transport.
Keywords: Phospholipids of nuclear membranes, Inner and outer nuclear membrane biogenesis and restitution, Phosphatidylinositol phosphates, Cytosol protein transport to nucleus.
1. Introduction
Transcriptome informs which genes are induced or repressed by the metabolic and physical status of the cell, but does not generate the roadmap that illustrates cellular homeostatic restitutions, or cellular changes leading to transformation. The largest void in the understanding of cellular signaling is created by an inadequate presentation of the correct spatial organization of all molecules that are crucial in carrying signals and traversing cellular, organellar, and nuclear membranes [1,2-5]. The general information on protein structure and modification is insufficient to explain when the protein traverses cellular membranes and uniquely intercalates into the membrane, but merely indicates potential to associate with outer- or inner membranous surface. While hydrophobic and hydrophilic domains of the protein indicate its native intra-membrane placement, the information on how and when the protein is placed within specialized membrane, or its intercalation into the specific membrane domain is assured remains unknown [6-9]. Hence, the quintessential event of the protein insertion into exact site of cellular membrane that governs fidelity of the homeostatic processes has not been identified [1, 6-9]. In the investigative paradigms of proteome, the studies also bypass the relevance of cell compartmentalization with specific membrane systems, the different rate of cellular compartments restitution and the significance of signal translation within the spaces delimited by membranous networks. Consequently, we only know that the genome responds to extracellular signals, the concentration of transcription factors, and that the rate of protein production and deposition in the specific sites or cell compartments must be ceaselessly reproduced [1,7-10].
In the terminally differentiated epithelial cell, the secretory activity of the apical surface of the cell requires steady and precise regeneration of cellular organelles, nucleus and the cell membrane. Thus, to sustain the designated and functional structure of the cell, the processes are engaged that generate new membranes and catabolize the membrane that is modified by signal-induced receptors [5,11-19]. Yet, this fundamental aspect of the whole cell reassembly is not investigated, and the synthesis of cellular membranes is presumed as deposition of native or extraneous lipids from cytosol, or the intracellular retrieval of the membranous transporters after release of the secretory cargo [7,8,20-26]. The provisional assumptions of the cell membranes regeneration are not challenged or argued that such processes would create indistinguishable cellular membranes, and not the membranes that provide discretely dissimilar profile for each intracellular organelle. Moreover, the receptors affinity to membrane due to hydrophobic amino acid sequence in the protein is also presumed as sufficient argument for the receptor intercalation into the membrane [20,23,27]. Hence, when the intercalation of membrane receptors takes place is not pursued.
Our investigations on biomembrane biogenesis concentrate on the initiation and restitution of the cellular membranes and provides evidence that in endoplasmic reticulum (ER), the membrane lipids and the specific membrane protein are intercalated and as ER transport vesicles delivered en bloc to Golgi [13-18]. In Golgi, the maturation/modification of the membrane ensures distinct composition of the transport vesicles that restore original proteins and lipids in Golgi, endosomes, and apical cell membrane [16-18]. In this report, we present evidence on the restitution of nuclear membranes which proceeds through the synthesis of lipids in the outer nuclear/ER membrane, and hypothesize that such a course of events is connected to the restitution of the inner nuclear membrane, transport of the cytosol-contained protein to nucleus, and the restitution of ER membrane. The finding that cell cytosol impacts organelle and nuclear membrane restitution provides important clue for designing media to redirect, impact or control cell growth, differentiation, and transformation.
2. Materials and Methods
Perfusion buffers
(a) buffered saline, with 10 mM potassium phosphate, pH 6.8, (b) buffered saline containing 0.5 mM MgCl2 and 0.5 mM MgS04, (c) buffered saline (100 ml) containing 66 mg collagenase, 80 mg hyaluronidase, and 2 g of albumin, (d) MSB, pH 6.9 buffer consisting of 0.1 Pipes, pH 6.9, 2.0 M glycerol, 1mM Mg acetate, 0.5 mM EGTA and mixture of protease inhibitors consisting of leupeptin, aprotinin and PMSF, (e) MSB buffer containing 0.2 % Triton X100, (f) 50 mM TRIS-HCl, pH 7.4 containing 0.25 M sucrose, 10 mM MgCl2 1 mM DTT,10 mg/ml leupeptin and 2 mM PMSF.
Perfusion of rat liver and isolation of hepatocytes
The abdomen of anestetized rat was open and liver cannulated and perfused with 50 ml of ice cold Hanks Balanced Salts (Sigma) without Ca2+containing 20 mg collagenase, type IV, 20 mg hyaluronidase, 1 g defatted albumin, and 3g of heparin. After initial perfusion, the liver was removed from animal abdomen, cut into slices and incubated with cold Hanks solution. Thus prepared slices were subjected to incubation in tissue incubator in 95% 02 and 5% C02 for 40 min, at 37°C. The slices were broken up with rubber policemen, incubated for additional 10 min and filtered through nylon mesh that separated single cells from larger debris. The cells were then centrifuged at 50xg for 2min, washed twice with the enzyme-free Hanks medium, twice with the Minimum Essential medium and counted in hemocytometer. Thus prepared cells were used for preparation of nuclei [28] subcellular organelles and cell cytosol [13-17] and lipid rafts [29]. In the experiments dedicated to the determination of lipid synthesis with and without cell cytosol, the preparations of intact nuclei were additionally rinsed with PBS and urea-PBS in order to remove the residual cytosolic proteins from the nuclear membranes. Thus prepared intact Nuclei (IN) were used for preparation of outer nuclear membrane (ONM) and the Inner Nuclear Membrane (INM). The DNAse treated nuclear matrix, the nuclear contents [28] was used in the set of experiments where instead of the CC, and the nuclear contents was checked for the phospholipids synthesizing activity. The synthesis of phosphatidylinositides and phospholipids was determined using radiolabeled [3H] inositol and arachidonate and [3H]choline [13-17]. The synthesis of transport vesicles (ER, Golgi transport vesicles) was performed in medium containing cytosol at concentration of 5 mg protein/ml of incubation mixture enriched with 50 mM ATP, 250 mM, 50 mM GTP, 5 mM creatine phosphate, 8.0 IU/ml creatine kinase, and where indicated 25 mg/ml RNase, 10 mM UDP-Glc and 10 mM palmitoyl CoA [13-18]. The same preparation of cytosol was used in the experiments with intact nuclei and nuclear membranes.
Isolation and separation of outer and inner nuclear membranes
One volume of the isolated hepatocytes or the IN was homogenized in 8 volumes of 10 mM potassium phosphate buffer, pH 6.8, 1.3 M sucrose and 1mM MgCl2 to rupture at least 80 % of nuclei [28]. The unbroken cells were removed by centrifugation at 50xg for 3 min, and the homogenate centrifuged for 15 min at 1000Xg. The soluble cellular material was saved for experiments evaluating its impact on membrane synthesis, and the nuclear pellet processed further [28]. The nuclear fractions were suspended in a minimum volume of buffered sucrose, adjusted to 2.2 M buffered sucrose with 2.4 M sucrose in the same buffer, followed by centrifugation at 100,000xg (27,000rpm in Beckman 45 Ti rotor) for 1h. The resultant pellet was suspended in 20 mM Tris-HCl, pH 7.5, centrifuged at 1000xg for 15 min, and the recovered nuclei suspended in a buffer (f) at concentration of 2 mg protein/ml. The preparation was then adjusted to 1% (w/v) with sodium citrate and incubated on ice with gentle stirring for 30 min and then centrifuged at 500xg for 15 min. The obtained supernatant contained the outer nuclear membranes ONM whereas the pellet contained the INM [28]. The pellet was suspended in buffer (f) at concentration of 5 mg/ml and digested with DNase 1 (250 mg/ml for 14 h at 4°C). The digested material was separated on sucrose gradient of 0.25-1.6- 2.4 M sucrose by centrifugation at 10,000xg for 2h. The inner nuclear membranes were recovered at 1.6 M sucrose boundary. The fraction of ONM was collected from citrate supernatant by centrifugation at 100,000xg for 20 min. The membrane pellet was suspended in buffer (f) and subjected to the same treatment as INM. The resultant 100 000Xg pellet of ONM was suspended in buffer (f). On the average, the preparation initiated with material obtained from two rat livers yielded 4.7 mg of inner and 3.1 mg outer nuclear membranes.
In the experiments employing IN, the incubation with CC followed by separation of the ONM and INM, the IN samples were subjected to additional sucrose gradient purification. The samples of IN were suspended in 40% buffered sucrose by adding 80% sucrose in 150 mM NaCl, 25 mM TRIS, pH 7.5 buffer and overlaid with 10-30% sucrose gradient in the same buffer and centrifuged at 29,000xg for 21 h. Following gradient centrifugation, the nuclei were recovered from 40 % bottom layer by diluting out sucrose with the buffer and centrifugation at 1,000xg for 10 min. The nuclear pellet was then suspended at a protein concentration of 2mg/ml in 1% citrate and proceeded with separation of outer and inner nuclear membranes. The material recovered from the gradient, was subjected to lipid extraction and SDS-PAGE. This treatment separated any cell membranes that were trapped with nuclei, and allowed us to determine whether incubation of IN with CC produced extranuclear radiolabeled membranes replacing ER membranes used in transport.
Cell surface and cell cytosol labeling with NIP
An aliquot of purified cells corresponding to 1 mg of cell membrane proteins or 1 mg of CC protein/ml of PBS, pH 7.4 was incubated with 10 μl 4-hydroxy-5-iodo-3-nitrophenyl acetate (NIP) in DMSO for 1h at 37°C. The excess of the reagent was removed on Sephadex G-25 column. The protein collected in three 1ml aliquots was used for incubation with individual lipids applied to microplates (ELISA) or with IN, ONM, INM or the respective lipid extracts. Where indicated the rat anti NIP IgG was also used. Protein recovery was calculated by multiplying the concentration of protein by the volume of the derivatized protein fraction recovered from the separation column. On the average, the concentration of NIP hapten was 1.0 x 10-5 mmoles/ml protein and 5.0 moles of NIP hapten/mol IgG. The absorbance of the derivatized protein was read at 280nm and 430nm.
3. Results
By choosing the cells which are extremely active in the synthesis and transport of the secretory proteins we were able to determine the specificity and lipid composition of the biomembrane that delivers the ER products to Golgi, determine the fine adjustments in the biomembrane composition for the restoration of the apical epithelial cell membrane, and the membrane which was provided for the restitution of endosomes [18]. Our previous studies have demonstrated that intracellular vesicular transport is dependent on the finely tuned synthesis of the cell-specific protein and lipids that assemble a precise vesicular biomembrane in ER [13-18]. Precise and explicit intercalation of the protein into the membrane can only take place when the translation of mRNA and synthesis of biomembrane proceed concomitantly within the ER space capable to generate specific lipids and translate specific mRNA [10,18]. Moreover, the composition of lipids and the membrane protein of a new membrane and the vesicular cargo determine the generation and directional transport of ER vesicles to Golgi. Thus, we found out that the synthesis of ceramides determines the quantity of transport vesicles that deliver protein to Golgi [18]. Concurrently, the substrates in the in situ medium (the cell cytosol) defined the process of transport vesicles synthesis and delivery to the next organelle, since depletion of mRNA from the cytosol reduced the overall transport to Golgi, and was rescued with reinstatement of the initial cytosolic mRNA [18]. Together, these results allowed us to hypothesize that synthesis of the intracellular biomembrane and the protein in ER must be initiated in a explicit site of the organelle, the site that is capable to assemble constitutive protein, specifically formulated biomembrane, and deliver it en bloc to the destination point.
In this report, our studies on nuclei and the inner and outer nuclear/ER membrane reveal feasible path associated with nuclear membrane restitution and transport of the cytosolic protein into the nucleus. While the previous studies [13-18] determined the requirements for the transport and restitution of biomembrane that constitute the outer apical portion of cell membrane and the membranes of Golgi and endosomes, the restitution of ER and the cell nucleus membranes were not apparent or consequential of the same pathway. Also, it was evident that ER is not renewed by the retrograde vesicular transport, since the lipids of ER and nucleus are not identical with the lipids of the apical cell membrane [25,26,29-32]. In addition, we could not substantiate a popular explanation of the subcellular membrane modification through the transferring of the individual lipids from cytosol into ER [33], or nuclear membrane [29,34], or bi-directional transport between ER and Golgi [26]. As shown in our previous investigation [18] and here in Fig. 1 (panel A, lanes 1,2,3), the incubation of ER, IN, ONM with cytosol enriched with sphingolipid extracts from Golgi vesicles, cell membrane rafts and caveolea, and apical cell membrane was not facilitating lipid integration into the membranes, and the lipids remained in cytosol (lanes 4,5). On the other hand, addition of lipid precursors ([3H]inositol, [3H]palmitate or [3H]choline) to CC produced membranes with newly synthesized integrated radiolabeled phospholipids and ceramides (lanes 7,8,9).
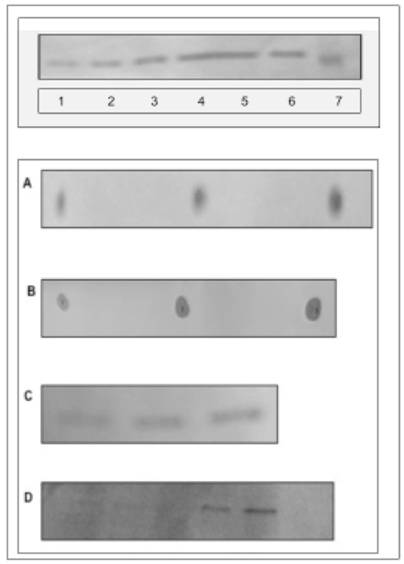
Requirements for biomembrane generation. Restitution of ER and nuclear membranes is accomplished by site-specific membrane biogenesis. and is not derived from retrograde transport of lipids, membrane rafts or caveolea from cell membrane. (A) Incubation of ER, IN, ONM with active CC enriched with ceramide-labeled sphingolipids isolated from Golgi vesicles (A-1, B-1), cell membrane rafts and caveolea (A-2, B-2) and cell apical membrane (A-3, B-3), has not produced the membranes labeled with sphingolipids. The lipid extracts from incubated ER, IN, ONM (A1-3), respectively were free of radiolabeled lipids provided in CC. The labeled sphingolipids from Golgi vesicles and membrane rafts (A4) and from apical membranes (A5) remained in the CC. The incubation of INM with nuclear contents enriched with radiolabeled precursors of membrane phospholipids consisting [3H] arachidonate and [3H]inositol has not generated radiolabeled INM (A-6). In contrast, the active CC enriched with radiolabeled arachidonate, and inositol generated radiolabeled membranes of IN (A-7), ONM (A-8), INM (A-9), ER (A-10) and ER transport vesicles (A-11). The lipid extracts from the membranes (IN, ONM, INM, ER, ER transport vesicles), from CC and nuclear contents were applied to HPTLC and scanned in Berthold Analyzer for 16 hours. The [3H]palmitate labeled ceramides and sphingolipids prepared from Golgi vesicles (A-1) cell membrane rafts and caveolea (A-2) and cell apical membrane (A-3) as depicted in B-1-3 were used. The amount of radiolabeled lipids added to CC corresponds to the one depicted in B. The lipid samples shown in A represents material applied to HPTLC and subjected to chromatography in the solvent system separating free fatty acids (ethyl ether/hexanes, 7:3, v/v), while sphingolipids, phosphosphingolipids and phosphoglycerides remain unseparated and close to the origin. The detection of radiolabeled lipid was performed in Berthold radioactivity analyzer for 16 h. (B) [3H]palmitate labeled sphingolipids corresponding to the amount used in incubation described in A following high performance thin layer chromatography (HPTLC) in chloroform/methanol/ water (65:35:8, v/v/v) mixture and scanning in Berthold analyzer for 16 h. The lanes correspond to sphingolipids extracted from preparations of 1-Golgi vesicles, 2-apical membrane rafts and caveolea, 3-apical membranes. (C) Synthesis of membrane lipids is affected by degradation of RNA in active CC. Synthesis of arachidonate and inositol-labeled lipids in IN, ONM, ER (C1-3) respectively, in the presence of RNase treated (C1-3), and IN, ONM in untreated (C-4, 5) CC. The scanning of the radiolabeled membranes was performed on membrane lipid extracts applied to HPTLC as described in A and scanned for 16 h in Berthold Analyzer. (D) The [3H]inositol labeled lipids of IN (D-1), ONM (D-2), INM (D-3) are more complex than those synthesized in ER (D-4). HPTLC of the lipid extracts was performed in chloroform/methanol/acetic acid/water, 65:35:8:4, by vol.) and scanned in Berthold radioactivity analyzer for 16 h.
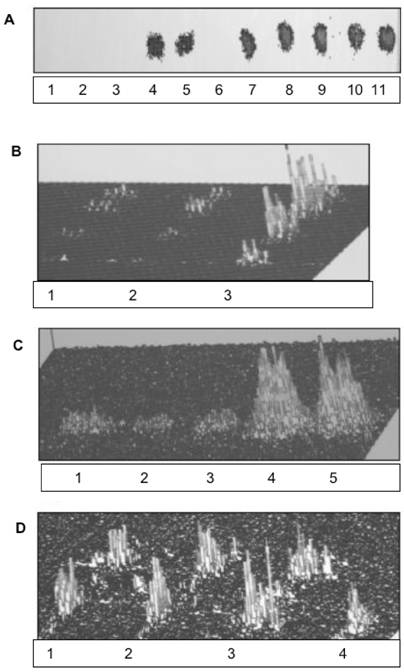

Based on an assumption that the impact of the extreme secretory activity must be also substantial and evident in the ER/nuclear membrane restitution, we investigated the impact of CC on ER/nuclear lipid synthesis and the potential to generate transport vesicles. The results of the studies on ER/nuclear membrane synthesis demonstrated that restitution of nuclear membranes is also associated with the ability of endoplasmic reticulum to synthesize membrane lipids and cellular protein (Fig. 1A, lane 10,11 and Fig. 1C). The RNase treated CC displayed minimal activity in biomembrane lipid synthesis (lanes 1,2,3), which was resumed in the presence of RNA-containing untreated CC (lanes 4,5). However, membrane lipids were not synthesized in the presence of nuclear contents (Fig. 1A, lane 6) and with or without RNA, the transport vesicles production by IN was not observed. Moreover, the lipid composition of ONM/ER was different than that of ER and ER transport vesicles Fig. 1D and Fig. 2).
Phosphatidylinositides, and phosphoglycerides comprised of phosphatidylcholine (PC) and phosphatidic acid generate ONM and INM of nuclear membranes. The two dimensional HPTLC was used to identify phospholipids of ONM (A), IN (B), INM (C), ER (D). The HPTLC was fist performed in solvent mixture consisting of chloroform/methanol/acetone/25% ammonium hydroxide 43:38:5:8, by vol. dried thoroughly to remove excess of moisture and ammonia and developed in second dimension in solvent system consisting of chloroform/methanol/acetic acid/water (65:35:8:4, by vol.). To identify different phospholipids, the HPTLC were subjected to iodine vapors, ninhydrin spray, phospholipids detecting spray and orcinol-carbohydrate-detecting spray. Based on the results of these experiments the lipids isolated from the membranes were identified as 1-phosphatidylinositol bisphosphate (PIP2), 1'-phosphatidylinositol monophosphate (PIP), 1''-phosphatidylinositol monophosphate (PIP), 2-phosphatidylcholine (PC), 3-phosphatidylcholine (PC), 4-phosphatidylinositol (PI), 5-phosphatidic acid (PA).
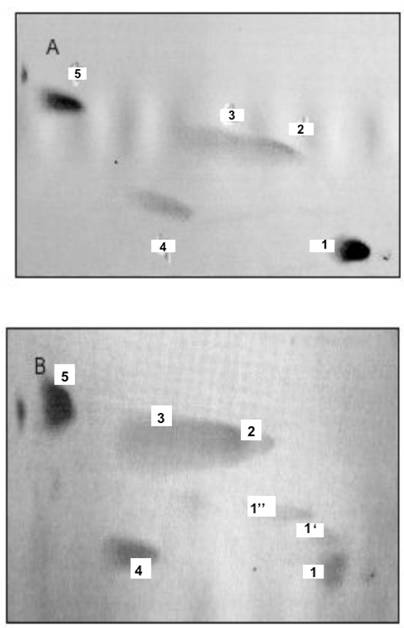

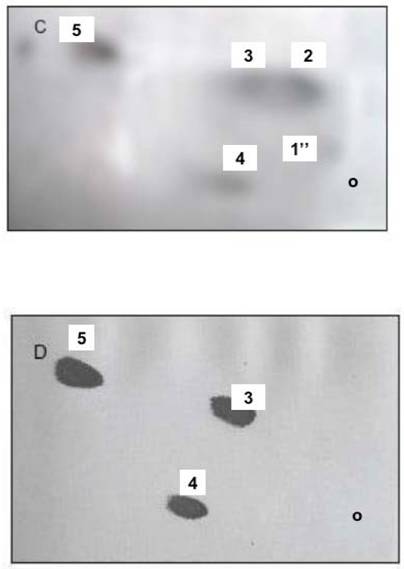
As shown in the figures, the incubation of ONM/ER with cytosol afforded synthesis of PC, PA, PI and PIPs, but the ceramides were not formed. In contrast, and as shown previously [30], the ER engaged in formation of ER transport vesicles was involved in synthesis of ceramides, and only phosphatidylinositol (PI) was representing phosphatidylinositides in their membrane (Fig. 2D). The ER/nuclear membrane- generated lipids consisted of PIPs, PI, PC and PA (Fig. 2 A-C) and all remained in the nuclear membrane. Hence, we rationalized that such restitution of ER membrane renewed ONM, compensated for the portions of ER membrane engaged in ceramides synthesis and exported in the form of vesicles delivered to Golgi. But, at the same time, the interpretation was not providing explanation for regeneration of physically distant INM. The presence of PIP2 on IN, its ONM and INM membrane could be explained if PIPs synthesis was supported by nuclear contents or ONM supplied new PIPs through lateral movement of the nuclear membrane. Such movement would allow the import of lipids and lipid- associated cytosolic protein to the nucleus.
To demonstrate whether membrane synthesis and transport to nucleus are concurrent with delivery of cytosolic protein to nucleus, the synthesis of ER/nuclear membrane was investigated with NIP-labeled cytosolic proteins. Here, we concentrated on transfer NIP-labeled 30kDa cytosolic protein with affinity for PIP2 (Fig. 3, and Fig. 4, top panel).
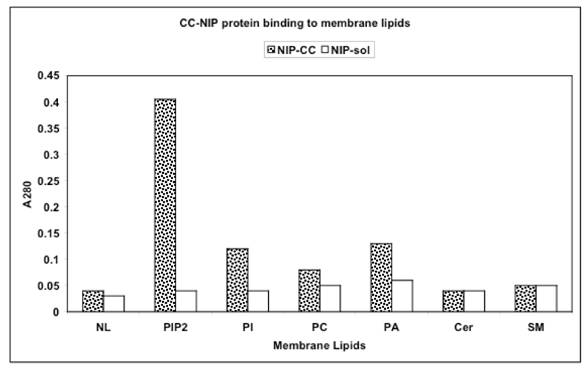
Interaction of NIP-labeled 30kDa protein of CC with lipids of IN, ONM, and INM represented by phosphatidylinositol bisphosphate (PIP2), phosphatidylinositol (PI), phosphatidylcholine (PC), phosphatidic acid (PA) and sphingolipids of ER-transport vesicles and Golgi represented by ceramides (Cer) and sphingomyelin (SM) and neutral lipids (NL) consisting of glycerides and cholesterol. Microtiter plates were coated with of with 0.5 to 20 μg lipids extracted from the IN, ONM, INM, ER transport vesicles, Golgi transport vesicles and cell apical membrane, reacted with 1.0-1.5 A280 NIP-labeled 30kDa cytosolic protein for 120 min, followed by incubation with CC proteins (1mg/ml). After incubation, the plates were rinsed with PBS containing 0.05% Tween and read at A280. Each reaction was performed in triplicate.
Identification of 30 kDa CC protein in IN, ONM, INM following incubation with NIP-labeled CC protein is depicted in the upper panel. The protein extracts from IN (lanes 1,4) ONM (lanes 2,5) and INM (lanes 3,6) incubated with NIP-CC protein for 30 min (lanes 1-3) and 60 min (lanes 4-6) and the purified 30 kDa NIP-protein (lane 7) were subjected to SDS-PAGE and western blot. The NIP labeled proteins retained on the membranes were identified with anti NIP IgG.
The IN incubated with NIP-labeled cytosol acquired 17.6mmoles of NIP-labeled cytosolic proteins on the 4.0-4.5 mg/ml IN (Fig. 4 A, B). After 30 min incubation in NIP-free cytosol, the IN-labeled protein decreased to 7.37 mmole/ml and to 4.46 mmole/ml after 120 min. Concurrently, the preparations of the INM and nuclear contents from NIP-labeled IN revealed the presence of NIP-labeled 30 kDa cytosolic protein in INM-bound and soluble form (Fig. 4 C, D). The incubation facilitated increase of NIP-hapten labeled protein on the INM (from 0.59 A430/2 A280 membranes to 0.95 after 30 min and to 1.15 after 120 min, and the appearance of labeled PIP2 in INM (Fig. 5). After 120 min incubation of the NIP-labeled IN in NIP-free cytosol, the INM-bound NIP-protein was mainly recovered from nuclear contents, the labeled phosphatidylinositides decreased in both membranes (Fig. 5 A, B) and PIP2 was practically undetectable in INM membrane, while the ONM retained, on the average, 0.18 A430 of NIP-coupled hapten per 2A280 of the protein, and contained labeled PI. Also, after 120 min IN incubation, the analysis of the material separating from IN membrane revealed fraction of membranes consisting of PC, PI and PA (Fig. 5D).
The NIP-labeled CC proteins association with PIP2 is demonstrated in panel (A), the NIP-labeled CC proteins at 5, 10, 20 μg/ dot are shown in (B), the increasing amount of NIP-labeled protein on INM, and in nuclear contents is shown in C and D, respectively.
The IN incubated for 30 min with NIP-CC were subjected to 60 and 120 min with unlabeled CC, followed by isolation of ONM, INM and nuclear contents. From each incubate the nuclear membranes and nuclear contents were separated by SDS-PAGE and the 30 kDa NIP-protein identified by western blotting. The NIP-labeled protein transfer from INM preparations is shown in C and the appearance of NIP-30 kDa protein in nuclear contents is demonstrated in (D). As evident, the NIP-30 kDa protein became detectable after 60-120 min incubation. The preparations of nuclear contents from shorter incubation periods contained undetectable concentration of NIP-30kDa protein.
Incubation of IN in active cytosol (CC) demonstrates movement of membrane lipids from ONM to INM and changes in phosphatidylinositides profile. During 60 min incubation with CC, the radiolabeled lipids decrease in the ONM and INM (A and B, respectively). The IN labeled with [3H]arachidonate and [3H]inositol were subjected to incubation with unlabeled CC, followed by inner and outer membrane separation and detection of the relative amount of radiolabeled lipid present in the ONM and INM of the IN after 30 and 60 min incubation. The level and the initial composition of the labeled phospholipids in ONM (Lane A) and INM (lane B) is depicted in panel C. Panel A demonstrates initial labeling of ONM (1), after 30 min (2) and 60 min (3) incubation of the IN. Panel B demonstrates the initial (1), 30 min (2) and 60 min (3) labeling of INM phospholipids. Following 60 min incubation of the labeled IN with CC, the IN were first subjected to the sucrose gradient purification and then to isolation of the ONM and INM and lipid extraction depicted in panels A and B. The lipids recovered from sucrose gradient-separated material from IN is depicted in panel D. The lipid extracts from the separated membranes were applied to HPTLC and subjected to two dimensional thin layer chromatography in solvent system consisting of chloroform/methanol/ammonia, 80:35:5, v/v/v, and after thorough drying to second dimension chromatography in chloroform/acetone/methanol/acetic acid/ water 30:40;10:10:5, by vol. The plates were dried and subjected to 16-h counting in Berthold Radioactivity Analyzer, to phospholipids detection, carbohydrate detection, ninhydrin-positive phospholipids detection and sulfuric acid charring. The combination of the screening allowed us to identify individual lipid of the membranes shown in panel D that consisted of phosphatidylcholine (PC) phosphatidylinositol (PI) and phosphatidic acid (PA).
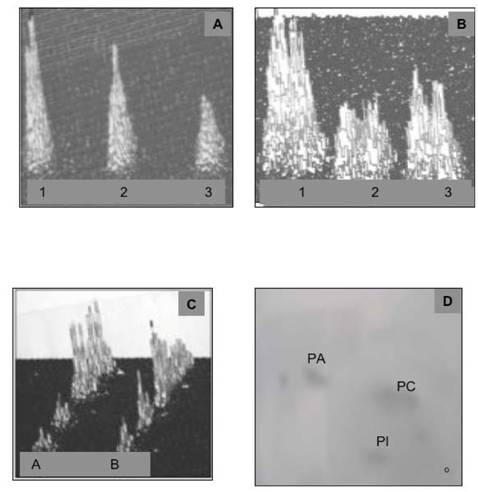

In our opinion, the separated membranes from IN subjected to prolonged incubation with CC represent the ER scaffold that reentered cytosol and is used for ceramides synthesis and mRNA translation.
Based on the obtained results, we formed hypothesis that the region of ER representing ONM is involved in the synthesis of membrane that generates outer and inner nuclear membrane. The enlargement of the outer area of the nuclear membrane, causes lateral movement of the nuclear membrane in-between nuclear pores and introduces the membrane with newly synthesized lipids to INM, which include PIPs (Fig. 2B) that import cytosol-derived protein into the nucleus. At the same time, continuation of the lateral movement out of the nucleus contributes to the membrane re-emergence from nucleus, transport of nuclear products to cytosol and, restitution of PI, PC, and PA-enriched ER membrane (Fig. 5D). Such movement would explain the presence of radiolabeled PIP2 lipids in the INM obtained from incubation of the IN with CC (Fig. 2A,B lack of lipid synthesis on INM facing nuclear contents, and appearance of the membranes that separate from purified and incubated IN that contain PI, PC and PA, the lipids which perhaps contribute to the restitution of ER membranes distant from ER/nuclear envelope.
Based on the lack of incorporation of NIP-labeled apical membranes and NIP-labeled caveolea into IN and ER (not shown since the NIP-labeled apical membrane proteins could not be detected in ER and nuclear membranes), we conclude that retrograde transport is neither contributing to the restitution of the ER/nuclear membrane nor to the transport of the phosphatidylinositides and cell cytosol protein to the nucleus. The incubation (up to 120 min) of the surface labeled hepatocytes with NIP has not introduced any NIP-labeled protein to either IN, ER or Golgi membrane. If any, the retrograde transport of biomembrane was not occurring in the detectable quantity, or the quantity that could restore the lipids and membranes used for constitutive secretory processes. The phosphatidylinositides detected in the nuclear envelope membranes were not derived from the pool of cell membrane, but were synthesized directly in the outer membrane of the nucleus. Their composition in INM phosphatidylinositides apparently is changing with PIPs hapten-associated protein, as the appearance of free NIP-30 protein in nuclear contents coincided with diminishing and changing PIPs profile in the INM (Fig.2C, Fig.4 C, D, Fig. 5).
Together, our studies suggest divergent and synchronous processes that restore cellular membranes and cell organelles. The CC impacts in synchrony the vesicular transport-assisted cellular and intracellular membranes synthesis restoring Golgi, endosomes and cell membrane, and the nuclear transport connected to nuclear membrane restitution, and ER membrane replacement. The divergent processes initiated in ER are concurrently impacted by the signaling clues deposited in the cell cytosol. And thus, the constitutive delivery of protein to the nucleus mediated through the synthesis of ER/ONM provides the link between genomic inherited design set in motion to restore cell components in homeostatic mode, as well as it may respond to signal-induced pathological cell differentiation in disease.
4. Discussion
In response to myriad of stimuli, the receptors expressed by the cell regulate cellular signaling pathways impacting cell metabolism, secretory properties, electrical activity, shape, motility, and virtually all functions within the realm of genome [1,30,31,35]. Several operative mechanisms to accomplish this goal have been delineated, and the concept that transmission of extracellular signals to the interior of the cell is one of the most fundamental processes of cell membrane receptors is used without reproach. Yet, within this global concept, there are still numerous factors to investigate and provide clear depiction of the cellular elements that contribute to appearance of the final cell and its function. One of them is the cell membrane lipid network, the membrane lipid biogenesis, and the fidelity of the processes charged with membrane restitution. It is not explored how the receptor proteins are delivered to cell membrane and incessantly provided with the same elemental environment, how the transmembrane architecture of the receptors is ceaselessly reproduced, or how the movement of the cytosolic proteins to nucleus is achieved [6,7,19,22,32,36,37]. Today, almost entire emphasis in cellular signaling is placed on the functional roles of RNAs in cytosol [10,28,38,39], and mutation-induced conformational changes in protein structure and function, simulated in most rudimentary lipid environment provided by the arbitrary cocktails of extraneous lipid mixtures or localized with aid of colored labels.
In our investigations we concentrated on the role of specific lipids in cellular membrane biogenesis that accompany the signaling processes, materialized in transcription, translation, posttranslational modification and homeostatic restitution of the cell [13-18]. Initially, we established that lipid biogenesis in ER is the elemental process initiating the intracellular membranes and vesicular transport from ER to Golgi [13-15]. The ER vesicles, consisting of newly synthesized lipids and proteins, transfer the synthesized cargo of proteins and newly assembled membranes from ER to Golgi. Further maturation processes of the delivered membrane, and the subcellular specificity of the membrane occurs in Golgi. The Cer of the membrane is used to synthesize sphingomyelin and glycosphingolipids, whereas PI is phosphorylated to PI3P and PI4P [14-18]. The transport vesicles with secretory proteins and the membrane containing PI3P and glycosphingolipids, displayed affinity for and fused with apical membranes, while the vesicles containing PI4P in their membrane were delivered to endosomes [16,18].
The specific biomembrane-protein assembly, and the lipid synthesized in ER and specifically modified in Golgi were in the entirety introduced to cell membrane or intracellular organelle [13-18]. Moreover, the vesicles capable of fusing with apical membrane would not fuse with ER or endosomes [13,14,18]. Further, the transport vesicles that remained integral part of endosomes would not integrate into the apical cell membrane, and in the experiments with apical membranes were found as portion of unattached vesicles remaining in the cytosol [16-18]. In both events, as determined by lipid composition of the products, the transport vesicles by fusing with membranes, delivered en bloc lipids and membrane proteins. The analysis of lipid composition of the products confirmed the specificity and irreversibility of the processes that allow ER membranes be free of SM and GLL. In our opinion, these data generate a solid support for our concept that transport is highly lipid membrane-specific, irreversible and that the retrograde transport is not returning empty transport vesicles to ER [26], since neither apical membrane specific lipids nor NIP-labeled apical membrane and caveolea [29,40] were identified in cell organelles.
If in the apical vesicular transport, the vesicles were recovered by the way of retrograde transport, fused with ER and resumed another round of transport, the composition of intracellular membranes would be uniform throughout, including cell and ER membrane. This is certainly not the case, since each organelle displays its specific membrane make-up [13-18]. It may be argued that the intracellular organelle modification is associated with delivery of lipids from the cytosol [5,33]. However, as we have shown in our reports, the lipids suspended in the cytosol do not intercalate into specific membranes, but some in a fashion similar to transport vesicles [16-18] may merely associate with membrane proteins. For instance, the addition of Cer to cytosol does not stimulate production of transport vesicles, yields more vesicles or vesicles with higher concentration of Cer [16,18]. With the use of some artificial lipid mixtures that encase protein receptors, some retention of the complex on the membrane may occur, but certainly not the genuine insertion or multiple intercalations into the membrane is taking place.
The process of membrane renewal is coordinated with protein and biomembrane synthesis in ER [18]. Only if both processes coincide, the specific intercalation of the protein is feasible. In keeping with our hypothesis that specific lipids and proteins are synthesized at designated sites of ER are the findings on cotranslational assembly of a protein complexes encoded by a family of colocalized mRNAs [9,10]. Local recruitment of mRNAs to coordinate synthesis of the complexes contributing to overall cell motility by supplying new protein (new actin monomers precisely at the sites where they are needed) is an example of the location-specific translation [39]. Although data on cotranslational assembly of a protein complexes encoded by a family of colocalized mRNAs are derived from completely unrelated to membrane biogenesis experimental paradigm [10], it firmly attests to the significance of the assembly of protein complexes at the specific sites of ER.
While the vesicular transport is subject of many reviews and highly imaginative interpretations are offered to explain secretory transport processes, the issue of the transport to nucleus remains ambiguous [1,20,23,27-31]. The question still is how the cytosol protein can be transported onto the inner nuclear membrane surface [1-5,11,12, 28,35,38]. In recent studies on nuclear receptors, the structural analyses revealed phosphatidylinositides as ligands for the NR5 orphan receptors and showed that PIP phospholipids are involved in the receptor transport [6,21,38]. While elegant structural paradigms present the activation of lipid ligands-regulated receptors, the conformation of the interacting molecules is deduced from studies with artificial liposomes [2].
Our study presented in this report addresses similar issue of protein delivery from cytosol to the nucleus, and show that ER membrane directly appose to nuclear membrane is involved. The results demonstrate that nuclear membrane synthesized by ER-initiated synthesis of new membrane lipids and the enlargement of the outer layer of the nuclear membrane must generate lateral movement of the membrane between nuclear pores. Such movement not only allows restitution of the nuclear inner membrane but also imports the lipid ligand-associated proteins from the cytosol to nucleus, and perhaps transfers the nuclear components to the cytosol. The fact that nuclear envelope consists of two layers of uninterrupted membrane connected with ER, provides most sensible explanation that lateral movement of the membrane is the most feasible process that connects nuclear, ER, and cytosolic events [11].
Interestingly, when the ER membranes, other than outer nuclear membrane were involved in the synthesis of lipids and generation of transport vesicles, the incubation of the ONM/ER with cytosol generated PC, PI, PIPs and PA but transport vesicles were not formed. The lipids were synthesized when the intact nuclei were incubated with the cytosol but not in the presence of the inner nuclear contents. On the other hand, the experiments performed with INM and ONM derived from intact nuclei but with cytosol afforded synthesis of new lipids in both membranes. The intriguing finding that INM is enriched with new lipids when incubated with the cytosol is in agreement with histochemical studies and the in vitro data suggesting the functional relationship between phospholipids and gene expression/transcription [19]. Yet, the existence of intranuclear synthesis of phospholipids is contradicted by the results of our experiments demonstrating that nuclear contents from purified nuclei is not capable to generate new phospholipids. Only, if the nuclear membrane movement is considered, our findings and those described above [19] can be reconciled by the interpretation that the appearance of new phospholipids in the INM is the consequence of ONM lateral movement-induced transfer. Such movement of the membrane would provide continuous transport of the lipid ligand-associated cytosolic protein into the nucleus, restitution of the nuclear INM phosphatidylinositides, and steady growth of ER membrane.
In keeping with the hypothesis is the fact that lipid ligand such as PIP2 interacts with cytosolic protein, but upon delivery to the nucleus the complex is dissociated. The NIP-30 initially in complex with PIP2 appears as free of lipid nuclear protein, whereas the PIP2 vanishes. At this stage we cannot provide evidence whether the PIPs degradation is taking place during release of the protein, or later before the membrane reenters cytosolic environment. The possibility exists that PIPs of INM are degraded by nuclear PLC and serve as a source of inositol phosphate in the nucleus. Recent study on RNA editing provide evidence that nuclear inositol phosphates are implicated in the formation of an active site of adenosine deaminase, and are important in the in vivo and in vitro deamination of adenosine [38].
The evidence that outer nuclear membrane/endoplasmic reticulum is involved in the synthesis of phospholipids and restitution of nuclear membranes, whereas the endoplasmic reticulum engaged in synthesis of transport vesicles containing ceramides is used for restitution of Golgi, endosomes and apical membranes, provides initial view of two highly dependent pathways responsible for reproduction of cell components and prevention of the cell demise. The dynamics of the pathways appear to control homeostatic restitution of the cell and its organelles, and all is linked by the cell endogenous cytosol. Hence, the reassembly of the specific units of the cell is determined by (1) the type of cell, (2) the protein induced, (3) the site of the protein function, and (4) the metabolic status of the cytosol. Therefore, the replacement of native cytosol with the one derived from different cells, may indeed impact cellular appearance, physiological functions, and the processes associated with transcription, mRNA translation and small RNAs functions [33,38,39].
As the transcriptome represents the transcripts of genes expressed in the cell and informs which genes are induced or repressed by the metabolic status and physical environment of such cell, the path between the signal translation into the gene transcription and the translation of the transcripts into cellular components was not spatially discerned heretofore. Our studies show that the intracellular and cellular membranes' biogenesis links the signaling events into a path that uniformly restores the cell structure and function. The signals from cell membrane through the cytosol link engage simultaneously the nuclear and organellar events [13-18]. Together, and in perfect synchrony, the ER-initiated processes contribute to the restitution of cell components, retention of the precisely controlled cell structure, and the designated function.
Conflict of interests
Declared none.
References
1. Schlessinger J. Common and distinct elements in cellular signaling via EGF and FGF receptors. Science. 2004 ;306:1506-1507
2. Loewen CJR, Gaspar ML, Jesch S.A, Delon C, Ktistakis NT, Henry SAM, Levine TP. Phospholipid metabolism regulated by a transcription factor sensing phosphatidic acid. Science. 2004 ;304:1644-1647
3. Vance JE, Vance DE. Metabolic insights into phospholipids function using gene-targeted mice. J Biol Chem. 2005 ;280:10877-10880
4. Shen X, Xiao H, Ranallo R, Wu W-H, Wu C. Modulation of ATP-dependent chromatin-remodeling complexes by inositol polyphosphates. Science. 2003 ;299:112-114
5. Czech MP. Dynamics of phosphoinositides in membrane retrieval and insertion. Annu Rev Physiol. 2003 ;65:791-815
6. Gozani O, Karuman P, Jones DR, Ivanov D, Cha J. The PHD finger of the chromatin-associated protein ING2 functions as a nuclear phosphoinositide receptor. Cell. 2003 ;11:99-111
7. Traub LM. Common principles in clathrin mediated sorting at the Golgi and the plasma membrane. Biochim Biophys Acta. 2005 ;1744:415-437
8. Rodriguez-Boulan E, Musch A. Protein sorting in the Golgi complex: Shifting paradigms. Biochim Biophys Acta. 2005 ;1744:455-464
9. Mukherjee S, Maxfield FR. Membrane domains. Annu Rev Cell Dev Biol. 2004 ;20:839-866
10. Moore MJ. From birth to death: The complex lives of eukaryotic mRNAs. Science. 2005 ;309:1514-1518
11. Ellenberg J, Siggia ED, Moreira JE, Smith CL, Presley JF, Worman HJ, Lippincott-Schwatz J. Nuclear membrane dynamics and reassembly in living cells: Targeting of inner nuclear membrane protein in interphase and mitosis. J Cell Biol. 1997 ;138:1193-1206
12. Steger GL, Haswell ES, Miller AL, Wente SR, O'Shea EK. Regulation of Chromatin remodeling by inositol polyphosphates. Science. 2003 ;299:114-116
13. Slomiany A, Grzelinska E, Kasinathan C, Yamaki K, Palecz D, Slomiany BL. Biogenesis of endoplasmic reticulum transport vesicles transferring gastric apomucin from ER to Golgi. Exp Cell Res. 1992 ;201:1669-1682
14. Slomiany A, Grzelinska E, Grabska M, Yamaki K, Tamura S, Slomiany BL. Intracellular processes associated with glycoprotein transport and processing. Arch Biochem Biophys. 1992 ;298:167-175
15. Slomiany A, Grabska M, Piotrowski E, Morita M, Slomiany BL. Intracellular processes associated with vesicular transport from endoplasmic reticulum to Golgi and exocytosis; Ethanol-induced changes in membrane biogenesis. Arch Biochem Biophys. 1994 ;310:247-255
16. Slomiany A, Nowak P, Piotrowski E, Slomiany BL. Effect of ethanol on intracellular vesicular transport from Golgi to apical cell membrane: Role of phosphatidylinositol 3-kinase and phospholipase A2 in Golgi vesicles association and fusion with the apical membrane. Alcoholism Clin Exp Res. 1998 ;22:167-175
17. Slomiany A, Slomiany BL. Lipidomic processes in homeostatic and LPS- modified cell renewal cycle. Role of phosphatidylinositol 3-kinase pathway in biomembrane synthesis and restitution of apical epithelial membrane. J Physiol Pharmacol. 2003 ;54:533-551
18. Slomiany A, Sano S, Grabska M, Yamaki K, Slomiany BL. Gastric mucosal cell homeostatic physiome. Critical role of ER-initiated membrane restitution in the fidelity of cell function renewal. J Physiol Pharmacol. 2004 ;55:837-860
19. Jackowski S, Fagone P. CDP: Phosphocholine cytidylyl transferase: Paving way from gene to membrane. J Biol Chem. 2005 ;280:853-856
20. Miller EA, Beilharz TH, Malcus PN, Lee MCS, Hamamoto S, Orci L, Schekman R. Multiple cargo binding sites on the CopII subunit Sec 24p ensure capture of diverse membrane proteins into transport vesicles. Cell. 2003 ;114:497-509
21. Gilmour RS, Mitchell MD. Nuclear lipid signaling: novel role of eicosanoids. Exp Biol Med. 2001 ;226:1-4
22. Ciufo LF, Barclay JW, Burgoyne RD, Morgan A. Munc18-1 regulates early and late stages of exocytosis via synthaxin-independent protein interactions. Mol Biol Cell. 2005 ;16:470-482
23. Mossessova E, Bickford LC, Goldberg J. SNARE selectivity of the COP II coat. Cell. 2003 ;114:483-495
24. John R, Lang T, Sudhof TC. Membrane fusion. Cell. 2003 ;112:519-533
25. Bonifacino JS, Glick BS. The mechanisms of vesicle budding and fusion. Cell. 2004 ;116:153-166
26. Lee MCS, Miller EA, Goldberg J, Orci L, Schekman R. Bi-directional protein transport between ER and Golgi. Annu Rev Cell Dev Biol. 2004 ;20:87-123
27. Gerst JE. SNARE regulators: matchmakers and matchbreakers. Biochim Biophys Acta. 2003 ;1641:99-110
28. Humbert JP, Matter N, Artault JC, Koppler P, Malvija AN. Inositol 1,4,5-trisphosphate receptor is located to the inner nuclear membrane vindicating regulation of nuclear calcium signaling by inositol 1,4,5-trisphosphate. J Biol Chem. 1996 ;271:478-485
29. Pike LJ. Lipid rafts: bringing order to chaos. J Lipid Res. 2003 ;44:655-667
30. Gorlich D, Kutay V. Transport between the cell nucleus and the cytoplasm. Annu Rev Cell Dev Biol. 1999 ;15:607-670
31. Roth MG. Phosphoinositides in constitutive membrane traffic. Physiol Rev. 2004 ;83:699-730
32. De Matteis MA, Di Campli A, Godi A. The role of the phosphoinositides at the Golgi complex. Biochim Biophys Acta. 2005 ;1744:396-405
33. Kolter T, Winau F, Schaible UE, Leippe M, Sandhoff K. Lipid binding proteins in membrane digestion, antigen presentation, and antimicrobial defense. J Biol Chem. 2005 ;280:41125-41128
34. Kooijman EE, Chupin V, de Kruijff B, Burger KN. Modulation of membrane curvature by phosphatidic acid and lysophosphatidic acid. Traffic. 2003 ;4:162-174
35. Steger DG, Haswell ES, Miller AL, Wente SR, O'Shea EK. Regulation of chromatin remodeling by inositol polyphosphates. Science. 2003 ;299:114-116
36. Izaurralde E, Adam S. Transport of macromolecules between nucleus and cytoplasm. RNA. 1998 ;4:351-364
37. Byrne RD, Barona TM, Garnier G, Koster G, Katan M, Poocia DL, Larijani B. Nuclear envelope assembly is promoted by phosphoinositide-specific phospholipase C with selective recruitment of phosphatidylinositol-enriched membranes. Biochem J. 2005 ;387:393-400
38. Macbeth MR, Schubert HL, VanDemark AP, Lingam AT, Hill CP, Bass BL. Inositol hexakisphosphate is bound in ADAR2 core and required for RNA editing. Science. 2005 ;310:1534-1539
39. Karp X, Ambros V. Encountering microRNAs in cell fate signaling. Science. 2005 ;310:1288-1289
40. Ostrom RS, Insel PA. The evolving role of lipid rafts and caveolae in G protein coupled receptor signaling: implications for molecular pharmacology. Brit J Pharmacol. 2004 ;143:235-245
Author contact
![]() Correspondence to: Amalia Slomiany, Research Center C-873, University of Medicine and Dentistry of New Jersey, New Jersey Dental School, 110 Bergen Street, Newark, NJ 07103. Phone: 973-972-7023, Fax: 973-972-7020, E-mail: slomiaamedu
Correspondence to: Amalia Slomiany, Research Center C-873, University of Medicine and Dentistry of New Jersey, New Jersey Dental School, 110 Bergen Street, Newark, NJ 07103. Phone: 973-972-7023, Fax: 973-972-7020, E-mail: slomiaamedu

 Global reach, higher impact
Global reach, higher impact