Impact Factor ISSN: 1449-2288
Int J Biol Sci 2008; 4(2):81-86. doi:10.7150/ijbs.4.81 This issue Cite
Research Paper
Interactions between SNP Alleles at Multiple Loci Contribute to Skin Color Differences between Caucasoid and Mongoloid Subjects
1. Shibaura Institute of Technology, 3-7-5 Toyosu, Koto-ku, Tokyo 135-8548 Japan
2. Nagahama Institute of Bio-Science and Technology, 1266 Tamura-cho, Nagahama-shi, Shiga 526-0829 Japan
3. Shimadzu Corporation, 1 Nishinokyo Kuwahara-cho, Chukyo-ku, Kyoto-shi, Kyoto 604-8511 Japan
Received 2008-1-14; Accepted 2008-3-31; Published 2008-3-31
Abstract
This study aimed to identify single nucleotide polymorphism (SNP) alleles at multiple loci associated with racial differences in skin color using SNP genotyping. A total of 122 Caucasians in Toledo, Ohio and 100 Mongoloids in Japan were genotyped for 20 SNPs in 7 candidate genes, encoding the Agouti signaling protein (ASIP), tyrosinase-related protein 1 (TYRP1), tyrosinase (TYR), melanocortin 1 receptor (MC1R), oculocutaneous albinism II (OCA2), microphthalmia-associated transcription factor (MITF), and myosin VA (MYO5A). Data were used to analyze associations between the 20 SNP alleles using linkage disequilibrium (LD). Combinations of SNP alleles were jointly tested under LD for associations with racial groups by performing a χ2 test for independence. Results showed that SNP alleles at multiple loci can be considered the haplotype that contributes to significant differences between the two population groups and suggest a high probability of LD. Confirmation of these findings requires further study with other ethnic groups to analyze the associations between SNP alleles at multiple loci and skin color variation among races.
Keywords: SNP, Polygene, Haplotype, Linkage disequilibrium
1. Introduction
The evolution of new traits has allowed humans to adapt to complicated and challenging environmental conditions. Human skin color variations, which are regulated by the expression of melanin, represent an environmental adaptation to different levels of exposure to ultraviolet (UV) rays. For example, people indigenous to Northern Europe have pale skin, while people indigenous to Africa have dark skin. At low latitudes, melanin production is increased to protect against continual exposure to UV irradiation. At high latitudes, melanin production is decreased, which increases the body's ability to synthesize vitamin D, offering a variety of health benefits, including protection against rickets (osteomalacia) [1-3].
Although variations in human skin color are associated with environmental factors such as levels of UV radiation, the genetic mechanism underlying the regulation of human skin color is still unclear. Several studies of skin pigmentation have reported polymorphisms in genes associated with skin color, such as the melanocortin 1 receptor (MC1R), oculocutaneous albinism II (OCA2), and agouti signaling protein (ASIP). Nevertheless, human skin color variation is thought to likely be controlled by interactions among multiple loci at alleles known to contain single nucleotide polymorphisms (SNPs) [4-11]. This study was undertaken to clarify the molecular basis for the genetic background of human skin color by investigating how interactions between SNP alleles at multiple loci contribute to variations in skin pigmentation.
2. Material and methods
This study compared Caucasoid and Mongoloid cohorts of different skin colors. The genetic data obtained from 122 Caucasian subjects from a previous study [1] was used for analysis in this report. Genetic data was also collected from 100 Mongoloid subjects living in Japan, and the sample collection and melanin measurements were conducted in accordance with a protocol approved by the Human Subjects Research Committee of Shibaura Institute of Technology, Japan (Research project #06-002).
Subjects provided informed consent for the collection of buccal samples. Samples were anonymously coded, and the melanin skin pigmentation index was determined using the Mobile Mexameter MSC100/MX18 (Integral Corporation, Shinjuku-ku, Tokyo, Japan), which uses photodiode arrays to measure the intensity of particular wavelengths of light. Two measurements were obtained from the back as an index of inherent skin color influenced by genetic factors, and two measurements were obtained from the cheek as an index of modified skin color influenced by environmental factors.
DNA was extracted from the buccal samples using the DNA extraction kit, ISOHAIR (NIPPON GENE Co., Ltd., Chiyoda-ku, Tokyo, Japan). To ensure sufficient genomic DNA for SNP genotyping, whole genomic DNA was amplified using the REPLI-g Kit (QIAGEN K. K., Chuo-ku, Tokyo, Japan). Candidate genes for human skin pigmentation were identified [4-16] and included the Agouti signaling protein (ASIP), tyrosinase-related protein 1 (TYRP1), tyrosinase (TYR), melanocortin 1 receptor (MC1R), oculocutaneous albinism II (OCA2), microphthalmia-associated transcription factor (MITF), and myosin VA (MYO5A). Twenty SNPs that had been registered for the database dbSNP [17] were selected from loci in the candidate genes: rs819136, rs1129414, rs2075508, rs10960756, rs3793976, rs2298458, rs3212363, rs1805008, rs3212371, rs2279727, rs4778182, rs1800419, rs2311843, rs1800414, rs1800404, rs7623610, rs704246, rs16964944, rs1724577, and rs4776053. Polymerase chain reaction (PCR) of the DNA samples was performed to amplify the regions containing the SNPs of interest, and the products were subjected to 2% agarose gel electrophoresis to verify the generation of a single-band product. The verified PCR products were purified with ExoSAP-IT (Amersham Pharmacia Biotech, USA).
The allele discrimination assay consisted of PCR amplification of multiple SNP alleles at a particular locus using specific primers with tags differing in molecular weight. For this assay, the purified PCR products were combined with two hemi-nested allele-specific primers and two universally tagged Masscode oligonucleotide primers. Each tag was covalently attached to the 5' end of an oligonucleotide primer via a photolabile linker. Following PCR amplification, the SNP-specific PCR products were passed through a QIAquick 96 silica-based filter membrane to remove unincorporated tagged primers. The filtered PCR products were exposed to a 254-nm mercury lamp to cleave the incorporated tags, and the tags were analyzed using an optimized Agilent 1100 single quadrupole mass spectrometer. The presence of a particular tag indicated the presence of the corresponding SNP allele in the genomic DNA sample. Genotype data were reported in a comma-delimited flat-file format that contained the SNP and sample identifiers for each allele detected. Alleles were reported using binary nomenclature, in which l represented wild-type alleles and 2 represented variant alleles. The SNP allele was classified into three types: wild-type homozygous, variant-type homozygous, and heterozygous. Thus, a homozygous wild-type allele was designated as 1,1 and a heterozygous allele was designated as 1,2 [18-19].
With the data obtained from the SNP genotyping of Caucasoid subjects from the previous study [1] and the Mongoloid subjects in this study, we conducted the following analyses. 1) We determined the genotype and allele frequencies of the 20 SNPs observed in the Caucasoid (n=122) and Mongoloid (n=100) subjects to determine racial differences in distribution. 2) We conducted cluster analysis for genetic differentiation of the SNP genotyping results by condensing the genotype assignment for each SNP allele into a single numeric value as follows: homozygous wild type 1,1=0, heterozygous 1,2=0.5, and homozygous variant 2,2=1. An UPGMA dendrogram was constructed based on the genotyping data using the euclidean distance. 3) For each melanin index measurement taken from a subject's back, we calculated the mean value of melanin ± 2σ and defined two groups in the Mongoloid cohort based on constitutive skin color (low and high melanin groups). To examine the contribution of non-random associations of SNP alleles at multiple loci to skin color variations (i.e. low/high melanin content), we examined the associations among the 20 SNP alleles at various loci in the genome (in candidate genes). Linkage disequilibrium (LD) served as a measure of gene-gene interaction between unlinked loci [20]. The p value of LD was determined with a χ2 test, and statistical significance was set at 0.05. Combinations of SNP alleles at multiple loci under LD were jointly tested for association with low/high melanin by performing a χ2 test for independence. 4) To examine the contribution of non-random associations of SNP alleles at multiple loci to racial differences, we examined the associations of the 20 SNP alleles at various loci in the genome (in candidate genes) by LD. The p value of LD was determined with a χ2 test, and statistical significance was set at 0.05. Combinations of SNP alleles at multiple loci under LD were jointly tested for association with Caucasoid/Mongoloid race by performing a χ2 test for independence. Only data that fit the Hardy-Weinberg equilibrium were used in the analysis.
3. Results
Table 1 shows the genotype and allele frequencies for the 20 SNPs in the Caucasoid and Mongoloid groups speculated to contribute to differences in skin color between races. Cluster analysis showed that each racial group formed a separate cluster, except for one Mongoloid subject that was included in the Caucasoid cluster (Figures 1 and 2).
Genotypes and allele frequencies for 20 SNPs in Caucasoid and Mongoloid populations.
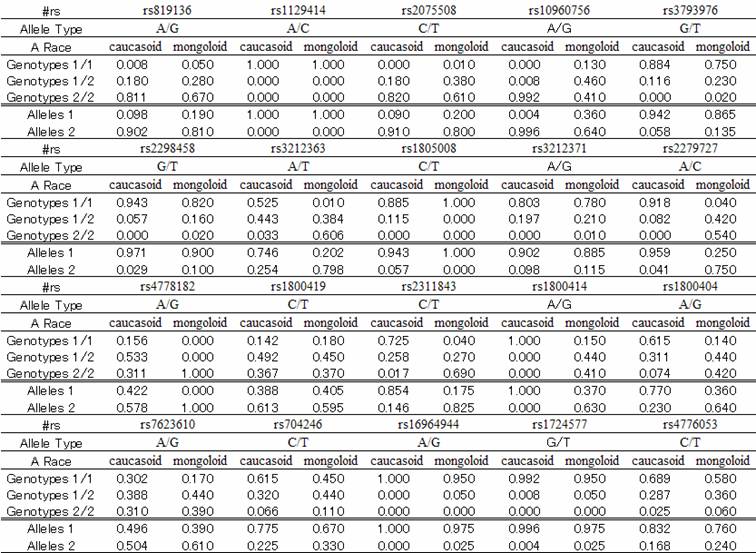
Dendrogram obtained from cluster analysis. Racial groups are separated by a red line. Left: Mongoloid subjects; right: Caucasoid subjects.
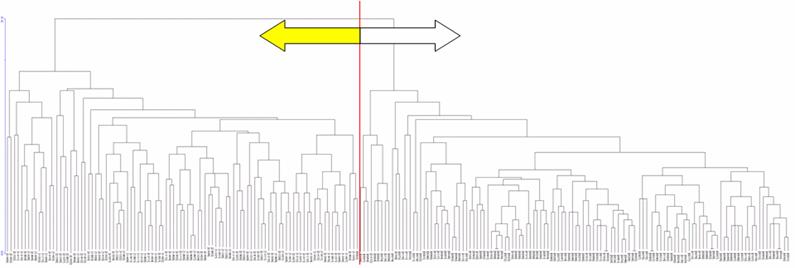
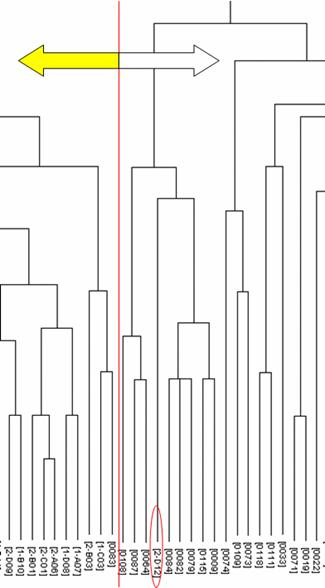
An enlarged figure of part of the dendrogram. Racial groups are separated by a red line except for one Mongoloid subject circled in red. Left: Mongoloid subjects; right: Caucasoid subjects.
The mean values of melanin for the back and the cheek in the Mongoloid group were 148.9 and 184.7, respectively, and the mean values of melanin for the back and the cheek in the Caucasoid group from the previous study were 115.6 and 155.6, respectively. The results obtained from analysis by linkage disequilibrium (LD) from the previous study showed that gene-gene interaction between SNP alleles at multiple loci contributed to skin color variation (i.e. low/high melanin content) in the Caucasoid cohort. However, it was not possible to analyze the two groups (low and high melanin groups) in the Mongoloid cohort by LD due to the low sample numbers obtained for each group.
The allele combination rs1800419-C/rs1800414-G/rs1800404-G was associated with the Mongoloid group (p = 5.39 x 10-20). These alleles are found in the gene for oculocutaneous albinism II on chromosome 15, and formed a haplotype. The protein encoded by this gene controls the transport of tyrosine, a precursor of melanin, into the melanosome for melanin synthesis.
The allele combination rs2311843-C/rs1800404-A/rs4776053-C was associated with the Caucasoid group (p = 5.51 x 10-33). These alleles are found in the genes for oculocutaneous albinism II and myosin VA on chromosome 15, and formed a haplotype. Myosin VA functions in vesicle transport, and mutations in this gene confer a lighter skin color due to defects in actin-based pigment granule transport within melanocytes; the pigmentation variation is believed to be due to abnormal distribution of melanosomes along the dendrites of melanocytes [21].
There were significant differences in the allele combinations (i.e., haplotypes) between the two racial groups. The allele combination rs2311843-C/rs1800404-A/rs4776053-C was associated with the Caucasoid group. The rs4776053 allele was in the myosin VA gene found only in the Caucasoid group, and the other alleles of the combination, rs2311843 and rs1800404, were both in the oculocutaneous albinism II gene, found in both groups. Thus, the rs4776053 allele that was only associated with the Caucasoid group could be considered to confer lighter skin color. The rs4776053 allele in the myosin VA gene exhibits a C/T variation, and the allele frequencies for this genotype were 83.2/16.8 (C/T) for Caucasoid and 76.0/24.0 for Mongoloid groups. The high frequency of the C variant in the Caucasoid group suggested that rs4776053-C could be a SNP allele that confers lighter skin color. This result suggests that the lighter skin pigmentation observed in Caucasoid populations is the result of positive selection on different loci in different human populations [4].
4. Discussion
As previously mentioned, it was not possible to analyze the two groups (low and high melanin groups) in the Mongoloid cohort by linkage disequilibrium (LD) due to the low sample numbers obtained for each group. On the other hand, the results of the LD analysis of the racial groups (Caucasoid and Mongoloid groups) showed that SNP alleles at multiple loci that contributed to racial differences were on the same chromosome and are likely to form haplotypes.
Every gene has variable SNPs that may constitute haplotypes; some haplotypes are present in all populations, and some are population-specific [22]. The haplotypes associated with skin color differences between the Caucasoid and Mongoloid races are population-specific haplotypes. For most genes, haplotypes represent an opportunity for functional adaptation and diversification [22]. Consistent with this idea, the haplotypes identified in this study are the result of adaptation to different UV ray intensities. This study adds to the growing evidence of genetic variability in different geographically isolated populations with regard to skin color.
In spite of recent advances in gene identification and functional analysis, much work remains to elucidate the underlying genetic mechanisms regulating normal human skin pigmentation [6]. Our previous study was undertaken to discover the genetic basis of the substantial differences in melanin content between groups with low and high melanin among 122 Caucasoid subjects. Our results demonstrated that gene-gene interactions between SNP alleles at multiple loci contributed to inter-individual skin color variation (i.e. low/high melanin content) while suggesting a high probability of linkage disequilibrium (LD) [1]. This study aimed to discover the genetic basis of the physiological trait differences in skin color between Caucasoid and Mongoloid subjects. Our results showed that SNP alleles at multiple loci that contributed to racial differences in skin color were statistically dependent, indicating a high probability of LD structure. Confirmation of these findings requires further study with other ethnic groups, such as with a Negroid population, to analyze the associations between SNP alleles at multiple loci and racial differences in skin color. Clarifying these associations not only unravels the mechanisms regulating an interesting physiological trait, but also provides a model or test system for gene discovery in other polygenic traits (like complex diseases) that exhibit greater environmental sources of variation [6]. There is a clear justification for adopting an evolutionary approach in exploring human pigmentation genetics. These have already revealed a trait with a highly dynamic and complex evolutionary past and pointed to the molecular mechanisms underlying phenotypic variability [6], the most recently identified by the signature of selection in the functionally important SLC24A5 and POMC genes [23-24]. The identification and analysis of additional genes involved in human skin pigmentation and the functional characterization of the allelic variants at the candidate loci presented here will help clarify the nature and extent of skin pigmentation adaptation in human populations [4].
Quantitative trait loci for a quantitative trait, such as skin pigmentation, will often manifest complex genotype-phenotype relationships, such as gene-gene interactions and pleiotropy [25-26]. Future studies await the development of analytical methods for identifying the variation in SNP allele combinations at multiple loci that determine the phenotypic variations in samples obtained from the back (an index of inherent skin color). Moreover, future studies using samples obtained from the cheek (an index of modified skin color) may clarify the diversity of the functional potential of melanin synthesis against UV rays.
Acknowledgements
This work is supported by the Ministry of Education, Culture, Sports, Science, and Technology grants.
Conflict of Interests
The authors have declared that they have no conflict of interest.
References
1. Anno S, Abe T, Sairyo K. et al. Interactions Between SNP Alleles at Multiple Loci and Variation in Skin Pigmentation in 122 Caucasians. Evol Bioinform Online. 2007;3:169-178
2. Rouzaud F, Kadekaro AL, Abdel-Malek ZA. et al. MC1R and the response of melanocytes to ultraviolet radiation. Mutat Res. 2005;571:133-152
3. Jablonski NG, Chaplin G. The evolution of human skin coloration. J Hum Evol. 2000;39:57-106
4. Myles S, Somel M, Tang K. et al. Identifying genes underlying skin pigmentation differences among human populations. Hum Genet. 2007;120:613-621
5. Izagirre N, Garcia I, Junquera C. et al. A scan for signatures of positive selection in candidate loci for skin pigmentation in humans. Mol Biol Evol. 2006;23(9):1697-1706
6. McEvoy B, Beleza S, Shriver MD. The genetic architecture of normal variation in human pigmentation: an evolutionary perspective and model. Hum Mol Genet. 2006;15(2):R176-181
7. Bonilla C, Boxill LA, Donald SA. et al. The 8818G allele of the agouti signaling protein (ASIP) gene is ancestral and is associated with darker skin color in African Americans. Hum Genet. 2005;116:402-406
8. Makova K, Norton H. Worldwide polymorphism at the MC1R locus and normal pigmentation variation in humans. Peptides. 2005;26:1901-1908
9. Naysmith L, Waterston K, Ha T. et al. Quantitative measures of the effect of the melanocortin 1 receptor on human pigmentary status. J Invest Dermatol. 2004;122(2):423-428
10. Ancans J, Flanagan N, Hoogduijn MJ. et al. P-locus is a target for the melanogenic effects of MC-1R signaling: a possible control point for facultative pigmentation. Ann N Y Acad Sci. 2003;994:373-377
11. Rees JL. Genetics of hair and skin color. Annu Rev Genet. 2003;37:67-90
12. Tadokoro T, Yamaguchi Y, Batzer J. et al. Mechanisms of Skin Tanning in Different Racial/Ethnic Groups in Response to Ultraviolet Radiation. J Invest Dermatol. 2005;124:1326-1332
13. Bonilla C, Shriver MD, Parra EJ. et al. Ancestral proportions and their association with skin pigmentation and bone mineral density in Puerto Rican women from New York city. Hum Genet. 2004;115:57-68
14. Shriver MD, Parra EJ, Dios S. et al. Skin pigmentation, biogeographical ancestry, and admixture mapping. Hum Genet. 2003;112:387-399
15. Hoggart CJ, Parra EJ, Shriver MD. et al. Control of confounding of genetic associations in stratified populations. Am J Hum Genet. 2003;72:1492-1504
16. Sturm RA, Teasdale RD, Box NF. Human pigmentation genes: identification, structure and consequences of polymorphic variation. Gene. 2001;277:49-62
17. Searchable NCBI site for Single Nucleotide Polymorphisms. National Library of Medicine. http://www.ncbi.nlm.nih.gov/projects/SNP/
18. Kokoris M, Dix K, Moynihan K. et al. High-throughput SNP genotyping with the Masscode system. Mol Diagn. 2000;5(4):329-340
19. Ogata K, Ikeda S, Ando E. QIAGEN Genomics Inc. A study of SNP genotyping using Masscode™ technology. Shimadzu Hyoka. 2002;58:125-129
20. Zhao J, Jin L, Xiong M. Test for interaction between two unlinked loci. Am J Hum Genet. 2006;79(5):831-845
21. Libby RT, Lillo C, Kitamoto J. et al. Myosin Va is required for normal photoreceptor synaptic activity. J Cell Sci. 2004;117:4509-4515
22. Stephens JC, Schneider JA, Tanguay DA. et al. Haplotype variation and linkage disequilibrium in 313 human genes. Science. 2001;293:489-493
23. Lamason RL, Mohideen MA, Mest JR. et al. SLC24A5, a putative cation exchanger, affects pigmentation in zebrafish and humans. Science. 2005;310:1782-1786
24. Millington GWM. Pro-opiomelanocortin (POMC): the cutaneous roles of its melanocortin products and receptors. Clin Exp Dermatol. 2006;31:407-412
25. Flanagan N, Healy E, Ray A. et al. Pleiotropic effects of the melanocortin 1 receptor (MC1R) gene on human pigmentation. Hum Mol Genet. 2000;9(17):2531-2537
26. Akey JM, Wang H, Xiong M. et al. Interaction between the melanocortin-1 receptor and P genes contributes to inter-individual variation in skin pigmentation phenotypes in a Tibetan population. Hum Genet. 2001;108(6):516-520
Author contact
![]() Correspondence to: Sumiko Anno, Tel: +81-3-5859-8363; Fax: +81-3-5859-8401; E-mail: annoushibaura-it.ac.jp
Correspondence to: Sumiko Anno, Tel: +81-3-5859-8363; Fax: +81-3-5859-8401; E-mail: annoushibaura-it.ac.jp

 Global reach, higher impact
Global reach, higher impact