10
Impact Factor
ISSN: 1449-2288
Int J Biol Sci 2009; 5(7):736-744. doi:10.7150/ijbs.5.736 This issue Cite
Research Paper
Cloning, Expression and Purification of an Acetoacetyl CoA Thiolase from Sunflower Cotyledon
1. Department of Chemistry and Biochemistry, Montclair State University, Montclair, NJ 07043 USA
2. Medizinisches Proteom-Center, Ruhr-Universtität Bochum, Bochum, Germany
3. PharmaCenter Bonn, Pharmaceutical Chemistry, Bonn, Germany
Received 2009-10-9; Accepted 2009-11-24; Published 2009-12-2
Abstract
Thiolase I and II coexist as part of the glyoxysomal β-oxidation system in sunflower (Helianthus annuus L.) cotyledons, the only system shown to have both forms. The importance of thiolases can be underscored not only by their ubiquity, but also by their involvement in a wide variety of processes in plants, animals and bacteria. Here we describe the cloning, expression and purification of acetoacetyl CoA thiolase (AACT) in enzymatically active form. Use of the extensive amount of sequence information from the databases facilitated the efficient generation of the gene-specific primers used in the RACE protocols. The recombinant AACT (1233 bp) shares 75% similarity with other plant AACTs. Comparison of specific activity of this recombinant AACT to a previously reported enzyme purified from primary sunflower cotyledon tissue was very similar (263 nkat/mg protein vs 220 nkat/mg protein, respectively). Combining the most pure fractions from the affinity column, the enzyme was purified 88-fold with a 55% yield of the enzymatically active, 47 kDa AACT.
Keywords: thiolase, sunflower, acetoacetyl CoA, cloning, expression
1. Introduction
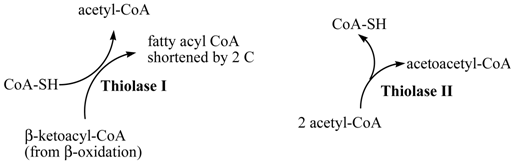
Both the metabolic degradation and synthesis of carbon chains can require at some point the involvement of a thiolase activity. These two types of activity have been designated as Thiolase I (EC 2.3.1.16, 3-ketoacyl-CoA thiolase KAT) and Thiolase II (EC 2.3.1.9, acetoacetyl CoA thiolase AACT). The accepted name for thiolase I is acetyl-CoA C-acyltransferase, while for thiolase II is acetyl-CoA C-acetyltransferase. Thiolase I (3-ketoacyl CoA thiolase), which exhibits broad substrate specificity, is also called the degradative thiolase as it mainly catalyzes the final step in the β-oxidation pathway, shortening an acyl-CoA by the two-C acetyl CoA, the other product of the reaction. Thiolase II (acetoacetyl CoA thiolase) uses only acetoacetyl Co A and is also called the synthetic thiolase as it catalyzes the condensation of two acetyl CoA to form the 4-C acetoacetyl CoA, an important starting molecule of hormone and cholesterol synthesis (Figure 1) [1].
Catabolism of fatty acids via the four-reaction β-oxidation cycle is a ubiquitous aspect of metabolism, and thiolase is the final enzyme in this cycle in both prokaryotes and eukaryotes [1]. In the final step of β-oxidation, thiolase catalyzes the CoA-dependent thiolytic cleavage of acyl-CoA esters resulting in the formation of acetyl-CoA and an acyl-CoA ester that is two carbon units shorter than the activated fatty acid that entered the cycle.
Fatty acids are converted to acyl-CoA esters and then are completely degraded to acetyl-CoA by repeatedly passing through the reactions of β-oxidation. Mammalian cells have both mitochondrial and peroxisomal thiolase, but controversy remains regarding subcellular location of thiolase in plants. Much evidence indicates that plant thiolase activity is peroxisomal [2,3]; however, some evidence for mitochondrial activity exists [4,5]. In plants, peroxisomal matrix proteins are synthesized in the cytosol and directed to the matrix by either peroxisomal targeting signal 1 (PTS1) or by PTS2 [6].
The crystal structure of Arabidopsis degradative thiolase I has been reported [7], and studies on mutant thiolase I indicate that it has a role in plant development [8,9]. More recent work has shown that the KAT2 thiolase is involved in the early onset of senescence as well as the catabolic processes of senescence [10].
The crystal structures of human cytosolic [11] and mitochondrial [12] AACT have been reported. This cytosolic thiolase appears to have an important role in mevalonate pathway, while the mitochondrial thiolase is involved in synthesis and degradation of ketone bodies as well as for the degradation of 2-methylacetoacetyl-CoA.
Extensive characterization of two thiolase enzymes from the glyoxysomal fraction of sunflower cotyledon tissue has shown that both the Thiolase I and Thiolase II activities occur in this tissue [13]. We have previously cloned and purified an enzymatically active 3-ketoacyl CoA thiolase from sunflower cotyledon [14], and here we report the cloning, expression and purification of the acetoacetyl CoA thiolase from the sunflower cotyledon.
Enzymatic activities of Thiolase I and Thiolase II. Acetyl CoA produced by Thiolase I (EC 2.3.1.16, 3-ketoacyl-CoA thiolase KAT) activity is used in the TCA cycle for energy production, while acetoacetyl CoA resulting from Thiolase II (EC 2.3.1.9, acetoacetyl CoA thiolase AACT) activity is used in anabolic processes including ketone body and sterol synthesis.

2. Materials and Methods
2.1 Plant material and growth conditions
Achenes of common sunflower (Helianthus annuus L.) were soaked overnight in tap water, planted in a beaker containing sterile vermiculite/perlite (2:1) moistened with sterile tap water loosely covered with aluminum foil. For germination and growth achenes were kept in the dark at 27°C for three days.
2.2 RNA extraction
All glassware was baked at 200°C for 8 hours to ensure removal of contaminating RNase. Total RNA was extracted from the cotyledons of the etiolated seedlings using RNeasy Plant Mini Kit (Qiagen). Two cotyledons (about 100 mg) from 3-day post imbibition etiolated seedlings were ground to a fine powder in liquid nitrogen using a mortar and pestle. The powder was transferred to a 1.5 mL microcentrifuge tube and the remaining kit protocol was followed. Total RNA was eluted from the column in 40 µL DEPC-treated water. The RNA concentration was determined spectrophotometrically. RNA integrity was determined by resolving 50 µg of the total RNA on a denaturing agarose gel and visualizing the intact 28S and 18S rRNA bands.
2.3 Gene-Specific Primer Design for 3'- and 5'-RACE
For the design of suitable primers for the 3'-RACE the conserved domain database (CCD) search tool was used [15]. The consensus sequence obtained from the alignments of the conserved C-terminal domains of several known thiolases was backtranslated into DNA [16]. From the resulting BLAST alignments an Arabidopsis thaliana sequence (NM_203172) was chosen as template for the primer design (see Figure 2). The primer Athal (5'-CTCAATGCGCATGGAGGGGC-3') starts at position 1184 in actual A. thaliana DNA sequence, so the predicted 3'RACE product would be ~ 200 bp.
Based on the sequence data from the partial sunflower AACT generated with the 3' RACE protocol, gene specific primers were made for use as the 3' primer for a 5'RACE strategy to obtain the remainder of the sunflower AACT. Two primers, Sun1 (5'-CCGAACCTTCCGTTATTTTGCC-3') and Sun2 (5'-CAACTTGGATAGGGTCCCTGC-3'), were the reverse complement of bases in the fragment generated in the 3'-RACE. The expected 5' RACE product for Sun1 is ~1225 bp and for Sun2 ~1317 bp.
Primer design for 3'-RACE. The conserved domain database (CCD) search tool was used to obtain the consensus sequence for the C-terminal domain of thiolases. This consensus sequence was backtranslated to DNA (C-termCD: C-terminal conserved domain) and a BLAST search was performed with the resulting sequence. A. thaliana sequence (NM_203172) was chosen as template for the primer design and the highlighted sequence was synthesized as 3'-RACE primer Athal.

2.4 cDNA synthesis and 3'-RACE
The 3 ́ RACE System for Rapid Amplification of cDNA Ends (Invitrogen) was used. First strand cDNA synthesis was catalyzed by SuperScript II reverse transcriptase (Invitrogen) using 5 µg of total RNA from sunflower cotyledon and the Adapter Primer that initiates cDNA synthesis at the poly (A) region of the mRNA as described in the kit manual. Gene-specific primers described above were used as the 5'-primer in tandem with the Invitrogen kit Abridged Universal Anchor Primer (AUAP) in the following PCR protocol: 2 min 95°C- initial denaturation 1x; 45 sec 95 °C, 50 sec 60°C-anneal; 45 sec 72°C-extension 25 x; 2 min 72°C- final extension. Reactions were 50 μL and used components of Promega GoTaq PCR Core I system. 10 µL of the PCR products were resolved on 1% agarose TAE gel and were stained with ethidium bromide and then visualized on a UV transilluminator.
2.5 5'RACE protocol
Total RNA from the sample used for the 3' strategy (RNA1) along with another total RNA sample also extracted from sunflower cotyledon (RNA2) was dephosphorylated, decapped and ligated to the 5' GeneRacer primer according to the 5'RACE manual (Invitrogen). This modified RNA was then reverse transcribed with Thermoscript reverse transcriptase (Invitrogen).
These two cDNA products were used as templates in a touchdown PCR protocol with Taq DNA polymerase. 94°C initial denaturation-2 min; 5X: 94°C, 30 sec; 72°C 90 sec; 5X: 94°C, 30 sec; 70°C 90 sec; 25 X: 94°C, 30 sec; 58°C 30 sec, 72°C 90 sec; 72 °C 10 min.
2.6 Cloning of PCR product from both 3'- and 5'-RACE
RACE reaction products were resolved on 0.8% low-melt agarose gels, and the bands of expected size were excised from gel and purified using a QiaQuick PCR purification kit (Qiagen). PCR product was eluted into a final volume of 30 µL H20, and 2 µL of each was used in TOPO-TA cloning reactions with the pCR 2.1 TOPO vector(Invitrogen). Two µL of the cloning reaction was used to do a heat-shock transformation of One-Shot TOP10 chemically competent cells (Invitrogen) and plated overnight on LB/AMP plates.
Individual colonies on the LB/AMP plate were inoculated into 5 mL of LB + 75 µg/mL ampicillin and allowed to incubate with shaking at 250 rpm at 37°C overnight. Plasmid DNA was isolated using a Promega plasmid miniprep kit and subjected to restriction enzyme digestion with EcoRI and resolved on 0.8% agarose gel.
Plasmids from digests giving the expected restriction pattern were sent for sequencing (Macrogen) using the T3 and T7 vector-based primers. Sequence analysis was done using BLASTn.
This analysis indicated that several clones contained DNA sequences that were consistent with them being portions of a putative acetoacetyl CoA thiolase.
Because the sequence analysis indicated that full-length sunflower AACT was obtained, but not all in one clone, the following strategy was developed to get the full-length sunflower AACT all in one clone. Two primers (5'FullThio1 ATGCAGATCATGGATAAAATTAACCC and 3'FullThio1 TTACAACTTGGTAGGGTCCCTGCAC) were synthesized to flank the full-length coding sequence of the sunflower AACT. 3'FullThio1 was used in a reverse transcriptase reaction using the same total RNA as before with ThermoScript RT (Invitrogen). This RT product served as the template in a touchdown PCR protocol as described above using the 5' and 3' FullThio1 primers, Platinum Pfx polymerase, and PCRx enhancer (Invitrogen).
The resulting 1236 bp PCR product was cloned into the pCR4 TOPO-TA vector (Invitrogen), transformed into TOP10 E. coli, and the plasmid DNA was isolated and characterized. Sequence analysis indicated that the full coding sequence including start and stop codons occurred in the clone designated pCR-TOPO AACT.
The full-length sunflower AACT was excised from the pCR4-TOPO AACT with EcoRI. This full-length AACT with EoRI sticky ends was ligated into the pBAD His B vector (Invitrogen) that had been cut with EcoRI and treated with calf intestinal alkaline phosphatase (Promega) to reduce religation of the pBAD His B vector. This also facilitates the in-frame cloning of the AACT with an N-terminal 6X His-tag encoded by the pBAD His B vector. The 10 µL ligation reaction followed the Promega protocol and consisted of 2 µL 5X buffer, 1µL PromegaT4 DNA ligase, 100 ng EcoRI digested/ CIAP treated pBAD-HisB vector, 50 ng full Thio1 with EcoRI sticky ends, water to 10 µL. After a 4 h incubation at 15 °C, 5 µL of the ligation reaction were transformed into TOP10 E. coli by heat shock, cells plated on LB/ampicillin plates and incubated overnight at 37 °C. Individual colonies from the overnight plate were then inoculated into 5 mL of LB/ampicillin and grown overnight at 37 °C with shaking.
2.7 Expression and Purification
A single colony of E. coli TOP 10 containing pBAD/HisB-AACT was inoculated into LB/ampicillin medium and grown overnight. This 5 mL culture was transferred into 95 mL of the same medium. The culture was split into two 50 mL cultures, and expression was induced in one of them at OD600 ~0.5 with 0.2% (w/v) arabinose. The other culture served as the uninduced control. Bacteria were grown at 37°C with shaking at 270 rpm for four hours. The following steps were all conducted at 4oC unless otherwise specified. Cultures were centrifuged at 5,000 rpm for 10 minutes, supernatant was discarded, and the pellet was resuspended in 10 mL of lysis buffer (50 mM NaH2PO4 pH 8.0, 300 mM NaCl, 2 mM DTT). After a 15 minute incubation during which lysozyme was present at a final concentration of 2 mg/mL, the bacteria were sonicated (10 second bursts followed by 10 seconds on wet ice) until a noticeable clearing of the bacterial suspension occurred. An Ultrasonic W-225 sonicator set at 70% was used. Sonicated bacteria were centrifuged at 10,000 x g in a JA 25-50 rotor in an Avanti J-30I centrifuge (Beckman) for 15 minutes. Supernatant was combined with 2 mL of a nickel (II)-nitrilotriacetic acid (Ni-NTA) agarose resin (Invitrogen) equilibrated in lysis buffer. Binding took place on a rotating wheel for 2 hours after which entire contents were poured into a Poly-Prep Bio-Rad mini-column. Flow-through was saved, and column was washed with four column volumes of wash buffer (lysis buffer plus 10 mM imidazole). Proteins were eluted with a step gradient of imidazole (10, 50, 100, 125, 150, 250 mM) in 2 mL increments.
SDS-PAGE analysis. 25 µL of each fraction were combined with 5 µL 6x SDS-loading buffer (300 mM Tris HCl, pH 6.8, 400 mM DTT, 20% SDS, 3% bromophenol blue, 50% glycerol), heated at 95oC for 5 minutes and resolved on a 0.75 mm 4% stacking/10% resolving minigel (BioRad). Proteins were visualized with Simply Blue stain (Invitrogen) and sizes estimated from broad range standards (Invitrogen).
2.8 Thiolase assay (340 nm assay)
Thiolase activity was determined spectrophotometrically by monitoring the rate of NADH formation at 340 nm in a coupled enzyme assay. In this assay, the product of the thiolase reaction, acetyl-CoA, was used as substrate for the condensation with oxaloacetate. This reaction, which is the first reaction in the TCA cycle, is catalyzed by citrate synthase (CS). The reaction that precedes the CS step in the TCA cycle, the oxidation of malate to oxaloacetate by malate dehydrogenase (MDH), produces NADH. By converting one molecule of oxaloacetate and one molecule of acetyl-CoA to citrate, the equilibrium of the MDH reaction, which is highly endergonic, is shifted towards the production of oxaloacetate and NADH. The ratio between NADH formation and acetoacetyl-CoA degradation is 2:1. The reaction mixture consisted of 175 mM Tris-HCl (pH 8.5), 0.12 mM CoA, 2.0 mM dithioerythritol (DTE), 2.6 mM malate, 0.14 mM NAD, 58 nkat malate dehydrogenase, 18 nkat citrate synthase, 0.05% (w/v) bovine serum albumin, and recombinant thiolase in a total volume of 1 mL. The reaction was initiated with 20μM acetoacetyl-CoA. All reagents were purchased from Sigma. For calculations of specific activities the protein concentration was determined using the Bradford dye-binding procedure with the BioRad protein assay kit and BSA as the standard.
3. Results and Discussion
5'- and 3'-RACE protocol was used to clone the full-length AACT cDNA from sunflower cotyledon tissue. The 5' primers used for the 3'RACE were designed based on the multiple sequence alignment of several putative AACT sequences from other species. Using the conserved domain database (CCD) search tool [15], the consensus sequence for the C-terminal domain of thiolases was determined (Figure 2). The DNA sequence obtained after backtranslating [16] this consensus sequence (C-termCD) was used in a BLAST search. Among the several plant and animal AACT sequences that were found, the A. thaliana sequence was chosen as template for designing for the 5'-primers used in the 3'-RACE (Figure 2).
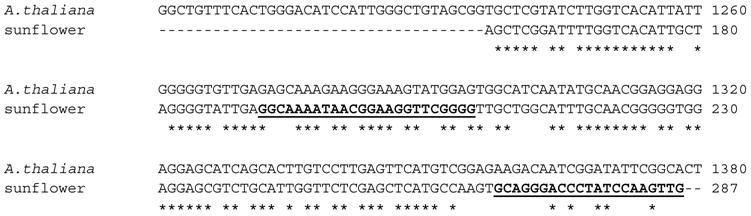
A thermoscript Reverse Transcriptase was used to produce the single-stranded cDNA using total RNA extracted from about 100 mg of sunflower cotyledon tissue. For the 3'-RACE, using the cDNA as template, the Athal primer as the 5' primer and the UAP (Universal Adapter Primer) kit primer as the 3' primer in a PCR protocol, a double-stranded fragment of the expected size of 500 bp was obtained. This fragment was cloned into a TOPA-TA vector and sequenced using the vector-based T3 and T7 primers. Results of a BLAST search with this sequence indicated that it shares up to 75% identity with AACTs from other plant species such as rubber tree, rice, castor bean and corn. The alignment of this product with the A. thaliana NM_203172 is shown in Figure 3.
Alignment of 3'-RACE PCR product with A. thaliana thiolase (NM_203172). The putative sunflower AACT shares 70% sequence identity with the Arabidopsis sequence. Reverse complements of the underlined bold sequences were the primer sequences Sun1 and Sun2, which were the 3'-primers for the 5'-RACE protocol.
Sequence information from the 3'RACE was used to generate the 3'- primers to be used in the 5'-RACE protocol—primers Sun1 and Sun2. The total RNA extracted from the sunflower cotyledons was treated as described in Materials and Methods and then reverse transcribed with a Thermoscript reverse transcriptase. Touchdown PCR was used to obtain the full 5' region of the sunflower AACT. Both Sun1 and Sun2 resulted in 5'-RACE product. Cloning and sequencing of these products verified their identities as AACT fragments, including the 5' start codon.
To obtain the complete and contiguous sunflower AACT, primers corresponding to the 5' and 3' ends of the sunflower coding sequence were synthesized and designated as 5'Fullthio and 3'Fullthio. 3'Fullthio was first used in a reverse transcriptase reaction with total sunflower RNA template, after which the resulting cDNA was used as a template with both 5'Fullthio and 3'Fullthio as the primers in a touchdown PCR protocol. Reaction products were resolved on a 1% agarose TAE gel, and the resulting ~1200bp fragment was gel-purified and cloned into the pCR4 TOPO-TA vector (Invitrogen). Sequence analysis confirmed that the full-length coding region for the sunflower AACT was present (Figure 4).
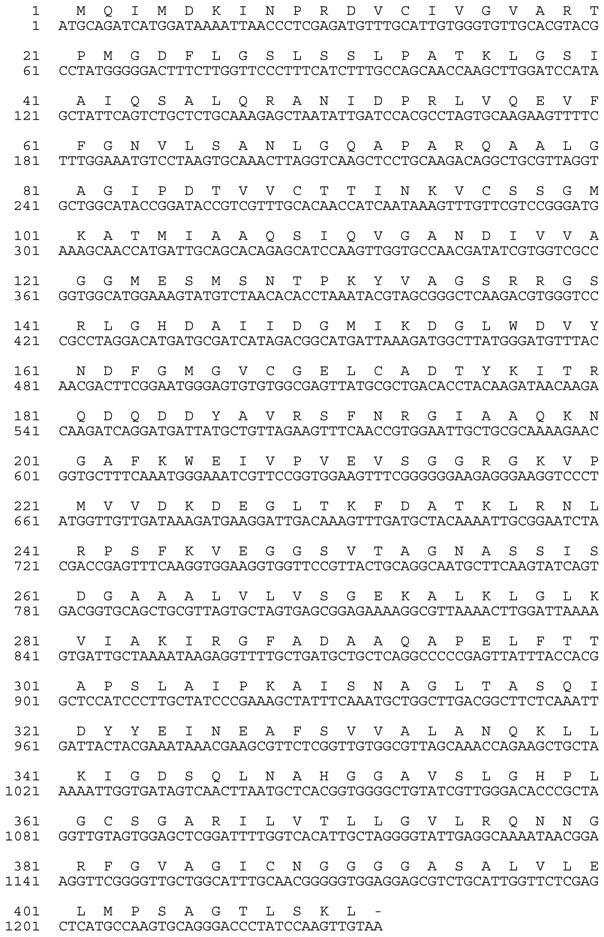
The full-length sunflower AACT was excised from the pCR4 TOPO vector with EcoRI, facilitating in-frame cloning into the EcoRI-digested bacterial expression vector pBAD His B (Invitrogen). The pBAD His B-full AACT was sequenced in both directions, confirming the integrity of the coding region and the 5' 6X His tag coded by the vector sequence. The 1233 bp AACT coding sequence is predicted to encode a 411 amino acid protein. This size corresponds well with the length of the other predicted AACTs corresponding to the Arabidopsis annotated gene products from At5g47720 and At5g48230: 415 and 403 amino acids, respectively [17].
Nucleotide and deduced amino acid sequences of the AACT (thiolase II) from sunflower (Helianthus annuus L.) cotyledon. The sequences have been submitted to GenBank (Accession No. GQ254017).

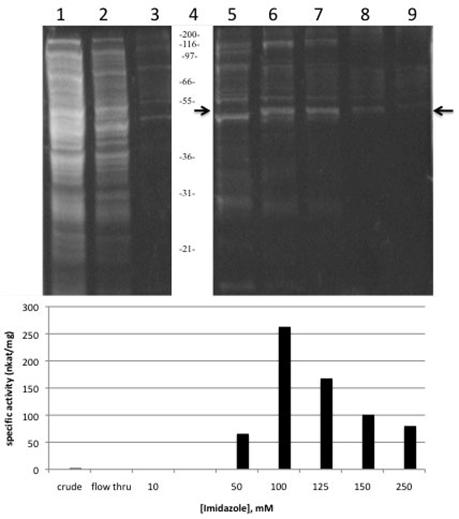
Sunflower AACT expression in bacteria harbouring the pBAD HisB-full AACT vector was induced with the addition of arabinose. L- arabinose turns on the expression of recombinant protein from the pBAD promoter, while in the absence of L-arabinose, expression levels are very low [18]. The optimal arabinose concentration was determined to be 0.2% (w/v). TOP10 E. coli used for this pilot expression are suitable because they are able to transport arabinose, but they can not metabolize it [19]. The His-tagged AACT was purified using the Ni-NTA affinity chromatography and a step-wise concentration gradient of imidazole to elute the bound His-tagged protein from the column. SDS-PAGE analysis under reducing conditions of the fractions from the column showed a distinct band at 48 kDa. Using the “Compute pI/Mw” tool on the ExPASy server [20], the calculated MW of the recombinant AACT including the four vector-encoded linker amino acids, 6 His tag and the additional 37 vector-encoded amino acids is 47711 Da. Over 70% of the proteins eluted from the column with imidazole concentration higher than 125 mM appear to be the 48 kDa thiolase (Figure 5, lanes 7, 8 and 9). At a concentration of 100 mM imidazole, the recombinant thiolase is also present but is more significantly contaminated with higher and lower molecular weight proteins (Figure 5, lane 6). The size of the AACT purified and characterized from the glyoxysomal fraction of sunflower cotyledons was reported as 63 kDa [13], which is significantly higher than the molecular weight reported here; however, another thiolase activity was reported in that paper and was not further characterized. This second thiolase perhaps corresponds to the thiolase reported here. The predicted size of the sunflower AACT is 42.6 kDa, while the size of other AACT subunits that have been purified or cloned range from 40-45 kDa [3,12,21,22,23].
Coomassie blue-stained SDS-PAGE and thiolase activity assay of recombinant sunflower AACT. (A) Recombinant thiolase expressed in E. coli and purified with nickel-affinity chromatography. Equal volumes of each fraction were prepared and resolved on a 0.75 mm 4% stacking/10% resolving minigel (BioRad). Lane 1: Lysate supernatant; Lane 2: flow-through from column; Lanes 3, 5-9: elution fractions (10, 50, 100, 125, 150, 250 mM imidazole); lane 4: Invitrogen Mark 12 standards , kDa. Arrow at 47 kDa indicates the AACT.
The enzyme assay for the thiolase activity is a coupled reaction assay in which the product of the thiolase reaction, acetyl-CoA, becomes the substrate for the reaction catalyzed by citrate synthase (CS). CS catalyzes the condensation reaction between acetyl-CoA and oxaloacetate, which is produced in this assay from the NAD-dependent oxidation of malate, a reaction catalyzed by malate dehydrogenase. Thus the extent to which NAD+ is reduced to NADH, which can be monitored spectrophotometrically at 340 nm, serves as an indicator of the level of thiolase activity.
The purified recombinant AACT was enzymatically active, and no endogenous thiolase activity was detectable in the uninduced controls. No thiolase activity was observable in the flow-through, so apparently all activity was bound to the column (Figure 5, lane 2), further indicating that no endogenous thiolase activity was present. Thiolase activity was distributed throughout the fractions eluted from the affinity column, with the highest specific activity eluting in the 100 mM fraction (Figure 5, lane 6). Specific activity of this fraction was 263 nkat/mg protein, corresponding to a 118-fold purification of the recombinant thiolase with a yield of 31%. These values are comparable to a specific activity of 220 nkat/mg protein obtained in the partial purification of the AACT from the glyoxysomal fraction of sunflower cotyledon [13]. Pooling of the 100-250 mM fractions resulted in an overall yield of 56% with a specific activity of 198 nkat/mg protein, which is sufficiently pure for further characterization of the AACT activity (Table 1).
Purification of recombinant sunflower acetoacetyl-CoA thiolase
| Purification step | Total protein (mg) | Total activity (nkat) | Specific activity (nkat/mg) | Yield (%) | Purification (-fold) |
|---|---|---|---|---|---|
| Lysate supernatant | 32.4 | 75 | 2.2 | 100 | 1 |
| 10 mM imidazole | 0.6 | 0.0 | 0.0 | 0 | 0 |
| 50 mM imidazole | 0.069 | 4.5 | 66 | 6.3 | 29 |
| 100 mM imidazole | 0.086 | 22.6 | 263 | 31.2 | 118 |
| 125 mM imidazole | 0.054 | 9.1 | 168 | 12.5 | 75 |
| 150 mM imidazole | 0.0445 | 4.5 | 101 | 6.2 | 45 |
| 250 mM imidazole | 0.051 | 4.1 | 80 | 5.6 | 36 |
| 100-250 mM pooled | 0.2355 | 40.2 | 198 | 55.5 | 88 |
Clustal W2 alignments of the deduced amino acid sequences of the AACT from sunflower cotyledons with the amino acid sequences of several other plant species resulted in about 50% amino acid sequence identity and 75% sequence similarity. The sunflower AACT had the C-terminus peroxisomal targeting sequence (PTS) consisting of the tripeptide SKL, while all the other sequences had some variant of this PTS1. Three of the AACT homologues (from Populos, Nicotiana and Oryza) had the recently identified PST1 SSL [24], which was confirmed in a leaf peroxisomal proteome study [25]. As seen in Figure 6, eight of the fourteen AACTs were truncated at the C-terminus and had no discernible N-termini and also had no PST2 which consists of the nonapeptide RX6(H/Q)(L/A/F) located close to the N-terminus [26].
Multiple sequence alignment of the closest hits of a BLAST search performed with the deduced amino acid sequence of Helianthus annuus AACT showing the C-termini only. Alignment was done using CLUSTAL W2. The accession numbers are as follows: AAU95619 (N. tabacum), XP_002522876 (R. communis), AF429383_1 (H. brasiliensis), BAF98276 (H. brasiliensis 2), ABC74567 (Picrorhiza kurrooa), BAD22334 (Oryza sativa Japonica Group), NP_199583 (A. thaliana), XP_002308755 (P. trichocarpa), ACN40771 (P. sitchensis), ACF85124 (Z. mays), XP_002320528 (P. trichocarpa 2), XP_002265690 V. vinifera), EEC69797 (O. sativa Indica Group).

Conflict of Interest
The authors have declared that no conflict of interest exists.
References
1. Igual JC, González-Bosch C, Dopazo J, Pérez-Ortín JE. Phylogenetic analysis of the thiolase family. Implications for the evolutionary origin of peroxisomes. Journal of Molecular Evolution. 1992;35:147-155
2. Frevert j, Kindl H. Purification of glyoxysomal acetyl-CoA acyltransferase. Hoppe Seylers Z Physiol Chem. 1980;361:537-542
3. Carrie C, Murcha MW, Millar AH, Smith SM, Whelan J. Nine 3-ketoacyl-CoA thiolases (KATs) and acetoacetyl-CoA thiolases (ACATs) encoded by five genes in Arabidopsis thaliana are targeted either to peroxisomes or cytosol but not to mitochondria. Plant Molecular Biology. 2007;63:97-108
4. Masterson C, Wood C. Mitochondrial β-oxidation of fatty acids in higher plants. Physiologia Plantarum. 2000;109:217-224
5. Masterson C, Wood C. Mitochondrial and peroxisomal β-oxidation capacities of organs from a non-oilseed plant. Proceedings of the Royal Society of London Series B: Biological Sciences. 2001;268:1949-1953
6. Eubel H, Meyer EH, Taylor NL, Bussell JD, O'Toole Heazlewood N, Castleden I, Small ID, Smith SM, Millar AH. Novel Proteins, Putative Membrane Transporters, and an Integrated Metabolic Network Are Revealed by Quantitative Proteomic Analysis of Arabidopsis Cell Culture Peroxisomes. Plant Physiol. 2008;148:1809-1829
7. Sundaramoorthy R, Micossi E, Alphey MS, Germain V, Bryce JH, Smith SM, Leonard GA, Hunter WN. The Crystal Structure of a Plant 3-Ketoacyl-CoA Thiolase Reveals the Potential for Redox Control of Peroxisomal Fatty Acid [beta]-Oxidation. Journal of Molecular Biology. 2006;359:347-357
8. Germain V, Rylott EL, Larson TR, Sherson SM, Bechtold N, Carde J-P, Bryce JHIAG, Smith SM. Requirement for 3-ketoacyl-CoA thiolase-2 in peroxisome development, fatty acid β-oxidation and breakdown of triacylglycerol in lipid bodies of Arabidopsis seedlings. The Plant Journal. 2001;28:1-12
9. Footitt S, Cornah JE, Pracharoenwattana I, Bryce JH, Smith SM. The Arabidopsis 3-ketoacyl-CoA thiolase-2. kat2-1;mutant exhibits increased flowering but reduced reproductive success. J Exp Bot. 2007;58:2959-2968
10. Castillo MC, Leon J. Expression of the β-oxidation gene 3-ketoacyl-CoA thiolase 2. KAT2;is required for the timely onset of natural and dark-induced leaf senescence in Arabidopsis. J Exp Bot. 2008;59:2171-2179
11. Kursula P, Sikkila H, Fukao T, Kondo N, Wierenga RK. High Resolution Crystal Structures of Human Cytosolic Thiolase. CT): A Comparison of the Active Sites of Human CT, Bacterial Thiolase, and Bacterial KAS I. Journal of Molecular Biology. 2005;347:189-201
12. Haapalainen AM, Merilainen G, Pirila PL, Kondo N, Fukao T, Wierenga RK. Crystallographic and Kinetic Studies of Human Mitochondrial Acetoacetyl-CoA Thiolase: The Importance of Potassium and Chloride Ions for Its Structure and Function. Biochemistry. 2007;46:4305-4321
13. Oeljeklaus S, Fischer K, Gerhardt B. Glyoxsomal acetoacetyl-CoA thiolase and 3-osoayl-CoA thiolase from sunflower cotyledons. Planta. 2002;214:597-607
14. Schiedel AC, Oeljeklaus S, Minihan P, Dyer JH. Cloning, expression and purification of glyoxysomal 3-oxoacyl-CoA thiolase from sunflower cotyledon. Protein Expression and Purification. 2004;33:25-33
15. Marchler-Bauer A, Bryant SH. CD-Search: protein domain annotations of the fly. Nucleic Acids Research. 2004;32(web server issue):W327-331
16. Backtranslation Tool. Fischer M. http://www.entelechon.com/backtranslation
17. Ahumada I, Cairo A, Hemmerlin A, Gonzalez V, Pateraki I, Bach TJ, Rodriguez-Concepcion M, Campos N, Boronat A. Characterisation of the gene family encoding acetoacetyl-CoA thiolase in Arabidopsis. Functional Plant Biology. 2008;35:1100-1111
18. Schleif R. Regulation of the -arabinose operon of Escherichia coli. Trends in Genetics. 2000;16:559-565
19. Bachmann BJ. Linkage map of Escherichia coli K-12, edition 8. Microbiol Mol Biol Rev. 1990;54:130-197
20. ExPASy. Compute pI/MW tool. 2009. http://www.expasy.ch/tools/pi_tool.html
21. Fukao T, Yamaguchi S, Kano M, Fujiki Y, Osumi T, Hashimoto T. Molecular cloning and sequence of the complementary DNA encoding human mitochondrial acetoacetyl-coenzyme A thiolase and study of the variant enzymes in cultured fibrobalsts form patients with 3-ketothiolase deficiency. Journal of Clinical Investigation. 1990;86:2086-2092
22. Meng Y, Li J. Cloning, expression and characterization of a thiolase gene from Clostridium pasteurianum. Biotechnology Letters. 2006;28:1227-1232
23. Reddick JJ, Williams JK. The mmgA gene from Bacillus subtilis encodes a degradative acetoacetyl-CoA thiolase. Biotechnology Letters. 2008;30:1045-1050
24. Goepfert S, Hiltunen JK, Poirier Y. Identification and Functional Characterization of a Monofunctional Peroxisomal Enoyl-CoA Hydratase 2 That Participates in the Degradation of Even cis-Unsaturated Fatty Acids in Arabidopsis thaliana. J Biol Chem. 2006;281:35894-35903
25. Reumann S, Babujee L, Ma C, Wienkoop S, Siemsen T, Antonicelli GE, Rasche N, Luder F, Weckwerth W, Jahn O. Proteome Analysis of Arabidopsis Leaf Peroxisomes Reveals Novel Targeting Peptides, Metabolic Pathways, and Defense Mechanisms. Plant Cell. 2007;19:3170-3193
26. Swinkels BW, Gould SJ, Bodnar AG, Rachubinski RA, Subramani S. A novel, cleavable peroxisomal targeting signal at the amino-terminus of the rat 3-ketoacyl-CoA thiolase. EMBO Journal. 1991;10:3255-3262
Author contact
![]() Corresponding author. FAX 973-655-7165 email address: dyerjmontclair.edu
Corresponding author. FAX 973-655-7165 email address: dyerjmontclair.edu

 Global reach, higher impact
Global reach, higher impact