Impact Factor ISSN: 1449-2288
Int J Biol Sci 2010; 6(1):51-67. doi:10.7150/ijbs.6.51 This issue Cite
Review
A manually curated network of the PML nuclear body interactome reveals an important role for PML-NBs in SUMOylation dynamics
1. Laboratory of Protein Chemistry, Proteomics and Signal Transduction, Department of Biomedical Sciences, University of Antwerp (Campus Drie Eiken), Universiteitsplein 1 - Building T, 2610 Wilrijk, Belgium.
2. Intelligent Systems Laboratory (ISLab) Department of Mathematics and Computer Science, University of Antwerp (Campus Middelheim), Middelheimlaan 1 - Building G, 2020 Antwerp, Belgium
Received 2009-10-20; Accepted 2010-1-9; Published 2010-1-12
Abstract
Promyelocytic Leukaemia Protein nuclear bodies (PML-NBs) are dynamic nuclear protein aggregates. To gain insight in PML-NB function, reductionist and high throughput techniques have been employed to identify PML-NB proteins. Here we present a manually curated network of the PML-NB interactome based on extensive literature review including database information. By compiling 'the PML-ome', we highlighted the presence of interactors in the Small Ubiquitin Like Modifier (SUMO) conjugation pathway. Additionally, we show an enrichment of SUMOylatable proteins in the PML-NBs through an in-house prediction algorithm. Therefore, based on the PML network, we hypothesize that PML-NBs may function as a nuclear SUMOylation hotspot.
Keywords: PML-NB, SUMOylation, Cytoscape, protein-protein interaction, network
Introduction
Nuclear domain 10 (also called Kremer bodies or PODs) was first observed almost 50 years ago by electron microscopy as nuclear dense granular bodies[1, 2]. These punctuate structures are now dubbed “PML nuclear bodies” (PML-NBs) after their scaffold protein Promyelocytic Leukaemia Protein (PML, also known as TRIM19 or MYL). The PML gene consists of nine exons and alternative splicing yields six nuclear PML isoforms (PML I-VI) and one smaller cytoplasmatic isoform (PMLVII). However, of these seven isoforms there are several alternative spliced variants yielding at least 18 different forms of PML[3]. At the conserved N-terminus, PML contains a motif that defines the TRIM (Tripartite Motif) family of proteins. It is characterized by an RBCC motif composed of a conserved RING domain, one or two B-boxes, and an α-helical coiled-coil domain [4, 5]. The RING domain is a cysteine rich domain with zinc bound in a cross-brace conformation. This structure is ideal for the formation of larger protein aggregates and therefore crucial in processes such as cell growth, oncogenesis, apoptosis, and RNA trafficking during viral infections [5, 6]. The B-box provides the correct orientation and alignment of the coiled-coil domain [7]. When correctly positioned, the coiled-coil domain is responsible for homo and hetero-dimer interactions between TRIM proteins[3]. The RBCC motif of the PML protein makes it possible to sustain a highly dynamic turnover of partner proteins during the cell cycle [8, 9] which could explain the functionally promiscuous nature of the PML-NBs. PML is not only responsible for maintaining the integrity of the complex but also for the recruitment and correct localization of all PML-NB proteins, e.g. p53, Sp100, Daxx and CBP [4]. SUMOylation plays an important role in the scaffold function of PML. SUMO-3 attachment has been reported to be responsible for the correct nuclear localization of PML [10], and without SUMO1 modification several protein partners are not recruited to the PML-NB [10-12]. However, unSUMOylatable mutants of PML (3KR) still show the typical nuclear speckle pattern [11] indicating that also other factors play a role in PML-NB formation. Recently, PML was identified as the first protein degraded by SUMO-dependent polyubiquitination after treatment with arsenic [12].
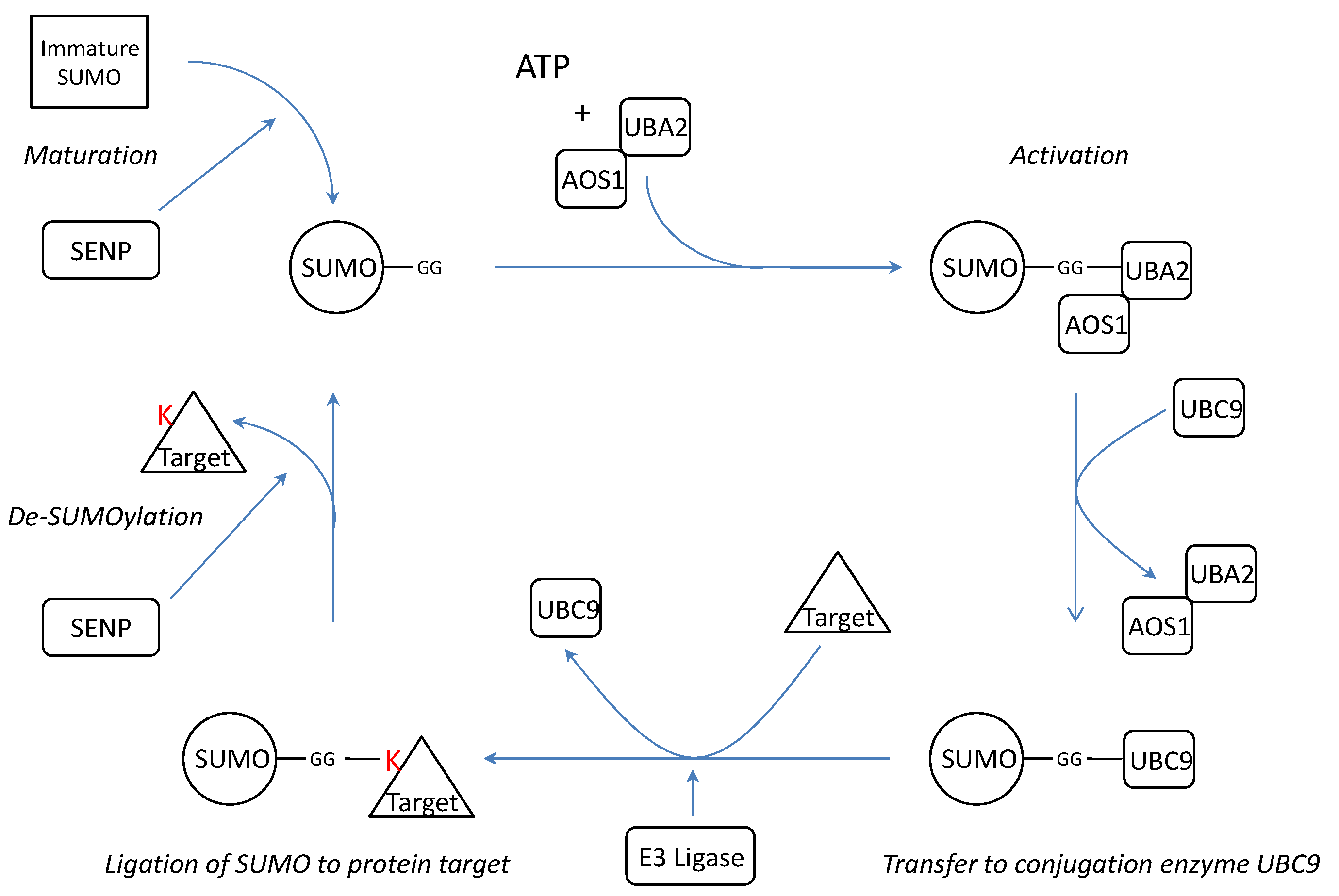
SUMOylation is a post-translational modification where one of the four isoforms of the Small Ubiquitin-like Modifier (SUMO) protein is conjugated to a target protein as a single SUMO molecule (SUMO1) or in polymeric chains (SUMO2/3)[13]. The enzymatic cascade is similar to the ubiquitin pathway[14] (Figure 1). However, unlike ubiquitination, which targets proteins for degradation, SUMOylation modifies proteins to interact in nuclear transport[10, 15], transcriptional regulation[14, 16], apoptosis[17, 18] and protein trafficking[19].
SUMO is synthesized as a precursor and its conjugation requires exposure of two C-terminal glycine residues. The processing of immature SUMO is one of the three functions of SUMO specific proteases (SENPs). In addition, SENPs are also capable of dismantling SUMO chains and reversing SUMOylation by cleaving the isopeptide bonds between SUMO and its protein targets or between the several SUMOs[20]. Thus far, there are six known SENP isoforms (SENP1-3 and SENP5-7)[20] with different assignments in the cascade (Table 1).
Schematic representation of the SUMOylation pathway. Before conjugation, immature SUMO is processed by SUMO specific proteases (SENPs) to expose two C-terminal glycine residues required for conjugation. After processing, mature SUMO is activated in an ATP dependent manner by E1 enzymes AOS1 and UBA2. Subsequently, SUMO is transferred to E2 conjugation enzyme UBC9 and finally conjugated to a target protein by an E3 ligase. SUMO conjugation can be reversed by protease activity of SENPs thereby releasing free SUMO and target. K= Lysine residue.
The SENP isoforms show different affinity for the several SUMO isoforms in maturation processing and de-conjugation.
| SENP isoform | Maturation | De-conjugation |
|---|---|---|
| SENP1 | SUMO1>SUMO2>>SUMO3[21] | SUMO1, SUMO2[22, 23] |
| SENP2 | SUMO2>SUMO1, poor activity towards SUMO3[24, 25] | SUMO2>SUMO1, poor activity towards SUMO3[24, 25] |
| SENP3 | No processing activity | SUMO2, SUMO3[26, 27] |
| SENP5 | SUMO3[26;27] | SUMO1[24, 25] |
| SENP6 | No processing activity[20, 28] | SUMO2, SUMO3[29] |
| SENP7 | No processing activity[20, 28] | SUMO2, SUMO3>>>SUMO1[28] |
After maturation, SUMO is activated in an ATP dependent manner by thio-ester bond formation with the E1 activation proteins UBA2/AOS1[14]. The activating E1 enzyme Uba2p was first identified in yeast where it is, together with Aos1, necessary and sufficient for Smt3p (yeast homologue of SUMO) activation in vitro [30]. The human homologue hUBA2 was shown to act in the same way[31]. Subsequently SUMO is transferred from UBA2 to the SUMO conjugating enzyme UBC9 [14, 32]. UBC9 recognizes the consensus sequence ΨKXE where Ψ is a large hydrophobic amino acid and K is the lysine for SUMO conjugation[33]. UBC9 is believed to be the only SUMO conjugating enzyme[32, 34] and SUMOylation of its binding partners is enhanced by interaction with p14ARF[35]. Finally, an isopropyl bond between the terminal glycine of SUMO and the ε-aminogroup of lysine of the target protein is formed by an E3 ligase. Several unrelated proteins have been attributed E3 SUMO ligase activity: nucleoporin protein RANBP2[36], TOPORS[37, 38], several PIAS proteins[39-41], polycomb group protein Pc2[42], and RNF4[43]. Recently an anti-SUMO ubiquitin-protein isopeptide ligase (E3) has been described : mel-18 interacts both with HSF2 and the SUMO E2 UBC9 and inhibits UBC9's activity which decreases HSF2 SUMOylation[44].
Because of the many associated proteins, PML-NBs are involved in a wide variety of cellular processes such as DNA damage and repair[45, 46], apoptosis[47-49], senescence[50, 51], cell cycle[52], transcription[53], carcinogenesis[54] and virus infection[55, 56]. Despite the involvement in such vital processes, PML deficient mice are normal, fertile and do not have a higher occurrence of tumors [57]. Although the actual function of the bodies still has to be determined, three non-mutually exclusive hypotheses about the function of PML-NBs have been proposed in literature. The first hypothesis gives PML-NBs a role as a nuclear depot thereby providing a mechanism for regulating nucleoplasmic levels of proteins in which PML bodies release proteins as they are needed[58]. Since nascent RNA has been found in the proximity of the PML-NBs[59] and PML-NBs seem to associate in regions with high transcriptional activity[60, 61], the second hypothesis proposes that PML-NBs are associated with nuclear activity such as gene transcription and DNA replication [62]. Indeed, most well-described partners of PML-NBs such as p53, Daxx, Mdm2 and Sp100, all have transcription activation or repression activity [4, 49, 63-66]. Furthermore, PML-NB structure is dependent on the integrity of the surrounding DNA. In fact, many factors which accumulate in PML-NBs are known histone methyltransferases, histone deacetylases or DNA methyltransferases, suggesting that PML-NBs may play an important role as epigenetic regulator of transcription and replication (for a review see [67] and references therein). The third hypothesis is that PML-NBs provide a nuclear platform for post-translational modification (PTM) such as phosphorylation [68, 69], acetylation[70] and SUMOylation[71].
These three hypotheses support the concept of dynamic interactions with PML and a high turnover of protein associates, instead of a static complex with a fixed number of partners. The involvement of PML-NBs in such a variety of processes has led to an enormous research interest in this enigmatic protein which has culminated in an overwhelming number of publications. In recent years, several techniques such as affinity capture, yeast two hybrid, co-localization and mass spectrometry have been employed to identify proteins of the PML complex which could lead to a better understanding of the PML-NB function. This reductionist approach has successfully identified several of the components of the PML-NBs but added more pieces to the puzzle instead of narrowing the spectrum of possible functions. Therefore an integrative “Systems Biology” approach could be essential to grasp the true complexity of the PML-NBs and to allow us to generate hypotheses based upon the vast amount of information available in distributed sources.
Not all PML partners described in literature are well represented in protein databases such as IntAct[72], Molecular Interactions Database (MINT)[73], BIOGRID[74], Human Protein Reference Database (HPRD)[75] and Nuclear Protein Database (NPD)[76]. This paper attempts to bridge the gap between (the lack of) database content and literature information. An integrated view of all the existing information is indispensable to gain true understanding of the functionality of large protein complexes such as PML-NBs.
We present a curated network model of PML-NBs based on information extracted from protein interaction databases (MINT, IntAct, BIOGRID, HPRD and NPD) and fragmented information obtained through extensive literature search. For the compilation of the network we used the open source network analysis software Cytoscape. It features high flexibility and the possibility to analyze protein networks with various (downloadable) plug-in software extensions allowing us to analyze and interpret the collected PML network. Based on this model we offer a system-wide perspective on the pathways and proteins that gather at the nuclear bodies. The network is publicly available (Proteomics division at http://www.ua.ac.be/ppse) to make it a central information resource for “PML-omics” researchers.
Compiling a manually curated PML-NB network
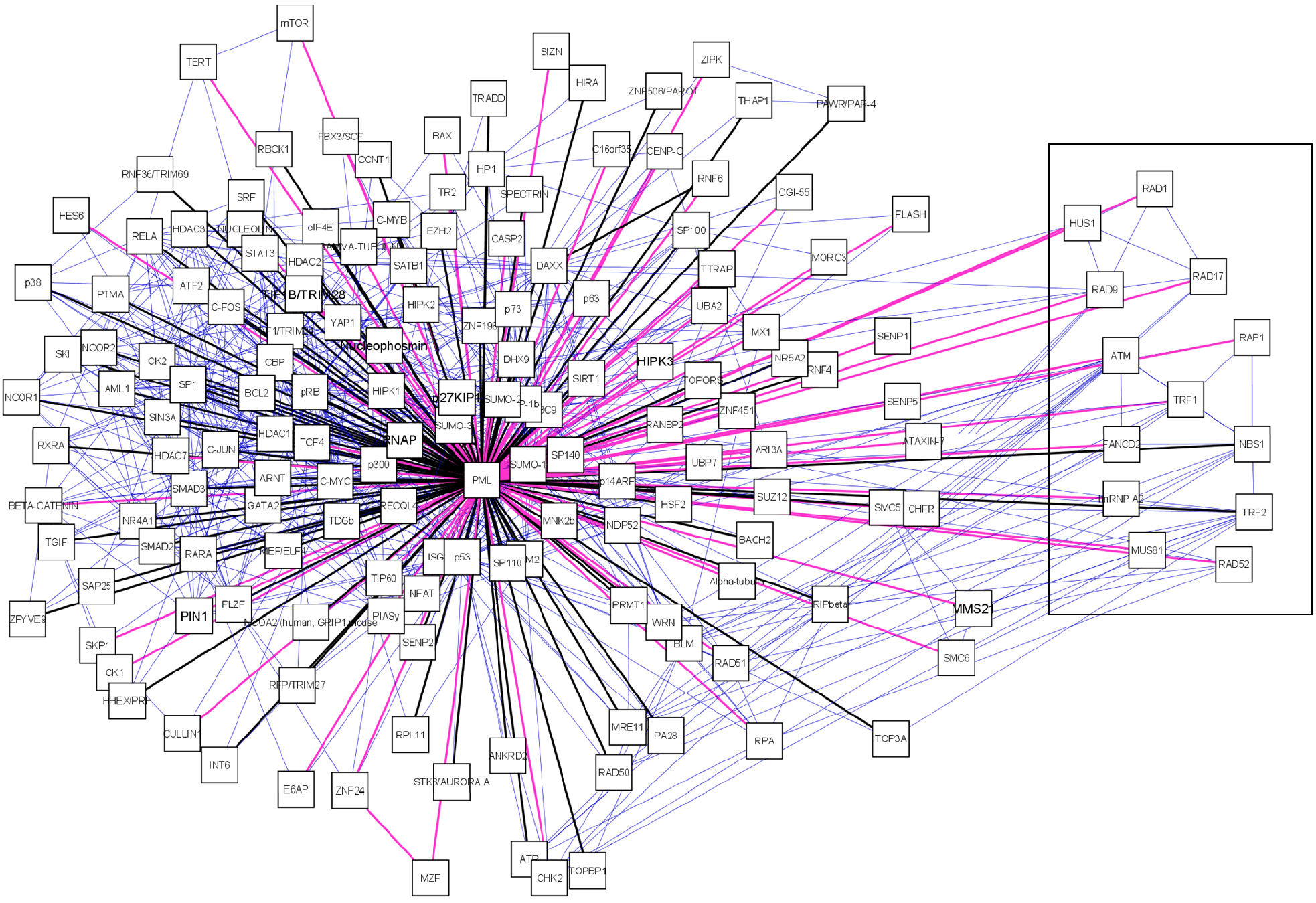
A manually curated PML-NB interaction network was established in Cytoscape[77] by integration of PML interaction data retrieved from databases and literature (figure 2).
First we assembled a primary PML-NB network established with data retrieved from protein databases i.e. MINT[73], BIOGRID[74], HPRD[75], NPD[76] and IntAct[72]. Second, we expanded the primary network by adding PML-NB components which were exclusively found through extensive literature (only peer-reviewed journals) search. Finally, to create a biologically relevant network, interactions between PML-NB associated proteins were retrieved with the aid of Cytoscape import tools. Via the UniProtID we searched the IntAct and cPATH database [78] to further complete the network. After importing all available data from these databases the information was confirmed or completed by searching the BIOGRID, HPRD and MINT databases also using the UniProtID. Interactions found by cPATH and IntAct exploration which could not be verified in other databases were manually checked in literature.
To compile the network, PML was the only protein used as PML-NB determinant. Often Sp100 or Daxx are used as PML-NB determinants but these proteins also dynamically move in and out of the PML-NBs[59]. This makes PML the only protein which always defines the PML-NBs.
Interesting is the presence of a sub-network displaying Alt-associated-PML nuclear bodies (APBs) (figure 2, boxed). These special PML bodies are only seen in a subset of interfase nuclei in telomerase-negative immortalized human cells. These cells maintain their telomeres by a mechanism known as alternative lengthening of telomeres (ALT). Thus far, APBs have not been observed in mortal cells or telomerase positive immortal cells [79, 80]. In the APB, PML co-localizes with telomere binding proteins TRF1 and TRF2, Replication Protein A, RAD50, RAD51, RAD52, RAP1, Mre11, TIN2 and NBS1/p95[79, 81-84]. For RAD51 and RAD52 there is no evidence yet but all other components, including PML are required for APB formation [85]. The function of these bodies remains unclear. It is suggested that PML is required for the transcriptional activation of p53 in ALT cells and thus induces apoptosis via the p53 pathway[86].
After integrating the PML-NB components from database search and literature the network contains 166 nodes (proteins) and 781 edges (interactions). To provide an overview of the characteristics of the PML-NB components and interactions which connect these we provide several edge and node attributes (Table 2). The criteria used to annotate these attributes are standardized and the order in which several sources are used is strictly maintained.
Whereas in most publications only the most common PML-NB partners are mentioned, this is the first time PML-NB components are brought together in a comprehensive network. Depending on the researcher's interest, the network can be analyzed from different angles, eventually using additional tools such as cytoscape plugins.
Analyzing the PML-NB interactome
We analyzed the general properties of the PML-NB components using the attributes described above in table 2. The network contains 166 protein associates of PML of which 70 are described in literature but are not available in any of the standard protein interaction databases.
The majority (57%) of the PML-NB proteins are nuclear, 33% of the PML-NB interactome shuttles actively between nucleus and cytoplasm, while 9% of the components are predominantly cytoplasmatic.
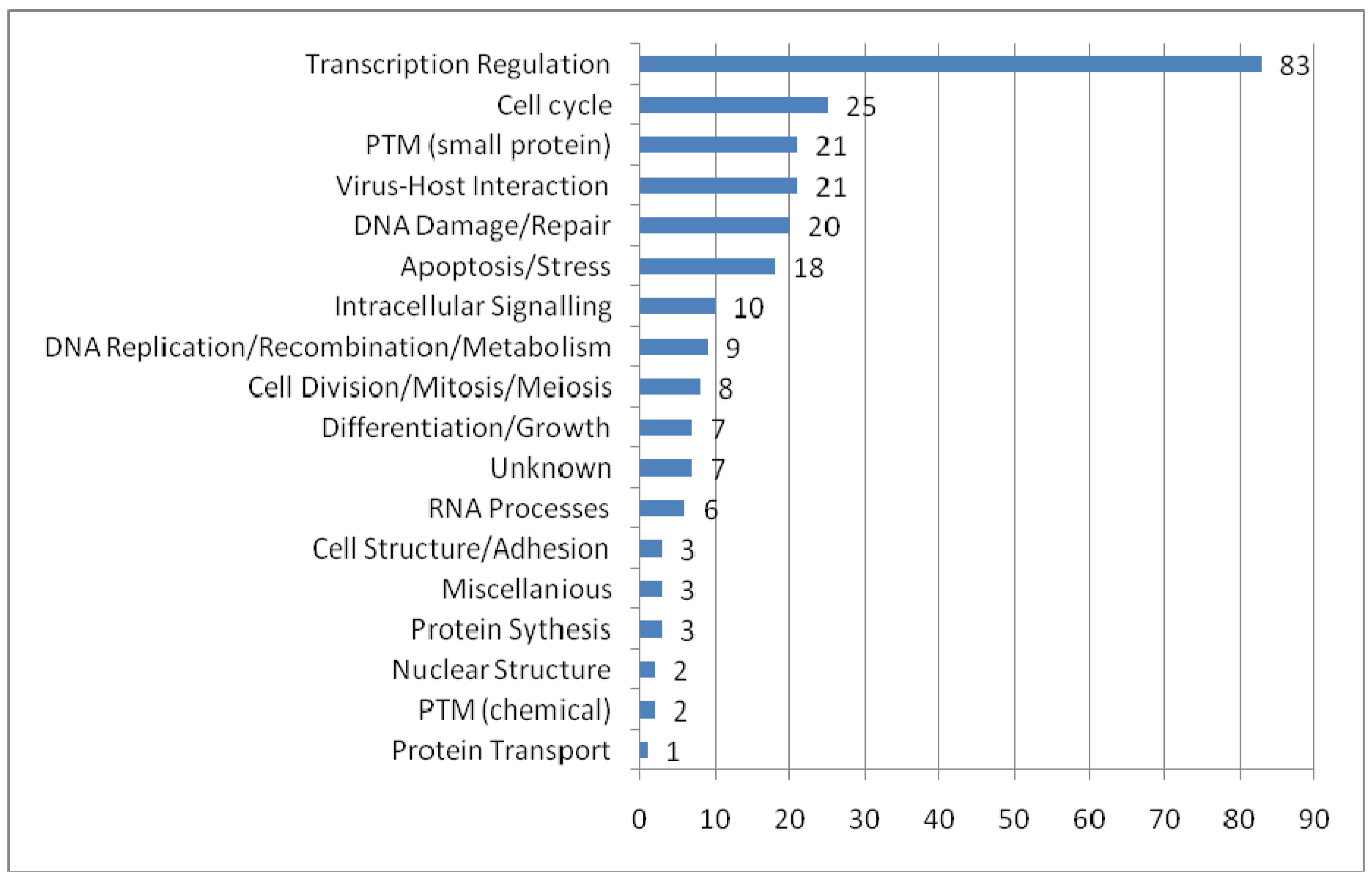
To provide an overview of the processes in which PML-NBs are involved, we categorized the UniProt function keywords in groups and distributed the 166 PML-NB partners in these groups. It is possible that one protein simultaneously belongs to several groups. Figure 3 shows the functional distribution. The representation of proteins involved in transcription regulation is remarkably high. As expected, most of these proteins appear to be nuclear or nucleus/cytoplasm shuttles. Another interesting and well represented group involves proteins which are part of the virus host interaction. Proteins belonging to this group are mostly shuttles between cytoplasm and nucleus. These shuttles emphasize once more the role of PML-NBs in viral infection[93]. In addition, PML nuclear bodies have been implicated in processes as apoptosis and cell cycle it is not surprising that many of the protein partners are directly involved in these processes. An interesting group is the one of the small protein conjugation, e.g. ubiquitin and SUMO, which is found under “PTM (small protein)” in the figure 3.
A manually curated PML-NB network. The central node is the scaffold protein PML. PML-NB components are connected to PML. Black edges represent protein components found in any of the databases (MINT, BIOGRID, IntAct, NPD, HRPD), pink edges were exclusively found in literature. We also represent the interactions between PML-NB components which are depicted as blue edges. For each node we provided the canonical name. Boxed components are those exclusively found in ALT cells.
Overview of the different node and edge attributes in the PML-NB interaction network. In the 'field' column we present the attribute as it is assigned in the network. In the 'description' column we provide a brief explanation of what is embodied by the attribute. Finally, in the 'criteria' column the different sources of information are mentioned.
| Field | Description | Criteria (in order) |
|---|---|---|
| Node Attributes | ||
| ID | UniProtID [87] | Retrieved from UniProt |
| Canonical Name | Common name | Most used in literature > First designation in UniProt |
| Cellular Localization | Subcellular localization differentiates amongst the several organelles in the nucleus and cytoplasm. | GO (cellular component ontology) > Locate [88] (marked with “*”) > Literature > predicted localization by Locate |
| Simplified Cellular Localization | Only differentiates between nucleus, cytoplasm and shuttles between these two compartiments. | 'nucleus' (all nuclear organelles); 'cytoplasm' (all cytoplasmatic organelles) and 'shuttle' between cytoplasm and nucleus. |
| Function | For this attribute we used the UniProt ontology as a simplified version of the GO terms to provide a comprehensible overview. To annotate the network with the complete GO annotations we refer to Cytoscape plugins such as BINGO[89]. | UniProt Ontologies > Panther database [90] |
| Connection with disease | This field is filled in when mutations, knockout or other modifications of the protein resulting in disease. If a MIM number was available it is mentioned, otherwise there is reference to specific publications. This attribute is additional information and does not mean that the PML-NBs are explicitly involved in the disease process. | UniProt (General Annotation) and literature |
| Induction | Information from UniProt and literature about regulation by several (bio)chemical stimuli. | UniProt (General Annotation) and literature |
| SUMO Consensus | We manually searched if any PML-NB components contained a predicted negatively charged amino acid-dependent sumoylation motif (NDSM)[91] or phosphorylation-dependent sumoylation motif (PDSM)[92] and present these in this attribute. Proteins predicted by us are labeled as 'IS Lab'. | Literature |
| SUMO Isoform | Here we provide an overview of the conjugated isoform described in literature. Proteins marked in this attribute with 'pathway' are enzymes or actors of the SUMO conjugation machinery. | Literature |
| Remarks | In the 'Remarks' attribute we provide potentially important details about the protein | Literature and/or UniProt |
| Edge Attributes | ||
| Author(s) | Reference to the first author, year of publication and PubMedID. | PubMed |
| Source | Indication of source where a certain interaction was found. | Literature and/or database (IntAct, HPRD, BIOGRID, MINT and NPD) search |
| Methods | Gives an overview of the methods used to detect the interaction. As a designation for these methods we used the same evidence code as used on BIOGRID. | Literature and/or database (IntAct, HPRD, BIOGRID, MINT and NPD) search |
| Fluorescence Microscopy | Gives details about the conditions for co-localization studies e.g. endogenous levels, overexpression data and or partial overlap (if mentioned in the publication) | Literature |
| Cell type | Cell type used for PML interaction studies. | Literature |
| PML isoform | For PML interactions it is important to make reference of the PML isoform used if this data is available. | Literature |
| Remarks | In the 'Remarks' attribute we provide potentially important details interaction | Literature |
Functional distribution of the PML-NB components. For each functional category the number of nuclear, cytoplasmatic and shuttle proteins are plotted. The categories were based on the keywords in UniProt Ontologies. Several related categories were merged to facilitate interpretation, such as 'intracellular signaling' (merge of Notch signaling, Wnt signaling, signal transduction and intracellular signaling ontologies), 'differentation and growth' (merge of growth, differentiation, muscle development, spermatogenesis and neurogenesis), 'PTM (chemical)' (acetylaction and phosphorylation), 'RNA processes' (RNA splicing, catabolism and processing), nuclear structure (chromatin and centrosome) and miscellaneous (lipid metabolism, oncogene and phagocytosis).
The information embedded in the edges illustrates the diversity of techniques underlying the integrated data that have been used to detect interactions with PML. Affinity capture, in vitro reconstitution of protein complexes and co-localization by immunofluorescence are the most commonly used methods to detect interactors of the PML-NBs. Further research will demonstrate if components found with these methods interact binary with PML. Less frequently used are the two-hybrid methodology or biochemical assays, two techniques which do in fact prove binary interactions with PML.
Of the total number of PML-NB components (166), 59% of the interactions were proven with only one technique. One must therefore keep in mind that false positives may be present in the network.
Experiments were performed in diverse cell types which range from commonly used immortal lines such as HeLa, MCF-7 and HEK293 to primary cell cultures. Using several cell lines gives an overall idea of the composition of the PML-NB which is relevant for integration in systems biology and contributes to the knowledge of PML-NB composition in different cell types. However, this generalization has to be treated with caution. The possibility exists that we adopt certain proteins as part of the complex while this architecture is unique for a certain cell type (e.g. ALT cells). Equally important to create a uniform view of the PML protein complex is the necessity to clearly state -if possible- which isoform was used. Since PML has 6 different splice variants with different kinetics [9], it is expected that these isoforms may (partially) bind other partner proteins. In only one third of the cases the PML isoform was clearly stated. Thus far the most used isoform is PMLIV. Therefore, to determine isoform-dependent differences in PML-NBs it is vital that researchers clearly state which PML variant is used in their experiments whenever possible.
Apart from the direct interactions with PML, we imported the links between the several PML partners from the IntAct database to show distinct biological pathways that meet at the PML-NBs. Hence, next to the direct edges (166) with PML, these 166 partners are interconnected by 615 edges (figure 2).
Upon analyses of the functions of the PML-NB components, a significant number of mostly nuclear proteins seems to be involved in ubiquitin-like modification processes such as SUMOylation and ubiquitination (figure 3, PTM (small protein)). Due to the close relationship between PML SUMOylation and NB formation[10, 11, 94, 95] and because of PML-NBs may act as a platform for post-translational modifications; we explored the possibility if the PML-NBs could be SUMOylation hotspot in the nucleus.
Are PML-NBs SUMOylation hotspots in the nucleus?
PML-NBs are a hub for numerous proteins and are involved in a range of functions. Moreover, the complex is highly dynamic and proteins such as CBP and Sp100 show a dynamic exchange between nuclear bodies and nucleoplasm[96].
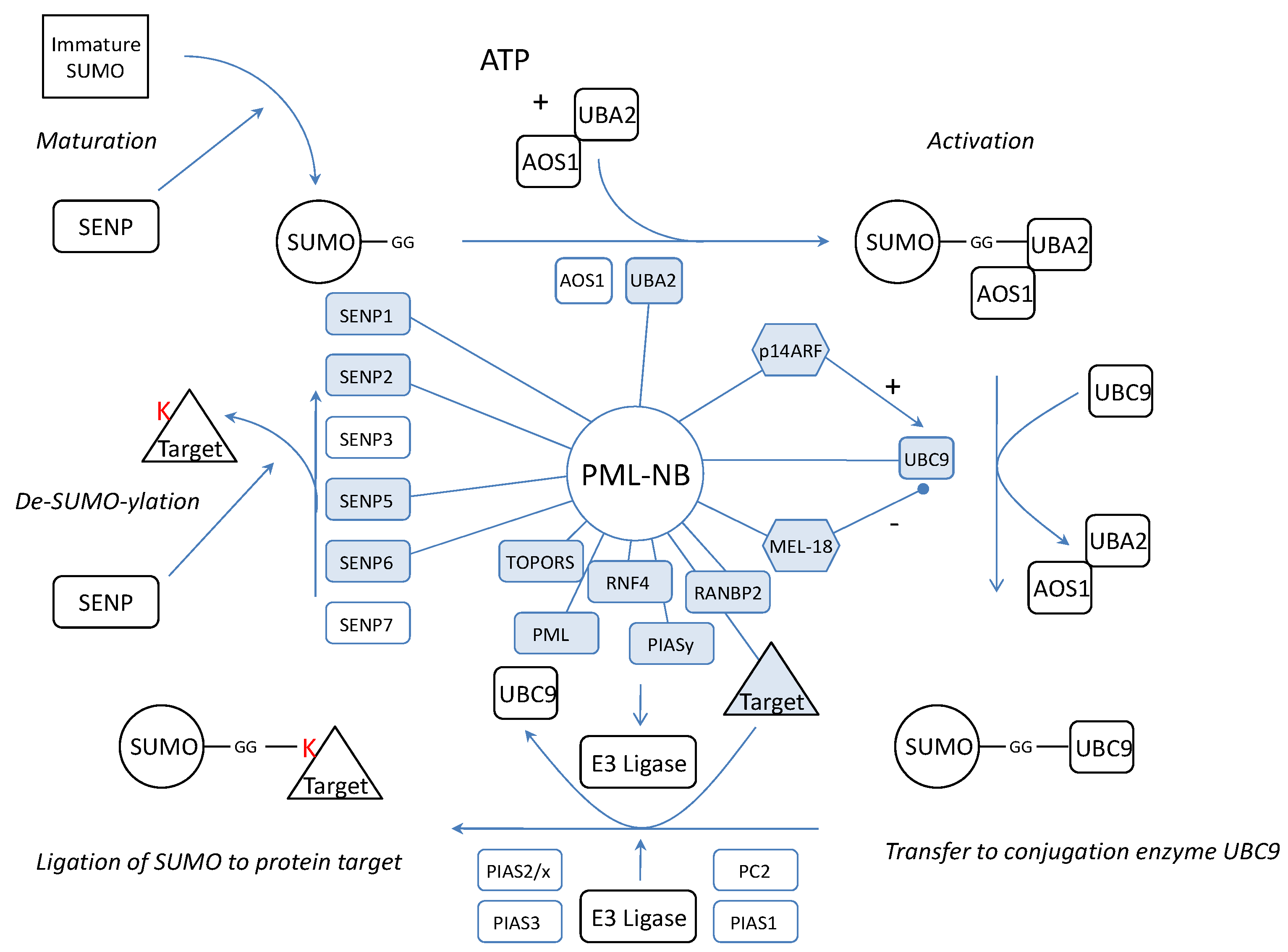
We wondered if the dynamics of the complex would be related to SUMOylation of NB proteins. Thus, the PML-NBs would be a (de)SUMOylation platform where SUMO is coupled or detached to/from its target protein. Possible evidence confirming this hypothesis lies in the observation that a large number of confirmed SUMOylated proteins localizes at the NBs. In fact, apart from SUMO4, all SUMO isoforms are found in the PML-NBs. Furthermore, not only several SENPs were co-localized with PML but also activating enzyme UBA2[97], the only SUMO conjugating enzyme UBC9[98, 99] and several SUMO ligases such as RANBP2 and PIASy [100-103] (Figure 4). One of these, RANBP2, promotes PML SUMOylation [104]. In addition, another PML-NB component, HDAC7, has been described to exert “SUMO E3 ligase-like” activity and also enhances SUMO modification of PML [105]. In fact, the most interesting SUMO E3 ligase at the PML-NBs could be PML itself. Indeed, in yeast, the RING domain of PML is sufficient for PML auto-SUMOylation and SUMOylation of other proteins [106]. Moreover, in transfected mammalian cells, PML RING mutants show a dramatical decrease in PML auto-SUMOylation [107]. In vitro, when purified PML is incubated with HeLa lysate, UBC9 and SUMO1, this leads to auto-SUMOylation [98]. PML was already described to mediate SUMOylation for proteins such as YAP1 [108]. However, one must keep in mind that, although all these observations suggest a SUMO E3 ligase activity for PML, indirect effects (e.g. stimulation or increased expression of enzymes from the SUMOylation machinery) may also be involved and further research is required to define PML as a novel SUMO E3 ligase.
Dynamics between SUMOylation and de-SUMOylation seems important for the architecture and function of the nuclear bodies. In fact, manipulations of SENPs show distinct effects on PML-NBs. It has been shown that IL6 cytokine induces the upregulation of mRNA expression of SENP1. Augmented SENP1 expression causes PML de-SUMOylation which abolishes the suppressor activity of PML on STAT3 [109]. On the other hand, suppression of SENP1 increases the number of SUMOylated proteins and the size of the PML-NBs[110]. SENP2 co-localizes with PML-NBs and PML enhances its effects. SENP2 hydrolyses SUMO1 from PML in vitro and changes the cellular localization of PML protein partners such as CBP, Daxx and p53[111]. Further on, truncated SENP5 co-localizes in the PML-NBs instead of in the nucleolus and both truncated- and wild type-SENP5 show the ability to remove SUMO2 or SUMO3 from PML Lys160 or Lys490[26]. Finally, SENP6 suppression results in the accumulation of SUMO2/3-modified species in PML bodies [29] but SENP6 has not been co-localized with PML yet. These observations suggest that not only PML is influenced by the presence of SENPs but that they also have an effect on other PML-NB targeted proteins.
All the necessary components to attach SUMO to a target are present at the PML-NBs but also the SENPs to remove SUMO again. This means that the complete cycle of SUMOylation and de-SUMOylation could take place at the PML-NBs (figure 4). In fact, PML contains a SUMO binding motif which forms a non-covalent interaction with SUMO attached to protein partners[107]. Possibly these proteins are recruited to make a functional switch at the PML-NBs by detachment of SUMO.
A large amount of PML partners have been described in literature as SUMOylated (38%). However, this percentage may well be an underestimation because SUMOylation of many PML partners has not been investigated yet. Therefore, to have an idea about other possible SUMOylation targets in the network we searched for PML-NB proteins with an negatively charged amino acid-dependent sumoylation motif (NDSM) motif as predicted by Yang et al.[91]. In fact, more than 56% of the PML-NB components contain an NDSM motif of which about half was already found to be SUMOylated in “wet lab” experiments.
Representation of the SUMOylation pathway in relation to the PML-NB. Indicated are the proteins which have been linked to PML (blue, linked with PML). SUMO is synthesized as an immature protein which needs processing by SENPs. SENP1, SENP2 and SENP5 exert processing activity and have been linked to the PML-NBs implying that maturation may take place at the NBs. After maturation, the SUMO protein is activated by two E2 enzymes of which UBA2 was co-localized with the PML-NBs. Also the only conjugating UBC9 can be found in the PML-NBs and molecules which influence UBC9 such as p14ARF and MEL-18 have been targeted to the PML-NBs. For final conjugation of SUMO to a target protein, an E3 ligase is needed and apart from PML which potentially contains E3 activity itself, four other ligases (TOPORS, RNF4, RANBP2, PIASy) have been targeted to the NBs. We propose that the target protein, the E3 ligase and UBC9-SUMO meet at the nuclear bodies for the completion of the SUMO pathway. Finally, SUMOylation can be reversed by SENPs. Only de-conjugation SENP3 and SENP7 have not been linked to the PML-NBs thus far.
In-house prediction of SUMOylation targets
Complementary to existing knowledge of SUMOylation sites, we employed an in-house developed machine learning approach to predict genome-wide SUMOylation sites.
We used the method based on Conditional Random Fields that previously proved successfully to predict kinase specific phosphorylation sites[112]. A 9 residue sequence window surrounding the annotated SUMOylation sites from UniProt was used to train a Conditional Random Field-based model as described in Dang et al. (2008)[112], with a feature function template optimized with a Genetic Algorithm as described in Dang et al. (2009) (in publication). Known SUMOylation sites were extracted from UniProt (downloaded august 18, 2009) based on the sequence feature "CROSSLINK" and a feature description containing the string "SUMO". Feature annotations containing the "probable" or "by similarity" qualifiers were removed. This data was used as training and test data.
The performance of the training step was assessed by k-fold (k=4) cross validations. In four fold cross validation, the protein set is partitioned into four subsets, of these subsets a single subset is retained as validation data for testing the model. The remaining three subsets are used as training data. This process is repeated four times with each of the four subsets used exactly once as the test set.
The resulting Receiver Operator (ROC) curve is shown in figure 5a. Its excellent performance is demonstrated by an Area Under the Curve (AUC) value of 0,91. Also in figure 5 is a Venn-diagram of the overlap of what is described in literature and what is predicted. In general there is a large overlap between our screening method and the search for NDSM motifs[91]. We determine fewer proteins as SUMOylation target compared to the NDSM paper, which is determined by the stringency of our threshold. We chose to include only few insecurities and high confidence in our predictions. Figure 5b demonstrates that the overlap between literature and our approach and the overlap between the NDSM predictions and literature is equal, even though Yang et al. found more unconfirmed targets. Interestingly, there appears to be a portion of targets which were not predicted by our method or the method of Yang et al., suggesting that there are other motifs and surrounding amino acids which characterize a SUMOylation target.
A. Receiver Operator Curve (ROC) of our in-house prediction algorithm with Area Under the Curve (AUC) value of 0,91. B. Overlap between literature, our predictions and the predictions performed by Yang et al.. Due to the different approach there are some unique prediction data for both our prediction method and the predictions performed by Yang et al.. C. Predicted percentages by our in-house algorithm (FPR =1%). Genome-wide we predict about one fourth of the proteins to be SUMOylated. Membrane and extracellular proteins are less likely to be SUMOylation targets then cytoplasmatic or nuclear proteins. Within the PML-NBs we see a clear enrichment of SUMOylatable proteins.
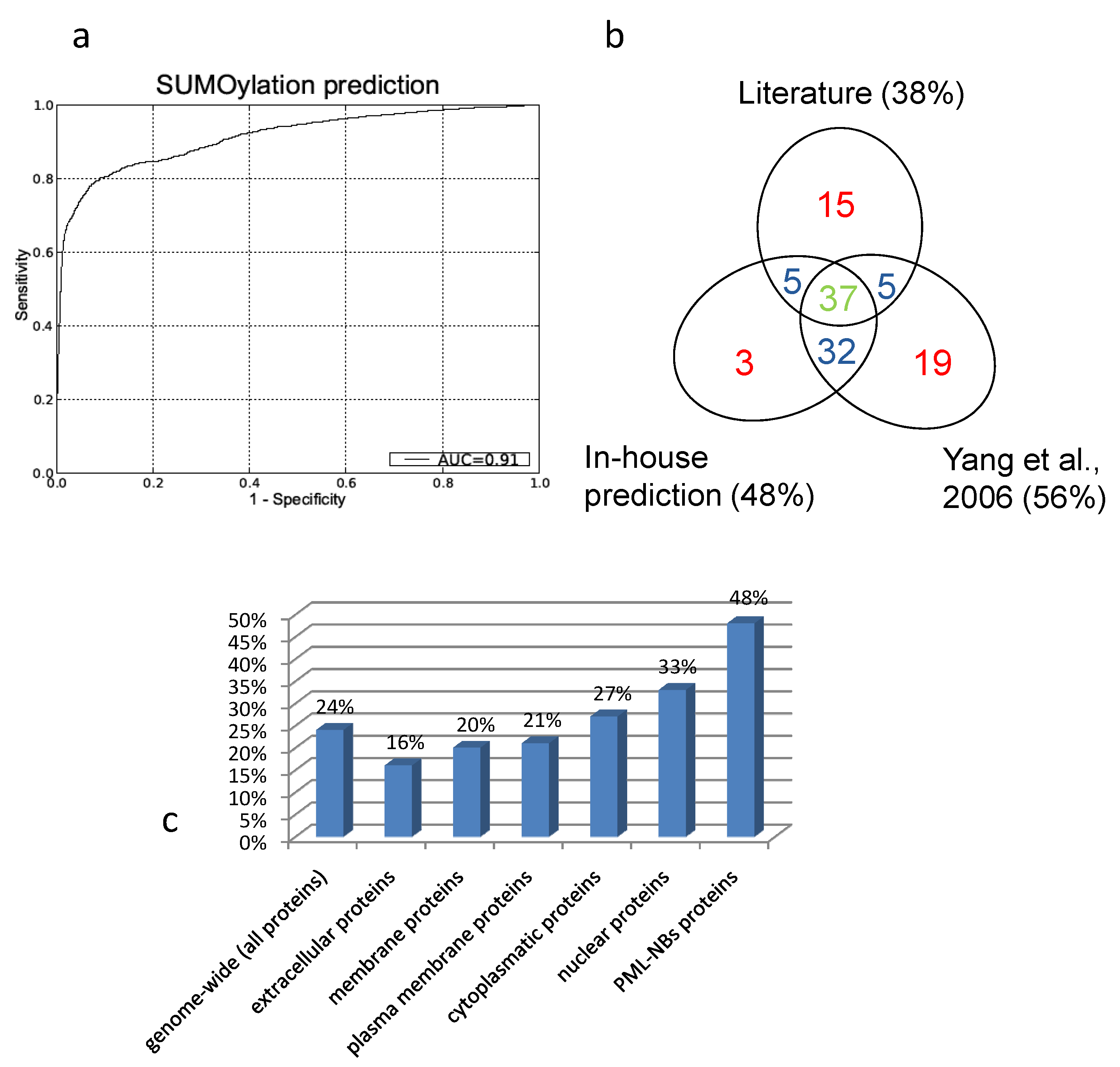
To asses a potential enrichment of SUMOylated proteins at the PML-NBs, percentages of predicted SUMOylated proteins were compared between the full human proteome and the PML-NB.
To accomplish this, the trained model was used to predict potential SUMOylation sites in the full human proteome (using the UniProt sequences), with a predefined maximum allowed false positive rate (FPR) of 1% as described in Dang et al.[112]. Figure 5c gives an overview of the predicted percentages of SUMOylatable proteins in several cellular compartments. Genome wide about 24% of the protein sequences contained at least one predicted potential SUMOylation site. Within proteins annotated with the GO Cellular component terms "cytoplasm" and "nucleus", respectively 26% and 33% were predicted to contain one or more SUMOylation sites. Within the assembled PML network, 48% contained 1 or more predicted SUMOylation motifs, demonstrating a clear overrepresentation of SUMOylation motifs in the PML network that cannot solely be attributed to its nuclear localization.
Since our predictor is trained with all possible known SUMOylation sites, its predictions are not focussing on one particular motif (such as the NDSM motif) but on all possible motifs. Together with the excellent performance demonstrated by the ROC curve and the stringent thresholds applied, we believe that this is the most reliable approach to be able to draw conclusions from the intracellular distribution of predicted SUMOylation sites. The observed discrepancies between our predictor and the NDSM results may, besides algorithmic performance differences, also reflect the diversity of the training sequences and the stringency of the performed prediction.
Conclusion
Dynamic complexes such as the PML-NBs require high-throughput methods as well as specialized reductionist approaches for characterization and identification of all the participating components. As much, the PML community should be cautious for scattered data making it difficult to maintain a general overview. We believe that our PML network helps in giving a comprehensive insight in the vast amount of PML data that has been generated.
Creating this overview focused the attention on a role of the PML-NBs in SUMOylation. The presence of the enzymes for both SUMOylation and de-SUMOylation suggests that both the attachment and detachment of SUMO can take place at the PML-NBs. In fact, the properties of the PML protein such as the RING domain and coiled coil motif are ideal to recruit unmodified proteins which are targets for SUMOylation. Amongst others, PML itself may act as an E3 SUMO ligase. In addition, PML contains a SUMO binding motif that can bind SUMOylated proteins which might be in turn de-SUMOylated.
To estimate if there was an enrichment of possibly SUMOylated proteins at the PML-NBs, independent of cellular localization, we used an in-house prediction method for a genome wide screen for SUMOylatable proteins. We show that 24% of the proteins genome wide is predicted to be SUMOylated. Although in the nucleus this number rises slightly to 33% we found that 48% of the PML-NB components contain one or more possible SUMOylation sites. This suggests that the PML-NBs are enriched sites for SUMOylated proteins.
Taken together the presence of (de)-SUMOylation enzymes combined with an enrichment in SUMOylatable proteins suggests that PML-NBs are a hotspot for nuclear SUMOylation where protein targets meet the components of the (de)-SUMOylation pathway.
The dynamic regulation of key point proteins at the PML-NBs supports this hypothesis since the regulatory function of SUMOylation on PML-NB related processes such as transcription [14, 113, 114], cell cycle[115], apoptosis[116], senescence[117] and viral infection [118-120] is slowly uncovered. As an example we present in table 3 the influence of SUMOylation on PML-NB related transcription factors.
Further, this hypothesis could be the glue between the three previously formulated hypotheses about PML-NB function. Since SUMOylation is often related to a functional switch, it is not unthinkable that by SUMOylation certain protein factors are retained in the nucleus until they are needed and are released by de-SUMOylation or vice versa. This SUMOylation dependent switching could meticulously regulate specific nuclear processes such as transcription and DNA related processes. In fact, our functional analysis reveals a very high number of transcription factors confirming the close relationship between PML-NBs and the modulation of transcription, possibly in a SUMO-dependent manner. Similarly, several protein partners are involved in DNA replication, damage sensing and repair which indicate a role in DNA related processes including epigenetic regulation [67].
The PML-NBs can thus be seen as a nuclear relay station where protein fractions involved in a variety of nuclear processes make a SUMOylation regulated functional switch. In addition, it is important to state that SUMOylation itself is regulated. Increasingly, the crosstalk between SUMOylation and other PTMs such as acteylation and phosphorylation (e.g. via the PDSM SUMOylation motif [92]) is uncovered [121, 122]. Since at the PML-NBs also kinases, acetyltransferases and methyltransferases gather, it is likely that we are just exploring the borders of the regulatory complexity within this protein complex.
Effect of SUMOylation on transcription factors present in the PML-NBs. Several transcription factors have a predicted PDSM motif but have not been shown to be SUMOylated (ARI3A, Ataxin7, FLASH, HDAC2, HHEX, HIRA, LBP1, mTOR, MZF, NCOR1, NR4A1, PAR4, PAX5, SIRT1, SKI, SP140, THAP1 and ZNF24). About the remainder of the transcription factors associated with PML-NBs there is no available data regarding SUMOylation.
| UniProtID | Name | Motif | Residue | Effect of SUMOylation | Positieve/negative Effect (+/-) |
|---|---|---|---|---|---|
| P27540 | ARNT | NDSM | K245 | SUMOylation affects the ability of ARNT to interact with cooperative molecules such as PML[123] | - |
| Q9BYV9 | BACH2 | NDSM | SUMOylation is an important regulatory system for the mobility of the nuclear domains formed by Bach2[124] | + | |
| Q92793 | CBP | NDSM | SUMO modification negatively modulates the transcriptional activity of CREB-binding protein[125] | - | |
| P01100 | C-FOS | NDSM | K299, K257 | SUMOylation down-regulates c-Fos/c-Jun AP-1 dimer activity [126] | - |
| P05412 | C-JUN | NDSM | K257, K 229 | SUMOylation down-regulates c-Fos/c-Jun AP-1 dimer activity [126] | - |
| P10242 | C-MYB | N/PDSM | K499, K523 | SUMO1-ylation regulates the transactivation function of c-Myb[127] | - |
| Q9UER7 | Daxx | K630, K631 | Role of Daxx SUMOylation unknown[125, 128] | ||
| Q15910 | EZH2 | NDSM | Modulation of polycomb repressive complex 2[129] | ||
| P23769 | GATA2 | NDSM | Negative regulation of transcriptional activity[130] | - | |
| Q13547 | HDAC1 | NDSM | K444, K476 | SUMOylation reduces HDAC1-mediated transcriptional repression[131] | - |
| Q86Z02 | HIPK1 | NDSM | SUMOylation induces a cytoplasmic translocation leading to ASK1-JNK activation[132] | + | |
| Q9H2X6 | HIPK2 | NDSM | K25 | SUMOylation inhibits HIPK2-induced JNK activation and p53-independent antiproliferative function[133] | - |
| Q03933 | HSF2 | NDSM | K82 | SUMOylation of HSF2 results in conversion to the active DNA binding form[134] | + |
| Q9UJU2 | LEF1 | SUMOylation induces potent repression of LEF1 activity[135] | - | ||
| Q99607 | MEF/ELF4 | K657 | SUMOylation down-regulates ELF4 activity[136] | - | |
| Q15596 | NCOA2 (human, GRIP1 mouse | NDSM | K239, K731, K788 | SUMOylation of GRIP1 (mouse equivalent of NCOA2) enhances binding to the androgen receptor[137] | + |
| Q9Y618 | NCOR2 | N/PDSM | unknown | ||
| O95644 | NFAT | NDSM | K684, K897 | SUMOylation of K684 is required for NFAT1 transcriptional activity, SUMOylation of K897 is only required for nuclear anchorage[138] | + |
| O00482 | NR5A2 | NDSM | K173,K289 | SUMOylation is associated with transcription repression[139] | - |
| Q8N726 | p14ARF | N/A | Not SUMOylated. Promotes SUMOylation by enhancing UBC9[35] | + | |
| P04637 | p53 | K386 | SUMOylation induces premature senescence and stress response[116] | + | |
| Q9H3D4 | p63 | K637 | SUMOylation has a negative effect on p63 driven transcription[140] | - | |
| O15350 | p73 | K627 | SUMO1-ylation of p73 is involved its proteasome-dependent degradation, subnuclear localization and modulation of interactions with other SUMO1 substrates[116] | - | |
| Q8N2W9 | PIASy | NDSM | K35 | SUMO1-ylation of PIASy is necessary for PIASy-dependent activation of Tcf-4[41] | + |
| Q05516 | PLZF | NDSM | K242, K387, K396 | SUMOylation enhances the transcriptional repression activity, correlating with a loss of PLZF-mediated growth suppression[141] | + |
| P29590 | PML | NDSM | K65, K160, K490 | SUMOylation is important for nuclear localization of PML and nuclear body formation[4] | + |
| P06400 | pRB | NDSM | SUMOylation induces premature senescence and stress response[142] | + | |
| P14373 | RFP/TRIM27 | SUMOylation strengthens the transcriptional repressive activity of RFP[143] | + | ||
| P78317 | RNF4 | N/A | SUMOylation of RNF4 not shown, RNF4 is a SUMO-dependent Ubiquitin ligase for poly-SUMOylated proteins[12] | ||
| Q01826 | SATB1 | NDSM | K744 | SUMOylation enhances caspase cleavage of SATB1[144] | - |
| Q96ST3 | SIN3A | NDSM | SUMOylation regulates tumorsupression of SIN3a[38] | ||
| Q6PEW1 | SIZN | SUMOylation mediates localization in PML-NBs[145] | |||
| P84022 | SMAD3 | SUMOylation affects the DNA-binding activity of Smad3 and induces nuclear export[146] | - | ||
| P08047 | SP1 | NDSM | K16 | SUMOylation inhibits cleavage of Sp1 N-terminal negative regulatory domain and inhibits Sp1-dependent transcription[147] | - |
| P11831 | SRF | NDSM | K147 | Activated SRF is suppressed by its SUMOylation[148] | - |
| Q15022 | SUZ12 | NDSM | K72, K73, K75 | SUMOylation modulates PRC2 repressive activity[129] | +/- |
| Q9NQB0 | TCF4 | NDSM | K297 | SUMOylation is involved in beta-catenin-dependent and Tcf-4-mediated gene expression in the Wnt signaling pathway[41] | + |
| O15164 | TIF1/TRIM24 | NDSM | K690, K708 | unknown | |
| Q92993 | TIP60 | NDSM | K430, K451 | SUMOylation of TIP60 augments its acetyltransferase activity in vitro and in vivo[149] | + |
| P13056 | TR2 | NDSM | SUMOylated TR2 recruits corepressor RIP140 to act as a repressor for its target Oct4[150] | - | |
| O95551 | TTRAP | Unknown | |||
| Q9UBW7 | ZNF198 | K963 | SUMOylation of ZNF198 is important for PML body formation[151] | + | |
| Q9Y4E5 | ZNF451 | NDSM | ZNF451 exerts its effects via SUMOylation and trafficking of transcription regulators between promyelocytic leukemia bodies and nucleoplasm[152] |
Acknowledgements
This work was supported by an NOI grant (BOF-UA-2005) from the University of Antwerp.
Research by EVD is funded by a Ph.D grant of the Institute for the Promotion of Innovation through Science and Technology in Flanders (IWT-Vlaanderen). KL and DTH are supported by SBO grant (IWT-600450) of the Flemish Institute supporting Scientific—Technological Research in industry (IWT).
We apologize to the authors whose work has been overseen in this publication. We welcome comments, corrections and additions to the network by any reader or PML-enthusiast. Please email your contributions to ellen.vandamme@ua.ac.be.
Conflict of Interest
The authors have declared that no conflict of interest exists.
References
1. de THE H, RIVIERE M, BERNHARD W. [Examination by electron microscope of the VX2 tumor of the domestic rabbit derived from the Shope papilloma.]. Bull Assoc Fr Etud Cancer. 1960Oct;47:570-84
2. HINGLAIS-GUILLAUD N, MORICARD R, BERNHARD W. [Ultrastructure of invasive pavement-cell cancers of the uterine cervix in women.]. Bull Assoc Fr Etud Cancer. 1961Apr;48:283-316
3. Jensen K, Shiels C, Freemont PS. PML protein isoforms and the RBCC/TRIM motif. Oncogene. 2001Oct29;20(49):7223-33
4. Ishov AM, Sotnikov AG, Negorev D, Vladimirova OV, Neff N, Kamitani T. et al. PML is critical for ND10 formation and recruits the PML-interacting protein daxx to this nuclear structure when modified by SUMO-1. J Cell Biol. 1999Oct18;147(2):221-34
5. Borden KL. RING fingers and B-boxes: zinc-binding protein-protein interaction domains. Biochem Cell Biol. 1998;76(2-3):351-8
6. Kentsis A, Dwyer EC, Perez JM, Sharma M, Chen A, Pan ZQ. et al. The RING domains of the promyelocytic leukemia protein PML and the arenaviral protein Z repress translation by directly inhibiting translation initiation factor eIF4E. J Mol Biol. 2001Sep28;312(4):609-23
7. Reymond A, Meroni G, Fantozzi A, Merla G, Cairo S, Luzi L. et al. The tripartite motif family identifies cell compartments. EMBO J. 2001May1;20(9):2140-51
8. Boisvert FM, Kruhlak MJ, Box AK, Hendzel MJ, Bazett-Jones DP. The transcription coactivator CBP is a dynamic component of the promyelocytic leukemia nuclear body. J Cell Biol. 2001Mar5;152(5):1099-106
9. Weidtkamp-Peters S, Lenser T, Negorev D, Gerstner N, Hofmann TG, Schwanitz G. et al. Dynamics of component exchange at PML nuclear bodies. J Cell Sci. 2008Aug15;121(Pt 16):2731-43
10. Fu C, Ahmed K, Ding H, Ding X, Lan J, Yang Z. et al. Stabilization of PML nuclear localization by conjugation and oligomerization of SUMO-3. Oncogene. 2005Aug18;24(35):5401-13
11. Lallemand-Breitenbach V, Zhu J, Puvion F, Koken M, Honore N, Doubeikovsky A. et al. Role of promyelocytic leukemia (PML) sumolation in nuclear body formation, 11S proteasome recruitment, and As2O3-induced PML or PML/retinoic acid receptor alpha degradation. J Exp Med. 2001Jun18;193(12):1361-71
12. Lallemand-Breitenbach V, Jeanne M, Benhenda S, Nasr R, Lei M, Peres L. et al. Arsenic degrades PML or PML-RARalpha through a SUMO-triggered RNF4/ubiquitin-mediated pathway. Nat Cell Biol. 2008May;10(5):547-55
13. Vertegaal AC. Small ubiquitin-related modifiers in chains. Biochem Soc Trans. 2007Dec;35(Pt 6):1422-3
14. Hay RT. SUMO: a history of modification. Mol Cell. 2005Apr1;18(1):1-12
15. Melchior F, Schergaut M, Pichler A. SUMO: ligases, isopeptidases and nuclear pores. Trends Biochem Sci. 2003Nov;28(11):612-8
16. Hamard PJ, Boyer-Guittaut M, Camuzeaux B, Dujardin D, Hauss C, Oelgeschlager T. et al. Sumoylation delays the ATF7 transcription factor subcellular localization and inhibits its transcriptional activity. Nucleic Acids Res. 2007;35(4):1134-44
17. Huen MS, Chen J. The DNA damage response pathways: at the crossroad of protein modifications. Cell Res. 2008Jan;18(1):8-16
18. Li T, Santockyte R, Shen RF, Tekle E, Wang G, Yang DC. et al. Expression of SUMO-2/3 induced senescence through p53- and pRB-mediated pathways. J Biol Chem. 2006Nov24;281(47):36221-7
19. Wilson VG, Rangasamy D. Intracellular targeting of proteins by sumoylation. Exp Cell Res. 2001Nov15;271(1):57-65
20. Mukhopadhyay D, Dasso M. Modification in reverse: the SUMO proteases. Trends Biochem Sci. 2007Jun;32(6):286-95
21. Xu Z, Au SW. Mapping residues of SUMO precursors essential in differential maturation by SUMO-specific protease, SENP1. Biochem J. 2005Mar1;386(Pt 2):325-30
22. Shen LN, Dong C, Liu H, Naismith JH, Hay RT. The structure of SENP1-SUMO-2 complex suggests a structural basis for discrimination between SUMO paralogues during processing. Biochem J. 2006Jul15;397(2):279-88
23. Shen L, Tatham MH, Dong C, Zagorska A, Naismith JH, Hay RT. SUMO protease SENP1 induces isomerization of the scissile peptide bond. Nat Struct Mol Biol. 2006Dec;13(12):1069-77
24. Reverter D, Lima CD. A basis for SUMO protease specificity provided by analysis of human Senp2 and a Senp2-SUMO complex. Structure. 2004Aug;12(8):1519-31
25. Reverter D, Lima CD. Structural basis for SENP2 protease interactions with SUMO precursors and conjugated substrates. Nat Struct Mol Biol. 2006Dec;13(12):1060-8
26. Gong L, Yeh ET. Characterization of a family of nucleolar SUMO-specific proteases with preference for SUMO-2 or SUMO-3. J Biol Chem. 2006Jun9;281(23):15869-77
27. Kuo HY, Chang CC, Jeng JC, Hu HM, Lin DY, Maul GG. et al. SUMO modification negatively modulates the transcriptional activity of CREB-binding protein via the recruitment of Daxx. Proc Natl Acad Sci U S A. 2005Nov22;102(47):16973-8
28. Shen LN, Geoffroy MC, Jaffray EG, Hay RT. Characterization of SENP7, a SUMO-2/3-specific isopeptidase. Biochem J. 2009Jul15;421(2):223-30
29. Mukhopadhyay D, Ayaydin F, Kolli N, Tan SH, Anan T, Kametaka A. et al. SUSP1 antagonizes formation of highly SUMO2/3-conjugated species. J Cell Biol. 2006Sep25;174(7):939-49
30. Johnson ES, Schwienhorst I, Dohmen RJ, Blobel G. The ubiquitin-like protein Smt3p is activated for conjugation to other proteins by an Aos1p/Uba2p heterodimer. EMBO J. 1997Sep15;16(18):5509-19
31. Okuma T, Honda R, Ichikawa G, Tsumagari N, Yasuda H. In vitro SUMO-1 modification requires two enzymatic steps, E1 and E2. Biochem Biophys Res Commun. 1999Jan27;254(3):693-8
32. Desterro JM, Thomson J, Hay RT. Ubch9 conjugates SUMO but not ubiquitin. FEBS Lett. 1997Nov17;417(3):297-300
33. Rodriguez MS, Dargemont C, Hay RT. SUMO-1 conjugation in vivo requires both a consensus modification motif and nuclear targeting. J Biol Chem. 2001Apr20;276(16):12654-9
34. Gong L, Kamitani T, Fujise K, Caskey LS, Yeh ET. Preferential interaction of sentrin with a ubiquitin-conjugating enzyme, Ubc9. J Biol Chem. 1997Nov7;272(45):28198-201
35. Rizos H, Woodruff S, Kefford RF. p14ARF interacts with the SUMO-conjugating enzyme Ubc9 and promotes the sumoylation of its binding partners. Cell Cycle. 2005Apr;4(4):597-603
36. Pichler A, Gast A, Seeler JS, Dejean A, Melchior F. The nucleoporin RanBP2 has SUMO1 E3 ligase activity. Cell. 2002Jan11;108(1):109-20
37. Weger S, Hammer E, Heilbronn R. Topors acts as a SUMO-1 E3 ligase for p53 in vitro and in vivo. FEBS Lett. 2005Sep12;579(22):5007-12
38. Pungaliya P, Kulkarni D, Park HJ, Marshall H, Zheng H, Lackland H. et al. TOPORS functions as a SUMO-1 E3 ligase for chromatin-modifying proteins. J Proteome Res. 2007Oct;6(10):3918-23
39. Yamamoto H, Ihara M, Matsuura Y, Kikuchi A. Sumoylation is involved in beta-catenin-dependent activation of Tcf-4. EMBO J. 2003May1;22(9):2047-59
40. Sachdev S, Bruhn L, Sieber H, Pichler A, Melchior F, Grosschedl R. PIASy, a nuclear matrix-associated SUMO E3 ligase, represses LEF1 activity by sequestration into nuclear bodies. Genes Dev. 2001Dec1;15(23):3088-103
41. Ihara M, Yamamoto H, Kikuchi A. SUMO-1 modification of PIASy, an E3 ligase, is necessary for PIASy-dependent activation of Tcf-4. Mol Cell Biol. 2005May;25(9):3506-18
42. Kagey MH, Melhuish TA, Wotton D. The polycomb protein Pc2 is a SUMO E3. Cell. 2003Apr4;113(1):127-37
43. Tatham MH, Geoffroy MC, Shen L, Plechanovova A, Hattersley N, Jaffray EG. et al. RNF4 is a poly-SUMO-specific E3 ubiquitin ligase required for arsenic-induced PML degradation. Nat Cell Biol. 2008May;10(5):538-46
44. Zhang J, Goodson ML, Hong Y, Sarge KD. MEL-18 interacts with HSF2 and the SUMO E2 UBC9 to inhibit HSF2 sumoylation. J Biol Chem. 2008Mar21;283(12):7464-9
45. Conlan LA, McNees CJ, Heierhorst J. Proteasome-dependent dispersal of PML nuclear bodies in response to alkylating DNA damage. Oncogene. 2004Jan8;23(1):307-10
46. Lijima K, Komatsu K, Matsuura S, Tauchi H. The Nijmegen breakage syndrome gene and its role in genome stability. Chromosoma. 2004Sep;113(2):53-61
47. Bernardi R, Papa A, Pandolfi PP. Regulation of apoptosis by PML and the PML-NBs. Oncogene. 2008Oct20;27(48):6299-312
48. Krieghoff-Henning E, Hofmann TG. Role of nuclear bodies in apoptosis signalling. Biochim Biophys Acta. 2008Nov;1783(11):2185-94
49. Everett RD, Earnshaw WC, Pluta AF, Sternsdorf T, Ainsztein AM, Carmena M. et al. A dynamic connection between centromeres and ND10 proteins. J Cell Sci. 1999Oct;112( Pt 20):3443-54
50. Ferbeyre G. PML a target of translocations in APL is a regulator of cellular senescence. Leukemia. 2002Oct;16(10):1918-26
51. Mallette FA, Goumard S, Gaumont-Leclerc MF, Moiseeva O, Ferbeyre G. Human fibroblasts require the Rb family of tumor suppressors, but not p53, for PML-induced senescence. Oncogene. 2004Jan8;23(1):91-9
52. Lai HK, Borden KL. The promyelocytic leukemia (PML) protein suppresses cyclin D1 protein production by altering the nuclear cytoplasmic distribution of cyclin D1 mRNA. Oncogene. 2000Mar23;19(13):1623-34
53. Shima Y, Shima T, Chiba T, Irimura T, Pandolfi PP, Kitabayashi I. PML activates transcription by protecting HIPK2 and p300 from SCFFbx3-mediated degradation. Mol Cell Biol. 2008Dec;28(23):7126-38
54. de The H, Lavau C, Marchio A, Chomienne C, Degos L, Dejean A. The PML-RAR alpha fusion mRNA generated by the t(15;17) translocation in acute promyelocytic leukemia encodes a functionally altered RAR. Cell. 1991Aug23;66(4):675-84
55. Chelbi-Alix MK, Quignon F, Pelicano L, Koken MH, de TH. Resistance to virus infection conferred by the interferon-induced promyelocytic leukemia protein. J Virol. 1998Feb;72(2):1043-51
56. Espert L, Degols G, Gongora C, Blondel D, Williams BR, Silverman RH. et al. ISG20, a new interferon-induced RNase specific for single-stranded RNA, defines an alternative antiviral pathway against RNA genomic viruses. J Biol Chem. 2003May2;278(18):16151-8
57. Wang ZG, Delva L, Gaboli M, Rivi R, Giorgio M, Cordon-Cardo C. et al. Role of PML in cell growth and the retinoic acid pathway. Science. 1998Mar6;279(5356):1547-51
58. Negorev D, Maul GG. Cellular proteins localized at and interacting within ND10/PML nuclear bodies/PODs suggest functions of a nuclear depot. Oncogene. 2001Oct29;20(49):7234-42
59. Boisvert FM, Hendzel MJ, Bazett-Jones DP. Promyelocytic leukemia (PML) nuclear bodies are protein structures that do not accumulate RNA. J Cell Biol. 2000Jan24;148(2):283-92
60. Wang J, Shiels C, Sasieni P, Wu PJ, Islam SA, Freemont PS. et al. Promyelocytic leukemia nuclear bodies associate with transcriptionally active genomic regions. J Cell Biol. 2004Feb16;164(4):515-26
61. Kiesslich A, von MA, Hemmerich P. Cell cycle-dependent association of PML bodies with sites of active transcription in nuclei of mammalian cells. J Struct Biol. 2002Oct;140(1-3):167-79
62. Ching RW, Dellaire G, Eskiw CH, Bazett-Jones DP. PML bodies: a meeting place for genomic loci? J Cell Sci. 2005Mar1;118(Pt 5):847-54
63. Fogal V, Gostissa M, Sandy P, Zacchi P, Sternsdorf T, Jensen K. et al. Regulation of p53 activity in nuclear bodies by a specific PML isoform. EMBO J. 2000Nov15;19(22):6185-95
64. Kurki S, Latonen L, Laiho M. Cellular stress and DNA damage invoke temporally distinct Mdm2, p53 and PML complexes and damage-specific nuclear relocalization. J Cell Sci. 2003Oct1;116(Pt 19):3917-25
65. Pearson M, Carbone R, Sebastiani C, Cioce M, Fagioli M, Saito S. et al. PML regulates p53 acetylation and premature senescence induced by oncogenic Ras. Nature. 2000Jul13;406(6792):207-10
66. Szostecki C, Guldner HH, Netter HJ, Will H. Isolation and characterization of cDNA encoding a human nuclear antigen predominantly recognized by autoantibodies from patients with primary biliary cirrhosis. J Immunol. 1990Dec15;145(12):4338-47
67. Torok D, Ching RW, Bazett-Jones DP. PML nuclear bodies as sites of epigenetic regulation. Front Biosci. 2009;14:1325-36
68. Moller A, Sirma H, Hofmann TG, Rueffer S, Klimczak E, Droge W. et al. PML is required for homeodomain-interacting protein kinase 2 (HIPK2)-mediated p53 phosphorylation and cell cycle arrest but is dispensable for the formation of HIPK domains. Cancer Res. 2003Aug1;63(15):4310-4
69. Yang S, Jeong JH, Brown AL, Lee CH, Pandolfi PP, Chung JH. et al. Promyelocytic leukemia activates Chk2 by mediating Chk2 autophosphorylation. J Biol Chem. 2006Sep8;281(36):26645-54
70. Langley E, Pearson M, Faretta M, Bauer UM, Frye RA, Minucci S. et al. Human SIR2 deacetylates p53 and antagonizes PML/p53-induced cellular senescence. EMBO J. 2002May15;21(10):2383-96
71. Zhong S, Muller S, Ronchetti S, Freemont PS, Dejean A, Pandolfi PP. Role of SUMO-1-modified PML in nuclear body formation. Blood. 2000May1;95(9):2748-52
72. Kerrien S, am-Faruque Y, Aranda B, Bancarz I, Bridge A, Derow C. et al. IntAct--open source resource for molecular interaction data. Nucleic Acids Res. 2007Jan;35(Database issue):D561-D565
73. Zanzoni A, Montecchi-Palazzi L, Quondam M, Ausiello G, Helmer-Citterich M, Cesareni G. MINT: a Molecular INTeraction database. FEBS Lett. 2002Feb20;513(1):135-40
74. Stark C, Breitkreutz BJ, Reguly T, Boucher L, Breitkreutz A, Tyers M. BioGRID: a general repository for interaction datasets. Nucleic Acids Res. 2006Jan1;34(Database issue):D535-D539
75. Peri S, Navarro JD, Amanchy R, Kristiansen TZ, Jonnalagadda CK, Surendranath V. et al. Development of human protein reference database as an initial platform for approaching systems biology in humans. Genome Res. 2003Oct;13(10):2363-71
76. Dellaire G, Farrall R, Bickmore WA. The Nuclear Protein Database (NPD): sub-nuclear localisation and functional annotation of the nuclear proteome. Nucleic Acids Res. 2003Jan1;31(1):328-30
77. Shannon P, Markiel A, Ozier O, Baliga NS, Wang JT, Ramage D. et al. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res. 2003Nov;13(11):2498-504
78. Cerami EG, Bader GD, Gross BE, Sander C. cPath: open source software for collecting, storing, and querying biological pathways. BMC Bioinformatics. 2006;7:497
79. Yeager TR, Neumann AA, Englezou A, Huschtscha LI, Noble JR, Reddel RR. Telomerase-negative immortalized human cells contain a novel type of promyelocytic leukemia (PML) body. Cancer Res. 1999Sep1;59(17):4175-9
80. Grobelny JV, Godwin AK, Broccoli D. ALT-associated PML bodies are present in viable cells and are enriched in cells in the G(2)/M phase of the cell cycle. J Cell Sci. 2000;113(Pt 24):4577-85
81. Wu G, Lee WH, Chen PL. NBS1 and TRF1 colocalize at promyelocytic leukemia bodies during late S/G2 phases in immortalized telomerase-negative cells. Implication of NBS1 in alternative lengthening of telomeres. J Biol Chem. 2000Sep29;275(39):30618-22
82. Wu G, Jiang X, Lee WH, Chen PL. Assembly of functional ALT-associated promyelocytic leukemia bodies requires Nijmegen Breakage Syndrome 1. Cancer Res. 2003May15;63(10):2589-95
83. Nabetani A, Yokoyama O, Ishikawa F. Localization of hRad9, hHus1, hRad1, and hRad17 and caffeine-sensitive DNA replication at the alternative lengthening of telomeres-associated promyelocytic leukemia body. J Biol Chem. 2004Jun11;279(24):25849-57
84. Lombard DB, Guarente L. Nijmegen breakage syndrome disease protein and MRE11 at PML nuclear bodies and meiotic telomeres. Cancer Res. 2000May1;60(9):2331-4
85. Jiang WQ, Zhong ZH, Henson JD, Reddel RR. Identification of candidate alternative lengthening of telomeres genes by methionine restriction and RNA interference. Oncogene. 2007Jul12;26(32):4635-47
86. Stagno DM, Mendez-Bermudez A, Foxon JL, Royle NJ, Salomoni P. Lack of TRF2 in ALT cells causes PML-dependent p53 activation and loss of telomeric DNA. J Cell Biol. 2007Dec3;179(5):855-67
87. Apweiler R, Bairoch A, Wu CH, Barker WC, Boeckmann B, Ferro S. et al. UniProt: the Universal Protein knowledgebase. Nucleic Acids Res. 2004Jan1;32(Database issue):D115-D119
88. Sprenger J, Lynn FJ, Karunaratne S, Hanson K, Hamilton NA, Teasdale RD. LOCATE: a mammalian protein subcellular localization database. Nucleic Acids Res. 2008Jan;36(Database issue):D230-D233
89. Maere S, Heymans K, Kuiper M. BiNGO: a Cytoscape plugin to assess overrepresentation of gene ontology categories in biological networks. Bioinformatics. 2005Aug15;21(16):3448-9
90. Thomas PD, Kejariwal A, Campbell MJ, Mi H, Diemer K, Guo N. et al. PANTHER: a browsable database of gene products organized by biological function, using curated protein family and subfamily classification. Nucleic Acids Res. 2003Jan1;31(1):334-41
91. Yang SH, Galanis A, Witty J, Sharrocks AD. An extended consensus motif enhances the specificity of substrate modification by SUMO. EMBO J. 2006Nov1;25(21):5083-93
92. Hietakangas V, Anckar J, Blomster HA, Fujimoto M, Palvimo JJ, Nakai A. et al. PDSM, a motif for phosphorylation-dependent SUMO modification. Proc Natl Acad Sci U S A. 2006Jan3;103(1):45-50
93. Tavalai N, Stamminger T. New insights into the role of the subnuclear structure ND10 for viral infection. Biochim Biophys Acta. 2008Nov;1783(11):2207-21
94. Heun P. SUMOrganization of the nucleus. Curr Opin Cell Biol. 2007Jun;19(3):350-5
95. Maul GG, Negorev D, Bell P, Ishov AM. Review: properties and assembly mechanisms of ND10, PML bodies, or PODs. J Struct Biol. 2000Apr;129(2-3):278-87
96. Boisvert FM, Kruhlak MJ, Box AK, Hendzel MJ, Bazett-Jones DP. The transcription coactivator CBP is a dynamic component of the promyelocytic leukemia nuclear body. J Cell Biol. 2001Mar5;152(5):1099-106
97. Lemos TA, Kobarg J. CGI-55 interacts with nuclear proteins and co-localizes to p80-coilin positive-coiled bodies in the nucleus. Cell Biochem Biophys. 2006;44(3):463-74
98. Duprez E, Saurin AJ, Desterro JM, Lallemand-Breitenbach V, Howe K, Boddy MN. et al. SUMO-1 modification of the acute promyelocytic leukaemia protein PML: implications for nuclear localisation. J Cell Sci. 1999Feb;112( Pt 3):381-93
99. Knipscheer P, Flotho A, Klug H, Olsen JV, van Dijk WJ, Fish A. et al. Ubc9 sumoylation regulates SUMO target discrimination. Mol Cell. 2008Aug8;31(3):371-82
100. Tatham MH, Kim S, Jaffray E, Song J, Chen Y, Hay RT. Unique binding interactions among Ubc9, SUMO and RanBP2 reveal a mechanism for SUMO paralog selection. Nat Struct Mol Biol. 2005Jan;12(1):67-74
101. Rasheed ZA, Saleem A, Ravee Y, Pandolfi PP, Rubin EH. The topoisomerase I-binding RING protein, topors, is associated with promyelocytic leukemia nuclear bodies. Exp Cell Res. 2002Jul15;277(2):152-60
102. Yamamoto H, Ihara M, Matsuura Y, Kikuchi A. Sumoylation is involved in beta-catenin-dependent activation of Tcf-4. EMBO J. 2003May1;22(9):2047-59
103. Ihara M, Yamamoto H, Kikuchi A. SUMO-1 modification of PIASy, an E3 ligase, is necessary for PIASy-dependent activation of Tcf-4. Mol Cell Biol. 2005May;25(9):3506-18
104. Saitoh N, Uchimura Y, Tachibana T, Sugahara S, Saitoh H, Nakao M. In situ SUMOylation analysis reveals a modulatory role of RanBP2 in the nuclear rim and PML bodies. Exp Cell Res. 2006May1;312(8):1418-30
105. Gao C, Ho CC, Reineke E, Lam M, Cheng X, Stanya KJ. et al. Histone deacetylase 7 promotes PML sumoylation and is essential for PML nuclear body formation. Mol Cell Biol. 2008Sep;28(18):5658-67
106. Quimby BB, Yong-Gonzalez V, Anan T, Strunnikov AV, Dasso M. The promyelocytic leukemia protein stimulates SUMO conjugation in yeast. Oncogene. 2006May18;25(21):2999-3005
107. Shen TH, Lin HK, Scaglioni PP, Yung TM, Pandolfi PP. The mechanisms of PML-nuclear body formation. Mol Cell. 2006Nov3;24(3):331-9
108. Lapi E, Di AS, Donzelli S, Gal H, Domany E, Rechavi G. et al. PML, YAP, and p73 are components of a proapoptotic autoregulatory feedback loop. Mol Cell. 2008Dec26;32(6):803-14
109. Ohbayashi N, Kawakami S, Muromoto R, Togi S, Ikeda O, Kamitani S. et al. The IL-6 family of cytokines modulates STAT3 activation by desumoylation of PML through SENP1 induction. Biochem Biophys Res Commun. 2008Jul11;371(4):823-8
110. Yates KE, Korbel GA, Shtutman M, Roninson IB, DiMaio D. Repression of the SUMO-specific protease Senp1 induces p53-dependent premature senescence in normal human fibroblasts. Aging Cell. 2008Oct;7(5):609-21
111. Best JL, Ganiatsas S, Agarwal S, Changou A, Salomoni P, Shirihai O. et al. SUMO-1 protease-1 regulates gene transcription through PML. Mol Cell. 2002Oct;10(4):843-55
112. Dang TH, Van LK, Verschoren A, Laukens K. Prediction of kinase-specific phosphorylation sites using conditional random fields. Bioinformatics. 2008Dec15;24(24):2857-64
113. Gill G. SUMO and ubiquitin in the nucleus: different functions, similar mechanisms? Genes Dev. 2004Sep1;18(17):2046-59
114. Lyst MJ, Stancheva I. A role for SUMO modification in transcriptional repression and activation. Biochem Soc Trans. 2007Dec;35(Pt 6):1389-92
115. Di BA, Gill G. SUMO-specific proteases and the cell cycle. An essential role for SENP5 in cell proliferation. Cell Cycle. 2006Oct;5(20):2310-3
116. Stehmeier P, Muller S. Regulation of p53 family members by the ubiquitin-like SUMO system. DNA Repair (Amst). 2009Apr5;8(4):491-8
117. Bischof O, Dejean A. SUMO is growing senescent. Cell Cycle. 2007Mar15;6(6):677-81
118. Boggio R, Chiocca S. Viruses and sumoylation: recent highlights. Curr Opin Microbiol. 2006Aug;9(4):430-6
119. Muller S, Dejean A. Viral immediate-early proteins abrogate the modification by SUMO-1 of PML and Sp100 proteins, correlating with nuclear body disruption. J Virol. 1999Jun;73(6):5137-43
120. Parkinson J, Everett RD. Alphaherpesvirus proteins related to herpes simplex virus type 1 ICP0 affect cellular structures and proteins. J Virol. 2000Nov;74(21):10006-17
121. Liu B, Shuai K. Regulation of the sumoylation system in gene expression. Curr Opin Cell Biol. 2008Jun;20(3):288-93
122. Anckar J, Sistonen L. SUMO: getting it on. Biochem Soc Trans. 2007Dec;35(Pt 6):1409-13
123. Oshima M, Mimura J, Sekine H, Okawa H, Fujii-Kuriyama Y. SUMO modification regulates the transcriptional repressor function of aryl hydrocarbon receptor repressor. J Biol Chem. 2009Apr24;284(17):11017-26
124. Kono K, Harano Y, Hoshino H, Kobayashi M, Bazett-Jones DP, Muto A. et al. The mobility of Bach2 nuclear foci is regulated by SUMO-1 modification. Exp Cell Res. 2008Feb15;314(4):903-13
125. Kuo HY, Chang CC, Jeng JC, Hu HM, Lin DY, Maul GG. et al. SUMO modification negatively modulates the transcriptional activity of CREB-binding protein via the recruitment of Daxx. Proc Natl Acad Sci U S A. 2005Nov22;102(47):16973-8
126. Bossis G, Malnou CE, Farras R, Andermarcher E, Hipskind R, Rodriguez M. et al. Down-regulation of c-Fos/c-Jun AP-1 dimer activity by sumoylation. Mol Cell Biol. 2005Aug;25(16):6964-79
127. Bies J, Markus J, Wolff L. Covalent attachment of the SUMO-1 protein to the negative regulatory domain of the c-Myb transcription factor modifies its stability and transactivation capacity. J Biol Chem. 2002Mar15;277(11):8999-9009
128. Lin DY, Huang YS, Jeng JC, Kuo HY, Chang CC, Chao TT. et al. Role of SUMO-interacting motif in Daxx SUMO modification, subnuclear localization, and repression of sumoylated transcription factors. Mol Cell. 2006Nov3;24(3):341-54
129. Riising EM, Boggio R, Chiocca S, Helin K, Pasini D. The polycomb repressive complex 2 is a potential target of SUMO modifications. PLoS ONE. 2008;3(7):e2704
130. Chun TH, Itoh H, Subramanian L, Iniguez-Lluhi JA, Nakao K. Modification of GATA-2 transcriptional activity in endothelial cells by the SUMO E3 ligase PIASy. Circ Res. 2003Jun13;92(11):1201-8
131. David G, Neptune MA, DePinho RA. SUMO-1 modification of histone deacetylase 1 (HDAC1) modulates its biological activities. J Biol Chem. 2002Jun28;277(26):23658-63
132. Li X, Luo Y, Yu L, Lin Y, Luo D, Zhang H. et al. SENP1 mediates TNF-induced desumoylation and cytoplasmic translocation of HIPK1 to enhance ASK1-dependent apoptosis. Cell Death Differ. 2008Apr;15(4):739-50
133. Hofmann TG, Jaffray E, Stollberg N, Hay RT, Will H. Regulation of homeodomain-interacting protein kinase 2 (HIPK2) effector function through dynamic small ubiquitin-related modifier-1 (SUMO-1) modification. J Biol Chem. 2005Aug12;280(32):29224-32
134. Goodson ML, Hong Y, Rogers R, Matunis MJ, Park-Sarge OK, Sarge KD. Sumo-1 modification regulates the DNA binding activity of heat shock transcription factor 2, a promyelocytic leukemia nuclear body associated transcription factor. J Biol Chem. 2001May25;276(21):18513-8
135. Sachdev S, Bruhn L, Sieber H, Pichler A, Melchior F, Grosschedl R. PIASy, a nuclear matrix-associated SUMO E3 ligase, represses LEF1 activity by sequestration into nuclear bodies. Genes Dev. 2001Dec1;15(23):3088-103
136. Suico MA, Nakamura H, Lu Z, Saitoh H, Shuto T, Nakao M. et al. SUMO down-regulates the activity of Elf4/myeloid Elf-1-like factor. Biochem Biophys Res Commun. 2006Sep29;348(3):880-8
137. Kotaja N, Karvonen U, Janne OA, Palvimo JJ. The nuclear receptor interaction domain of GRIP1 is modulated by covalent attachment of SUMO-1. J Biol Chem. 2002Aug16;277(33):30283-8
138. Terui Y, Saad N, Jia S, McKeon F, Yuan J. Dual role of sumoylation in the nuclear localization and transcriptional activation of NFAT1. J Biol Chem. 2004Jul2;279(27):28257-65
139. Yang FM, Pan CT, Tsai HM, Chiu TW, Wu ML, Hu MC. Liver receptor homolog-1 localization in the nuclear body is regulated by sumoylation and cAMP signaling in rat granulosa cells. FEBS J. 2009Jan;276(2):425-36
140. Ghioni P, D'Alessandra Y, Mansueto G, Jaffray E, Hay RT, La MG. et al. The protein stability and transcriptional activity of p63alpha are regulated by SUMO-1 conjugation. Cell Cycle. 2005Jan;4(1):183-90
141. Chao TT, Chang CC, Shih HM. SUMO modification modulates the transrepression activity of PLZF. Biochem Biophys Res Commun. 2007Jun29;358(2):475-82
142. Binda O, Roy JS, Branton PE. RBP1 family proteins exhibit SUMOylation-dependent transcriptional repression and induce cell growth inhibition reminiscent of senescence. Mol Cell Biol. 2006Mar;26(5):1917-31
143. Matsuura T, Shimono Y, Kawai K, Murakami H, Urano T, Niwa Y. et al. PIAS proteins are involved in the SUMO-1 modification, intracellular translocation and transcriptional repressive activity of RET finger protein. Exp Cell Res. 2005Aug1;308(1):65-77
144. Tan JA, Sun Y, Song J, Chen Y, Krontiris TG, Durrin LK. SUMO conjugation to the matrix attachment region-binding protein, special AT-rich sequence-binding protein-1 (SATB1), targets SATB1 to promyelocytic nuclear bodies where it undergoes caspase cleavage. J Biol Chem. 2008Jun27;283(26):18124-34
145. Cho G, Lim Y, Golden JA. SUMO interaction motifs in Sizn1 are required for promyelocytic leukemia protein nuclear body localization and for transcriptional activation. J Biol Chem. 2009Jul17;284(29):19592-600
146. Imoto S, Ohbayashi N, Ikeda O, Kamitani S, Muromoto R, Sekine Y. et al. Sumoylation of Smad3 stimulates its nuclear export during PIASy-mediated suppression of TGF-beta signaling. Biochem Biophys Res Commun. 2008May30;370(2):359-65
147. Spengler ML, Brattain MG. Sumoylation inhibits cleavage of Sp1 N-terminal negative regulatory domain and inhibits Sp1-dependent transcription. J Biol Chem. 2006Mar3;281(9):5567-74
148. Matsuzaki K, Minami T, Tojo M, Honda Y, Uchimura Y, Saitoh H. et al. Serum response factor is modulated by the SUMO-1 conjugation system. Biochem Biophys Res Commun. 2003Jun20;306(1):32-8
149. Cheng Z, Ke Y, Ding X, Wang F, Wang H, Wang W. et al. Functional characterization of TIP60 sumoylation in UV-irradiated DNA damage response. Oncogene. 2008Feb7;27(7):931-41
150. Gupta P, Ho PC, Huq MM, Ha SG, Park SW, Khan AA. et al. Retinoic acid-stimulated sequential phosphorylation, PML recruitment, and SUMOylation of nuclear receptor TR2 to suppress Oct4 expression. Proc Natl Acad Sci U S A. 2008Aug12;105(32):11424-9
151. Kunapuli P, Kasyapa CS, Chin SF, Caldas C, Cowell JK. ZNF198, a zinc finger protein rearranged in myeloproliferative disease, localizes to the PML nuclear bodies and interacts with SUMO-1 and PML. Exp Cell Res. 2006Nov15;312(19):3739-51
152. Karvonen U, Jaaskelainen T, Rytinki M, Kaikkonen S, Palvimo JJ. ZNF451 is a novel PML body- and SUMO-associated transcriptional coregulator. J Mol Biol. 2008Oct10;382(3):585-600
Author contact
![]() Correspondence to: Prof. Dr. Xaveer Van Ostade, xaveer.vanostadeac.be, tel: 0032 (0)3 365 23 19
Correspondence to: Prof. Dr. Xaveer Van Ostade, xaveer.vanostadeac.be, tel: 0032 (0)3 365 23 19

 Global reach, higher impact
Global reach, higher impact