10
Impact Factor
ISSN: 1449-2288
Int J Biol Sci 2011; 7(7):1068-1076. doi:10.7150/ijbs.7.1068 This issue Cite
Review
Global Gene Expression during the Human Organogenesis: From Transcription Profiles to Function Predictions
1. College of Life Sciences, Wuhan University, Wuhan 430072, P.R. China
2. College of Life Sciences, South-Central University for Nationalities, Wuhan 430074, P.R. China
3. Section on Molecular Morphogenesis, Laboratory of Gene Regulation and Development, NICHD, National Institutes of Health, Bethesda, MD 20892, USA
* These authors contribute equally to this work.
Received 2011-7-19; Accepted 2011-8-24; Published 2011-9-7
Abstract
Human embryogenesis includes an integrated set of complex yet coordinated development of different organs and tissues, which is regulated by the spatiotemporal expression of many genes. Deciphering the gene regulation profile is essential for understanding the molecular basis of human embryo development. While molecular and genetic studies in mouse have served as a valuable tool to understand mammalian development, significant differences exists in human and mouse development at morphological and genomic levels. Thus it is important to carry out research directly on human embryonic development. Here we will review some recent studies on gene regulation during human embryogenesis with particular focus on the period of organogenesis, which had not been well studied previously. We will highlight a gene expression database of human embryos from the 4th to the 9th week. The analysis of gene regulation during this period reveals that genes functioning in a given developmental process tend to be coordinately regulated during human embryogenesis. This feature allows us to use this database to identify new genes important for a particular developmental process/pathway and deduce the potential function of a novel gene during organogenesis. Such a gene expression atlas should serve as an important resource for molecular study of human development and pathogenesis.
Keywords: Human embryonic development, Gene expression profile, Gene regulation database, Organogenesis, Microarrays, Maternal genes
Introduction
Elucidating the molecular mechanisms governing human embryogenesis is a challenge even in the post-genomic era. Largely due to the obstacle in obtaining suitable human embryos for biomedical research, for over 50 years, the mouse (Mus musculus) has played a premier role as a model for studying early human development, which has led to enormous progress in our understanding of mammalian development. Elaborate genetic studies have revealed the function of many genes in development and genome-wide expression profile analyses have further identified many genes that are regulated during the earliest stages of mouse embryogenesis. On the other hand, significant developmental differences exist between human and model organisms including mouse at both morphological and functional levels. More importantly, these differences appear to be genetically determined, making it critical to carry out studies directly on human development. Here, we highlight some distinct features between human and mouse development and review some recent gene expression studies on human and mouse embryogenesis. We focus particularly on the recent studies on the gene regulation during organogenesis, a period that had not been previously analyzed at genome-wide level even in mouse.
Some distinct morphological and molecular features of mouse and human embryogenesis
In the mammal, fertilization initiates a series of well-coordinated developmental events, including morulation (preimplantation stage), gastrulation, and organogenesis (postimplantation stage). Figure 1 compares the developmental stages of mouse and human embryos. The Theiler Stages (TS 1-28) for mouse embryos are based in part on the somite number and some other identifiable characteristics [1, 2].
Comparison of the developmental stages of mouse and human embryos. Mouse embryonic stages (Theiler stages or TS) are based on somite number and characteristics and consists of 28 stages from fertilized egg to birth, up to 20 days post conception (dpc) (the middle panel). The criteria for staging human embryos were described by the Carnegie Institution of Washington, which are based on the development of structures, not by size or the number of days of development. The corresponding stages of mouse and human are indicated by dashed lines. The lower panels indicate the TS or CS with the corresponding developmental time (dpc) for the mouse and human embryos, respectively. The 23 Carnegie stages (CS) only covers the first 60 days of human embryo development, thereafter that the term embryo is replaced with fetus. The numbers in brackets above the large brackets for mouse and below the large brackets for human indicate the references that report previous human and mouse embryo transcriptome data of the indicated stages.
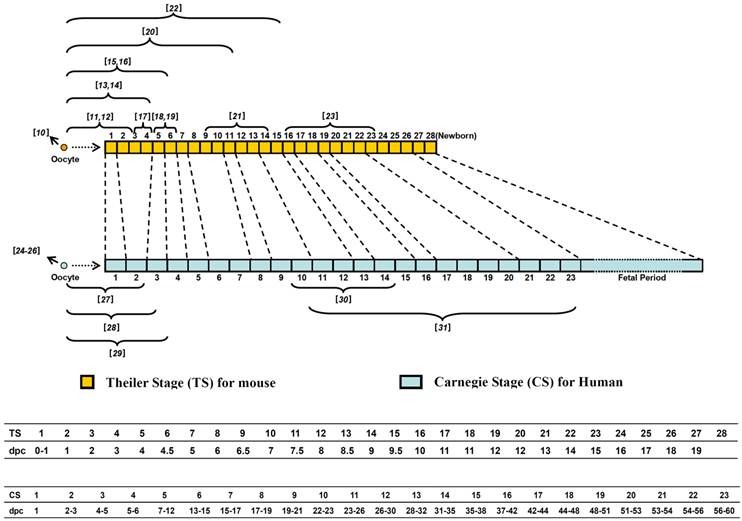
The Carnegie Stages (CS 1-23) for human embryos are based on the development of structures, not in the size or the number of days of development and it covers the first 60 days of development, at the end of which the term embryo is usually replaced with the term fetus [3]. Considering approximately 90 million years of independent evolution, it is not surprising that significant morphological differences exist in human and mouse embryogenesis, especially some important characteristics developed during organogenesis. For example, the facial organs of human and mouse embryo develop in a very different sequence and form distinct features. While the optic pit is the earliest facial organ for mouse (a signature for all rodent animal), essentially all human facial organs appear around the same time. The contralateral limbs of human embryo rotate to the proper positions ventrally with flexible joints while the mouse limbs show little rotation and the joints are less flexible. Mouse embryo's tail elongates and thins from TS17 while human embryo's tail regresses during CS 23 (about 9th week of embryogenesis) (Fig. 1). Another obvious difference between mouse and human embryogenesis is the time of birth. The mouse embryo is born almost immediately after all the organs are developed (around TS27-TS28, 19-20 days post conception). In contrast, at the end of organogenesis (CS23, corresponding to TS26, Fig. 1), the human embryo continues to stay in the uterus for a few more months: the fetal period of ongoing growth of many organs (Fig. 1).
While the anatomical differences between human and mouse embryogenesis have been studied for over 100 years, systematic molecular analyses are becoming available in the last decade, thanks to the advances in genome sequencing and microarray technologies. Human-mouse genome-wide comparisons have revealed a number of differences related to the species-specific adaptations especially in reproduction, immunity, and olfaction systems. These include many mouse genes missing in human and vice versa [4, 5] and distinct alternative splicing of orthologous genes associated with increased evolutionary change [6-8], etc. Interestingly, the most significant differences in mice and human are not in the number of genes that each has but in the expression of genes and the activities of their protein products [9]. Below, we will briefly summarize some of the microarray studies on gene expression in mouse and human embryos [10-31].
Gene expression studies of mouse and human embryos
Genome-wide expression analyses, especially microarray profiling, of human and mouse embryogenesis have provided valuable information on the deciphering of the molecular network underlining morphological changes during development and insights into mammalian evolution. During the past decades, most of the research on human and mouse embryos focused mainly on the transcriptomes of oocytes and preimplantation embryos [10-19, 24-29], largely due to the relative ease to obtain such embryos. Within this period, although there are little morphological differences between the human and mouse embryo, the global gene expression patterns show dissimilarities. A characteristic difference is the timing of transition from maternal transcripts to zygotic transcripts. Human zygotic gene expression first occurs between the four- and eight-cell stages of preimplantation development while major mouse zygotic genome activation (ZGA) occurs at the two-cell stage [32, 33]. Furthermore, human preimplantation development consists of two main phases: one from the meiosis II (MII) oocyte to 4-cell embryo when the maternal genes predominate, and the other from the 8-cell embryo to the blastocyst when maternal genes are downregulated while zygotic genes are upregulated [29]. While in mouse, there are two major transcriptional waves during preimplantation. One group of zygotic genes are activated at the 2-cell stages and appear to be preferentially those involved in transcription and RNA processing [15]. The other group of genes are activated in 4-cell to 8-cell embryos and are involved mostly in morphological and functional changes of the embryos [13].
Despite these differences, there are also a number of common signaling processes during early mouse and human development, suggesting that during preimplantation, many molecular mechanisms are evolutionarily conserved between human and mouse [16, 28]. These include the Notch, TGFβ/BMP, and Wnt pathways etc. The same ligands and receptors of these signaling pathways are temporally and spatially regulated to determine zygote asymmetry, cell fate, cell polarity, communication, adhesion and migration, and they are important for both hESCs (human embryonic stem cells) and mESCs (mouse embryonic stem cells) [34, 35]. For example, expression of type I and type II receptors for TGF-β both in mouse and human fertilized oocytes and blastocysts suggests a role for TGF-β in early preimplantation development [36].
After implantation, especially during organogenesis, notable morphological differences exist between mouse and human development, suggesting the existence of different gene expression profiles. Indeed, significant differences in embryonic expression patterns were reported between human and mouse development for a variety of genes such as Wnt7a and CAPN3, particularly in the formation of neural crest, midbrain, lens, heart, and smooth muscle [37, 38]. Unfortunately, few genome-wide gene expression analyses have been carried out on either mouse or human post-implantation embryos.
Recently, Fang et al. and Yi et al. independently reported the global gene expression patterns of human embryos covering the whole organogenesis period (CS9-CS23) [30, 31]. In the study by Fang et al. human embryos were grouped into six embryonic developmental stages (CS9-CS14) and microarray analysis was carried out for every stage. On the other hand, Yi et al. isolated total RNA from human embryos for each week of 4th-9th weeks (CS10-23) for expression profiling. Both studies employed the Human Genome U133 Array by Affymetrix (Santa Clara, CA, USA). Although different methods were used for data analysis, both groups found that the regulated genes during the organogenesis period could be divided into two major groups. Fang et al. defined a gradual decrease in 'stemness' concomitant, cell cycle, and metabolism-related genes, and an increase in organogenesis specific genes, including muscle, fat, and connective tissue specific genes and differentiation-associated gene, etc. Similarly, Yi et al. discovered an up-regulation of gene categories enriched with genes involved in multi-cellular organismal development/processes, cell adhesion, cell surface receptor-linked signaling, and cell-cell signaling, which correlate well with the differentiation and development of organs that takes place during the 4th-9th weeks, and observed the down-regulation of gene categories associated with various metabolic processes and transcriptional regulation that are likely involved in the transition of the embryo from mainly cell proliferation in the 4th week to mostly organ development by the 8th-9th weeks. In addition, Yi et al. identified many genes with an arch-shaped gene expression pattern during the 4th-9th week of development, which were enriched with a number of GO categories, such as those related to eye development with the peak level expression of the genes correlated well with the formation of the lens and the development of retinal pigmentation during the 5th-7th weeks of embryogenesis. These studies by Fang et al. and Yi et al. using embryos covering overlapping developmental periods (CS9-14 and CS10-23, respectively) not only confirm the findings but also complement each other to offer a more complete molecular profile of early human embryogenesis.
These studies also enable us to do comparative study to reveal distinct features in human embryo development. For instance, given the differences and similarities between mouse and human embryogenesis, an important question is to what extent the gene expression profiles are conserved between mouse and human embryos. The period of human embryonic developmental stages CS9-CS14 [30] resembles that from gastrulation to early organogenesis of mouse embryos (from TS12-TS17) [21]. Consistently, our analysis showed that dramatic changes in gene expression profiles occurred at the beginning of organogenesis in both mouse (around TS 12, embryonic day 8) and human (around CS9-CS10, or embryonic week 3/4) [21, 30]. Fang et al. further found out that the human-mouse homologs in the down-regulated group are primarily linked to embryonic lethality and abnormal lipid circulation. They believed that these genes should be important for the initiation of mammalian embryo organogenesis. In addition, genes related to myogenesis, osteogenesis, heart development, and neurogenesis were also found to be similarly regulated in human and mouse during organogenesis, displaying a gradually increase in expression. Interestingly, transcriptome comparison has also identified a set of genes that are expressed early in human embryonic development but have not yet been implicated in early mouse development [30]. For example, PAX6 appears to be a human neuroectoderm cell fate determinant and regulate forebrain development but it is not required in mouse [39]. Such differences argue that obtaining the panorama of regulation gene profile in human embryo is indispensible for us to fully understand human development.
The embryonic processes that occur between fertilization and gastrulation (0-4th week) are anatomically distinct from the subsequent organogenesis and histogenesis (4th-9th week). However, the underlying molecular basis of this change remains elusive. By comparing their gene regulation data with those on human oocytes and very early embryos (up to day 3) [26, 27], Yi et al. found that the developmentally regulated genes during the 4-9th week fell into several functional categories/pathways, which are distinct from those associated with the maternal genes that function in early embryonic processes between fertilization and gastrulation. The traditionally highly expressed maternal transcripts could be divided into three groups, absent by the 4th week (about 26.7%), regulated during the 4-9th week (about 14.9%, mostly are down-regulated), and constitutively expressed during the 4-9th week (the rest) [31]. The first two groups of maternal genes are significantly enriched with genes involved in membrane utilization, metabolism, and cell cycle regulation, all of which are important for early embryogenesis and are down-regulated subsequently. Genes in the third group are more likely housekeeping genes, as they are constitutively expressed at least until 9th week of embryogenesis. These genes seem not to be specific for any developmental processes, at least up to the 9th week. These results indicate that the very early phase of embryogenesis utilizes maternal genes in the GO categories related to membrane, metabolism, and cell cycle while the subsequent organ development utilizes mainly zygotically transcribed genes belonging to distinct GO categories. Similarly, Fang et al. compared their data with the hESC genes and differentiation genes defined by Assou et al. [40] and found that the genes with reduced expression during CS9-CS14 significantly overlapped with the stemness-specific genes, in agreement with Yi et al.'s observation that most maternal genes are not detected or down-regulated by the 4th week of human embryogenesis (CS10). Some notable examples of these genes include the core stem cell identity controlling genes OCT4, NANOG, and SOX2 [30, 31, 41]. While the potential binding sites of these genes were found to be significantly enriched in hESC D group (the overlapping genes between decreasing clusters of CS9-CS14 and hESC genes), suggesting that the expression of these genes were no longer needed in the organogenesis period. This is in accordance with that the expression of OCT4, NANOG, and SOX2 were both down-regulated seriously or even undetected during the 4-9th week of embryogenesis.
Aside from the common findings, Yi et al. have also identified some unique features in human embryo development. For instance, they revealed that approximate half of all human genes were expressed and 18.6% of the expressed genes were differentially regulated during the human organogenesis period (CS10-23). Surprisingly, only 1,099 of the regulated transcripts (26.1% of the 4,203 genes) had previously been implicated to be related to development. Thus, most of the genes regulated during this important period of human development represent novel development-related genes. More importantly, by analyzing how genes involved in a given biological/developmental process or GO category are regulated during the 4th-9th week of human embryogenesis, they have observed strong co-regulation of these genes and that their regulation correlates well with the developmental changes associated with this period [31]. For example, the vast majority of the genes associated with stem cells are highly expressed during the 4th or 5th weeks but are down-regulated by the 9th week, consistent with their roles in stem cells since the number of stem cells or undifferentiated cells is reduced when more cells begin to differentiate into different organ/tissue-specific cell types toward the 8th-9th week of development. Likewise, the vast majority of the genes likely involved in organ development, such as genes associated with muscle, fat, and connective tissues or those associated with internal organs or blood and lymph tissues are coordinately upregulated, with the highest levels of expression occurring in the 9th week, in agreement with the ongoing organogenesis. Such findings suggest that genes participating in the same developmental processes tend to be regulated coordinately during the 4th-9th week of development.
A novel in-silico resource for identifying genes involved in and predicting gene function during a developmental process
The co-regulation of genes in a given GO category during the 4th -9th week of human embryogenesis has led us to propose that our gene expression database [31] can be used as a searchable resource. That is, one can use it to search for genes likely involved in a particular developmental process or inferring the biological functions of a gene of interest, if it is among the regulated genes, during this developmental period. This is important because the developmental functions of many genes are unknown and that many genes involved in different developmental process also remain to be identified.
The gene expression atlas is a simple searchable database (http://vmolab.whu.edu.cn:8080/HumanGeneSearch/). Fig. 2 outlines the basic steps to use the database.
A. To predict the potential developmental role of an unknown gene X (Fig. 2, left panel)
- Search the database to obtain the expression profile of Gene X
- Cluster the expression profile of Gene X together with all developmentally regulated genes (the list can be downloaded from the database) by hierarchical clustering methods
- Identify the gene cluster most closely co-regulated with Gene X
- Analyze the gene cluster with Gene Ontology or GenMaPP to determine potential roles of or potential pathways involving Gene X (user can adjust the coefficient to define the size of the gene cluster in step 2, in order to get satisfied GO function categories)
- Experimental validation (e.g., confirmation of the expression profile by RT-PCR, in situ hybridization, and/or immunohistochemistry, functional analysis in cultured cells or in vivo by using gene knock-out or transgenesis, etc.)
Schematic diagram for using the expression database to predict the potential roles of unknown genes and identify genes involved in a particular biological process during the 4th-9th week of human embryogenesis.
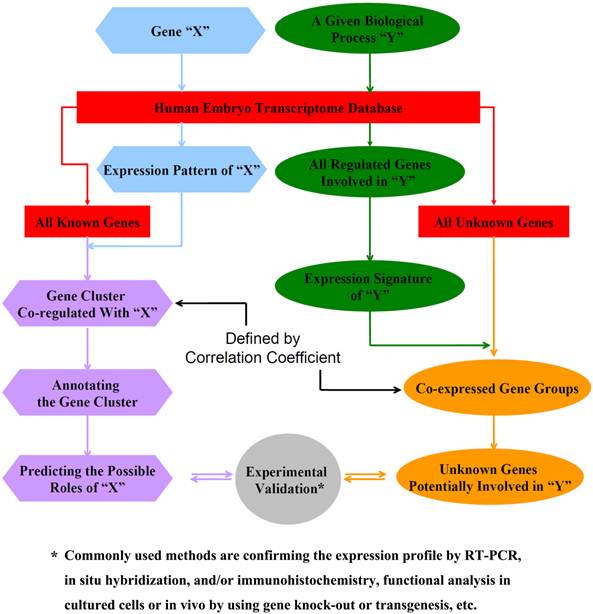
B. To identify genes involved in a particular biological process or cellular pathways (Fig. 2, right panel)
- Search the database to obtain the expression profiles of all genes known to be involved in a given biological/developmental process Y (e.g., a particular GO category)
- Identify the expression signature of these genes (may get more than one featured cluster)
- Cluster the expression patterns of these genes with all unknown but developmentally regulated genes
- Identify the unknown genes that are co-regulated with the genes involved in the biological/developmental process Y. (again, user can adjust the coefficient to define the size of the signature gene cluster in step 2, or use different signature cluster in order to get enough unknown genes)
- Experimental validation (e.g., confirmation of the expression profile by RT-PCR, in situ hybridization, and/or immunohistochemistry, functional analysis in cultured cells or in vivo by using gene knock-out or transgenesis, etc.)
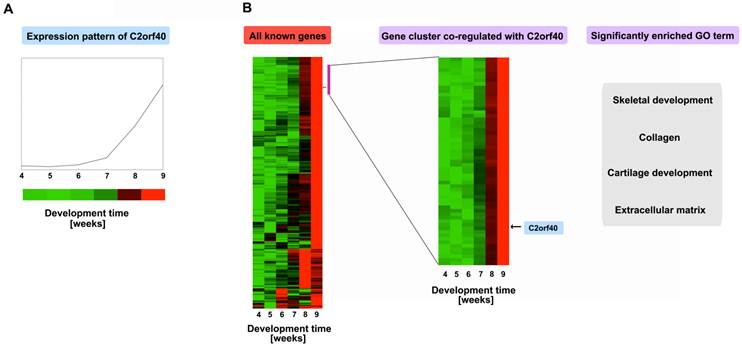
As a test and an example for using this atlas with pipeline A, a novel developmentally regulated genes, C2orf40 (chromosome 2 open reading frame 40), was analyzed as described above [31]. Firstly searching the microarray database revealed that it is upregulated throughout the developmental period (Fig. 3A), with an obvious increasing trend in the 7th week. Then we clustered its expression profile with all 5,358 developmentally regulated genes during the 4th-9th week of human embryogenesis, showing that its temporal regulation pattern is similar to a small cluster of 59 genes (Fig. 3B, right panel, indicted by the purple-red bar). These genes are highly enriched with GO categories involved in skeletal development, collagen, cartilage development, and the extracellular matrix, suggesting that C2orf40 is likely involved in skeletal development. Indeed, in situ hybridization revealed that C2orf40 has little expression in the 4th week but is upregulated in the skeletal system by the 7th-9th week. In particular, it is highly expressed in the vertebrae by the 7th week. These spatial patterns provide further support that the C2orf40 gene is likely involved in skeletal and cartilage development, although further experimentation is required to demonstrate its function. Actually, we can also use pipeline B to confirm the inferring of pipeline A. We combined the well-known 148 skeletal genes to combine with the 1,218 developmentally regulated genes with no known GO categories and clustered them based on their expression profiles. We identified four signature non-overlapping clusters with some unknown genes (see the Figure 5 of Reference 31), while C2orf40 was exactly in one of them. These and other analyses suggest that the gene expression atlas can be a valuable resource for understanding the molecular pathways governing mammalian, especially human, organogenesis.
Gene expression profiles suggest a potential role in skeletal development for a novel developmentally regulated gene C2orf40. A, The C2orf40 expression profile as obtained from the expression database. B, The expression profile of C2orf40 clustered with all developmentally regulated genes. A cluster of genes that most closely co-regulated with C2orf40 was identified by their correlation efficient and indicated with a purple bar on the right, and was expanded (right panel, with the location of C2orf40 indicated). Some significant GO categories associated with these clusters are shown on the right; they are all associated with skeletal development.
Conclusions and perspectives
The advances in genetics and genome sequence information in the past two decades has made mouse an extremely valuable model for understanding mammalian development. However, the divergence in morphological and molecular changes during mouse and human embryogenesis argues that it is important to carry out, to the extent possible, research directly on human embryos. On the other hand, because of obvious ethical concerns, human embryonic tissues are hard to obtain. Thus, the knowledge on human embryonic development has been mostly on oocytes, hESCs, and very early human embryos. The two recent microarray analyses on human embryogenesis have provided the first glimpse on the global gene expression profiles during human organogenesis and revealed important information for understanding the molecular pathways of human embryonic development. More importantly, the database for the gene expression profiles of the 4th to 9th weeks of human development appears to function as a valuable resource for identifying novel genes involved in and predicting the potential roles of developmentally regulated genes during organogenesis, a critical period with little prior molecular knowledge. A pressing task is to further improve and expand our database as more information on human development is collected with the development of new technologies such as next-generation seq, RNA-seq, etc. This should enhance the value of the database for studying the molecular basis of human development. Finally, it is worth mentioning that the most significant morphological differences between mouse and human development appear during organogenesis. The human gene expression atlas should also aid in the determination of the molecular basis underlying the conservations and evolutionary divergences between mouse and human development once the transcriptome analyses of corresponding period of mouse embryos are carried out. Given the advances in mouse genetics, such information will be extremely useful for future functional dissections of the molecular mechanisms underlying mammalian organogenesis.
Acknowledgements
This work was supported by National High Technology Research and Development Program of China (863 Program) Grant 2006AA02A306, National Natural Science Foundation of China Grant 30900265 and 31101047). Y.-B. Shi was supported by NICHD Intramural Research Program, National Institutes of Health.
Conflict of Interests
The authors have declared that no conflict of interest exists.
References
1. Bard JL, Kaufman MH, Dubreuil C. et al. An internet-accessible database of mouse developmental anatomy based on a systematic nomenclature. Mech Dev. 1998;74:111-20
2. Downs KM, Davies T. Staging of gastrulating mouse embryos by morphological landmarks in the dissecting microscope. Development. 1993;118:1255-66
3. Hill MA. Early human development. Clin Obstet Gynecol. 2007;50:2-9
4. Lauw FN, Caffrey DR, Golenbock DT. Of mice and man: TLR11 (finally) finds profilin. Trends Immunol. 2005;26:509-11
5. Zhang X, Firestein S. The olfactory receptor gene superfamily of the mouse. Nat Neurosci. 2002;5:124-33
6. Modrek B, Lee CJ. Alternative splicing in the human, mouse and rat genomes is associated with an increased frequency of exon creation and/or loss. Nat Genet. 2003;34:177-80
7. Church DM, Goodstadt L, Hillier LW. et al. Lineage-specific biology revealed by a finished genome assembly of the mouse. PLoS Biol. 2009;7:e1000112
8. Hughes AL. Genomes of mice and men. Heredity. 2003;90:115-7
9. Su AI, Cooke MP, Ching KA. et al. Large-scale analysis of the human and mouse transcriptomes. Proc Natl Acad Sci U S A. 2002;99:4465-70
10. Hamatani T, Falco G, Carter MG. et al. Age-associated alteration of gene expression patterns in mouse oocytes. Hum Mol Genet. 2004;13:2263-78
11. Li XY, Cui XS, Kim NH. Transcription profile during maternal to zygotic transition in the mouse embryo. Reprod Fertil Dev. 2006;18:635-45
12. Zeng F, Schultz RM. RNA transcript profiling during zygotic gene activation in the preimplantation mouse embryo. Dev Biol. 2005;283:40-57
13. Hamatani T, Carter MG, Sharov AA. et al. Dynamics of global gene expression changes during mouse preimplantation development. Dev Cell. 2004;6:117-31
14. Wang S, Cowan CA, Chipperfield H. et al. Gene expression in the preimplantation embryo: in-vitro developmental changes. Reprod Biomed Online. 2005;10:607-16
15. Zeng F, Baldwin DA, Schultz RM. Transcript profiling during preimplantation mouse development. Dev Biol. 2004;272:483-96
16. Wang QT, Piotrowska K, Ciemerych MA. et al. A genome-wide study of gene activity reveals developmental signaling pathways in the preimplantation mouse embryo. Dev Cell. 2004;6:133-44
17. Tanaka TS, Ko MS. A global view of gene expression in the preimplantation mouse embryo: morula versus blastocyst. Eur J Obstet Gynecol Reprod Biol. 2004;115(Suppl 1):S85-91
18. Hamatani T, Daikoku T, Wang H. et al. Global gene expression analysis identifies molecular pathways distinguishing blastocyst dormancy and activation. Proc Natl Acad Sci U S A. 2004;101:10326-31
19. Chen HW, Chen JJ, Yu SL. et al. Transcriptome analysis in blastocyst hatching by cDNA microarray. Hum Reprod. 2005;20:2492-501
20. Tanaka TS, Jaradat SA, Lim MK. et al. Genome-wide expression profiling of mid-gestation placenta and embryo using a 15,000 mouse developmental cDNA microarray. Proc Natl Acad Sci U S A. 2000;97:9127-32
21. Mitiku N, Baker JC. Genomic analysis of gastrulation and organogenesis in the mouse. Dev Cell. 2007;13:897-907
22. Sharov AA, Piao Y, Matoba R. et al. Transcriptome analysis of mouse stem cells and early embryos. PLoS Biol. 2003;1:E74
23. Zhang W, Morris QD, Chang R. et al. The functional landscape of mouse gene expression. J Biol. 2004;3:21
24. Kocabas AM, Crosby J, Ross PJ. et al. The transcriptome of human oocytes. Proc Natl Acad Sci U S A. 2006;103:14027-32
25. Zhang P, Kerkela E, Skottman H. et al. Distinct sets of developmentally regulated genes that are expressed by human oocytes and human embryonic stem cells. Fertil Steril. 2007;87:677-90
26. Assou S, Cerecedo D, Tondeur S. et al. A gene expression signature shared by human mature oocytes and embryonic stem cells. BMC Genomics. 2009;10:10
27. Dobson AT, Raja R, Abeyta MJ. et al. The unique transcriptome through day 3 of human preimplantation development. Hum Mol Genet. 2004;13:1461-70
28. Adjaye J, Huntriss J, Herwig R. et al. Primary differentiation in the human blastocyst: comparative molecular portraits of inner cell mass and trophectoderm cells. Stem Cells. 2005;23:1514-25
29. Zhang P, Zucchelli M, Bruce S. et al. Transcriptome profiling of human pre-implantation development. PLoS One. 2009;4:e7844
30. Fang H, Yang Y, Li C. et al. Transcriptome analysis of early organogenesis in human embryos. Dev Cell. 2010;19:174-84
31. Yi H, Xue L, Guo MX. et al. Gene expression atlas for human embryogenesis. FASEB J. 2010;24:3341-50
32. Braude P, Bolton V, Moore S. Human gene expression first occurs between the four- and eight-cell stages of preimplantation development. Nature. 1988;332:459-61
33. Hamatani T, Ko M, Yamada M. et al. Global gene expression profiling of preimplantation embryos. Hum Cell. 2006;19:98-117
34. Sato N, Sanjuan IM, Heke M. et al. Molecular signature of human embryonic stem cells and its comparison with the mouse. Dev Biol. 2003;260:404-13
35. Zhang Y, Yang Z, Wu J. Signaling pathways and preimplantation development of mammalian embryos. FEBS J. 2007;274:4349-59
36. Roelen BA, Mummery CL. Transforming growth factor-betas in pre-gastrulation development of mammals (minireview). Mol Reprod Dev. 2000;56:220-6
37. Fougerousse F, Bullen P, Herasse M. et al. Human-mouse differences in the embryonic expression patterns of developmental control genes and disease genes. Hum Mol Genet. 2000;9:165-73
38. Trainor P, Krumlauf R. Development. Riding the crest of the Wnt signaling wave. Science. 2002;297:781-3
39. Zhang X, Huang CT, Chen J. et al. Pax6 is a human neuroectoderm cell fate determinant. Cell Stem Cell. 2010;7:90-100
40. Assou S, Le Carrour T, Tondeur S. et al. A meta-analysis of human embryonic stem cells transcriptome integrated into a web-based expression atlas. Stem Cells. 2007;25:961-73
41. Boyer LA, Lee TI, Cole MF. et al. Core transcriptional regulatory circuitry in human embryonic stem cells. Cell. 2005;122:947-56
Author contact
![]() Corresponding author: Wen-Xin Li: Phone: +86 (27) 6875-2831; Fax: +86 (27) 6875-2146; Email: liwxlabedu.cn. Yun-Bo Shi: Phone: +1 (301) 402-1004; Fax: +1 (301) 402-1323; Email: shinih.gov
Corresponding author: Wen-Xin Li: Phone: +86 (27) 6875-2831; Fax: +86 (27) 6875-2146; Email: liwxlabedu.cn. Yun-Bo Shi: Phone: +1 (301) 402-1004; Fax: +1 (301) 402-1323; Email: shinih.gov

 Global reach, higher impact
Global reach, higher impact