10
Impact Factor
ISSN: 1449-2288
Int J Biol Sci 2012; 8(5):761-777. doi:10.7150/ijbs.4036 This issue Cite
Research Paper
Differential Expression and Functional Analysis of the Tristetraprolin Family during Early Differentiation of 3T3-L1 Preadipocytes
1. Institute of Biological Chemistry, Academia Sinica, 128, Academia Road Sec 2, Nankang, Taipei 115, Taiwan;
2. Graduate Institute of Biochemical Sciences, College of Life Science, National Taiwan University, 1 Roosevelt Road Sec 4, Taipei 106, Taiwan.
* The three authors contributed equally to this work.
Received 2012-1-2; Accepted 2012-5-15; Published 2012-5-31
Abstract
The tristetraprolin (TTP) family comprises zinc finger-containing AU-rich element (ARE)-binding proteins consisting of three major members: TTP, ZFP36L1, and ZFP36L2. The present study generated specific antibodies against each TTP member to evaluate its expression during differentiation of 3T3-L1 preadipocytes. In contrast to the inducible expression of TTP, results indicated constitutive expression of ZFP36L1 and ZFP36L2 in 3T3-L1 preadipocytes and their phosphorylation in response to differentiation signals. Physical RNA pull-down and functional luciferase assays revealed that ZFP36L1 and ZFP36L2 bound to the 3′ untranslated region (UTR) of MAPK phosphatase-1 (MKP-1) mRNA and downregulated Mkp-1 3′UTR-mediated luciferase activity. Mkp-1 is an immediate early gene for which the mRNA is transiently expressed in response to differentiation signals. The half-life of Mkp-1 mRNA was longer at 30 min of induction than at 1 h and 2 h of induction. Knockdown of TTP or ZFP36L2 increased the Mkp-1 mRNA half-life at 1 h of induction. Knockdown of ZFP36L1, but not ZFP36L2, increased Mkp-1 mRNA basal levels via mRNA stabilization and downregulated ERK activation. Differentiation induced phosphorylation of ZFP36L1 through ERK and AKT signals. Phosphorylated ZFP36L1 then interacted with 14-3-3, which might decrease its mRNA destabilizing activity. Inhibition of adipogenesis also occurred in ZFP36L1 and TTP knockdown cells. The findings indicate that the differential expression of TTP family members regulates immediate early gene expression and modulates adipogenesis.
Keywords: tristetraprolin, 3T3-L1, ZFP36L1, ZFP36L2, MKP-1, AU-rich element.
Introduction
The tristetraprolin (TTP) family, which regulates mRNA decay, comprises three members in humans and four in rodents [1,2]. The three related proteins are TTP (also named ZFP36, TIS11), ZFP36L1 (TIS11B, BRF1), and ZFP36L2 (TIS11D, BRF2). Zfp36l3 is a rodent-specific gene [3], only expressed in the placenta and extraembryonic tissues of mice. All TTP family members have a highly conserved (70% identity) tandem zinc finger (TZF) domain, which is responsible for high-affinity binding to AU-rich elements (AREs) of mRNAs. Their N-terminal and C-terminal mRNA decay activation domains are <40% conserved [4]. The optimal RNA sequence for TTP binding is the nonamer UUAUUUAUU [5,6]. The TTP family members all share TTP-like activities and promote the deadenylation and destruction of ARE-containing target sequences in co-transfection and in vitro deadenylation assays [7,8].
TTP family proteins contain a high percentage of serines and threonines, which can undergo phosphorylation [9,10]. Phosphorylation of TTP and ZFP36L1 can regulate their subcellular localization, protein stability, and mRNA destabilization function [11-13]. Numerous studies have shown that mitogen-activated protein kinase (MAPK)-activated protein kinase 2 (MK2) phosphorylates TTP to allow binding of 14-3-3 adaptor proteins and to inhibit deadenylase complex recruitment, thereby inhibiting the mRNA destabilizing activity of TTP [14-19]. Protein kinase B (also known as AKT) phosphorylates ZFP36L1 at Ser92 and Ser203, resulting in binding to 14-3-3 and impairment of mRNA decay activity [9,20]. The effects of ZFP36L2 phosphorylation remain unclear.
A variety of studies have indicated that the TTP family proteins have important physiological roles. In macrophages, inflammatory signals induce TTP, which then acts as an anti-inflammatory factor that negatively regulates the expression of proinflammatory mediators [21]. Mice lacking TTP spontaneously develop erosive arthritis, cachexia, alopecia, dermatitis, autoantibodies, and myeloid hyperplasia [22] owing to the overproduction of TNFα resulting from its prolonged mRNA half-life [23]. Zfp36l1-deficient mice fail to undergo chorioallantoic fusion, and the embryos die in utero [24]. Deletion of the first exon of Zfp36l2 results in complete infertility of female mice [25], whereas Zfp36l2 knockout mice display defects in hematopoiesis [26]. A recent report showed that mice lacking both ZFP36L1 and ZFP36L2 during thymopoiesis develop T-cell acute lymphoblastic leukemia [27]. Previous research has identified several mRNA targets of TTP [2], including cytokines, oncogenes, growth factors, and immediate early gene (IEG) mRNAs such as Mkp-1 (encoding a MAPK phosphatase, or MKP) and Ttp itself [28-32]. The TTP-sensitive targets identified in knockout cells, such as granulocyte-macrophage colony-stimulating factor, polo-like-kinase 3, and immediate early response 3, do not respond to knockout of ZFP36L1 (and ZFP36L2) [24,33,34].
Although TTP family proteins contain conserved RNA-binding domains, they have differential expression patterns and different mRNA targets, and thus they have distinct biological functions. Differentiation of 3T3-L1 preadipocytes involves a series of events such as mitotic clonal expansion, which involves transient expression of IEGs. Our group previously demonstrated the inducible expression of TTP during differentiation of 3T3-L1 preadipocytes [30], and we identified Mkp-1 mRNA as one of its targets [29]. The present study includes further characterization of the expression and regulation of ZFP36L1 and ZFP36L2 in 3T3-L1 preadipocytes and the generation of specific antibodies against each member to demonstrate its expression. Short hairpin RNA (shRNA) or small interfering (siRNA) knockdown of the TTP family proteins were used to characterize their functions in modulating the stability of ARE-containing Mkp-1 mRNA, demonstrating that ZFP36L1 and TTP sequentially regulate the level of Mkp-1 mRNA during early differentiation of 3T3-L1 preadipocytes.
Materials and Methods
Plasmid constructs
The coding sequences of mouse ZFP36L1, ZFP36L2, and 14-3-3zeta were PCR-amplified from 2 h lipopolysaccharide-treated RAW264.7 cDNAs using primers 5′-ATGACCACCACCCTCGTGTC-3′ and 5′-TTAGTCATCTGAGATGGAGAG-3 for Zfp36l1, 5′-ATGTCGACCACACTTCTGTCAC-3′and 5′-TCAGTCGTCGGAGATGGAGAGGCG-3′ for Zfp36l2, and 5′-ATGGATAAAAATGAGCTGG-3′ and 5′-CCAGCTCATTTTTATCCAT-3′ for 14-3-3 zeta. The PCR fragments were ligated into pCRII-TOPO vector (Invitrogen, Carlsbad, CA, USA). After sequences were confirmed, recombinant genes were then subcloned into pCMV-Tag-2B (Stratagene, La Jolla, CA, USA) and pEGFP-C2 (Clontech Laboratories, Mountain View, CA, USA) via EcoRI sites for FLAG-tagged and green fluorescent protein (GFP) expression, respectively, in mammalian cells. 14-3-3 zeta was also cloned into pGEX-3 (GE Healthcare BioScience, Chalfont St. Giles, UK) for glutathione-S-transferase (GST) fusion protein production. TTP expression plasmid and the 3′UTR of Mkp-1 mRNA were cloned as described [29]. For luciferase reporter constructs, Mkp-1 3′UTR was subcloned into pCMV-Tag-2C-Luciferase (Stratagene) reporters using the ApaI and KpnI sites.
Expression of recombinant TTP family proteins and production of antisera
The mouse cDNA fragments corresponding to amino acid residues 185-306 of TTP, residues 225-324 of ZFP36L1, and residues 300-448 of ZFP36L2 were amplified by PCR and cloned into the pMAL-c4x expression vector (New England Biolabs, Ipswich, MA, USA). Additional 6×His tag sequences were added to their 3' primers. The recombinant proteins were expressed in Escherichia coli STBL2 (Invitrogen) and purified using Ni-NTA resin (QIAGEN, Germantown, MD, USA). The antisera were induced according to general procedures. Briefly, approximately 250 µg maltose-binding protein fusion protein was diluted in 0.5 ml phosphate-buffered saline (PBS), mixed with 0.5 ml Freund's complete adjuvant, and injected intradermally into New Zealand white rabbits. The rabbits were injected with the same amount of antigen in Freund's incomplete adjuvant every 4 weeks following the primary injection. After the second round of injections, antisera were collected and immunoglobulins were purified for immunoblotting analysis. All procedures adhered to institutional guidelines for the ethical use of animals in research.
Cell culture
The 3T3-L1 preadipocytes were grown in Dulbecco's modified Eagle's medium (Gibco, Grand Island, NY, USA) containing 1.5 g/l NaHCO3 and supplemented with 10% bovine serum (Gibco), 100 U/l penicillin, and 100 mg/l streptomycin (Gibco) in a humidified 37°C incubator containing 5% CO2. Once 60% to 70% confluency had been reached, 3T3-L1 cells were washed once in PBS and subcultured by trypsinization. To induce differentiation, 2-day post-confluent 3T3-L1 cells were stimulated by changing the culture medium to fresh medium containing 10% fetal bovine serum (HyClone Laboratories, Logan, UT, USA) with a hormonal cocktail containing 0.5 mM methylisobutylxanthine (Sigma-Aldrich, St. Louis, MO, USA), 5 µM dexamethasone (Sigma-Aldrich), and 0.17 µM bovine insulin (Sigma-Aldrich) (cocktail together named FMDI inducer). Human embryonic kidney (HEK) 293T cells were grown in Dulbecco's modified Eagle's medium containing 3.7 g/l NaHCO3, 10% fetal bovine serum, 100 U/l penicillin, and 100 mg/l streptomycin, and were subcultured by flushing the cells with culture medium.
Preparation of cellular extracts and immunoblotting
For preparation of whole-cell extracts (WCEs), cells were washed once in PBS in a 100-mm culture dish and resuspended in 400 μl of WCE buffer (25 mM HEPES, pH 7.7, 1.5 mM MgCl2, 0.2 mM EDTA, 0.5 mM DTT (Dithiothreitol), 0.1% (v/v) NP-40, 0.3 M NaCl) with protease inhibitor cocktails (Sigma-Aldrich) and phosphatase inhibitors (0.01 M β-glycerol phosphate, 0.1 mM Na2MoO4, 0.1 mM Na3VO4, pH 10.0, and 0.01 M NaF). The cell suspension was shaken at 4°C for 30 min, centrifuged at 13,000 rpm, 4°C for 5 min, and the supernatant was then collected. Proteins in cell lysates were quantified using Bradford reagent (Bio-Rad, Hercules, CA, USA), separated by 10% SDS-PAGE, and transferred to a polyvinylidene difluoride membrane (Millipore, Billerica, MA, USA) via a semi-dry transfer system for immunoblotting with anti-TTP, anti-ZFP36L1, anti-ZFP36L2, and anti-β-tubulin. Commercial antibodies against ERK, MKP-1, and AKT (each from Santa Cruz Biotechnology, Santa Cruz, CA, USA) and phosphorylated ERK (p-ERK) and p-AKT (Ser473) (each from Cell Signaling Technology, Beverly, MA, USA),and anti-FABP4 (aP2)(Epitomics, Burlingame, CA, USA). To examine protein phosphorylation, the WCEs were incubated with 20 U calf intestinal alkaline phosphatase (CIP, New England Biolabs) at 37°C for 1 h, and then subjected to immunoblotting.
RNA extraction, reverse-transcription, and quantitative PCR
Total RNA was harvested using TRIzol reagent (Invitrogen) Then, 3-5 µg of total RNA was reverse transcribed to produce cDNA using M-MLV reverse transcriptase (Promega, Madison, WI, USA). Real-time PCR was performed in a total volume of 20 µl using the Applied Biosystems 7300 Real-Time PCR System (Applied Biosystems, Foster City, CA, USA). Ttp, Zfp36l1, Zfp36l2, and Mkp-1 expression was analyzed using SYBR Green PCR Master Mix (Applied Biosystems) with 50 ng of cDNA and 400 nM of each of the forward and reverse primers 5′-TGAGCGAAGTTTTATGCAAGGG-3′ and 5′-GCTGGGCAGAGTGACCGAG-3′ for Zfp36l1, 5′-GATGTCGACTTGTTGTGCAAGACG-3′ and 5′-GCGTCCCTACCGCCTTCT-3′ for Zfp36l2, and primers described previously [29] for Ttp, Mkp-1, and β-actin. β-actin was used as an internal control. The real-time PCR amplification conditions were 40 cycles of 95°C for 15 s and 60°C for 1 min. Real-time PCR data were analyzed using the 2-△△dCt relative quantification method according to the manufacturer's instructions.
RNA pull-down assays
Biotin-labeled RNAs containing Mkp-1 3′UTR or 18S rRNA were transcribed in vitro using the T7-MEGAshortscriptTM High Yield Transcription kit (Ambion, Austin, TX, USA). Briefly, 2 μg of BamHI-linearized pCRII-TOPO-MKP-1 3′UTR plasmid and pT7-RNA-18S control template DNA were transcribed in vitro using T7 polymerase in the presence of 1 mM biotin-CTP (Invitrogen) at 37°C for 2 h. After DNase I digestion, the riboprobes were purified using phenol-chloroform extraction, and the total amount of transcribed mRNA was determined by measuring optical density. Cytoplasmic extracts of 3T3-L1 cells were prepared in binding buffer containing 10 mM HEPES, pH 7.5, 90 mM potassium acetate, 1.5 mM magnesium acetate, 2.5 mM DTT, 0.05% NP-40, and protease inhibitor cocktails. After the addition of 0.1 U/µl RNase inhibitor (Promega) and 20 µg/µl yeast tRNA (Ambion), lysates were absorbed by heparin-agarose (Sigma-Aldrich) and streptavidin-Sepharose (Invitrogen) beads for approximately 1 h at 4°C. After brief centrifugation, the supernatant was collected and total protein was quantified using the Bradford reagent. The same amount of total protein was incubated with biotin-labeled RNA probe and streptavidin-Sepharose for 3 h at 4°C. After washing five times with the binding buffer, the pulled-down RNA-protein complexes were subjected to immunoblotting with anti-TTP, anti-ZFP36L1, and anti-ZFP36L2.
Transfection and luciferase reporter assays
HEK293T cells were seeded in six-well plates and co-transfected using the calcium phosphate precipitation method with 0.5 µg of pCMV-Luc-Mkp-1 3′UTR or pCMV-Luc control vectors, together with 1 µg of pSV-β-galactosidase as an internal control. pCMV-Flag-TTP, -ZFP36L1, or -ZFP36L2 (0.2 μg or 0.5 μg) were also transfected into the cells. After 24 h, the cells were harvested and assayed for luciferase and β-galactosidase activities. Luciferase activity was determined using a luminometer (Packard, Downers Grove, IL, USA) with luciferin (Promega) as the substrate. β-galactosidase activity was determined using a standard colorimetric assay with 2-nitrophenyl β-D-galactopyranoside (Sigma-Aldrich) as the substrate. WCEs were then prepared and analyzed by immunoblotting with specific antibodies. Luciferase activity was primarily normalized with β-galactosidase activity and secondarily normalized with control sets. Each treatment group contained triplicate cultures, and each experiment was independently performed three times.
GST pull-down assays
Glutathione-Sepharose 4B beads (approximately 8 µl, GE Healthcare Life Sciences, Piscataway, NJ, USA) were incubated with bacterially expressed GST or GST-14-3-3 proteins in PBS containing 1% (v/v) Triton X-100 on a rotary shaker for 20 min at room temperature. After washing three times with the same buffer, the beads were combined with 300 µg cell lysates from 3T3-L1 cells that had undergone various treatments in a final volume of 200 µl of buffer containing 20 mM HEPES, pH 7.9, 100 mM NaCl, 2.5 mM MgCl2, 0.1 mM EDTA, 0.05% NP-40, and 1% Triton X-100, along with 1 mM DTT, and 1 mM PMSF (Phenylmethylsulfonyl fluoride), on a rotary shaker. The mixtures were incubated at 4°C for 2 h, and then the beads were washed four times with the latter buffer lacking DTT and PMSF but containing 0.2 M NaCl and once with 50 mM Tris, pH 6.8. Bound proteins were eluted by boiling in SDS-PAGE sample buffer and analyzed by immunoblotting. To block ZFP36L1 phosphorylation, 3T3-L1 cells were induced with FMDI in the presence of 10 μM U0126 (Sigma-Aldrich), 100 nM wortmannin (Sigma-Aldrich), or both together for 30 min.
Gene knockdown assays
shRNAs against mouse Zfp36l1, Zfp36l2, and Luciferase were obtained from the national RNAi core facility located at the Institute of Molecular Biology, Genomic Research Center, Academia Sinica. The target sequences were 5′-CTTCGAAATGTCCGTTCGGTT-3′ (TRCN0000072243) for Luciferase, 5′-CCGCTGCCACTTCATTCATAA-3′ (TRCN0000123470) for Zfp36l1, and 5′-CCAAACACTTAGGTCTCAGAT-3′ (TRCN0000173172) for Zfp36l2. For virus production, HEK293T cells were co-transfected with vectors necessary for lentivirus packaging, including 14 µg of pCMV∆R8.91, 2 µg of pMD.G, and 14 µg of hairpin-pLKO.1 plasmids for cells cultured in 100-mm dishes. Between 6 and 8 h after transfection, the transfection media were replaced by fresh 3T3-L1 culture media, and the virus-containing media were collected twice every 24 h, added to 8 μg/ml polybrene, and stored at 4°C for less than 2 weeks. Before infection, the virus-containing media were briefly centrifuged at 1,200 × g for 5 min at room temperature to remove 293T cell debris.
For virus infection in 3T3-L1 cells, 40-50% confluent cells in a 60-mm dish were infected with 3 ml virus-containing medium (the primary infection). After 24 h, another infection was conducted using the same procedure (super infection). The virus-containing medium was replaced by fresh culture medium 1 day later, followed by 2 µg/ml puromycin selection the subsequent day. The culture media containing 2 µg/ml puromycin were changed every 2 days until the infected 3T3-L1 cells reached 100% confluency. siRNAs used to knock down TTP and MKP-1 were purchased from Dharmacon (ON-TARGETplus SMARTpool against mouse Zfp36 and Mkp-1, Lafayette, CO, USA). Specific siRNAs, or non-targeting control siRNA, were delivered into 50% confluent 3T3-L1 cells using Dharmafect 3 transfection reagent (Dharmacon). At 48 h post-transfection, confluent 3T3-L1 cells were induced to differentiate. Total lysates were then immunoblotted. Total RNA was purified and subjected to real time-PCR analysis using specific primers.
Oil red O staining
After 6 days of adipogenic differentiation, 3T3L1 cells were washed once in PBS and fixed with 3.7% formaldehyde in PBS for 1.5 h at 37°C. Cells were then rinsed three times with distilled water and stained with 0.1% oil red O solution (in 60% isopropanol) for 3 h at 37°C. The oil red O was extracted with isopropanol and quantified by measuring the optical density at 510 nm.
Statistical analysis
All of the data are presented as the mean ± SD of at least three independent experiments. The statistically significant values (*P<0.05, **P<0.01 or *** P<0.001) were determined by one-tailed Student's t-test.
Results
Generation of specific antibodies against individual TTP family members
The TTP family members have highly conserved amino acid sequences in the TZF domains and C-terminal ends (Fig. 1A). Commercial antisera against TTP family members recognize the TZF domain, causing cross-reactions among members. To distinguish these members clearly, peptides in non-conserved regions among TTP family members were used to produce antisera (Fig. 1A, underlined sequences). These non-conserved regions (residues 185-306 in TTP, 225-324 in ZFP36L1, and 300-448 in ZFP36L2) were expressed as recombinant proteins with a maltose-binding protein tag at the N terminus and a His-tag at the C terminus (Fig. 1B). After purifying these recombinant proteins and immunizing rabbits, specificity tests on induced antisera, conducted with GFP fusion proteins expressed in 293T cells, indicated that only the corresponding antiserum could recognize the transfected cell lysate that transiently overexpressed GFP-TTP, GFP-ZFP36L1, or GFP-ZFP36L2 (Fig. 1C). These antisera could, therefore, distinguish the TTP family proteins.
Analysis of ZFP36L1, ZFP36L2, and TTP expression profiles during 3T3-L1 early differentiation
Our previous studies demonstrated that induction of TTP during early differentiation of 3T3-L1 preadipocytes downregulates IEG expression [29,30]. However, these investigations did not evaluate ZFP36L1 and ZFP36L2 expression in 3T3-L1 cells. We therefore used immunoblotting to determine the expression profiles of ZFP36L1 and ZFP36L2 in early differentiation of 3T3-L1 preadipocytes (Fig. 2A). In contrast to the inducible expression of TTP, ZFP36L1 and ZFP36L2 were constitutively expressed. Analyses identified multiple bands of ZFP36L1 and ZFP36L2 in 3T3-L1 preadipocytes, which shifted to higher molecular weight positions following induction of differentiation. After CIP treatment of the samples from 0 and 30 min of induction, the multiple ZFP36L1 and ZFP36L2 bands returned to the lower-migrating positions, suggesting the presence of phosphorylated products (Fig. 2B). However, CIP treatment did not change the mobility of TTP (Fig. 2B). The mRNA expression profiles of the three TTP members differed during differentiation (Fig. 2C). Zfp36l1 mRNA slightly increased to 1.8 fold at 30 min of induction and decreased to ~0.2 fold after 2 h of induction. Zfp36l2 mRNA level changed only minimally (1.9 to 0.6 fold) during the differentiation period. Ttp mRNA level underwent rapid induction (a 50-fold increase), then rapidly decreased to basal level after 4 h of induction (Fig. 2C). Although ZFP36L1 and ZFP36L2 protein levels did not completely correlate with their mRNA levels, they showed minimal changes in expression and underwent phosphorylation in response to differentiation signals.
ZFP36L1 and ZFP36L2 bind to Mkp-1 AREs and negatively control Mkp-1 mRNA expression
ZFP36L1 and ZFP36L2 contain an RNA-binding TZF domain similar to that of TTP. We therefore evaluated the interaction of ZFP36L1 and ZFP36L2 with Mkp-1 mRNA, a TTP target. The 3′UTR of Mkp-1 mRNA contains three AREs [29]. RNA pull-down assays demonstrated the RNA-protein interaction. Incubation of the biotinylated Mkp-1 3′UTR with 3T3-L1 cell lysates, followed by immunoblotting of the associated proteins, identified ZFP36L1 and ZFP36L2 in the complexes pulled down from the control and induced cell lysates. The levels of pulled-down proteins generally correlated with their expression levels (Fig. 3A). TTP expressed in 1 h- and 2 h-induced cell lysates pulled down by Mkp-1 ARE served as a positive control. To evaluate the mRNA destabilization activities of ZFP36L1 and ZFP36L2, we co-expressed their expression plasmids with the Mkp-1 3′UTR-mediated Luciferase reporter. As shown in Figure 3B, ectopic expression of ZFP36L1 greatly suppressed luciferase activity. Furthermore, compared to vector control, overexpression of ZFP36L1 and ZFP36L2 decreased the half-life of Luc-Mkp-1 mRNA via the Mkp-1 3′UTR from 217.7 min to 82.5 min and 70.8 min, respectively (Fig. 3C). In the immunoblot shown in Figure 3B, the FLAG-ZFP36L2 expression level seemed to be higher than that of FLAG-ZFP36L1, and the higher dose of ZFP36L2 (0.5 μg) increased the ARE-mediated luciferase activity as was reported for TTP [28].
Characterization of specific antibodies against TTP family proteins. (A) Alignment of the amino acid sequences of the three mouse TTP family members. The TZF domain and the C terminus are highly conserved. Black dots denote CCCH residues in the TZF domain. Underlining indicates regions used to produce antiserum for each protein. The amino acids which are the same in three family proteins are highlighted in black, and those only the same in two members are highlighted in gray. (B) Coomassie blue staining of purified TTP family members expressed in E. coli with a maltose-binding protein tag at the N terminus and a 6×His tag at the C terminus for antiserum production. L1: ZFP36L1; L2: ZFP36L2. (C) HEK293T cells expressed GFP-TTP, GFP-ZFP36L1 (L1), and GFP-ZFP36L2 (L2). WCEs (40 µg) were applied to each lane for immunoblotting with anti-TTP, anti-ZFP36L1, and anti-ZFP36L2. Antiserum dilution was 1:1000.
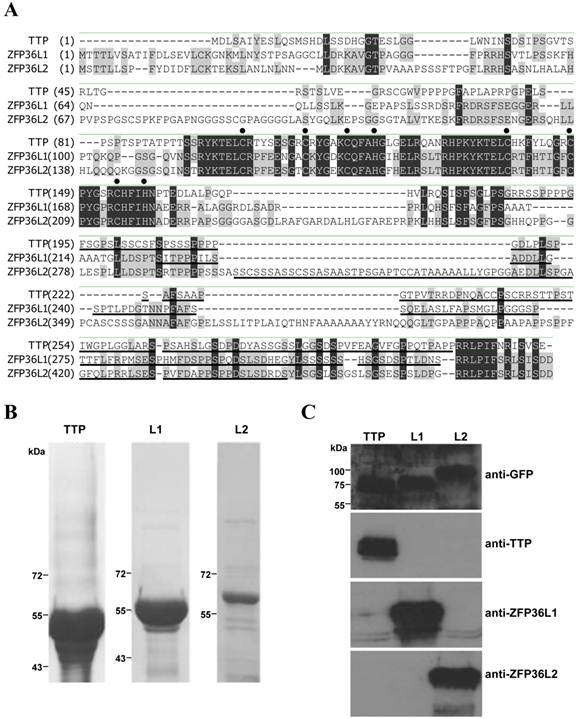
Protein and mRNA expression profiles of the TTP family during 3T3-L1 early differentiation. Two-day post-confluent 3T3-L1 cells were induced to differentiate by FMDI for the indicated times. (A) WCEs were used for immunoblotting using anti-TTP, -ZFP36L1, and -ZFP36L2. β-tubulin served as a protein loading control. (B) WCEs from 0-min and 30-min induction were subjected to in vitro CIP treatment for 1 h at 37°C, and then immunoblotting was performed as (A). Data are representative of three independent experiments with similar results. (C) RNA was isolated for quantitative PCR analysis to measure Ttp, Zfp36l1, and Zfp36l2 mRNA levels. Bars, mean ± SD of three independent samples normalized to β-actin mRNA levels.
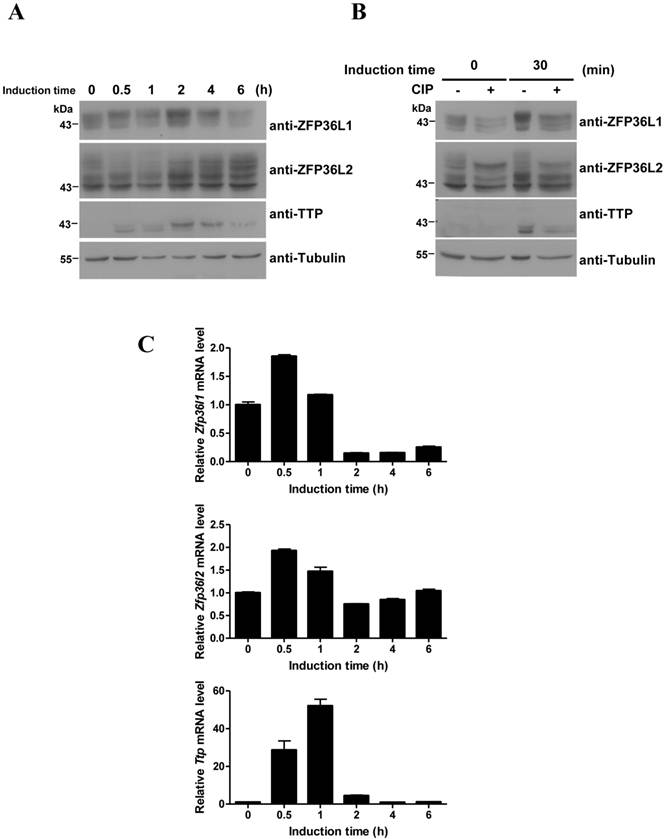
Physical and functional interaction of ZFP36L1 and ZFP36L2 with 3′UTR of Mkp-1 mRNA (A) RNA pull-down shows the interaction between the TTP family and Mkp-1 mRNA 3′UTR. Biotinylated Mkp-1 3′UTR was incubated with cytosolic extracts from 3T3-L1 cells induced with FMDI for 0, 0.5, 1, and 2 h, and then brought down by streptavidin-Sepharose beads. Biotinylated 18S rRNA served as a negative control. The RNA-protein complexes were separated by SDS-PAGE and immunoblotted with anti-TTP, anti-ZFP36L1, and anti-ZFP36L2. β-tubulin served as a negative control. A representative of three independent experiments with similar results is shown. (B) ZFP36L1 and ZFP36L2 downregulate Mkp-1 3′UTR-driven luciferase activity. HEK293T cells were co-transfected with 0.2 μg or 0.5 μg of FLAG-tagged ZFP36L1 or ZFP36L2 expression plasmid and a pCMV-Luciferase reporter construct containing the Mkp-1 3′UTR. The ARE-derived luciferase assay results were normalized with β-galactosidase activity to correct for variations in transfection efficiency, and also normalized with the activity of a luciferase reporter lacking the ARE to compensate for transcriptional activation (upper panel). The lower panel shows the ectopic expression of the proteins as revealed by immunoblotting with antibodies against the indicated proteins. Each reaction contained duplicate cultures, and each experiment was repeated 3 or 4 times independently. **P < 0.01; ns, not significant. (C) ZFP36L1 and ZFP36L2 downregulate Mkp-1 3′UTR-mediated Luciferase RNA stability. HEK293T cells were co-transfected with pCMV-Luciferase-MKP-1 3′UTR, FLAG-ZFP36L1, or FLAG-ZFP36L2 expression plasmid, and pCMV-β-galactosidase. After 24 h, the cells were left untreated or treated with 10 μg/ml actinomycin D to block transcription for 30 min or 60 min. RNA was subjected to quantitative PCR analysis using primers directed to the luciferase and β-galactosidase genes. The relative Luciferase mRNA levels are shown. The mRNA half-life (t1/2) was calculated and indicated. *P < 0.05, **P < 0.01.
ZFP36L1and TTP temporally modulate Mkp-1 mRNA expression during early differentiation of 3T3-L1 cells
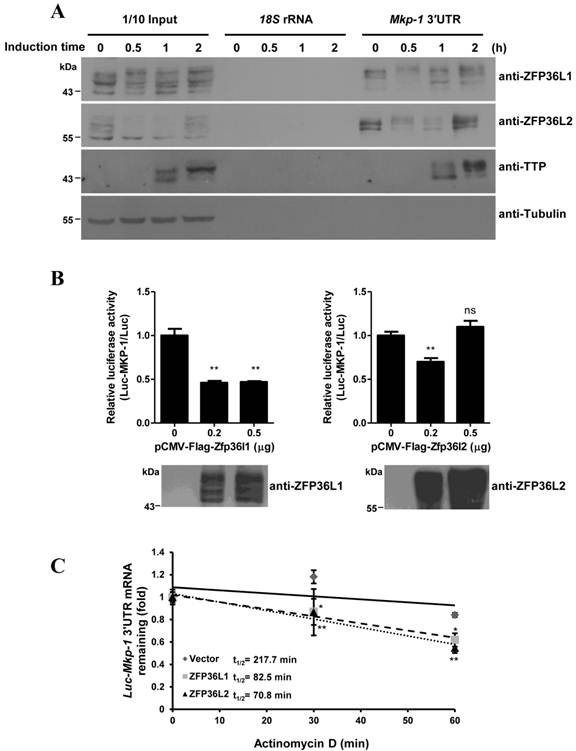
Mkp-1 mRNA was barely detectable in 3T3-L1 preadipocytes. After induction of differentiation, Mkp-1 mRNA rapidly increased to its highest level at 40 min of induction and rapidly decreased thereafter (Fig. 4A, upper graph). MKP-1 was induced at 60 min and gradually decreased after 3 h (Fig. 4A, lower panel). The mRNA half-life during early differentiation was determined by treatment with an RNA synthesis inhibitor (Fig. 4B). Mkp-1 mRNA was more stable at 30 min of induction (t1/2=285.1 min), when Mkp-1 mRNA levels in Figure 4A increased. However, its half-life decreased at 1 h and 2 h of induction (t1/2=28.5 min and 23.0 min, respectively), when Mkp-1 mRNA markedly decreased. TTP was knocked down by siRNA, and then regulation of Mkp-1 mRNA expression during 3T3-L1 differentiation was evaluated. Following TTP knockdown, the half-life of Mkp-1 mRNA increased at 1 h of induction (increased from control 16.6 min to 39.0 min), and MKP-1 protein levels increased at 3 h of induction (Fig. 4C and D), suggesting that inducible TTP controls the rapid decay of Mkp-1 mRNA.
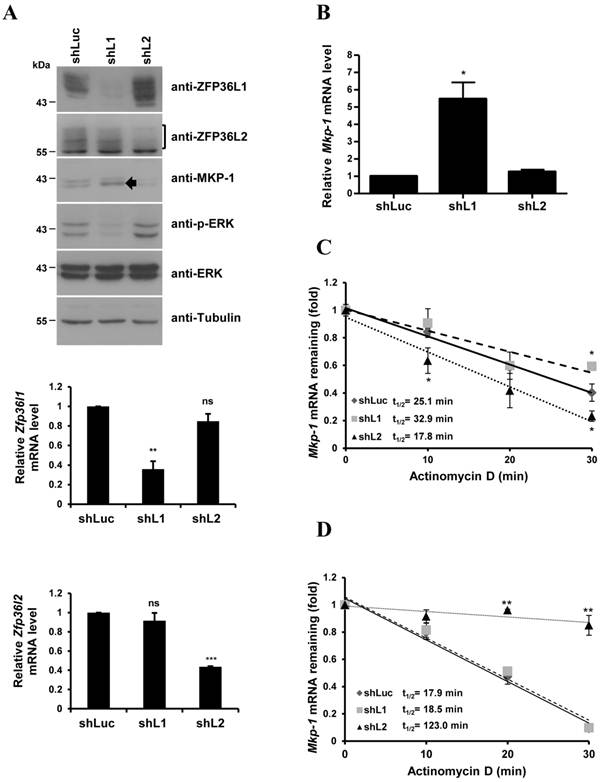
Lentiviruses carrying specific shRNAs were used to knockdown ZFP36L1 and ZFP36L2 (Fig. 5A). The knockdown efficiency was examined in the decrease of protein levels by immunoblotting (Fig.5A, upper panel) and the decrease of mRNA levels by quantitative PCR (Fig.5A, lower panel). ZFP36L1 knockdown, but not ZFP36L2 knockdown, upregulated MKP-1 protein and downregulated ERK activation (Fig. 5A). The basal Mkp-1 mRNA level was increased with ZF36L1 shRNA compared with the knockdown of control and ZFP36L2 shRNAs (Fig. 5B). IEG mRNAs (c-Myc, c-Jun, c-Fos, and Ttp) were also increased with ZFP36L1 shRNA (Supplementary Material: Fig. S1). As shown in Figure 5C, knockdown of ZFP36L1 extended the Mkp-1 mRNA half-life (from control 25.1 min to 32.9 min) under the non-differentiation condition. In contrast, knockdown of ZFP36L1 did not alter the Mkp-1 mRNA half-life at 1 h of induction, whereas knockdown of ZFP36L2 increased the half-life (Fig. 5D). Thus, the TTP family proteins collectively contribute to the temporal modulation of Mkp-1 mRNA stability in 3T3-L1 cells.
Phosphorylated ZFP36L1 interacts with 14-3-3
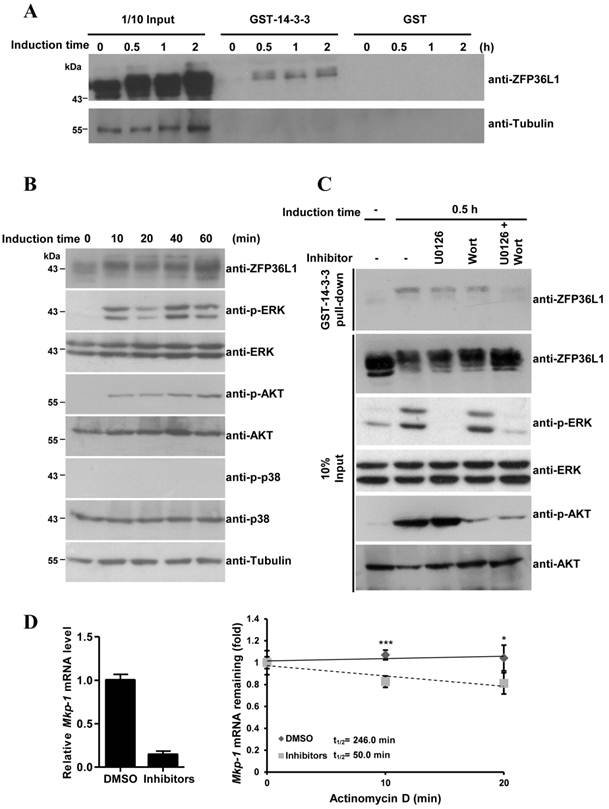
As described above, constitutive ZFP36L1 bound to Mkp-1 mRNA 3′UTR to destabilize its mRNA and reduce basal Mkp-1 mRNA levels. To explain the stabilization of Mkp-1 mRNA at 30 min of induction (Fig. 4B), we examined whether ZFP36L1 phosphorylation could inhibit its function by interacting with the signal adaptor 14-3-3, as described [20]. GST-fused 14-3-3 was incubated with 3T3-L1 cell lysates at 0, 30 min, 1 h, and 2 h post-induction of differentiation, and immunoblotting was used to analyze the associated protein complexes (Fig. 6A). GST-14-3-3 pulled down higher-molecular weight ZFP36L1 in differentiation-induced cell lysates but not in control lysates, suggesting that GST-14-3-3 might associate with phosphorylated ZFP36L1. To explore which kinases phosphorylate ZFP36L1 during 3T3-L1 differentiation, antibodies against activated kinases were used to assess the cell lysates from the 3T3-L1 differentiation time course (Fig. 6B). Phospho-ERK and phospho-AKT, but not phospho-p38, were detected in the differentiation-induced cell lysates, indicating the activation of both ERK and AKT signaling pathways. GST-14-3-3 pull-down assays, performed using cell lysates treated with an inhibitor of ERK and/or AKT signaling, were used to evaluate the regulation of the interaction between 14-3-3 and ZFP36L1 by these signaling pathways. As shown in Figure 6C, blockage of both ERK and AKT signaling pathways (by U0126 and wortmannin, respectively) inhibited ZFP36L1 phosphorylation and decreased the GST-14-3-3 pull-down levels. Subsequently, in the presence of U0126 and wortmannin, the expression level and half-life of Mkp-1 mRNA was decreased at 15 min of differentiation induction (Fig. 6D). This result suggested that differentiation-induced signaling caused the phosphorylation of ZFP36L1, which then interacted with 14-3-3 to diminish its mRNA destabilization activity. Owing to the decreased activities of ZFP36L1, its mRNA target (Mkp-1) was more stable at 30 min of induction than at longer differentiation times.
Functional analysis of TTP family proteins in adipogenesis
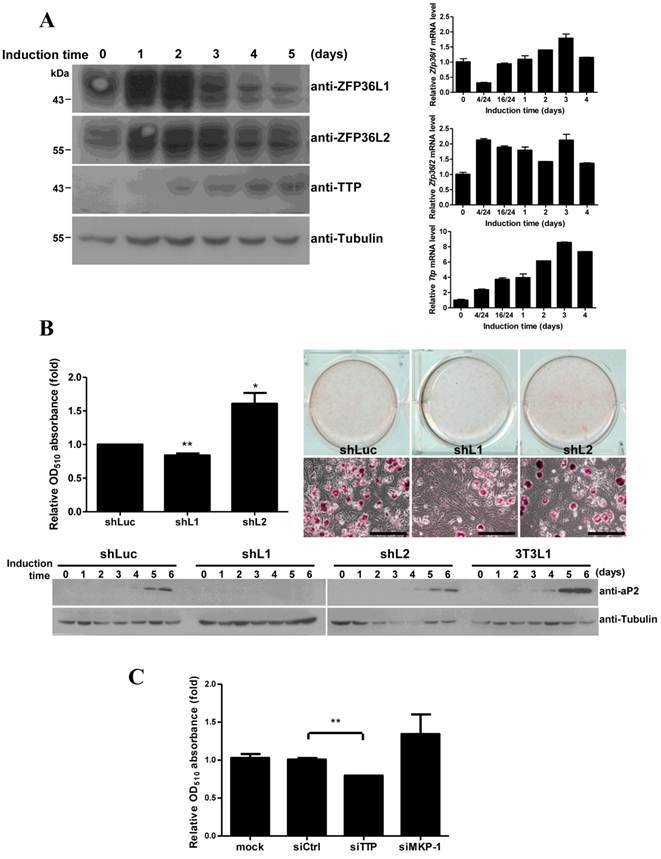
To evaluate the roles of TTP family proteins in adipogenesis, we investigated their expression during longer periods of 3T3-L1 differentiation. Analysis of mRNA and protein levels revealed high expression of both ZFP36L1 and ZFP36L2 proteins and slight expression of TTP during differentiation over 5 days (Fig. 7A). Knockdown of ZFP36L1 blocked adipogenesis, as evaluated using oil red O staining (Fig. 7B), whereas knockdown of ZFP36L2 increased adipogenesis. Immunoblotting revealed that the adipogenic marker aP2 was not induced in ZFP36L1 knockdown cells (Fig. 7B, lower panel). Cell number determination showed that knockdown of ZFP36L1 downregulated mitotic clonal expansion (Supplementary Material: Fig. S2). Knockdown of TTP by siRNA also inhibited adipogenesis (Fig. 7C). MKP-1 knockdown increased adipogenesis (Fig. 7C), which was consistent with the downregulation of MKP-1 by TTP and ZFP36L1. TTP family proteins differentially regulate MKP-1 expression during early differentiation of 3T3-L1 cells, and this regulation is critical for cell differentiation.
Knockdown of TTP family proteins affects Mkp-1 mRNA stability during 3T3-L1 differentiation. (A) Expression profiles of Mkp-1 mRNA (upper panel) and protein (lower panel) during the early differentiation stage of adipogenesis. Two-day post-confluent 3T3-L1 preadipocytes were induced with FMDI for the indicated times. RNA was isolated for quantitative PCR analysis. The graph shows the mean ± SD of relative Mkp1 mRNA levels normalized to β-actin mRNA levels from three independent samples. WCEs were immunoblotted with anti-MKP-1 anti-β-tubulin. (B) Analysis of Mkp-1 mRNA half-life after differentiation induction for 0.5 h, 1 h, and 2 h. Actinomycin D (10 µg/ml) was added for 0, 10 min, 20 min, and 30 min to stop transcription. RNA was isolated for quantitative PCR analysis. The half-life of Mkp-1 mRNA (t1/2) was determined and indicated. *P<0.05, **P<0.01 and *** P<0.001. (C) and (D) 3T3-L1 cells at 50% confluency were transfected with TTP-specific siRNA or with control non-targeting siRNA (siCtrl). After 48 h, cells were induced with FMDI for 0, 3, and 6 h. WCEs were isolated and then immunoblotted with anti-TTP, anti-MKP-1, and anti-β-tubulin (C). (D) Mkp-1 mRNA half-life analysis following TTP knockdown. After the addition of FMDI for 1 h, cells were exposed to 10 µg/ml actinomycin D for the indicated times, and then Mkp-1 mRNA was analyzed using quantitative RT-PCR. The relative amounts of Mkp-1 mRNA are presented after normalizing to β-actin mRNA values. The mRNA half-life (t1/2) was calculated and indicated.*P < 0.05.
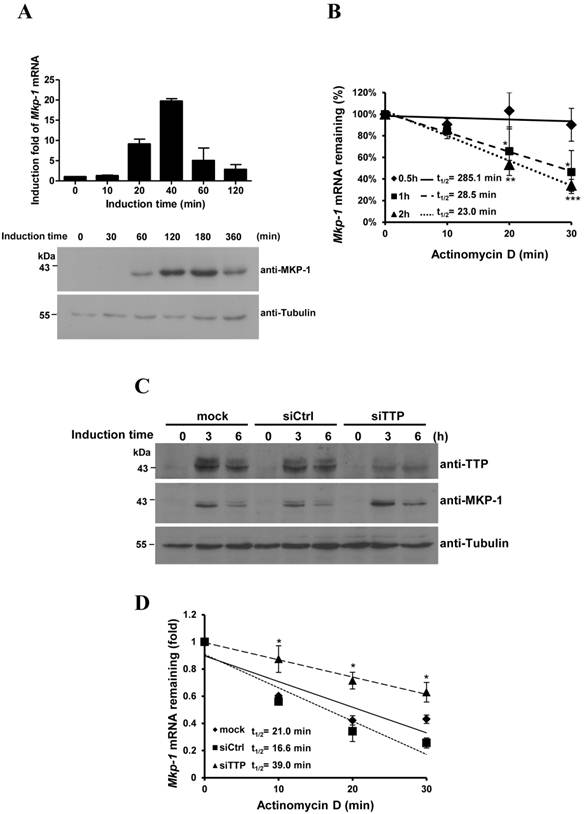
Knockdown of ZFP36L1 affects Mkp-1 mRNA stability in preadipocytes. (A) 3T3-L1 preadipocytes were infected with lentivirus carrying shRNA against ZFP36L1 (shL1), ZFP36L2 (shL2), or luciferase (shLuc, negative control) mRNA. WCEs from ZFP36L1 or ZFP36L2 knockdown cells were immunoblotted with anti-ZFP36L1, anti-ZFP36L2, anti-MKP-1, anti-p-ERK, anti-ERK, and anti-β-tubulin (upper panel). The bracket indicates the multiple forms of ZFP36L2. The arrow indicates the increased MKP-1 protein under knockdown of ZFP36L1. RNA was isolated for quantitative PCR to analyze the expression of Zfp36l1 and Zfp36l2 (lower panel). **P < 0.01; ***P < 0.001; ns, not significant. (B) ZFP36L1 knockdown by shL1 increases the basal level of Mkp-1 mRNA. RNA was isolated from knockdown cells for quantitative PCR with primers to Mkp-1 and β-actin. The relative amounts of Mkp-1 mRNA are presented after normalizing to β-actin mRNA values as the mean ± SD from three independent samples. *P < 0.05. (C) Mkp-1 mRNA half-life is increased in ZFP36L1 knockdown cells. 3T3-L1 cells were treated with shLuc, shL1, or shL2, and actinomycin D was added for the indicated times to stop RNA synthesis. RNA was isolated for quantitative PCR. Each experiment was performed three to five times independently. The mRNA half-life (t1/2) was calculated and indicated. *P < 0.05. (D) Determination of Mkp-1 mRNA half-life at 1 h of differentiation in control and knockdown cells. Two-day post-confluent 3T3-L1 cells knocked down with control shLuc, shL1, or shL2 were induced to differentiate for 1 h followed by actinomycin D treatment for 0 min (untreated), 10 min, 20 min, and 30 min. RNA was isolated for quantitative PCR with primers directed to Mkp-1 and β-actin. The mRNA half-life (t1/2) was calculated and indicated. **P < 0.01.
Phosphorylated ZFP36L1 interacts with 14-3-3. (A) GST pull-down analysis. E. coli-expressed GST-14-3-3 or GST were bound on glutathione-Sepharose beads. The beads were incubated with 3T3-L1 cell lysates after induction of differentiation for 0, 30 min, 1 h, and 2 h. The associated proteins were separated using SDS-PAGE and immunoblotted with anti-ZFP36L1 and anti-β-tubulin. (B) Activation of ERK and AKT during 3T3-L1 differentiation. Two-day post-confluent 3T3-L1 cells were induced with FMDI for 0, 10, 20, 40, and 60 min. The WCEs were immunoblotted with anti-p-ERK, anti-p-AKT, anti-p-p38, anti-ERK, anti-AKT, anti-p38 and anti-β-tubulin. (C) Interaction between ZFP36L1 and 14-3-3 is blocked by inhibitors of ERK and AKT signaling pathways. Two-day post confluent 3T3-L1 cells were induced with FMDI in the presence of 10 μM U0126, 100 nM wortmannin, or both for 30 min. The cell extracts were isolated for GST pull-down assays, and the pulled-down protein complexes were analyzed by immunoblotting with anti-ZFP36L1. One-tenth input of cell extracts was assessed with anti-ZFP36L1, anti-p-ERK, anti-ERK, anti-p-AKT, and anti-AKT. A representative of three independent experiments with similar results is shown. (D) Inhibition of ERK and AKT signaling pathways downregulates Mkp-1 RNA stability. Two-day post-confluent 3T3-L1 cells were untreated or pretreated with U0126 and wortmannin for 30 min, and then differentiation was induced for 15 min in the presence or absence of U0126 and wortmannin. Subsequently, 10 μg/ml of actinomycin D was added for 0, 10 min, and 20 min to block transcription, and RNA was isolated for quantitative PCR. The expression level of Mkp-1 mRNA under non-treatment of actinomycin D was shown (left panel). The regression curves and Mkp-1 mRNA half-lives were presented (right panel). *P < 0.05, ***P < 0.001.
Functional analysis of ZFP36L1 and ZFP36L2 in adipogenesis. (A) Expression profiles of TTP family proteins and mRNAs during adipogenesis. Two-day post-confluent 3T3-L1 cells were induced to differentiate for 1 to 5 days, and WCEs were prepared at the indicated times and immunoblotted with anti-TTP, anti-ZFP36L1, anti-ZFP36L2. Tubulin served as a loading control (left panel). RNA was isolated at indicated time for quantitative PCR (right panel). (B) The effects of ZFP36L1 and ZFP36L2 knockdown on adipogenesis. Lentivirus was used to infect 3T3-L1 with shRNA against mouse ZFP36L1, ZFP36L2, or luciferase (control). Infected 3T3-L1 cells were induced to differentiate for 6 days. The cells were stained with oil red O and observed with microscope, scale bar, 50 μm (right panel). Oil red O was extracted using isopropanol, and OD510 was measured to quantify triglyceride accumulation (left panel). The lower panel shows the expression of the adipogenic marker aP2 after differentiation for 0 to 6 days. *P < 0.05, **P < 0.01. (C) The effect of TTP and MKP-1 knockdown on adipogenesis. 3T3-L1 cells were transfected with siRNA against TTP, MKP-1, or non-targeting control. After induction for 6 days, the cells were stained with oil red O to quantify the differentiation. The graph shows the mean ± SD of relative OD510 from three independent experiments. **P < 0.01.
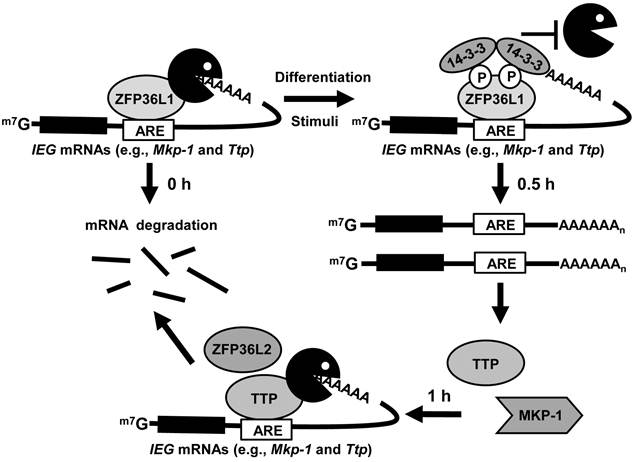
Schematic depicting the regulation of TTP family proteins in 3T3-L1 cells. TTP family proteins sequentially and post-transcriptionally regulate expression of ARE-containing IEGs during early differentiation of 3T3-L1 preadipocytes. In preadipocytes, ZFP36L1 bound to IEG mRNAs to limit their basal levels. Following induction of differentiation, phosphorylated ZFP36L1 associated with 14-3-3 to inhibit its mRNA destabilization activity and then stabilized IEG mRNAs. The newly produced TTP then rapidly degraded the induced IEG mRNAs. ZFP36L2, like TTP, may function under induction conditions.
Discussion
In this study, we generated antibodies against individual TTP family members to determine their protein expression profiles during 3T3-L1 differentiation. In preadipocytes, ZFP36L1 bound to Mkp-1 mRNA to limit its levels. Following induction of differentiation, phosphorylated ZFP36L1 associated with 14-3-3 to inhibit its mRNA destabilization activity and then stabilized Mkp-1 mRNA. The newly produced TTP then rapidly degraded the induced Mkp-1 mRNA (Fig.8). Knockdown of ZFP36L1 and TTP resulted in increased Mkp-1 mRNA at basal and induction levels, respectively, and suppressed adipogenesis.
Although previous reports have described expression of the three TTP family member mRNAs in adipocytes, macrophages, keratinocytes, human tissue, and cancer cell lines [35-38], no prior investigation has clearly characterized TTP protein expression patterns. We generated antibodies against each member of the TTP family, thus providing a tool with which to study their functions. Immunoblotting confirmed the specificity of these antisera and verified that there was no cross reactivity. Immunoblotting of the endogenous proteins using these antisera revealed different protein expression patterns during 3T3-L1 differentiation. This is the first study to describe the protein expression of endogenous ZFP36L2.
SDS-PAGE revealed multiple forms of ZFP36L1 and ZFP36L2 detectable in 3T3-L1 cell lysates. TTP, ZFP36L1, and ZFP36L2 contain numerous serines and threonines (mouse TTP contains 57 serines and 17 threonines in 319 residues, ZFP36L1 contains 49 serines and 21 threonines in 338 residues, and ZFP36L2 contains 73 serines and 21 threonines in 484 residues), suggesting their potential for extensive phosphorylation. However, phosphatase treatment only partially altered the protein mobility, suggesting that other protein modifications were present or that association with other proteins prevented phosphatase function [39]. The ERK and AKT signaling pathways were activated during 3T3-L1 differentiation, demonstrating their involvement in ZFP36L1 phosphorylation. According to recent reports, TTP interacts with the Ccr4-Caf1-Not deadenylase complex to trigger mRNA destabilization [17-19], whereas TTP phosphorylated by MK2 decreases deadenylase recruitment [17,18]. In addition, phosphorylated TTP interacts with 14-3-3 to prevent TTP from dephosphorylation by protein phosphatase 2A [39]. Our observation of differentiation-induced phosphorylation of ZFP36L1 and interaction with 14-3-3 thus suggests the downregulation of mRNA destabilizing activity, causing Mkp-1 mRNA stabilization at 30 min of induction (Fig.4B). Consistent with this result, when both ERK and AKT signaling pathways were inhibited by pharmaceutical inhibitors, the expression level and half-life of Mkp-1 mRNA were decreased (Fig.6D). Previous reports demonstrated that ZFP36L1 is phosphorylated by AKT at Ser92 and Ser203 [9,20], whereas ERK-mediated phosphorylation has not been adequately defined. The expression profiles of ZFP36L1 mRNA and protein did not fully correlate. ZFP36L1 mRNA decreased to one-fifth of its initial level at 2 h of induction, whereas its protein level showed minimal change. This might be caused by ZFP36L1 phosphorylation and stabilization that maintains an adequate ZFP36L1 level to regulate mRNA stability in response to alterations in environmental conditions [9]. The Zfp36l1 mRNA level recovered after 1 day of differentiation (Fig. 7 A), indicating that a complex mechanism governs the transcriptional regulation of this mRNA.
ZFP36L1 and ZFP36L2 contain conserved RNA-binding TZF domains. In vitro RNA pull-down analysis revealed that both proteins can associate with Mkp-1 mRNA. However, the functional reporter assay indicated that ZFP36L1 downregulated Mkp-1 ARE-mediated luciferase activity more effectively than ZFP36L2 did. ZFP36L1 knockdown cells, but not ZFP36L2 knockdown cells, showed changes in Mkp-1 mRNA expression and mRNA half-life in non-differentiated 3T3-L1 cells. However, the Mkp-1 mRNA half-life was increased at 1 h of differentiation in ZFP36L2 knockdown cells (Fig. 5D). These results indicate that ZFP36L1 downregulates Mkp-1 mRNA expression to control its basal level, whereas ZFP36L2, like TTP, may function under induction conditions, although the detailed mechanism by which ZFP36L2 regulates mRNA stability is unclear. The interacting proteins of the TTP family might, therefore, exert different functional effects on specific mRNA targets. Except for the TZF domain, the regions of the TTP family are not highly conserved, suggesting that they support different protein interaction networks. For example, the Cbl-interaction protein CIN85 interacts with the C terminus of human TTP but not of other members [40], and TTP, but not ZFP36L1, specifically interacts with the Tax protein from human T lymphotropic virus 1 via the C-terminal domain [41]. The C terminus of ZFP36L2 contains a unique proline-rich domain that may be involved in specific protein-protein interactions [38].
This study showed that knockdown of ZFP36L1 or TTP decreased adipogenesis (Fig. 7). Knockdown of ZFP36L1 expression also impaired mitotic clonal expansion, which is required for completion of differentiation [42]. The TTP family proteins are ARE-binding and RNA-destabilizing proteins and might, therefore, regulate some ARE-containing mRNAs with tightly controlled expression in adipogenesis. Target mRNAs of the TTP family were not systematically identified in 3T3-L1 cells. Mkp-1 mRNA might represent one of the ZFP36L1 and TTP targets in 3T3-L1 cells. In this study, induction of TTP controlled the transient expression of Mkp-1 mRNA, causing a reduction in its half-life at 1 h of induction. The constitutive expression of ZFP36L1 in 3T3-L1 cells served to control the basal expression of Mkp-1 mRNA at low levels. Bost et al. showed that differentiation inducers of preadipocytes activate MAPKs and that this activation is reversible [43]. Previous studies have also described an emerging family of dual-specificity protein phosphatases—the MKPs—which inactivate MAPKs via dephosphorylation of both threonine and tyrosine residues [44-46]. MKP-1, also referred to as DUSP1, 3CH134, CL100, or ERP, is the archetype of the MKP family. Mkp-1 is an IEG that is highly inducible by a broad variety of extracellular stimuli that are related to cell proliferation, differentiation, and apoptosis [47-50]. ERK is dephosphorylated and inactivated by MKP-1 [51,52]. Moreover, ERK activation is required for 3T3-L1 differentiation [53-55]. In the present study, the shRNA-mediated decrease in ZFP36L1 expression caused an increase in basal Mkp-1 mRNA levels, which may inhibit ERK activation and thereby downregulate adipogenesis. ERK signaling stimulates Mkp-1 transcription [56,57]; therefore, inactivation of ERK signaling caused by ZFP36L1 knockdown might decrease the induction of Mkp-1 transcripts in response to differentiation. Consistent with this, our findings indicate that knockdown of MKP-1 weakly promoted adipogenesis. It is unknown if ZFP36L1 has a similar biological function in vivo because ZFP36L1-knockout mice die at day 11 [24]. A range of results have shown the involvement of ZFP36L1 in myogenic differentiation. For example, a previous study showed that siRNA inhibition of ZFP36L1 expression in mouse myoblasts inhibits cell differentiation [58]. However, a different investigation showed that induction of shRNA specific for ZFP36L1 strongly stimulates cardiomyocyte formation within embryoid bodies [59]. In contrast, ZFP36L2 knockdown promoted adipogenesis compared with the shRNA control, indicating that TTP family proteins might target different mRNAs and play different functional roles under physiological conditions.
Manipulation of adipocyte differentiation might provide a means to combat obesity. Although a hierarchy of transcription factors control adipogenesis [60,61], little is known regarding the involvement of post-transcriptional regulation in adipocyte differentiation. Our findings indicate that the temporal expression of TTP family proteins controls the expression of ARE-containing IEGs, such as Mkp-1, during differentiation of 3T3-L1 cells. This is a novel regulation mode in adipogenesis.
Supplementary Material
Figure S1: Knockdown of ZFP36L1 in 3T3-L1 cells results in increasing IEG mRNAs. Figure S2: Mitotic clonal expansion is inhibited in ZFP36L1-knockdown 3T3-L1 cells.
Acknowledgements
We thank Yi-Li Liu and I-Ching Huang for technical support with DNA sequencing, supported in part by the Department of Medical Research in NTUH (National Taiwan University Hospital), and Dr. Sheng-Wei Lin for protein purification support. This work was supported by Academia Sinica, National Taiwan University (10R70602B1 to G. D. Chang) and National Science Council (grant number: NSC 97-2311-B-001-019-MY3 to C. J. Chang).
Competing Interests
The authors have declared that no competing interest exists.
References
1. Blackshear PJ. Tristetraprolin and other CCCH tandem zinc-finger proteins in the regulation of mRNA turnover. Biochem Soc Trans. 2002;30(Pt 6):945-52
2. Baou M, Jewell A, Murphy JJ. TIS11 family proteins and their roles in posttranscriptional gene regulation. J Biomed Biotechnol. 2009;2009:634520
3. Blackshear PJ, Phillips RS, Ghosh S. et al. Zfp36l3, a rodent X chromosome gene encoding a placenta-specific member of the Tristetraprolin family of CCCH tandem zinc finger proteins. Biol Reprod. 2005;73(2):297-307
4. Lykke-Andersen J, Wagner E. Recruitment and activation of mRNA decay enzymes by two ARE-mediated decay activation domains in the proteins TTP and BRF-1. Genes Dev. 2005;19(3):351-61
5. Blackshear PJ, Lai WS, Kennington EA. et al. Characteristics of the interaction of a synthetic human tristetraprolin tandem zinc finger peptide with AU-rich element-containing RNA substrates. J Biol Chem. 2003;278(22):19947-55
6. Brewer BY, Malicka J, Blackshear PJ. et al. RNA sequence elements required for high affinity binding by the zinc finger domain of tristetraprolin: conformational changes coupled to the bipartite nature of Au-rich MRNA-destabilizing motifs. J Biol Chem. 2004;279(27):27870-7
7. Lai WS, Kennington EA, Blackshear PJ. Tristetraprolin and its family members can promote the cell-free deadenylation of AU-rich element-containing mRNAs by poly(A) ribonuclease. Mol Cell Biol. 2003;23(11):3798-812
8. Lai WS, Blackshear PJ. Interactions of CCCH zinc finger proteins with mRNA: tristetraprolin-mediated AU-rich element-dependent mRNA degradation can occur in the absence of a poly(A) tail. J Biol Chem. 2001;276(25):23144-54
9. Benjamin D, Schmidlin M, Min L. et al. BRF1 protein turnover and mRNA decay activity are regulated by protein kinase B at the same phosphorylation sites. Mol Cell Biol. 2006;26(24):9497-507
10. Cao H, Deterding LJ, Venable JD. et al. Identification of the anti-inflammatory protein tristetraprolin as a hyperphosphorylated protein by mass spectrometry and site-directed mutagenesis. Biochem J. 2006;394:285-97
11. Brook M, Tchen CR, Santalucia T. et al. Posttranslational regulation of tristetraprolin subcellular localization and protein stability by p38 mitogen-activated protein kinase and extracellular signal-regulated kinase pathways. Mol Cell Biol. 2006;26(6):2408-18
12. Hitti E, Iakovleva T, Brook M. et al. Mitogen-activated protein kinase-activated protein kinase 2 regulates tumor necrosis factor mRNA stability and translation mainly by altering tristetraprolin expression, stability, and binding to adenine/uridine-rich element. Mol Cell Biol. 2006;26(6):2399-407
13. Maitra S, Chou CF, Luber CA. et al. The AU-rich element mRNA decay-promoting activity of BRF1 is regulated by mitogen-activated protein kinase-activated protein kinase 2. RNA. 2008;14(5):950-9
14. Chrestensen CA, Schroeder MJ, Shabanowitz J. et al. MAPKAP kinase 2 phosphorylates tristetraprolin on in vivo sites including Ser178, a site required for 14-3-3 binding. J Biol Chem. 2004;279(11):10176-84
15. Johnson BA, Stehn JR, Yaffe MB. et al. Cytoplasmic localization of tristetraprolin involves 14-3-3-dependent and -independent mechanisms. J Biol Chem. 2002;277(20):18029-36
16. Stoecklin G, Stubbs T, Kedersha N. et al. MK2-induced tristetraprolin:14-3-3 complexes prevent stress granule association and ARE-mRNA decay. EMBO J. 2004;23(6):1313-24
17. Marchese FP, Aubareda A, Tudor C. et al. MAPKAP kinase 2 blocks tristetraprolin-directed mRNA decay by inhibiting CAF1 deadenylase recruitment. J Biol Chem. 2010;285(36):27590-600
18. Clement SL, Scheckel C, Stoecklin G. et al. Phosphorylation of Tristetraprolin by MK2 Impairs AU-Rich Element mRNA Decay by Preventing Deadenylase Recruitment. Mol Cell Biol. 2011;31(2):256-66
19. Sandler H, Kreth J, Timmers HT. et al. Not1 mediates recruitment of the deadenylase Caf1 to mRNAs targeted for degradation by tristetraprolin. Nucleic Acids Res. 2011;39(10):4373-86
20. Schmidlin M, Lu M, Leuenberger SA. et al. The ARE-dependent mRNA-destabilizing activity of BRF1 is regulated by protein kinase B. EMBO J. 2004;23(24):4760-9
21. Carrick DM, Lai WS, Blackshear PJ. The tandem CCCH zinc finger protein tristetraprolin and its relevance to cytokine mRNA turnover and arthritis. Arthritis Res Ther. 2004;6(6):248-64
22. Taylor GA, Carballo E, Lee DM. et al. A pathogenetic role for TNF alpha in the syndrome of cachexia, arthritis, and autoimmunity resulting from tristetraprolin (TTP) deficiency. Immunity. 1996;4(5):445-54
23. Carballo E, Lai WS, Blackshear PJ. Feedback inhibition of macrophage tumor necrosis factor-alpha production by tristetraprolin. Science. 1998;281(5379):1001-5
24. Stumpo DJ, Byrd NA, Phillips RS. et al. Chorioallantoic fusion defects and embryonic lethality resulting from disruption of Zfp36L1, a gene encoding a CCCH tandem zinc finger protein of the Tristetraprolin family. Mol Cell Biol. 2004;24(14):6445-55
25. Ramos SB, Stumpo DJ, Kennington EA. et al. The CCCH tandem zinc-finger protein Zfp36l2 is crucial for female fertility and early embryonic development. Development. 2004;131(19):4883-93
26. Stumpo DJ, Broxmeyer HE, Ward T. et al. Targeted disruption of Zfp36l2, encoding a CCCH tandem zinc finger RNA-binding protein, results in defective hematopoiesis. Blood. 2009;114(12):2401-10
27. Hodson DJ, Janas ML, Galloway A. et al. Deletion of the RNA-binding proteins ZFP36L1 and ZFP36L2 leads to perturbed thymic development and T lymphoblastic leukemia. Nat Immunol. 2010;11(8):717-24
28. Brooks SA, Connolly JE, Rigby WF. The role of mRNA turnover in the regulation of tristetraprolin expression: evidence for an extracellular signal-regulated kinase-specific, AU-rich element-dependent, autoregulatory pathway. J Immunol. 2004;172(12):7263-71
29. Lin NY, Lin CT, Chang CJ. Modulation of immediate early gene expression by tristetraprolin in the differentiation of 3T3-L1 cells. Biochem Biophys Res Commun. 2008;365(1):69-74
30. Lin NY, Lin CT, Chen YL. et al. Regulation of tristetraprolin during differentiation of 3T3-L1 preadipocytes. FEBS J. 2007;274(3):867-78
31. Kuwano Y, Kim HH, Abdelmohsen K. et al. MKP-1 mRNA stabilization and translational control by RNA-binding proteins HuR and NF90. Mol Cell Biol. 2008;28(14):4562-75
32. Emmons J, Townley-Tilson WH, Deleault KM. et al. Identification of TTP mRNA targets in human dendritic cells reveals TTP as a critical regulator of dendritic cell maturation. RNA. 2008;14(5):888-902
33. Horner TJ, Lai WS, Stumpo DJ. et al. Stimulation of polo-like kinase 3 mRNA decay by tristetraprolin. Mol Cell Biol. 2009;29(8):1999-2010
34. Lai WS, Parker JS, Grissom SF. et al. Novel mRNA targets for tristetraprolin (TTP) identified by global analysis of stabilized transcripts in TTP-deficient fibroblasts. Mol Cell Biol. 2006;26(24):9196-208
35. Cao H, Urban JF Jr, Anderson RA. Insulin increases tristetraprolin and decreases VEGF gene expression in mouse 3T3-L1 adipocytes. Obesity (Silver Spring). 2008;16(6):1208-18
36. Hacker C, Valchanova R, Adams S. et al. ZFP36L1 is regulated by growth factors and cytokines in keratinocytes and influences their VEGF production. Growth Factors. 2010;28(3):178-90
37. Carrick DM, Blackshear PJ. Comparative expression of tristetraprolin (TTP) family member transcripts in normal human tissues and cancer cell lines. Arch Biochem Biophys. 2007;462(2):278-85
38. Liang J, Song W, Tromp G. et al. Genome-wide survey and expression profiling of CCCH-zinc finger family reveals a functional module in macrophage activation. PLoS One. 2008;3(8):e2880
39. Sun L, Stoecklin G, Van Way S. et al. Tristetraprolin (TTP)-14-3-3 complex formation protects TTP from dephosphorylation by protein phosphatase 2a and stabilizes tumor necrosis factor-alpha mRNA. J Biol Chem. 2007;282(6):3766-77
40. Kedar VP, Darby MK, Williams JG. et al. Phosphorylation of human tristetraprolin in response to its interaction with the Cbl interacting protein CIN85. PLoS One. 2010;5(3):e9588
41. Twizere JC, Kruys V, Lefebvre L. et al. Interaction of retroviral Tax oncoproteins with tristetraprolin and regulation of tumor necrosis factor-alpha expression. J Natl Cancer Inst. 2003;95(24):1846-59
42. Tang QQ, Lane MD. Adipogenesis: From stem cell to adipocyte. Annu. Rev. Bioch. 2012;81:21.1-21.22
43. Bost F, Aouadi M, Caron L. et al. The role of MAPKs in adipocyte differentiation and obesity. Biochimie. 2005;87(1):51-6
44. Keyse SM. Protein phosphatases and the regulation of mitogen-activated protein kinase signalling. Curr Opin Cell Biol. 2000;12(2):186-92
45. Pouyssegur J, Lenormand P. Fidelity and spatio-temporal control in MAP kinase (ERKs) signalling. Eur J Biochem. 2003;270(16):3291-9
46. Tamura S, Hanada M, Ohnishi M. et al. Regulation of stress-activated protein kinase signaling pathways by protein phosphatases. Eur J Biochem. 2002;269(4):1060-6
47. Keyse SM, Emslie EA. Oxidative stress and heat shock induce a human gene encoding a protein-tyrosine phosphatase. Nature. 1992;359(6396):644-7
48. Liu Y, Gorospe M, Yang C. et al. Role of mitogen-activated protein kinase phosphatase during the cellular response to genotoxic stress. Inhibition of c-Jun N-terminal kinase activity and AP-1-dependent gene activation. J Biol Chem. 1995;270(15):8377-80
49. Brondello JM, Brunet A, Pouyssegur J. et al. The dual specificity mitogen-activated protein kinase phosphatase-1 and -2 are induced by the p42/p44MAPK cascade. J Biol Chem. 1997;272(2):1368-76
50. Xu Q, Fawcett TW, Gorospe M. et al. Induction of mitogen-activated protein kinase phosphatase-1 during acute hypertension. Hypertension. 1997;30:106-11
51. Kassel O, Sancono A, Kratzschmar J. et al. Glucocorticoids inhibit MAP kinase via increased expression and decreased degradation of MKP-1. EMBO J. 2001;20(24):7108-16
52. Ito A, Suganami T, Miyamoto Y. et al. Role of MAPK phosphatase-1 in the induction of monocyte chemoattractant protein-1 during the course of adipocyte hypertrophy. J Biol Chem. 2007;282(35):25445-52
53. Sale EM, Atkinson PG, Sale GJ. Requirement of MAP kinase for differentiation of fibroblasts to adipocytes, for insulin activation of p90 S6 kinase and for insulin or serum stimulation of DNA synthesis. EMBO J. 1995;14(4):674-84
54. Benito M, Porras A, Nebreda AR. et al. Differentiation of 3T3-L1 fibroblasts to adipocytes induced by transfection of ras oncogenes. Science. 1991;253(5019):565-8
55. Porras A, Nebreda AR, Benito M. et al. Activation of Ras by insulin in 3T3 L1 cells does not involve GTPase-activating protein phosphorylation. J Biol Chem. 1992;267(29):21124-31
56. Li J, Gorospe M, Hutter D. et al. Transcriptional induction of MKP-1 in response to stress is associated with histone H3 phosphorylation-acetylation. Mol Cell Biol. 2001;21(23):8213-24
57. Lasa M, Abraham SM, Boucheron C. et al. Dexamethasone causes sustained expression of mitogen-activated protein kinase (MAPK) phosphatase 1 and phosphatase-mediated inhibition of MAPK p38. Mol Cell Biol. 2002;22(22):7802-11
58. Busse M, Schwarzburger M. et al. Strong induction of the Tis11B gene in myogenic differentiation. Eur J Cell Biol. 2008;87(1):31-8
59. Wegmuller D, Raineri I, Gross B. et al. A cassette system to study embryonic stem cell differentiation by inducible RNA interference. Stem Cells. 2007;25(5):1178-85
60. MacDougald OA, Lane MD. Transcriptional regulation of gene expression during adipocyte differentiation. Annu Rev Biochem. 1995;64:345-73
61. Farmer SR. Transcriptional control of adipocyte formation. Cell Metab. 2006;4(4):263-73
Author contact
![]() Corresponding author: Ching-Jin Chang, Graduate Institute of Biochemical Sciences, National Taiwan University, No.1 Sec 4 Roosevelt Rd. Taipei 106, Taiwan. Tel: 886-2-23620261ext 5583, Fax: 886-2-23635038. E-mail: chingjinsinica.edu.tw
Corresponding author: Ching-Jin Chang, Graduate Institute of Biochemical Sciences, National Taiwan University, No.1 Sec 4 Roosevelt Rd. Taipei 106, Taiwan. Tel: 886-2-23620261ext 5583, Fax: 886-2-23635038. E-mail: chingjinsinica.edu.tw

 Global reach, higher impact
Global reach, higher impact