Impact Factor ISSN: 1449-2288
Int J Biol Sci 2013; 9(10):1013-1020. doi:10.7150/ijbs.7191 This issue Cite
Review
Roles of Eukaryotic Initiation Factor 5A2 in Human Cancer
1. Sun Yat-sen University Cancer Center; State Key Laboratory of Oncology in South China. Collaborative Innovation Center of Cancer Medicine.
2. Department of Clinical Oncology, the University of Hong Kong, Hong Kong, China.
Received 2013-7-18; Accepted 2013-9-26; Published 2013-10-12
Abstract
Eukaryotic initiation factor 5A (eIF5A), the only known cellular protein containing the amino acid hypusine, is an essential component of translation elongation. eIF5A2, one of the two isoforms in the eIF5A family, is reported to be a novel oncogenic protein in many types of human cancer. Both in vitro and in vivo studies showed that eIF5A2 could initiate tumor formation, enhance cancer cell growth, and increase cancer cell motility and metastasis by inducing epithelial-mesenchymal transition. Accumulatied evidence suggests that eIF5A2 is a useful biomarker in the prediction of cancer prognoses and serves as an anticancer molecular target. In this review, we will focus on updating current knowledge of the EIF5A2 gene in human cancers. The molecular mechanisms of EIF5A2 related to tumorigenesis will also be discussed.
Keywords: Eukaryotic initiation factor 5A2, Cancer, Epithelial-mesenchymal transition, Metastasis.
Introduction
Amplification of 3q26.2 is one of the most frequent genetic alternations found in solid tumors. In 2000, our group isolated a putative oncogene, eukaryotic translation initiation factor 5A2 (EIF5A2), from 3q26.2, using the chromosome microdissection- hybrid selection technique [1]. In humans, eIF5A2 and its isoform eIF5A1 are the only two eukaryotic proteins that contain a unique hypusine residue [2, 3]. Hypusine is a polyamine-derived amino acid that is generated in eIF5A by a post-translational enzymatic modification that occurs in two steps. The first step involves polyamine spermidine cleavage and the transfer of its 4-aminobutyl group to a specific lysine residue of the eIF5A precursor by deoxyhypusine synthase (DHS), thus forming a deoxyhypusine residue [4, 5]. Subsequent hydroxylation at carbon 2 of the transferred 4-aminobutylmoiety intermediated by deoxyhypusine hydroxylase (DOHH) produces the hypusine residue, a mature and active form of eIF5A [6, 7]. Initially, eIF5A was thought to act in the final stage of the initiation phase of protein synthesis by promoting the formation of the first peptide bond, however, rencent reports have also suggested that eIF5A participates in translation elongation [8, 9]. To date, two eIF5A isoforms have been identified and have been found to share about 80% of their cDNA sequences and 94% of their proteins. Both human eIF5A1 and eIF5A2 were shown to complement growth of eIF5A2 null yeast, indicating similar functions between the two human isoforms in terms of eukaryotic cell survival [10, 11]. However, the genes encoding eIF5A1 and eIF5A2 are located on different chromosomes. The EIF5A gene resides on 17p12-p13, a genetically stable chromosome. eIF5A1 protein is ubiquitously expressed in almost all cells and tissues and plays crucial roles in translation elongation and RNA metabolism [9]. In contrast, eIF5A2 expression is tissue and cell-type-specific, and is mainly found in testis, brain and several cancer cell lines and tissues [10]. Our previous studies showed that over-expression of eIF5A2 could initiate tumor formation, promote cancer cell growth and enhance cell invasion/metastasis by inducing epithelial-mesenchymal transition (EMT) both in vitro and in vivo, suggesting it might have an oncogenic role in mammals. In this article, we will focus on EIF5A2-related studies that have been published over the last ten years, particularly those articles pertaining to its modification, sub-cellular location, upstream regulation and roles in cancer cell proliferation, invasion/metastasis, prognosis and prospects as a form of cancer treatment.
Protein Modification and Sub-cellular Location
eIF5A2 is a small (approximately 17kDa) universally conserved acidic protein classified in the eIF family. The eIF family represents a group of proteins that are involved in the initiation step of the protein translation. Each member of the eIF family plays a unique role in the initiation process by interacting with ribosomal subunits and mRNAs to form an elongation competent complex [12]. With robust molecular genetics and biochemical studies, Saini et al. verify that eIF5A promotes translation elongation [9]. Different from other eIF members, eIF5A2 and its isoform eIF5A1 have a polyamine-derived amino acid, hypusine, in their primary structures. Hypusine modification at the lysine-50 by DHS and DOHH is mandatory for the maturation of eIF5A2 protein [11]. Previous studies have shown that many kinds of cytokines and enzymes can enhance or decrease cell growth by regulating hypusine synthesis [13-17]. For example, transglutaminases (TGase), calcium-dependent enzymes, can catalyze formation of a cross-link between eIF5A hypusine residue and dimethylcasein [13, 15]. However, the impacts of the aforementioned cytokines and enzymes on eIF5A2 warrants further investigation. The amino acid sequences of the human eIF5A isoforms are conserved in the areas where hypusination takes place. The divergent residues between the human eIF5A1 and eIF5A2 are mainly located on the C-terminal domain [18], suggesting that different functions between the two isoforms may be linked to the C-terminal domain. The function of the C-terminal domain of eIF5A1 is shown by the finding that the mutation of Ser 149 to Pro in yeast eIF5A1 decreases general protein synthesis by 30% and increases mRNA stability [19]. However the biological effects of eIF5A2 C-terminal region remain unclear. As to the structure of eIF5A2, in brief, the N-terminal domain is dominated by β-strands and the C-terminal domain consists of a three-turn α-helix α2 and five strands of β7-β11 [18].
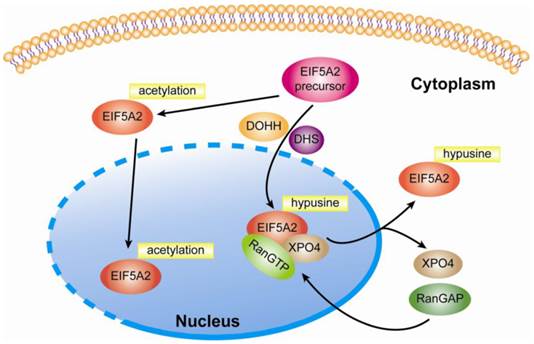
In addition to unique hypusine modification, Ishfaq et al. find that both eIF5A isoforms are acetylated at lysine-47, and hypusination of K50 can reduce acetylation in eIF5A2. Moreover, both HDAC6 and SIRT were shown to deacetylate eIF5A2 in SW480 cells when treated with three HDAC inhibitors, trichostatin A (TSA), nicotinamide (NA), and SCOP402. The authors further investigated the impact of hypusination and acetylation on the subcellular localization of eIF5A2 via its expression of Flag-tagged eIF5A2 wild type and the mutants in Hela cells. The results showed that acetylation induced nuclear accumulation and hypusination induced cytoplasmice localization of eIF5A2 [20, 21]. Additionally, in 1995, Hannelore Klier et al. purified and characterized one major and three minor isoforms of human eIF5A from Hela cells. The main form, which accounts for approximately 95% of all eIF5A, carries hypusine at position 50 and is amino-terminally acetylated as determined by amino acid composition analysis and electrospray ionization mass spectrometry. In contrast to the main form, all three minor isoforms of eIF5A are characterized by acetylation of lysine at position 47. Furthermore, no phosphorylation was found in any of the purified human eIF5A isoforms [22]. With regards to the cellular location of eIF5A, Lipowsky et al. found that exportin 4 (Exp4 or XPO4) mediated the nuclear export of eIF5A1 by means of the trimetric eIF5A1-Exp4-RanGTP complex, which required eIF5A1 hypusine modification [23]. In line with these findings, Zender and colleagues identified that exportin 4 (XPO4) was the nuclear export mediator of eIF5A2 in that nuclear accumulation of eIF5A1 or eIF5A2 was found in XPO4 deficient cells [24]. Taken together, this evidence indicates that eIF5A2 is a shuttling protein responsible for regulating genes' protein elongation in the cytoplasm (Fig 1.). Until now, eIF5A2 was still thought to be a cytoplasm protein and no studies have shown that it can act as a transcriptional factor in the nucleus.
Modification and subcellular location of eIF5A2. After modification by DHS and DOHH, eIF5A2 protein is transformed into a mature and active form. Nuclear eIF5A2 in complex with RanGTP and XPO4 mediates the export of mature eIF5A2 like eIF5A1. However, if acetylation at lysine-47 occurs, eIF5A2 might also be accumulated at the nucleus. Since it is not clear whether the hypusination and acetylation of eIF5A occur in cytoplasm or nuclei, a dotted line is applied to suggest the possible occurrence of the two processes in both compartments.
Upstream Regulation
The EIF5A2 gene resides on chromosome 3q26, a region that is frequently amplified in different human malignancies, including pancreatic [25], esophageal [26, 27], prostate [28], lung [29, 30], gastric [31, 32], ovarian [1, 33], colorectal [10, 34], liver [35], bladder [36], breast [37], and nasopharyngeal carcinoma [38] (Table 1). However, over-expression of eIF5A2 protein does not always necessarily mean the amplification of the gene. For instance, in the ovarian cancer cell line UACC-1598, which has a high fold amplification of the EIF5A2 gene, the level of EIF5A2 mRNA is much higher than that of EIF5A, yet the amount of eIF5A2 protein is merely comparable to that of eIF5A1 and no modified eIF5A2 is detectable in most cells (even when EIF5A2 mRNA is present) [11]. Using semi-quantitative RT-PCR and real-time PCR methods, we have shown that over-expression of eIF5A2 occurred in 50/81 (61.7%, P <0.0001, independent Student's t test) of HCCs. However, we did not detect a significant copy number change in these HCC samples when using semi-quantitative genomic PCR [35]. In 16 bladder urothelial carcinoma tissue samples, amplification of EIF5A2 was not observed even when eIF5A2 protein was over-expressed [36]. These results, taken together, suggest that other mechanisms, such as transcriptional regulation and posttranscriptional regulation, might play an important role in eIF5A2 regulation. Clement et al. found that the efficiency of eIF5A2 mRNA translation mainly affected the expression level of eIF5A2 protein since there was little difference in stability between eIF5A1 and eIF5A2 protein [39]. Furthermore, they observed that the open reading frames (ORFs) of the two isoforms were expressed almost equally in 293T cells, suggesting that the negative elements that inhibit the translation of EIF5A2 mRNA reside in the 5'-UTR or 3'-UTR. Previous studies also showed that poly (A) tail length played a critical role in regulating translation. Since the long mRNA containing 3.8-kb 3'-UTR is the major form of EIF5A2 mRNAs in most cells, 3'-UTRs of EIF5A2 may contribute to its variations in its protein expression among different cells [39]. Recently, a large body of studies demonstrated that microRNA can regulate gene expression at the posttranscriptional level by base-pairing with the 3'-untranslated region (UTR) of target mRNAs, which leads to translational repression or message degradation [40]. We therefore speculate that microRNA may take part in the regulation of EIF5A2 gene expression. However, the role of non-coding RNA (eg, microRNA, lncRNA) in the regulation of eIF5A2 expression is still enigmatic. Additionally, upstream signals, such as extracellular cytokine, membrane receptor and intracellular factor, which regulate the expression of EIF5A2 have not been sufficiently studied.
Summary of gene amplification and protein expression of EIF5A2 in human cancer.
| Reference | Materials | Methods | Main high level eIF5A2-related conclusions |
|---|---|---|---|
| Jenkins et al. [10] | 76 kinds of human tissues and cell lines | NB | Expression of EIF5A2 is limited to testis and parts of the brain |
| Guan et al. [1] | primary ovarian cancer samples and cell lines | SB, FISH | Amplification and over-expression of EIF5A2 in ovarian cancer |
| Clement et al. [11] | human cancer cell lines | RT-PCR, WB | Amplification of EIF5A2 in ovarian cancer and CRC cell lines |
| Guan et al. [33] | ovarian cancer TMA and cell lines | IHC, SB, NB, WB | Oncogenic role in the development of ovarian cancer |
| Clement et al. [39] | normal human cells and cancer cell lines | WB, NB | The expression of EIF5A2 is cell-specific |
| Xie et al. [45] | CRC TMA ,CRC tissues | IHC, FISH | Associated with CRC metastasis |
| Yang et al. [63] | ovarian tissues | IHC, FISH | Predicting outcome of ovarian cancer |
| Luo et al. [36] | paraffin and fresh bladder tissues | IHC, FISH, RT-PCR, WB | Predicting pTa/pT1 UC recurrence and progression |
| Lee et al. [47] | liver tissues and cell lines | IHC, qPCR, WB | Associated with prognosis of HCC |
| Chen et al. [64] | bladder cancer tissues and TMA | IHC, FISH, RT-PCR | Predicting outcome of UC |
| He et al. [65] | NSCLC TMA | IHC, FISH | Adverse prognosis marker of stage I NSCLC |
| Tang et al. [35] | HCC tissue TMA and cell lines | IHC, IF, RT-PCR, qPCR, WB | Promoting tumor metastasis in HCC |
| Zhu et al. [49] | CRC TMA and cell lines | IHC, WB, IF | Inducing EMT in CRC |
FISH, fluorescence in situ hybridization; IHC, immunohistochemistry; NB, northern blot; RT-PCR, reverse transcription and PCR; SB, southern blot; WB; western blot; IF, immunofluorescence; TMA: tissue microarray; UC: urothelial carcinoma; NSCLC, non-small cell lung cancer; HCC, hepatocellular carcinoma; CRC, colorectal cancer.
Function
Cell Proliferation
Accumulating evidence suggests that protein synthesis and its translational factors are involved in the regulation of cell proliferation and transformation. The IF (initiation factor) family plays a virtual role in the process of protein translation, and a growing body of evidence, which was excellently summarized by Caraglia et al., has shown that over-expression of these factors is involved in carcinogenesis, tumor cell proliferation and apoptosis [41]. It has been reported that eIF5A protein is essential for the sustained proliferation of mammalian cells [42]. Other studies have demonstrated the strong anti-proliferative effects of inhibitors of hypusine synthesis, such as N1-guanyl-1,7-diaminoheptane (GC7), a DHS inhibitor [43, 44] in mammalian cells, including various lines of human cancer cells. Moreover, IFN-α was found to inhibit human oropharyngeal epidermoid tumor cell growth by modulating hypusine synthesis, a process which might be accomplished through affecting eIF5A protein translation [14]. Such findings indicate that eIF5A is intimately involved in cell proliferation and may therefore contribute to malignant cell transformation. In agreement with these observations, our previous studies have shown that over-expression of eIF5A2 in NIH3T3 cells causes cell transformation and that EIF5A2 stably transfected LO2 cells (immortalized human liver cell line) displayed increased colony formation in soft agar and xenograph formation in nude mice. Similarly, the reduction of EIF5A2 in UACC-1598 inhibits cell growth; and the oncogenic ability of EIF5A2 can also be blocked by eIF5A2 silencing [1, 33, 39, 45]. In colorectal cancer, over-expression of eIF5A2 was significantly associated with the tumor cell proliferation rate detected by Ki-67 staining on tissue slices. Other groups demonstrated that eIF5A2 is required for proliferation of XPO4-deficient tumor cells and promotes hepatocellular carcinoma (HCC) in mice [24]. These results revealed the oncogenic role of eIF5A2 in the control of cancer cell proliferation, but the underling mechanism has not been elaborated upon. Perhaps in a role similar to that of eIF5A1, high levels of eIF5A2 protein may also enhance oncogenic translation.
Cancer Invasion/Metastasis
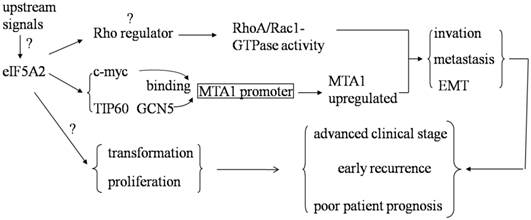
Metastasis is one of the main causes of mortality in the vast majority of solid tumors. Ramaswamy et al. found that 17 genes were associated with cancer metastasis by comparing the gene expression profiles of metastatic adenocarcinomas and unmatched primary adenocarcinomas in several types of human tumors. One of the 17 genes in the metastasis signature is deoxyhypusine synthase (DHPS), which is required for the hypusination of eIF5A2 [46]. In addition, a previous study showed that over-expression of eIF5A2 was significantly associated with lymphovascular invasion in colorectal carcinoma (CRC), venous infiltration in HCC, and a higher risk of lymph node metastasis in human gastric adenocarcinomas [45, 47, 48]. These results suggest that eIF5A2 may be responsible for the metastasis of human malignancies. We further investigated the role of eIF5A2 in the aggressive processes of HCC and CRC both in vitro and in vivo by using either loss-of-function or gain-of-function studies. We thus identified that eIF5A2 plays a pivotal role in promoting HCC cell metastasis by enhancing cell motility, promoting invasiveness, regulating the cytoskeleton, and activating EMT through Rho/Rac GTPases [35]. However, despite a lack of changes in Rho-GTPase activity, we also found that eIF5A2 enhances CRC cell invasion/metastasis and EMT mainly by up-regulating MTA1 expression through promoting c-myc, TIP60 and GCN5 binding with MTA1 promoter [49] (Fig 2.). These findings indicate that the metastasis-regulating mechanism of eIF5A2 may be tumor-type specific and is dependent on different cellular contexts. Clearly, the precise role of eIF5A2 in the regulation of cancer cell invasion and/or metastasis in different human cancers requires further investigation.
Clinical values of eIF5A2 in human cancer. Over-expression of eIF5A2 enhances RhoA/Rac1-GTPase activity in HCC and promotes c-myc, TIP60 and GCN5 to bind with MTA1 promoter and then up-regulate MTA1 expression in colon cancer. Both of these pathways can increase tumor cell invasion and metastasis by inducing EMT. Furthermore, eIF5A2 over-expression has the ability to transform NIH-3T3 fibroblasts, initiate tumor formation and enhance tumor cell growth. Based on its function and clinical data, eIF5A2 might be a pivotal human cancer oncogene.
Other Functions
To investigate the physiological and pathological effect of eIF5A2, Chen et al. have generated transgenic mice that over-express eIF5A2. Surprisingly, without spontaneous tumor formation, overexpression of eIF5A2 enhances the aging process of adult transgenic mice, including decreased growth rate and body weight, shortened life span, kyphosis, osteoporosis, delayed of wound healing and ossification. The authors also observed that eIF5A2 activation repressed p19, destabilized p53 and led to accumulation of chromosomal instability [50]. Previous studies have shown that the fate of oncogene activation is cell-specific and context dependent. For instance, c-Myc has been reported to enhance cell proliferation and malignant transformation in several kinds of human cancers since it was first reported by Bishop et al. in the 1970s [51-55]. But aberrant expression of c-myc can also induce apoptosis [56, 57]. Other studies have also shown that cellular senescence can be triggered by oncogene activation, as well as agents that damage DNA or alter chromatin structure [58, 59]. Future studies to investigate and confirm the other biological functions of EIF5A2 are warranted, and mice with EIF5A2 deficiency might be a good choice for such studies. In yeast, the eIF5A isoforms, eIF5A1 and eIF5A2, with 92% identity, are encoded by the two eIF5A genes, TIF51A and TIF51B respectively [60]. Although these 2 genes are reciprocally regulated by oxygen availability in yeast, they are functionally identical, and expression of either gene is sufficient for supporting yeast growth. Under aerobic conditions, only TIF51A is transcriptionally active, thus leading to eIF5A1 synthesis. In contrast, as oxygen is depleted and anaerobic conditions are established, TIF51B transcription is turned on when TIF51A is turned off, thereby ensuring the exclusive synthesis of eIF5A2 [61]. In a recent study, Vu et al. show that the hydroxylase activity of human DOHH is positively related to oxygen availability. In hypoxia, eIF5A cannot be fully hypusined [61]. Based on this evidence, we can speculate that the sub-cellular location of eIF5A2 may partly depend upon the availability of oxygen, but how oxygen affects the location eIF5A2 in mammals is not yet clear.
eIF5A2 and Prognosis of Human Cancers
Over the past ten years, many tissue-based studies have evaluated the expression of EIF5A2 mRNA and its encoded protein in human cancers. The majority of these studies demonstrated that eIF5A2 could be used as a biomarker for predicting prognosis. Over-expression of eIF5A2 was found to be correlated with a more advanced cancer stage and poor prognosis in patients with ovarian cancer [63]. In HCC, over-expression of eIF5A2 was reported to be associated with tumor features that indicate poor prognosis, such as the presence of tumor metastasis and venous infiltration [47]. A recent study from our group also showed similar results in CRC patients. Briefly summarized, a high level of eIF5A2 expression was found to be related to both lymph node and distant metastasis, as well as a more advanced clinical stage. In addition, eIF5A2 expression was an independent prognostic factor for poor CRC patients survival [48]. eIF5A2 may also play a role in the prognosis of patients with early stage tumors. For instance, a study assessing patients with bladder urothelial carcinoma showed that the overexpression of eIF5A2 as detected by IHC was able to predict the recurrence and progression of stage pTa/pT1 tumors [36]. In one of our previous studies, we concluded that the up-regulated expression of eIF5A2 might play an important role in the acquisition of a recurrence phenotype in superficial bladder cancer [64]. We reached a similar conclusion in one of our studies concerned with non-small cell lung cancer, which indicated that the overexpression of eIF5A2 was an adverse prognostic biomarker for the survival of stage I non-small cell lung cancer patients [65]. These studies demonstrate that eIF5A2 is a potential prognostic indicator in many solid cancers. However, as a cytoplasmic protein, eIF5A2 can only be assessed in tissue samples by PCR or IHC [66].
Implications for Therapy
eIF5A is of considerable interest as a potential therapeutic target in many human disorders. eIF5A is important in protein translation since disruption of the hypusination process by the DHS inhibitor, N1-guanyl- 1,7-diaminoheptane (GC7), has been shown to inhibit the growth of endothelial cells [67]. Moreover, mutants of eIF5A that cannot be hypusinated induce apoptosis in numerous cancer cell types including colon, cervical, skin, and lung cancer [68-70], thus inhibiting the growth of solid tumors [3]. eIF5A and syntenin can induce p53-dependent apoptosis collaboratively [71], but eIF5A has also been shown to initiate apoptosis through the mitochondrial pathway via the activation of caspases in p53-deficient cells [69]. In addition, the clinical drugs ciclopirox and deferiprone could impair transcription from HIV-1 promoters and decrease HIV-1 gene expression by inhibiting eIF5A hypusination [72]. Furthermore, eIF5A hypusination inhibition was demonstrated to have an anti-tumorigenic effect on leukemia cells when cells given either CPX alone or in combination with imatinib [73]. Our previous study showed that a combined treatment of eIF5A2 siRNA and GC7, an inhibitor of DHS, on HCC cells results in the synergistic inhibition of cell migration [35]. It was also shown that IFNα can inhibit tumor growth by decreasing hypusine synthesis and induce human lung epidermoid tumor cell apoptosis when combined with GC7 [14, 16]. Since the function of eIF5A is specifically regulated by hypusine modification [16, 41], a combination therapy of pharmacological agents which aim to inhibit the process of hypusine modification, such as GC7, ciclopirox (CPX) and cytokines, seems to be a reasonable recommendation for the future preclinical or clinic trials.
Conclusion and Perspectives
In summary, basic studies and clinical evidence show that eIF5A2 is a bona fide oncogene that may also be an important biomarker for the prognosis of many kinds of human tumors. In addition, antisense DNA against EIF5A2 or EIF5A2 specific siRNA effectively inhibits tumor cell growth and reduces tumor cells' ability to migrate. Therefore, eIF5A2 holds the potential to become a novel target for anticancer therapy. However, the underlying molecular mechanisms that regulate eIF5A2 expression remain enigmatic, such as how does non-coding RNA regulates the UTR of EIF5A2 and how its promoter is epigenetically modified. With regard to the downstream pathway, the exact mechanism of eIF5A2 in modulating and regulating its target as well as whether or not it can act as a transcriptional factor has not been elucidated. To further investigate the physiological and pathological role of eIF5A2 in mammals, eIF5A2 knockout mice may be needed.
Abbreviations
eIF5A2: eukaryotic initiation factor 5A2; EMT: epithelial-mesenchymal transition; DHPS: deoxyhypusine synthase; DOHH: deoxyhypusine hydroxylase; cDNA: complementary deoxyribonucleic acid; mRNA: message ribonucleic acid; HDAC: histonedeacetylase; SIRT2: sirtuin 2; TSA: trichostatin A; NA: nicotinamide; Exp4/XPO4: exportin 4; RT-PCR: reverse transcription polymerase chain reaction; HCC: hepatocellular carcinoma; ORF: open reading frame; UTR: untranslation region; lncRNA: long non-coding ribonucleic acid; siRNA: small interfering RNA; GC7: N1-guanyl-1,7-diaminoheptane; CRC: colorectal carcinoma; TIF: translation initiation factor; IHC: immunohistochemistry; HIV: human immunodeficiency virus. IFN α, Interferon α; TGase, transglutaminases; CPX, ciclopirox; DEF, deferiprone; EGF, epidermal growth factor.
Acknowledgements
This work was supported by grants from the Nature Science Foundation of China (No. 81225018) and the PhD Programs Foundation of the Ministry of Education of China (No 20110171110078).
Competing Interests
The authors have no financial conflicts of interest to disclose.
References
1. Guan XY, Sham JS, Tang TC. et al. Isolation of a novel candidate oncogene within a frequently amplified region at 3q26 in ovarian cancer. Cancer Res. 2001;61:3806-3809
2. Park MH, Nishimura K, Zanelli CF. et al. Functional significance of eIF5A and its hypusine modification in eukaryotes. Amino Acids. 2010;38:491-500
3. Caraglia M, Park MH, Wolff EC. et al. eIF5A isoforms and cancer: two brothers for two functions? Amino Acids. 2013;44:103-109
4. Park MH, Wolff EC, Folk JE. Hypusine: its post-translational formation in eukaryotic initiation factor 5A and its potential role in cellular regulation. Biofactors. 1993;4:95-104
5. Chen KY, Liu AY. Biochemistry and function of hypusine formation on eukaryotic initiation factor 5A. Biol Signals. 1997;6:105-109
6. Abbruzzese A. Developmental pattern for deoxyhypusine hydroxylase in rat brain. J Neurochem. 1988;50:695-699
7. Abbruzzese A, Park MH, Beninati S. et al. Inhibition of deoxyhypusine hydroxylase by polyamines and by a deoxyhypusine peptide. Biochim Biophys Acta. 1989;997:248-255
8. Caraglia M, Budillon A, Vitale G. et al. Modulation of molecular mechanisms involved in protein synthesis machinery as a new tool for the control of cell proliferation. Eur. J. Biochem. 2000;267:3919-3936
9. Saini P, Eyler DE, Green R. et al. Hypusine-containing protein eIF5A promotes translation elongation. Nature. 2009;459:118-121
10. Jenkins ZA, Haag PG, Johansson HE. Human eIF5A2 on chromosome 3q25-q27 is a phylogenetically conserved vertebrate variant of eukaryotic translation initiation factor 5A with tissue-specific expression. Genomics. 2001;71:101-109
11. Clement PM, Henderson CA, Jenkins ZA. et al. Identification and characterization of eukaryotic initiation factor 5A-2. Eur J Biochem. 2003;270:4254-4263
12. Jackson RJ, Hellen CU, Pestova TV. The mechanism of eukaryotic translation initiation and principles of its regulation. Nat Rev Mol Cell Biol. 2010;11:113-127
13. Beninati S, Nicolini L, Jakus J. et al. Identification of a substrate site for transglutaminases on the human protein synthesis initiation factor 5A. Biochem J. 1995;305:725-728
14. Caraglia M, Passeggio A, Beninati S. et al. Interferon a2 recombinant and epidermal growth factor modulate proliferation and hypusine synthesis in human epidermoid cancer KB cells. Biochem J. 1997;324:737-741
15. Beninati S, Gentile V, Caraglia M. et al. Tissue transglutaminase expression a¡ects hypusine metabolism in BALB/c 3T3 cells. FEBS Letters. 1998;437:34-38
16. Caraglia M, Marra M, Giuberti G. et al. The eukaryotic initiation factor 5A is involved in the regulation of proliferation and apoptosis induced by interferon-a and EGF in human cancer cells. J Biochem. 2003;133:757-765
17. Lalande M, Hanauske-Abel HM. A new compound which reversibly arrests T lymphocyte cell cycle near the G1/S boundary. Exp Cell Res. 1990;188:117-121
18. Tong Y, Park I, Hong BS. et al. Crystal structure of human eIF5A1: insight into functional similarity of human eIF5A1 and eIF5A2. Proteins. 2009;75:1040-1045
19. Zuk D, Jacobson A. A single amino acid substitution in yeast eIF-5A results in mRNA stabilization. EMBO J. 1998;17:2914-2925
20. Ishfaq M, Maeta K, Maeda S. et al. The role of acetylation in the subcellular localization of an oncogenic isoform of translation factor eIF5A. Biosci Biotechnol Biochem. 2012;76:2165-2167
21. Ishfaq M, Maeta K, Maeda S. et al. Acetylation regulates subcellular localization of eukaryotic translation initiation factor 5A (eIF5A). FEBS Lett. 2012;586:3236-3241
22. Klier H, Csonga R, Joao HC. et al. Isolation and structural characterization of different isoforms of the hypusine-containing protein eIF-5A from HeLa cells. Biochemistry. 1995;34:14693-14702
23. Lipowsky G, Bischoff FR, Schwarzmaier P. et al. Exportin 4: a mediator of a novel nuclear export pathway in higher eukaryotes. EMBO J. 2000;19:4362-4371
24. Zender L, Xue W, Zuber J. et al. An oncogenomics-based in vivo RNAi screen identifies tumor suppressors in liver cancer. Cell. 2008;135:852-864
25. Griffin CA, Morsberger L, Hawkins AL. et al. Molecular cytogenetic characterization of pancreas cancer cell lines reveals high complexity chromosomal alterations. Cytogenet Genome Res. 2007;118:148-156
26. Yen CC, Chen YJ, Pan CC. et al. Copy number changes of target genes in chromosome 3q25.3-qter of esophageal squamous cell carcinoma: TP63 is amplified in early carcinogenesis but down-regulated as disease progressed. World J Gastroenterol. 2005;11:1267-1272
27. Pack SD, Karkera JD, Zhuang Z. et al. Molecular cytogenetic fingerprinting of esophageal squamous cell carcinoma by comparative genomic hybridization reveals a consistent pattern of chromosomal alterations. Genes Chromosomes Cancer. 1999;25:160-168
28. Sattler HP, Lensch R, Rohde V. et al. Novel amplification unit at chromosome 3q25-q27 in human prostate cancer. Prostate. 2000;45:207-215
29. Viegas-Pequignot E, Flury-Herard A, De Cremoux H. et al. Recurrent chromosome aberrations in human lung squamous cell carcinomas. Cancer Genet Cytogenet. 1990;49:37-49
30. Shen H, Zhu Y, Wu YJ. et al. Genomic alterations in lung adenocarcinomas detected by multicolor fluorescence in situ hybridization and comparative genomic hybridization. Cancer Genet Cytogenet. 2008;181:100-107
31. Guan XY, Fu SB, Xia JC. et al. Recurrent chromosome changes in 62 primary gastric carcinomas detected by comparative genomic hybridization. Cancer Genet Cytogenet. 2000;123:27-34
32. Takada H, Imoto I, Tsuda H. et al. Screening of DNA copy-number aberrations in gastric cancer cell lines by array-based comparative genomic hybridization. Cancer Sci. 2005;96:100-110
33. Guan XY, Fung JM, Ma NF. et al. Oncogenic role of eIF-5A2 in the development of ovarian cancer. Cancer Res. 2004;64:4197-4200
34. He QJ, Zeng WF, Sham JS. et al. Recurrent genetic alterations in 26 colorectal carcinomas and 21 adenomas from Chinese patients. Cancer Genet Cytogenet. 2003;144:112-118
35. Tang DJ, Dong SS, Ma NF. et al. Overexpression of eukaryotic initiation factor 5A2 enhances cell motility and promotes tumor metastasis in hepatocellular carcinoma. Hepatology. 2010;51:1255-1263
36. Luo JH, Hua WF, Rao HL. et al. Overexpression of EIF-5A2 predicts tumor recurrence and progression in pTa/pT1 urothelial carcinoma of the bladder. Cancer Sci. 2009;100:896-902
37. Forozan F, Mahlamaki EH, Monni O. et al. Comparative genomic hybridization analysis of 38 breast cancer cell lines: a basis for interpreting complementary DNA microarray data. Cancer Res. 2000;60:4519-4525
38. Fang Y, Guan X, Guo Y. et al. Analysis of genetic alterations in primary nasopharyngeal carcinoma by comparative genomic hybridization. Genes Chromosomes Cancer. 2001;30:254-260
39. Clement PM, Johansson HE, Wolff EC. et al. Differential expression of eIF5A-1 and eIF5A-2 in human cancer cells. FEBS J. 2006;273:1102-1114
40. Bartel DP. MicroRNAs: genomics, biogenesis, mechanism, and function. Cell. 2004;116:281-297
41. Caraglia M, Marra M, Giuberti G. et al. The role of eukaryotic initiation factor 5A in the control of cell proliferation and apoptosis. Amino Acids. 2001;20:91-104
42. Wolff EC, Kang KR, Kim YS. et al. Posttranslational synthesis of hypusine: evolutionary progression and specificity of the hypusine modification. Amino Acids. 2007;33:341-350
43. Nishimura K, Murozumi K, Shirahata A. et al. Independent roles of eIF5A and polyamines in cell proliferation. Biochem J. 2005;385:779-785
44. Jasiulionis MG, Luchessi AD, Moreira AG. et al. Inhibition of eukaryotic translation initiation factor 5A (eIF5A) hypusination impairs melanoma growth. Cell Biochem Funct. 2007;25:109-114
45. Xie D, Ma NF, Pan ZZ. et al. Overexpression of EIF-5A2 is associated with metastasis of human colorectal carcinoma. Hum Pathol. 2008;39:80-86
46. Ramaswamy S, Ross KN, Lander ES. et al. A molecular signature of metastasis in primary solid tumors. Nat Genet. 2003;33:49-54
47. Lee NP, Tsang FH, Shek FH. et al. Prognostic significance and therapeutic potential of eukaryotic translation initiation factor 5A (eIF5A) in hepatocellular carcinoma. Int J Cancer. 2010;127:968-976
48. Marchet A, Mocellin S, Belluco C. et al. Gene expression profile of primary gastric cancer: towards the prediction of lymph node status. Ann Surg Oncol. 2007;14:1058-1064
49. Zhu W, Cai MY, Tong ZT. et al. Overexpression of EIF5A2 promotes colorectal carcinoma cell aggressiveness by upregulating MTA1 through C-myc to induce epithelial-mesenchymaltransition. Gut. 2012;61:562-575
50. Chen M, Huang JD, Deng HK. et al. Overexpression of eIF-5A2 in mice causes accelerated organismal aging by increasing chromosome instability. BMC Cancer. 2011;11:199
51. Amati B, Alevizopoulos K, Vlach J. Myc and the cell cycle. Front Biosci. 1998;3:250-268
52. Bishop JM. Retroviruses and cancer genes. Adv Cancer Res. 1982;37:1-32
53. Bouchard C, Staller P, Eilers M. Control of cell proliferation by Myc. Trends Cell Biol. 1998;8:202-206
54. Dang CV. c-Myc target genes involved in cell growth, apoptosis, and metabolism. Mol Cell Biol. 1999;19:1-11
55. Nesbit CE, Tersak JM, Prochownik EV. MYC oncogenes and human neoplastic disease. Oncogene. 1999;18:3004-3016
56. Evan GI, Wyllie AH, Gilbert CS. et al. Induction of apoptosis in fibroblasts by c-myc protein. Cell. 1992;69:119-128
57. Shi Y, Glynn JM, Guilbert LJ. et al. Role for c-myc in activation-induced apoptotic cell death in T cell hybridomas. Science. 1992;257:212-214
58. Campisi J. Aging and cancer: the double-edged sword of replicative senescence. J Am Geriatr Soc. 1997;45:482-488
59. Campisi J, d'Adda di Fagagna F. Cellular senescence: when bad things happen to good cells. Nat Rev Mol Cell Biol. 2007;8:729-740
60. Kang HA, Schwelberger HG, Hershey JW. The two genes encoding protein synthesis initiation factor eIF-5A in Saccharomyces cerevisiae are members of a duplicated gene cluster. Mol Gen Genet. 1992;233:487-490
61. Schwelberger HG, Kang HA, Hershey JW. Translation initiation factor eIF-5A expressed from either of two yeast genes or from human cDNA. Functional identity under aerobic and anaerobic conditions. J Biol Chem. 1993;268:14018-14025
62. Vu VV, Emerson JP, Martinho M. et al. Human deoxyhypusine hydroxylase, an enzyme involved in regulating cell growth, activates O2 with a nonheme diiron center. Proc Natl Acad Sci. 2009;106:14814-14819
63. Yang GF, Xie D, Liu JH. et al. Expression and amplification of eIF-5A2 in human epithelial ovarian tumors and overexpression of EIF-5A2 is a new independent predictor of outcome in patients with ovarian carcinoma. Gynecol Oncol. 2009;112:314-318
64. Chen W, Luo JH, Hua WF. et al. Overexpression of EIF-5A2 is an independent predictor of outcome in patients of urothelial carcinoma of the bladder treated with radical cystectomy. Cancer Epidemiol Biomarkers Prev. 2009;18:400-408
65. He LR, Zhao HY, Li BK. et al. Overexpression of eIF5A-2 is an adverse prognostic marker of survival in stage I non-small cell lung cancer patients. Int J Cancer. 2011;129:143-150
66. Shek FH, Fatima S, Lee NP. Implications of the Use of Eukaryotic Translation Initiation Factor 5A (eIF5A) for Prognosis and Treatment of Hepatocellular Carcinoma. Int J Hepatol. 2012;2012:760-928
67. Lee Y, Kim HK, Park HE. et al. Effect of N1-guanyl-1,7-diaminoheptane, an inhibitor of deoxyhypusine synthase, on endothelial cell growth, differentiation and apoptosis. Mol Cell Biochem. 2002;237:69-76
68. Taylor CA, Sun Z, Cliche DO. et al. Eukaryotic translation initiation factor 5A induces apoptosis in colon cancer cells and associates with the nucleus in response to tumour necrosis factor alpha signalling. Exp Cell Res. 2007;313:437-449
69. Sun Z, Cheng Z, Taylor CA. et al. Apoptosis induction by eIF5A1 involves activation of the intrinsic mitochondrial pathway. J Cell Physiol. 2010;223:798-809
70. Moore CC, Martin EN, Lee G. et al. Eukaryotic translation initiation factor 5A small interference RNA-liposome complexes reduce inflammation and increase survival in murine models of severe sepsis and acute lung injury. J Infect Dis. 2008;198:1407-1414
71. Li AL, Li HY, Jin BF. et al. A novel eIF5A complex functions as a regulator of p53 and p53-dependent apoptosis. J Biol Chem. 2004;279:49251-49258
72. Hoque M, Hanauske-Abel HM, Palumbo P. et al. Inhibition of HIV-1 gene expression by Ciclopirox and Deferiprone, drugs that prevent hypusination of eukaryotic initiation factor 5A. Retrovirology. 2009;6:90
73. Balabanov S, Gontarewicz A, Ziegler P. et al. Hypusination of eukaryotic initiation factor 5A (eIF5A):a novel therapeutic target in BCR-ABL-positive leukemias identified by a proteomics approach. Blood. 2007;109:1701-1711
Author contact
Corresponding author: Dan Xie, M.D. Ph.D. Sun Yat-Sen University Cancer Center, State key laboratory of oncology in South China, Collaborative Innovation Center of Cancer Medicine, No. 651, Dongfeng Road East, 510060 Guangzhou, China. Tel: 86-20-87343192 Fax: 86-20-87343170 Email: xiedanorg.cn.

 Global reach, higher impact
Global reach, higher impact