10
Impact Factor
ISSN: 1449-2288
Int J Biol Sci 2014; 10(1):109-118. doi:10.7150/ijbs.8198 This issue Cite
Review
Preliminary Success in the Characterization and Management of a Sudden Breakout of a Novel H7N9 Influenza A Virus
1. Lab of Molecular Immunology, Virus Inspection Department, Zhejiang Provincial Center for Disease Control and Prevention, 630 Xincheng Road, Hangzhou, 310051, PR China.
2. Lab of Chemical Biology and Molecular Drug Design, College of Pharmaceutical Science, Zhejiang University of Technology, 18 Chaowang Road, Hangzhou, 310014, PR China.
3. Center for Innovation in Immunoregulative Technology and Therapeutics, Graduate School of Medicine, Kyoto University, Kyoto, 606-8501, Japan.
Received 2013-11-22; Accepted 2013-12-5; Published 2014-1-9
Abstract
Influenza has always been one of the major threats to human health. The Spanish influenza in 1918, the pandemic influenza A/H1N1 in 2009, and the avian influenza A/H5N1 have brought about great disasters or losses to mankind. More recently, a novel avian influenza A/H7N9 broke out in China and until December 2, 2013, it had caused 139 cases of infection, including 45 deaths. Its risk and pandemic potential attract worldwide attention. In this article, we summarize epidemiology, virology characteristics, clinical symptoms, diagnosis methods, clinical treatment and preventive measures about the avian influenza A/H7N9 virus infection to provide a reference for a possible next wave of flu outbreak.
Keywords: H7N9 virus, avian influenza A, infection, diagnosis, treatment, prevention.
Background
During February 19 through March 7, 2013, three patients with severe pneumonia and pulmonary dysfunction in Shanghai and Anhui of China were admitted to hospitals successively. All cases had typical influenza-like symptoms such as cough, fever in an early phase, and severe pneumonia and acute respiratory distress with lethal outcome several days after the onset of the initial symptoms. Before too long, many analogous cases were also found in Yangtze River Delta region of China. The following etiological studies revealed that the patients were infected with a novel reassortant avian influenza virus, A/H7N9 virus [1, 2]. On March 31, 2013, China National Health and Family Planning Commission informed that this new avian influenza A/H7N9 virus had caused human deaths in China [3]. It was the first report that the avian influenza A/H7N9 virus could infect human and that a low pathogenic avian influenza virus could cause human deaths [4, 5]. Thus, it has aroused global public health concerns about its risk and pandemic potential [5-7].
Epidemiology
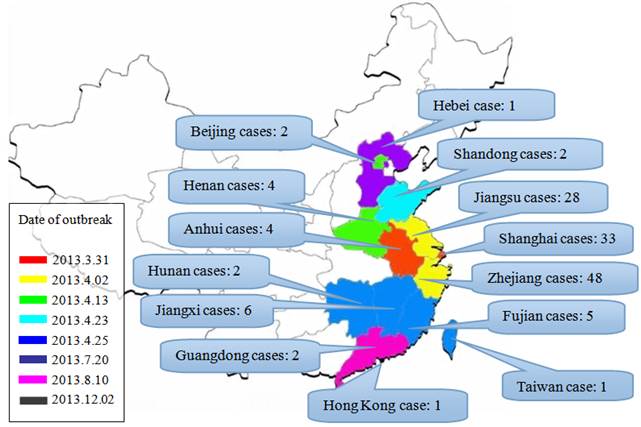
Until October 25, 2013, the World Health Organization (WHO) has reported a total of 137 cases of human infection including 45 fatal cases from ten provinces, two municipalities of China and Taiwan; Afterwards, other two cases was reported in China. Nearly 90% of the infection took place between March 19 and April 23 [8]. The infection was diffused from Yangtze River Delta region to the surrounding areas and inland (Fig. 1). So far, the case-mortality rate (33%) of the avian influenza A/H7N9 virus infection is significantly higher than that of previous influenza pandemic infection (range, 0.1%-2.5%) [9]. However, a study during March 4 to April 28, 2013 in China showed that only 6 of 20739 (0.03%) patients with influenza-like symptoms were confirmed as cases of infection with the A/H7N9 virus [10]. Accordingly, compared to previous assessment, this avian influenza A/H7N9 epidemic may cause high mortality rate, but it is unlikely to cause a large-scale outbreak [11].
Human infection with the A/H7N9 virus seems to occur through direct contact to infected poultry, most of cases have a history of exposure to live poultry [11, 12]. Zhang et al. investigated 10703 samples from different sources, and the survey report showed that all samples with the A/H7N9 virus were collected from live poultry markets except for one from a homing pigeon farm and another from a wild pigeon [13], suggesting that live poultry markets are the major origins of the A/H7N9 virus infection.
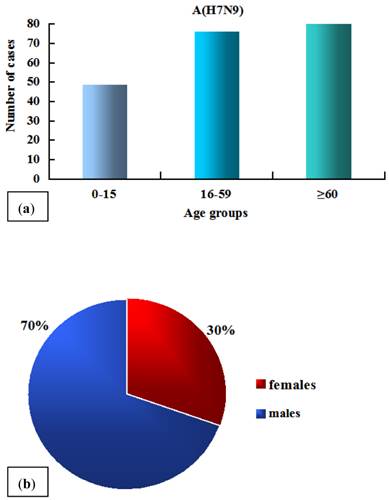
The A/H7N9 virus infection shows two typical epidemiological characteristics: Firstly, the elderly are more susceptible than the young (Fig. 2a) [11, 14, 15]. The average age of 125 infected cases was 62 [11, 16]. This may be explained by the facts that the aged spend more time in live poultry markets and therefore were more likely to be exposed to live poultry, and that they generally have weak immunity levels related to aging or comorbidities [11, 14, 17, 18]. Secondly, sex differences exist in the A/H7N9 virus infection (Fig. 2b), with the number of males being significantly greater than that of females [9, 11], whereas both of the avian influenza H5N1 and the pandemic H1N1 influenza virus infections exhibit no sex bias [15, 19]. A study conducted by Cowling group revealed that the preference for males occurs only in urban areas but not in countryside, implying that the current sex bias is attributed to different customs or habits which are associated with gender in different regions rather than some previous speculations about physiological differences between males and females [4, 15].
Although most of the patients appear to be sporadic and most of close contacts are negative for A/H7N9 virus [12], researchers found that human H7N9 virus isolates could efficiently spread among ferrets via direct contact [20] and respiratory droplets [13]. However, so far, human-to-human transmission remains very uncommon at this time. In fact, in a family cluster with two patients, the second patient developed symptoms after providing care for the index patient without protection, and genome sequences of the strains isolated from the two patients were almost identical, demonstrating that person-to-person transmission is feasible under certain conditions [21].
Geographical distribution of the A/H7N9 infection and the date when the first case of infection was reported in each region. Note: the statistical data are up until December 2, 2013.
The age (a) and sex (b) distribution of 125 cases. The data are obtained from references [11] and [16].
Virology
Phylogenetic analysis. The new A/H7N9 virus is a reassortant of earlier H7N9, H7N3, and H9N2 viruses [22-24]. Its hemagglutinin (HA), neuraminidase (NA), nonstructural protein (NS) may come from A/duck/Zhejiang/12/2011-H7N3, A/wild bird/Korea/A14/2011-H7N9 and A/chicken/Dawang/1/2011-H9N2, respectively, whereas the other five internal genes matrix (M), nucleoprotein (NP), acidic polymerase (PA), basic polymerase 1 (PB1) and basic polymerase 2 (PB2) appear to be originated from more than one source [25]. For M, PA, PB1 and PB2, each of them has a cluster together with the A/brambling/Beijing/16/2012-H9N2 and the other H9N2 clusters previously found in chickens in Yangtze River Delta, indicating that they might stem from the A/brambling/Beijing/16/2012-H9N2, and have reassorted and evolved rapidly into several clusters [26, 27]. Based on these results, it is speculated that the reassortment path of the new A/H7N9 virus is as follows (Fig. 3): the earlier A/H7N9 virus in migratory birds reassorted with H9N2 virus in northern China, so the reassortant virus retained the NA of H7N9 virus and obtained five internal gene fragments (PB2, PB1, PA , NP and M) from H9N2; when birds migrated to Jiangsu, the reassortant virus HXN9 (x = 7/9) reasserted with H9N2 (A/chicken/Dawang/1/2011-H9N2) again and gained its NS segment; meanwhile, the A/duck/Zhejiang/12/2011-H7N3 virus reassorted with some H9N2 strains in Yangtze River Delta and formed H7NX (x = 2/3), which retained H7 of the H7N3 and obtained PB2, PB1, PA, NP and M from the H9N2 viruses; at last, HXN9 reassorted with H7NX, consequently generating this new human-pathogenic A/H7N9 virus.
The reassortment path of a new A/H7N9 virus. BJ, Beijing; DW, Dawang; ZJ, Zhejiang; SH, Shanghai; JS, Jiangsu.
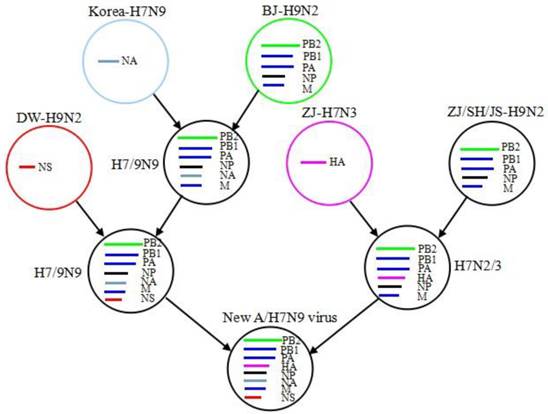
A series of reassortments enable the viruses to exchange their gene fragments very rapidly and facilitate a virus adaptation to human cells. Thus, it is necessary to care about ongoing reassortments and analyze the latest data timely, which will help us take prompt and effective measures to prevent potential infection outbreaks [28, 29].
Molecular signatures of new A/H7N9 viruses. Gene reassortments promote the formation of new virus strains while gene mutations facilitate the emergence of mammal adaptation and the increase in virulence. The new avian influenza A/H7N9 virus has possessed several signatures related to mammal adaptation and other characteristics so far (Table 1) [29-45]. The A/H7N9 virus could bind to both avian-type (α-2, 3 sialic acid) and human-type (α-2, 6 linked sialic acid) receptors. This property may attribute to several amino acid changes occurring in the receptor-binding site (RBS) of HA, such as Q226L (H3 numbering system) [23]. The single Q226L amino acid substitution could modulate the receptor-binding preference of viruses by generating a non-polar binding site for C-6 and the ring carbons of galactose-2 for the human receptor, thus providing a more hydrophobic binding site which favors combination with human receptor [46-50]. Watanabe et al. [51] reported that the A/H7N9 virus bearing L226 appears to be more efficient in infecting mammalian cells and releasing from infected cells. Further analysis revealed that the A/H7N9 virus bearing Q226 preferentially bound to the avian-type receptor; while the A/H7N9 virus bearing L226, exhibited an increased binding to both avian-type and human-type receptors [47, 52, 53]. The findings demonstrated that the viruses retained the preference for the avian receptor, which is not conducive to human-to-human transmission, at the same time, acquired other receptor-binding specificities characteristic of typical pandemic influenza viruses.
PB2 is another important determinant of the host range and virulence of influenza viruses. A critical mutation in PB2 of A/H7N9 virus, E627K, which was previously found in A/H5N1 [54] and A/H7N7 viruses [55], is considered contributing to the improvement of viral polymerase activity at human upper respiratory tract temperature (32-33oC) and thus facilitate the adaptation of the viruses to human. Notably, the E627K mutation can be detected in almost all the A/H7N9 isolates from human cases, but was not detected in any poultry or environmental isolates, suggesting that the E627K mutation took place after virus transmission from poultry to human [13, 29, 30, 51].
In addition, another interesting molecular feature of the A/H7N9 virus is the absence of multibasic amino acid motif in the HA cleavage site. The multibasic cleavage site is a sign of high pathogenicity for birds, but not for human, which may be the reason why the A/H7N9 virus infection causes human deaths while only mild symptoms in poultry [23, 50]. These important molecular signatures indicate that nascent A/H7N9 viruses may be readily transmitted to human, and they are, therefore, more difficult to deal with.
Clinical characteristics. The most common clinical symptom of patients infected with A/H7N9 viruses is severe pneumonia [56]. Most of cases manifested fever, cough with sputum production, wheeze, headache, myalgia and general malaise during early stage and later severe illnesses characterised by bilateral pneumonia, rapid progression to acute respiratory distress syndrome (ARDS), multiorgan dysfunction (impaired liver or renal function), septic shock, rhabdomyolysis and encephalopathy [29, 30, 57-61]. Bilateral ground-glass opacities, consolidation and pleural effusion are the typical radiologic findings [30, 56, 62]. Critically ill patients also generally exhibit lymphopenia and thrombocytopenia [49, 56], along with the increase of some serum cytokines or chemokines, including lactate dehydrogenase, creatinine kinase, aspartate aminotransferase, C-reactive protein, IP-10, MIG, MIP-1β, MCP-1, IL-6, IL-8 and IFN-α [46, 50, 62].
Molecular signatures of the A/H7N9 virus.
| Gene | Molecular signatures | Effect |
|---|---|---|
| HA | Q226L/I, G228S, T160A, A138S, G186V, G195V, D285N | Increased affinity to human-type receptors |
| PEIPKGR*G in HA cleavage site | Low pathogenic to birds | |
| NA | R292K, R152K | Resistance to neuraminidase inhibitors |
| 69-73 deletion | Adapted to terrestrial birds and increased virulence | |
| PB2 | L89V, E627K, N701D, A44S | Improved replication and virulence in mammals |
| PB1 | H99Y, I368V | Related to H5 virus transmission among ferrets |
| M1 | N30D, T215A | Increased virulence in mice |
| M2 | S31N | Amantadine resistance |
| NS1 | P42S | Increased virulence in mice |
| PA | V100A, K356R, S409N | Increased virulence to mammals |
Diagnosis
The diagnosis of patients infected with the A/H7N9 virus should follow the diagnosis and treatment scheme of human infection with H7N9 avian influenza (Version 2, 2013) [63]. It is noteworthy that a sputum specimen is more likely to show A/H7N9 virus positive than that of throat swab [50, 64-66]. Infected cases can be confirmed by means of virus isolation, serologic testing, and real time reverse transcriptase polymerase chain reaction (RT-PCR) assays [67-69]. Among these methods, the RT-PCR assay is the most common method and the gold standard of influenza virus detection [12, 70].
Baas et al. examined the ability of six antigen detection-based rapid influenza point-of-care tests (POCTs) in detecting the A/H7N9 virus. The low Ct values suggested that these POCTs could not be employed in the detection for the A/H7N9 infection [71]. In resource-limited regions, a subtype-specific reverse transcription loop-mediated isothermal amplification assay (LAMP) could be used as a reliable method for the early diagnosis. The procedure of this reaction only contains one step, 63°C for 60 min in a single tube containing hydroxynaphthol blue dye, using a strand displacement DNA polymerase and 4 primers designed for six regions of target genes. Its detection limits are comparable to corresponding H7 and N9 RT-PCR assays [72, 73].
Treatment and prevention
Treatment recommendations. Due to the acute progresses and severe outcomes of the A/H7N9 virus infection, it is suggested that all the confirmed cases and suspected cases received antiviral treatment as soon as possible [74-75]. Mounting evidence indicated that the early treatment is probably the major decisive factor for survival in the A/H1N1 and A/H5N1 virus infection [76, 77]. Therefore, poor outcomes in some A/H7N9 virus-infected patients may be attributed to the delayed antiviral treatment [78]. As for severe cases with respiratory dysfunction, mechanical ventilation should be supported without delay.
Therapeutic use. Preliminary data suggest that the A/H7N9 virus is resistant to adamantane because of the S31N mutation in the virus matrix2 (M2) protein which has been found in all isolates [79, 80], however, it remains sensitive to neuraminidase inhibitors, such as oseltamivir and zanamivir [50, 78, 81].
It is worth noting that R292K in NA, a mutation in a seasonal H3N2 influenza virus, which is considered to confer resistance to oseltamivir [82], has emerged in avian influenza A/H7N9 viruses. The R292K mutation causes a significant expansion in the active site cavity and increases in its solvent accessible surface, thereby leading to a remarkable loss of total binding free energy for NA-oseltamivir and an unfavorable shift in van der Waals interactions which may contribute to drug resistance [83]. Even though patients, who are infected with the A/H7N9 virus carrying R292K mutation, are treated with neuraminidase inhibitors timely, some of them fail to respond to the treatment, exhibiting persistent or rebound virus titer and poor outcomes [78, 84]. Nevertheless, neuraminidase inhibitors are still the front-line therapeutic choices against the A/H7N9 virus infection.
In addition, a study on glucocorticoids used against the A/H7N9 virus infection was reported [30]. Results showed that although glucocorticoids can generally be used as anti-inflammatory agents and immunomodulators [85, 86], they have been proved no benefit to the A/H7N9 virus infection [30]. It is noteworthy that two isolates from patients gained R292K mutation after administration of corticosteroids [78]. It remains to be seen whether corticosteroids could accelerate the emergence of R292K mutation [87]. At least, corticosteroids should be avoided for the treatment of A/H7N9 virus infection at present [64, 88].
Advances in the development of anti-flu drugs. Gene mutations of the A/H7N9 virus associated with drug resistance emphasize the importance of developing new drugs with different action mechanisms [88]. Currently novel anti-flu agents that target viral key proteins, host factors have achieved varying degrees of progress [89-96]. Several novel anti-influenza virus drugs are currently in clinical trials, including favipiravir, laninamivir, DAS181, Vertex VX-787, AVI-7100, apomivir, ergoferon and nitazoxanide [97]. Some of the screening drugs studied will probably be effective to A/H7N9 influenza even though none of them have been specifically designed for the A/H7N9 influenza.
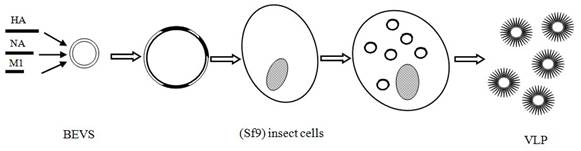
Vaccines for avian influenza A (H7N9). Vaccine is considered as the most important potential means for controlling the A/H7N9 influenza transmission [98]. Scientists around the world have been working on the development of vaccines specific to A/H7N9 virus in order to prevent a potential pandemic bird flu. Previous reports have demonstrated that H7 protein is poorly immunogenic in humans [5, 99-101] because of a very low T cell epitope content in the HA sequences and poor potential for cross-reactivity with the T cell epitope of currently circulating strains of influenza [101]. It, therefore, seems difficult to develop an effective vaccine for the A/H7N9 virus. Even so, at least four companies have succeeded in developing A/H7N9 candidate vaccines, including Greffex, Protein sciences, Medicago, and Novavax [102, 103]. All four companies adopted modern genetic engineering techniques to prepare key protein-containing vaccine for the A/H7N9 virus infection. Two virus-like particles (VLP) vaccines developed by Medicago and Novavax yielded promising results (pre-) clinical trials (Fig. 4) [103, 104].
The A/H7N9 VLP vaccine developed by Novavax consists of HA (A/Anhui/1/2013-H7N9), NA (A/Anhui/1/2013-H7N9) and M1 (A/Indonesia/05/2005-H5N1). This type of VLP not only elicits hemagglutination-inhibition (HAI) antibody titers of ≥1:64 and anti-neuraminidase (NA) antibody against the A/H7N9 virus, but also provides cross-protection for other H7 subtype virus, such as the H7N3 virus. In experimental mouse models, all animals receiving the A/H7N9 vaccine survived, while no survival was observed in control groups receiving an A/H5N1 vaccine or placebo [103]. These promising results may be ascribable to a fact that VLP can be presented not only by the MHC-II molecules for stimulation of CD4 helper T cells, but also by the MHC-I molecules for activation of cytotoxic T-lymphocytes (CTLs) [105]. Moreover, the path of VLP production is simple and time-saving. In fact, it needs only 26 days for the genetic construction of the vaccine for the production of an A/H7N9 VLP. The resulting VLP preparation does not contain genetic materials, which are essential for viral replication and infection. Therefore, compared with traditional inactivated or attenuated vaccines, the preparation of VLP is more convenient and safer when pandemic bird flu takes place suddenly. Other studies including phase I clinical trials in human are ongoing and anticipated be completed in August 2014 [97].
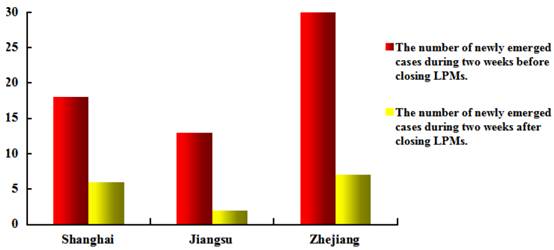
Prevention and control. Owing to profound lessons from SARS and experience from the A/H5N1 and A/H1N1 epidemics, responses of the Chinese Government to the A/H7N9 epidemic are swift and transparent [106-108]. Relevant departments quickly identified the cause of the disease and the source of the A/H7N9 virus, followed by a series of control measures, including: (1) closed live poultry markets, culled live poultry and prohibited the entry of adventive live poultry; (2) publicized preventive knowledge through various kinds of media; (3) strengthened surveillance on epidemic dynamics. These measures were proved effective as a prominent reduction occurs in the number of newly emerged cases since the closure of live poultry markets (Fig. 5) [111-113]. However, it remains a great challenge for the Chinese Government to control the epidemic. Because of the low pathogenicity of A/H7N9 virus to poultry and wild birds [4, 5], the virus may spread silently among poultry without being noticed and can infect human irregularly. Although measures currently taken by the government such as closing live poultry markets and culling live poultry have made great progress merely for disease control rather than prevention, these strategies can only control the transmission of the A/H7N9 epidemic but not block the formation of the epidemic. Therefore, searching for specific preventive measures to the A/H7N9 influenza is imperative.
The method of virus-like particles (VLP) vaccine production.
Impacts of closing live poultry markets (LPMs) on A/H7N9 epidemic.
Conclusion
The new reassortant A/H7N9 virus primarily infected human and has caused several deaths in China. Although there is no sufficient evidence to indicate sustained human-to-human transmission yet, this risk will increase as a result of frequent gene reassortants and mutations. Typical symptoms of the avian influenza A/H7N9 virus infection include fever, cough, even severe pneumonia, and multi-organ dysfunction syndrome. So far, real-time PCR is still the main diagnostic method, and neuraminidase inhibitors (oseltamivir, zanamivir, and peramivir) are used for clinical treatment but the M2 ion channel blockers (amantadine and rimantadine) are invalid; at the same time, the continuing resistance mutations have been hindering the treatment. It is thus urgent to explore new drugs and therapy strategies. Whereas closing live poultry markets, culling live poultry, and other control measures have prevented the transmission of the epidemic successfully, the pathologic characteristics of the A/H7N9 virus (low pathogenic to avian but high to human) may lead to insidious outbreak of deadly avian flu, and the molecular characteristics of the A/H7N9 virus make it difficult to develop effective vaccines. Along with a decrease in absolute humidity in autumn and winter, the survival and transmission abilities of influenza viruses will increase apparently [114]. If the A/H7N9 virus infection follows a similar epidemic pattern to that of the A/H5N1 or other pandemics, it will recur in autumn and winter. Thus, continually intensive surveillance and prepared measures for the prevention of major outbreaks remain quite important.
Acknowledgements
We gratefully acknowledge the support by the Program for Zhejiang Leading Team of Science and Technology Innovation (2011R50021), Zhejiang Provincial Natural Science Foundation of China (LY12B02019) and Social Development project of Zhejiang Province (2011C23004).
Competing Interests
The authors have declared that no competing interest exists.
References
1. Wen Y, Klenk HD. H7N9 avian influenza virus-search and re-search. Emerg Microbes Infect. 2013;2:e18. doi:10.1038/emi.2013.18
2. Number of confirmed human cases for influenza A(H7N9) reported to WHO. World Health Organization (WHO). http://www.who.int/influenza/human_animal_interface/influenza_h7n9/01_ReportWebH7N9Number.pdf
3. National Health and Family Planning Commission of China (NHFPC): Shanghai, Anhui occurred three cases of human infection with H7N9 avian influenza confirmed cases. http://www.moh.gov.cn/mohwsyjbgs/s3578/201303/44f25bd6bed14cf082512d8b6258fb3d.shtml
4. Arima Y, Zu R, Murhekar M. et al. Human infections with avian influenza A(H7N9) virus in China: preliminary assessments of the age and sex distribution. WPSAR. 2013 doi:10.5365/wpsar.2013.4.2.005
5. Uyeki TM, Cox NJ. Global concerns regarding novel influenza A(H7N9) virus infections. N Engl J Med. 2013;368(20):1862-4
6. Jonges M, Meijer A, Fouchier RA. et al. Guiding outbreak management by the use of influenza A(H7Nx) virus sequence analysis. Euro Surveill. 2013;18(16):20460
7. Fouchier RA, Kawaoka Y, Cardona C. et al. Gain-of-Function experiments on H7N9. Science. 2013;341:612-3
8. World Health Organization (WHO): Number of confirmed human cases for influenza A(H7N9) reported to WHO. http://www.who.int/influenza/human_animal_interface/influenza_h7n9/09_ReportWebH7N9Number.pdf
9. Ke Y, Wang Y, Liu S. et al. High severity and fatality of human infections with avian influenza A(H7N9) infection in China. Clin Infect Dis. 2013 doi:10.1093/cid/cit371
10. Xu C, Havers F, Wang L. et al. Monitoring avian influenza A(H7N9) virus through National Influenza-like illness Surveillance, China. Emerg Infect Dis. 2013;19(8):1289-92
11. Yu H, Cowling BJ, Feng L. et al. Human infection with avian influenza A H7N9 virus: an assessment of clinical severity. Lancet. 2013;382(9887):138-45
12. Li Q, Zhou L, Zhou M. et al. Preliminary report: epidemiology of the avian influenza A(H7N9) outbreak in China. N Engl J Med. 2013 doi:10.1056/NEJMoa1304617
13. Zhang Q, Shi J, Deng G. et al. H7N9 influenza viruses are transmissible in ferrets by respiratory droplet. Science. 2013;341(6144):410-4
14. Cowling BJ, Freeman G, Wong JY. et al. Preliminary inferences on the age-specific seriousness of human disease caused by avian influenza A(H7N9) infections in China, March to April 2013. Euro Surveill. 2013;18(19):20475
15. Cowling BJ, Jin L, Lau EHY. et al. Comparative epidemiology of human infections with avian influenza AH7N9 and H5N1 viruses in China: a population-based study of laboratory-confirmed cases. Lancet. 2013;382(9887):129-37
16. Latest news about human infection with influenza A(H7N9) Virus. http://roll.news.sina.com.cn/s_qlg2013_all/1/index.shtml
17. Guan Y, Farooqui A, Zhu H. et al. H7N9 incident, immune status, the elderly and a warning of an influenza pandemic. J Infect Dev Ctries. 2013;7(4):302-7
18. Bermejo-Martin JF, Almansa R, Lejarazu RO. et al. Weakened immunity in aged hosts with comorbidities as a risk factor for the emergence of influenza A H7N9 mutants. J Infect Dev Ctries. 2013;7(6):497-8
19. Zhang Q, Wang S, Wu M. et al. Epidemiological and risk analysis of the H7N9 subtype influenza outbreak in China at its early stage. Chin Sci Bull. 2013 doi:10.1007/s11434-013-5880-5
20. Zhu H, Wang D, Kelvin DJ. et al. Infectivity, transmission, and pathology of human-isolated H7N9 influenza virus in ferrets and pigs. Science. 2013;341(6142):183-6
21. Qi X, Qian Y, Bao C. et al. Probable person to person transmission of novel avian influenza A(H7N9) virus in eastern China. BMJ. 2013;347:f4752
22. Liu D, Shi W, Shi Y. et al. Origin and diversity of novel avian influenza A H7N9 viruses causing human infection: phylogenetic, structural, and coalescent analysis. Lancet. 2013;381(9881):1926-32
23. Gao R, Cao B, Hu Y. et al. Human infection with a novel avian-origin influenza A(H7N9) virus. N Engl J Med. 2013;368(20):1888-97
24. Ranst M, Lemey P. Genesis of avian-origin H7N9 influenza A viruses. Lancet. 2013;381(9881):1883-5
25. Wu A, Su C, Wang D. et al. Sequential reassortments underlie diverse influenza H7N9 genotypes in China. Cell Host Microbe. 2013; dx.doi.org/10.1016/j.chom. 2013 09.001
26. Zhang L, Zhang Z, Weng Z. Rapid reassortment of internal genes in avian influenza A H7N9 virus. Clin Infect Dis. 2013 doi:10.1093/cid/cit414
27. Bao C, Cui L, Zhou M. et al. Live-animal markets and influenza A(H7N9) virus infection. N Engl J Med. 2013;368(24):2337-9
28. Zhang B, Huang K, Guo J. Origin and the recombinant model of H7N9 virus prevailing in China. Nan Fang Yi Ke Da Xue Xue Bao. 2013;33(7):1017-21
29. Kageyama T, Fujisaki S, Takashita E. et al. Genetic analysis of novel avian A(H7N9) influenza viruses isolated from patients in China, February to April 2013. Euro Surveill. 2013;18(15):pii =20453
30. Li J, Yu X, Pu X. et al. Environmental connections of novel avian-origin H7N9 influenza virus infection and virus adaptation to the human. Sci China Life Sci. 2013;56(6):485-92
31. Wang W, Lu B, Zhou H. et al. Glycosylation at 158N of the hemagglutinin protein and receptor binding specificity synergistically affect the antigenicity and immunogenicity of a live attenuated H5N1 A/Vietnam/1203/2004 vaccine virus in ferrets. J Virol. 2010;84(13):6570-7
32. Zhang Y, Zhang Q, Gao Y. et al. Key molecular factors in hemagglutinin and PB2 contribute to efficient transmission of the 2009 H1N1 pandemic influenza virus. J Virol. 2012;86(18):9666-74
33. Chen Z, Zhou H, Kim L. et al. The receptor binding specificity of the live attenuated influenza H2 and H6 vaccine viruses contributes to vaccine immunogenicity and protection in ferrets. J Virol. 2012;86(5):2780-6
34. Naeve C, Hinshaw V, Webster R. Mutations in the hemagglutinin receptor-binding site can change the biological properties of an influenza virus. J Virol. 1984;51(2):567-9
35. Herfst S, Schrauwen E, Linster M. et al. Airborne transmission of influenza A/H5N1 virus between ferrets. Science. 2012;336(6088):1534-41
36. Imai M, Watanabe T, Hatta M. et al. Experimental adaptation of an influenza H5 HA confers respiratory droplet transmission to a reassortant H5 HA/H1N1 virus in ferrets. Nature. 2012;486(7403):420-8
37. Tharakaraman K, Jayaraman A, Raman R. et al. Glycan receptor binding of the influenza A virus H7N9 hemagglutinin. Cell. 2013;153(7):1486-93
38. Han J, Niu F, Jin M. et al. Clinical presentation and sequence analyses of HA and NA antigens of the novel H7N9 viruses. Emerg Micro Infect. 2013;2:e23 doi:10.1038/emi.2013.28
39. Xiong C, Zhang Z, Jiang Q. et al. Evolutionary characteristics of A/Hangzhou/1/2013 and source of avian influenza virus H7N9 subtype in China. Clin Infect Dis. 2013 doi:10.1093/cid/cit294
40. Matrosovich MN, Matrosovich TY, Gray T. et al. Human and avian influenza viruses target different cell types in cultures of human airway epithelium. Proc Natl Acad Sci USA. 2004;101(13):4620-4
41. Srinivasan K, Raman R, Jayaraman A. et al. Quantitative description of glycan-receptor binding of influenza A virus H7 hemagglutinin. PLoS One. 2013;8(2):49597
42. Liu Q, Lu L, Sun Z. et al. Genomic signature and protein sequence analysis of a novel influenza A(H7N9) virus that causes an outbreak in humans in China. Microbes Infect. 2013;15:432-9
43. Chen G, Lai MMC, Wu S. et al. Is avian influenza A(H7N9) virus staggering its way to humans? J Formos Med Assoc. 2013;112(6):312-8
44. Xu W, Sun Z, Liu Q. et al. PA-356R is a unique signature of the avian influenza A(H7N9) viruses with bird-to-human transmissibility: potential implication for animal surveillances. J Infect. 2013 doi:10.1016/j.jinf.2013.08.001
45. Yen H, McKimm-Breschkin JL, Choy KT. et al. Resistance to neuraminidase inhibitors conferred by an R292K mutation in a human influenza virus H7N9 isolate can be masked by a mixed R/K viral population. MBio. 2013;4(4):e00396-13 doi:10.1128/mBio.00396-13
46. Zhou J, Wang D, Gao R. et al. Biological features of novel avian influenza A(H7N9) virus. Nature. 2013;499(7459):500-3
47. Xiong X, Martin SR, Haire LF. et al. Receptor binding by an H7N9 influenza virus from humans. Nature. 2013;499(7459):496-499
48. Belser JA, Gustin KM, Pearce MB. et al. Pathogenesis and transmission of avian influenza A(H7N9) virus in ferrets and mice. Nature. 2013 doi:10.1038/nature12391
49. Rogers GN, Paulson JC, Daniels RS. et al. Single amino acid substitutions in influenza haemagglutin in change receptor binding specificity. Nature. 1983;304(5921):76-78
50. Chen Y, Liang W, Yang S. et al. Human infections with the emerging avian influenza A H7N9 virus from wet market poultry: clinical analysis and characterization of viral genome. Lancet. 2013;381(9881):1916-25
51. Watanabe T, Kiso M, Fukuyama S. et al. Characterization of H7N9 influenza A viruses isolated from humans. Nature. 2013 doi:0.1038/nature12392
52. Wei Hu. Receptor binding specificity and sequence comparison of a novel avian-origin H7N9 virus in China. J Biomed Sci Eng. 2013;6:533-42
53. Shi Y, Zhang W, Wang F. et al. Structures and Receptor Binding of Hemagglutinins from Human-Infecting H7N9 Influenza Viruses. Science. 2013 doi:10.1126/science.1242917
54. Hatta M, Gao P, Halfmann P. et al. Y. Molecular basis for high virulence of Hong Kong H5N1 influenza A viruses. Science. 2001;293(5536):1840-2
55. Munster VJ, de Wit E, van Riel D. et al. The molecular basis of the pathogenicity of the dutch highly pathogenic human influenza A H7N7 viruses. J Infect Dis. 2007;196(2):258-65
56. Gao H, Lu H, Cao B. et al. Clinical findings in 111 cases of influenza A(H7N9) virus infection. N Engl J Med. 2013;368(24):2277-85
57. Wu S, Wu F, He J. Emerging risk of H7N9 influenza in China. Lancet. 2013;381(9877):1539-40
58. Arden KE, Mackay IM. Avian influenza A(H7N9) virus: can it help us more objectively judge all respiratory viruses? J Clin Virol. 2013;58(1):388-9
59. Tang R, Chen H. An overview of the recent outbreaks of the avian-origin influenza A (H7N9) virus in the human. J Chin Med Assoc. 2013;76(5):245-8
60. Shen P, Zong Y, Shu J. et al. Dynamic assessment of lung injury by ultrasound in a case with H7N9 inluenza. Crit Care. 2013;17(3):438
61. Liu X, Li T, Zheng Y. et al. Poor responses to oseltamivir treatment in a patient with influenza A(H7N9) virus infection. Emerg Microbes Infect. 2013;2:e27 doi:10.1038/emi2013.30
62. Cao B. What clinicians should know to fight against the novel avian-origin influenza A (H7N9) virus? Chin Med J. 2013;126(12):2205-6
63. The diagnosis and treatment scheme of human infection with H7N9 avian influenza (Version 2, 2013). National Health and Family Planning Commission of China (NHFPC). http://www.moh.gov.cn/ewebeditor/uploadfile/2013/04/20130410212136993.doc
64. Wang Q, Yao K. A (H5N1) and A(H7N9) avian influenza: the H7N9 avian influenza outbreak in 2013. Chin J Contemp Pediatr. 2013;15(6):401-4
65. Chang S, Lin P, Tsai JC. et al. The first case of H7N9 influenza in Taiwan. Lancet. 2013;381(9878):1621
66. Lo Y, Chen W, Huang W. et al. Surveillance of avian influenza A(H7N9) virus infection in humans and detection of the first imported human case in Taiwan, 3 April to 10 May 2013. Euro Surveill. 2013;18(20):pii =20479
67. Real-time RT-PCR protocol for the detection of A-vian influenza A (H7N9) virus. World Health Organization (WHO). http://www.who.int/entity/influenza/gisrs_laboratory/cnic_realtime_rt_pcr_protocol_a_h7n9.pdf
68. Serological detection of avian influenza A (H7N9) virus infections by turkey heamagglutination-inhibition assay. World Health Organization (WHO). http://www.who.int/influenza/gisrs_laboratory/cnic_serological_diagnosis_hai_a_h7n9.pdf
69. Serological detection of avian influenza A(H7N9) infections by microne utralization assay. World Health Organization (WHO). http://www.who.int/influenza/gisrs_laboratory/cnic_serological_diagnosis_microneutralization_a_h7n9.pdf
70. Corman VM, Eickmann M, Landt O. et al. Specific detection by real-time reverse-transcription reaction assays of a novel avian influenza A(H7N9) strain associated with human spillover infections in China. Euro Surveill. 2013;18(16):20461
71. Baas C, Barr IG, Fouchier RA. et al. A comparison of rapid point-of-care tests for the detection of avian influenza A(H7N9) virus, 2013. Euro Surveill. 2013;18(21):20487
72. Nie K, Zhao X, Ding X. et al. Visual detection of human infection with influenza A(H7N9) virus by subtype-specific reverse transcription loop-mediated isothermal amplification with hydroxynaphthol blue dye. Clin Microbiol Infect. 2013;19(8):E372-5
73. Ge Y, Wu B, Qi X. et al. Rapid and sensitive detection of novel avian-origin influenza A(H7N9) virus by reverse transcription Loop-Mediated Isothermal Amplification combined with a Lateral-Flow device. PLoS one. 2013;8(8):e69941
74. Rapid risk assessment: severe respiratory disease associated with a novel influenza A virus, A (H7N9) China. European Centre for Disease Prevention and Control (ECDC). http://www.ecdc.europa.eu/en/publications/publications/ah7n9-china-rapid-risk-assessment.pdf
75. World health organization (WHO). Frequently asked questions on human infection with influenza A (H7N9) virus. http://www.who.int/influenza/human_animal_interface/faq_H7N9/en/.
76. Chan PK, Lee N, Zaman M. et al. Determinants of antiviral effectiveness in influenza virus A subtype H5N1. J Infect Dis. 2012;206(9):1359-66
77. Yang S, Cao B, Liang L. et al. Antiviral therapy and outcomes of patients with pneumonia caused by influenza A pandemic (H1N1) virus. PLoS One. 2013;7(1):e29652. doi:10.1371/journal.pone.0029652
78. Hu Y, Lu S, Song Z. et al. Association between adverse clinical outcome in human disease caused by novel influenza A H7N9 virus and sustained viral shedding and emergence of antiviral resistance. Lancet. 2013;381(9885):2273-9
79. Noah DL, Noah JW. Adapting global influenza management strategies to address emerging viruses. Am J Physiol Lung Cell Mol Physiol. 2013;305(2):108-17
80. Qin G, Yu K, Shi T. et al. How does influenza virus a escape from amantadine? J Phys Chem B. 2010;114(25):8487-93
81. Bernstein HH. As seasonal influenza wanes, be aware of H7N9. AAP News. 2013 doi:10.1542/aapnew.20130426-1
82. Whitley RJ, Boucher CA, Lina B. et al. Global assessment of resistance to neuraminidase inhibitors, 2008-2011: the influenza resistance information study (IRIS). Clin Infect Dis. 2013;56(9):1197-205
83. Kar P, Knecht V. Mutation-induced loop opening and energetics for binding of tamiflu to influenza N8 neuraminidase. J Phys Chem B. 2012;116(21):6137-49
84. Hay AJ, Hayden FJ. Oseltamivir resistance during treatment of H7N9 infection. Lancet. 2013;381(9885):2230-2
85. Barnes PJ. Anti-inflammatory actions of glucocorticoids: molecular mechanisms. Clin Sci Lond. 1998;94(6):557-72
86. Confalonieri M, Kodric M, Santagiuliana M. et al. To use or not to use corticosteroids for pneumonia? A clinician's perspective. Monaldi Arch Chest Dis. 2012;77(2):94-101
87. Bermejo-Martin JF, Almansa R, Lejarazu RO. et al. Weakened immunity in aged hosts with comorbidities as a risk factor for the emergence of influenza A H7N9 mutants. J Infect Dev Ctries. 2013;7(6):497-8
88. Lu S, Xi X, Zheng Y. et al. Analysis of the clinical characteristics and treatment of two patients with avian influenza virus (H7N9). BioSci Trends. 2013;7(2):109-12
89. Fiore AE, Fry A, Shay D. et al. Antiviral agents for the treatment and chemoprophylaxis of influenza recommendations of the Advisory Committee on Immunization Practices (ACIP). MMWR Recomm Rep. 2011;60(1):1-24
90. Lee SM, Yen HL. Targeting the host or the virus: current and novel concepts for antiviral approaches against influenza virus infection. Antiviral Res. 2012;96(3):391-404
91. Sailen B. New treatments for influenza. BMC Medicine. 2012;10:104
92. Liu Q, Liu DY, Yang ZQ. Characteristics of human infection with avian influenza viruses and development of new antiviral agents. Acta Pharmacol Sin. 2013;34(10):1257-69
93. Hayden FG. Newer influenza antivirals, biotherapeutics and combinations. Influenza Other Respir Viruses. 2013;7(Suppl. 1):63-75
94. Fedson DS. Treating influenza with statins and other immunomodulatory agents. Antiviral Res. 2013;99(3):417-35
95. Eyer L, Hruska K. Antiviral agents targeting the influenza virus: a review and publication analysis. Veterinarni Medicina. 2013(3):113-185
96. Edinger TO, Pohl MT, Stertz S. Entry of influenza A virus: host factors and antiviral targets. J Gen Virol. 2013 doi:10.1099/vir.0.059477-0
97. http://www.clinicaltrials.gov
98. Vaccine response to the avian influenza A (H7N9) outbreak. World Health Organization (WHO). http://www.who.int/influenza/vaccines/virus/CandidateVaccineVirusesH7N9_02May13.pdf
99. Couch RB, Patel SM, Wade-Bowers CL. et al. A randomized clinical trial of an inactivated avian influenza A (H7N7) vaccine. PLoS One. 2012;7(12):e49704. doi:10.1371/journal.pone.0049704119
100. Cox RJ, Madhun AS, Hauge S. et al. A phase I clinical trial of a PER. C6 cell grown influenza H7 virus vaccine. Vaccine. 2009;27(13):1889-97
101. De Groot AS, Ardito M, Terry F. et al. Low immunogenicity predicted for emerging avian-origin H7N9. Hum Vaccin Immunother. 2013;9(5):1-7
102. Riedmann EM. Human vaccines & immunotherapeutics: news. Hum Vaccin Immunother. 2013;9(6):1187
103. Smith GE, Flyer DC, Raghunandan R. et al. Development of influenza H7N9 virus like particle (VLP) vaccine: Homologous A/Anhui/1/2013(H7N9) protection and heterologous A/chicken/Jalisco/CPA1/2012(H7N3) cross-protection in vaccinated mice challenged with H7N9 virus. Vaccine. 2013
104. Fries LF, Smith GE, Glenn GM. A Recombinant viruslike particle influenza A (H7N9) vaccine. N Engl J Med. 2013 13
105. Song H, Wittman V, Byers A. et al. In vitro stimulation of human influenza specific CD8+ Tcells by dendritic cells pulsed with an influenza virus-like particle (VLP) vaccine. Vaccine. 2010;28(34):5524-32
106. Ahmad A, Krumkamp R, Reintjes R. Controlling SARS: a review on China's response compared with other SARS-affected countries. Trop Med Int Health. 2009;14:36-45
107. From SARS to H7N9. will history repeat itself? Lancet. 2013;381(9875):1333
108. Mei L, Song P, Tang Q. et al. Changes in and shortcomings of control strategies, drug stockpiles, and vaccine development during outbreaks of avian influenza A H5N1, H1N1, and H7N9 among humans. BioSci Trends. 2013;7(2):64-76
109. National Health and Family Planning Commission deploy prevention and control work about human infection with the A/H7N9 avian influenza. National Health and Family Planning Commission of China (NHFPC). http://www.moh.gov.cn/wsb/pxwfb/201304/99a9ebd290dd4e86a388fff6679c45df.shtml
110. Notice of strengthen epidemic prevention and control for human infection with H7N9 avian influenza. National Health and Family Planning Commission of China (NHFPC). http://www.moh.gov.cn/zhuzhan/wsbmgz/201307/56b09d2fa50e4a518b5a83d8defac4f9.shtml
111. Han J, Jin M, Zhan P. et al. Epidemiological link between exposure to poultry and all influenza A (H7N9) confirmed cases in Huzhou city, China, March to May 2013. Euro Surveill. 2013;18(20):20481
112. Xu J, Lu S, Wang H. et al. Reducing exposure to avian influenza H7N9. Lancet. 2013;381:1815-6
113. Murhekar M, Arima Y, Horby P. et al. Avian influenza A (H7N9) and the closure of live bird markets. WPSP. 2013 doi:10.5365/wpsar.2013.4.2.008
114. Shaman J, Kohn M. Absolute humidity modulates influenza survival, transmission, and seasonality. PNAS. 2009;106(10):3645-6
Author biography
 Dr. Yan-Ling Wu is a professor in Molecular Immunology and now heads the Cellular and Molecular Immunology Research Group. She received Master and Doctoral degrees in Applied Life Science in 2003 and in Medicine Science in 2006, respectively, from Tohoku University, Japan. After that, she entered to Professor Minato's group of School of Medicine, Kyoto University, Japan, as a senior researcher working in the field of molecular immunology. Her current research mainly focuses on understanding the molecular mechanisms of gene regulation related to diseases by immune inhibitory receptors. Dr. Wu have given oral presentations in international conferences and published related papers.
Dr. Yan-Ling Wu is a professor in Molecular Immunology and now heads the Cellular and Molecular Immunology Research Group. She received Master and Doctoral degrees in Applied Life Science in 2003 and in Medicine Science in 2006, respectively, from Tohoku University, Japan. After that, she entered to Professor Minato's group of School of Medicine, Kyoto University, Japan, as a senior researcher working in the field of molecular immunology. Her current research mainly focuses on understanding the molecular mechanisms of gene regulation related to diseases by immune inhibitory receptors. Dr. Wu have given oral presentations in international conferences and published related papers.
 Li-Wen Shen is a postgraduate majoring in pharmacy. He obtained the Bachelor's degree in Biological Engineering in 2012 from Shanxi Datong University, China. After that, he entered Prof. Zhang's group of College of Pharmaceutical Science, Zhejiang University of Technology, China, working with small molecules to regulate disease-related gene to explore gene-targeted drugs under the direction of Profs W. Zhang and Y.-L. Wu.
Li-Wen Shen is a postgraduate majoring in pharmacy. He obtained the Bachelor's degree in Biological Engineering in 2012 from Shanxi Datong University, China. After that, he entered Prof. Zhang's group of College of Pharmaceutical Science, Zhejiang University of Technology, China, working with small molecules to regulate disease-related gene to explore gene-targeted drugs under the direction of Profs W. Zhang and Y.-L. Wu.
 Yan-Ping Ding received her bachelor's degree in biological sciences from Zhejiang University of Chinese Traditional Medicine, China. Since graduation, she has been working in Zhejiang Center of Disease Control and Prevention, China. Her research mainly focuses on the field of Molecular Immunology under the guidance of Prof. Yanling Wu.
Yan-Ping Ding received her bachelor's degree in biological sciences from Zhejiang University of Chinese Traditional Medicine, China. Since graduation, she has been working in Zhejiang Center of Disease Control and Prevention, China. Her research mainly focuses on the field of Molecular Immunology under the guidance of Prof. Yanling Wu.
 Yoshimasa Tanaka received his Ph. D in Hokkaido University Graduate School of Agriculture with a specialization in Enzymology and Biochemistry. After graduation, he continued his research in the field of Immunobiology. Since 2008, he is an associate professor and works in the Center for Innovation in Immunoregulative Immunology and Therapeutics that belongs to the Kyoto University Graduate School of Medicine.
Yoshimasa Tanaka received his Ph. D in Hokkaido University Graduate School of Agriculture with a specialization in Enzymology and Biochemistry. After graduation, he continued his research in the field of Immunobiology. Since 2008, he is an associate professor and works in the Center for Innovation in Immunoregulative Immunology and Therapeutics that belongs to the Kyoto University Graduate School of Medicine.
 Dr. Wen Zhang is a full professor with 25 years of research and teaching experience in Bioorganic Chemistry and Chemical Biology. Dr. Zhang earned his doctorate degree in Bioorganic Chemistry from East China University of Science and Technology, China. Then, he entered Professor Ohrui's Lab of Tohoku University, Japan, working in the field of molecular recognition as a JSPS postdoctoral fellow. After that, he moved to Kyoto University, Japan, to join Professor Sugiyama's Chemical Biology group as a COE and JST research fellow working on biology and chemistry of polyamide-nucleic acids interaction. Now, Dr. Zhang has a special interest in elucidating the gene regulation mechanisms with small organic molecules and the development of gene-targeted drug. His group formed in 2008 and established an extremely fruitful collaboration with Prof. Sugiyama's Group in order to better pursue aspects of gene-targeted drug research. To date, Dr. Zhang has published better papers as the first/corresponding author in excellent Journals including JACS, JASN, ChemBioChem, Chem & Biol, Int J Biol Sci etc.
Dr. Wen Zhang is a full professor with 25 years of research and teaching experience in Bioorganic Chemistry and Chemical Biology. Dr. Zhang earned his doctorate degree in Bioorganic Chemistry from East China University of Science and Technology, China. Then, he entered Professor Ohrui's Lab of Tohoku University, Japan, working in the field of molecular recognition as a JSPS postdoctoral fellow. After that, he moved to Kyoto University, Japan, to join Professor Sugiyama's Chemical Biology group as a COE and JST research fellow working on biology and chemistry of polyamide-nucleic acids interaction. Now, Dr. Zhang has a special interest in elucidating the gene regulation mechanisms with small organic molecules and the development of gene-targeted drug. His group formed in 2008 and established an extremely fruitful collaboration with Prof. Sugiyama's Group in order to better pursue aspects of gene-targeted drug research. To date, Dr. Zhang has published better papers as the first/corresponding author in excellent Journals including JACS, JASN, ChemBioChem, Chem & Biol, Int J Biol Sci etc.
![]() Corresponding author: Yanling Wu, Lab of Molecular Immunology, Virus Inspection Department of Zhejiang Provincial Center for Disease Control and Prevention, 630 Xincheng Road, Hangzhou, 310051, PR China; Tel: +86-571-87115282; Fax: +86-571-87115282; e-mail: ylwuzj.cn. Wen Zhang, Lab of Chemical Biology and Molecular Drug Design, College of Pharmaceutical Science, Zhejiang University of Technology, 18 Chaowang Road, Hangzhou, 310014, PR China; Tel: +86-571-88871507; Fax: +86-571-88871507; e-mail: wzhang63edu.cn.
Corresponding author: Yanling Wu, Lab of Molecular Immunology, Virus Inspection Department of Zhejiang Provincial Center for Disease Control and Prevention, 630 Xincheng Road, Hangzhou, 310051, PR China; Tel: +86-571-87115282; Fax: +86-571-87115282; e-mail: ylwuzj.cn. Wen Zhang, Lab of Chemical Biology and Molecular Drug Design, College of Pharmaceutical Science, Zhejiang University of Technology, 18 Chaowang Road, Hangzhou, 310014, PR China; Tel: +86-571-88871507; Fax: +86-571-88871507; e-mail: wzhang63edu.cn.

 Global reach, higher impact
Global reach, higher impact