Impact Factor ISSN: 1449-2288
Int J Biol Sci 2014; 10(8):846-860. doi:10.7150/ijbs.9297 This issue Cite
Research Paper
Identification of Candidate Olfactory Genes in Chilo suppressalis by Antennal Transcriptome Analysis
1. State Key Laboratory for Biology of Plant Diseases and Insect Pests, Institute of Plant Protection, Chinese Academy of Agricultural Sciences, Beijing, China
2. Laboratory of Pesticide, College of Plant Science & Technology, Huazhong Agricultural University, Wuhan, China
3. Swedish University of Agricultural Sciences, Department of Plant Protection Biology, Chemical Ecology Research Group, Alnarp, Sweden
# These authors contributed equally to this work.
Received 2014-4-4; Accepted 2014-5-25; Published 2014-7-26
Abstract
Antennal olfaction, which is extremely important for insect survival, mediates key behaviors such as host preference, mate choice, and oviposition site selection. In insects, odor detection is mediated by multiple proteins in the antenna, especially the odorant receptors (ORs) and ionotropic receptors (IRs), which ensure the specificity of the olfactory sensory neuron responses. In this study, we identified the olfactory gene repertoire of the rice stem borer, Chilo suppressalis, an economically important agricultural pest, which inflicts great damage to the rice yield in south and east part of Asia, especially in Southern China. By Illumina sequencing of male and female antennal transcriptomes, we identified 47 odorant receptors, 20 ionotropic receptors, 26 odorant binding proteins, 21 chemosensory proteins and 2 sensory neuron membrane proteins. Our findings make it possible for future research of the olfactory system of C. suppressalis at the molecular level.
Keywords: C. suppressali, antennal olfaction, olfactory gene
Introduction
Chemical sensing is critically important to insect survival. For insects, olfaction, which is the primary sensory perception modality, is used to detect odor molecules in the environment. Olfaction guides the insect towards food, mating partners, and oviposition sites and also to facilitate detection of predators and toxic compounds [1]. The antenna is a specialized organ for insect sensing, especially for olfaction. Several types of sensilla, which are specialized hair-like, multi-pore structures, cover the surface of the antennae. Olfactory receptor neurons (ORNs) and auxiliary structures are housed within the antennae, positioned at the sensilla root [2]. For most of the olfactory sensilla, each hosts 1-4 ORNs, which extend their dendrites up into the sensilla and project their axons into the antennal lymph on towards the brain [3]. The ORNs convert ecologically relevant volatile chemicals into an electrical impulse, which is transported to the primary olfactory center of the brain, the antennal lobe [4]. Within the sensilla-ORN structure, a number of gene families have been identified to play active roles in olfaction. These include the odorant binding proteins (OBPs), chemosensory proteins (CSPs), odorant receptors (ORs) ionotropic receptors (IRs), and the sensory neuron membrane proteins (SNMPs).
OBPs are hydrophilic soluble proteins that are secreted by the accessory cells around the ORNs and accumulate in the sensilla lymph [5]. OBPs are thought to be the first proteins that participate in the olfactory signal transduction procedure [6]. It is postulated that as the odor molecules diffuse through pores on sensilla, the soluble OBPs in sensillum lymph fluid selectively bind the liposoluble odor molecules [7] and transport them through the sensillum lymph to the surface of ORN dendrites [8, 9]. OBPs are also thought to be directly involved in the activation of ORx/Orco complex in the recognition of some special odors [10, 11]. Like OBPs, the CSPs are small soluble proteins that are enriched in the sensillum lymph, but also expressing broadly in non-olfactory tissues. The olfactory properties of CSPs are quite clear, for they bind odorant or pheromone compounds [12], but little is known about how CSPs function in olfactory system. Moreover, the broad tissues expression of CSPs implies unknown roles in non-olfactory procedures.
ORs are trans-membrane proteins located in the dendrite membrane of ORNs. Insect ORs are seven-transmembrane domain proteins [13] with a reversed membrane topology (intracellular N-terminus) compared to the G-protein coupled vertebrate ORs [14]. In chemosensory signal transduction process, ORs play a central role as a bio-transducer, facilitating the conversion of the chemical message to an electrical signal. In this system, it is generally thought that in disparate individual OSNs, a single variable, ligand-binding ORx and a highly conserved, non-ligand binding Orco protein make up a stand-alone heteromeric structure that functions as a ligand-gated ion channel [14-17].
IRs make up a recently discovered ionotropic glutamate receptor (iGluR) -like protein family which has been shown to be involved in chemosensation [18]. The insect IRs contain structural regions that are conserved in iGluRs, namely, three transmembrane domains (M1, M2 and M3), a bipartite ligand-binding domain with two lobes (S1 and S2) and one ion channel pore (P). But the conserved iGluR glutamate binding residues in S1 and S2 lobes are not retained in IRs, indicating their atypical binding characters [19]. Unlike the exclusive ORs, two or three IR genes were always co-expressed with one or both of the conserved IR8a and IR25a in one IR-expressing neuron [18]. Furthermore, IRs are thought to be a class of receptors far more ancient than OR families that animals use for sensing chemicals in the surrounding environment; this gene family has an extensive distribution, as it is found in mollusks, annelid and nematodes [20], and it displays a relatively high homology across species [21].
In this study, we sequenced and analyzed Chilo suppressalis adult antennal transcriptomes using Illumina sequencing. Our goals were to identify olfaction-related genes of this pest insect species, which is destructive to the rice farming in China, across Asia and in the Pacific. We report the results including sequencing, gene annotation, GO annotation and specifically, a set of 47 ORs, 20 IRs, 26 OBPs, 21 CSPs and 2 SNMPs.
Results
Transcriptome overview
With utilization of a 90PE RNA-Seq strategy by Illumina HiSeq 2000, about 56.4 million and 58.8 million raw-reads were obtained respectively from the libraries of male and female antenna. After filtering, 53.4 million and 55.3 million clean-reads comprised of 4.8 and 4.9 gigabases were generated for male and female antenna. Assemblies led to the generation of 79,706 and 77,404 unigenes separately for male and female. After merging and clustering, a final transcript dataset was revealed, with 66,560 unigenes consisting of 15,462 distinct clusters and 51,098 distinct singletons. The dataset was 50.63 megabases in size and with a mean length of 761nt and N50 of 1,271nt. 11,849 unigenes were larger than 1,000nt in length, which comprised 17.80% of all unigenes (Table 1).
Assembly summary of C. suppressalis antenna transcriptome
| Sample | Total Number | Total Length(nt) | Mean Length(nt) | N50 | Consensus Sequences | Distinct Clusters | Distinct Singletons | |
|---|---|---|---|---|---|---|---|---|
| Contig | Female | 130,229 | 44,138,907 | 339 | 569 | - | - | - |
| Male | 133,394 | 44,421,350 | 333 | 543 | - | - | - | |
| Unigene | Female | 77,404 | 45,204,675 | 584 | 969 | 77,404 | 12,254 | 65,150 |
| Male | 79,706 | 44,793,753 | 562 | 916 | 79,706 | 11,670 | 68,036 | |
| Merge | 66,560 | 50,635,660 | 761 | 1271 | 66,560 | 15,462 | 51,098 |
Through annotation by blastx, 30,232 (45.4%) unigenes matched to known proteins; the remaining unigenes failed to match against any sequence with an e-value < 1e-5 in neither of the nr nor SwissProt databases. Among the annotated unigenes, 70.4% had a best blast match to Lepidopteran sequences, primarily Danaus plexippus (59.2%), and Bombyx mori (7.4%) (Figure 1A). 52.0% of the annotated unigenes showed strong homology, with e-value < 1e-45.
Annotation summary of C. suppressalis antenna unigenes. (A) Species distribution of unigenes' best-hit annotation term in nr database. (B) Gene ontology classifications of the C. suppressalis unigenes.
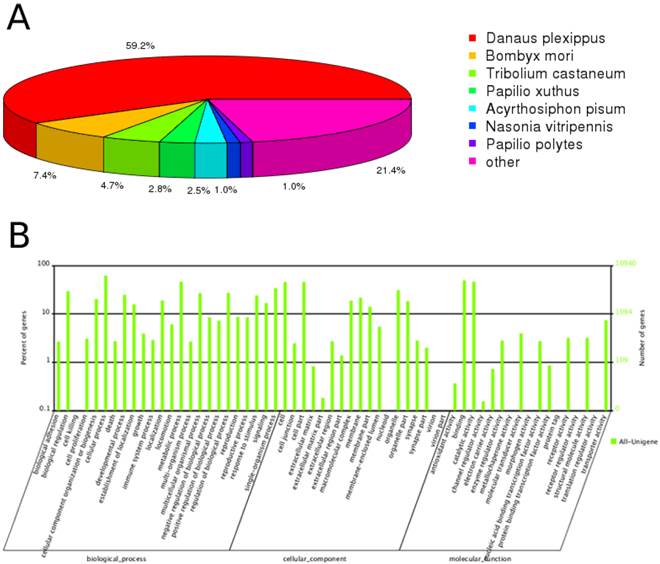
Gene ontology (GO) annotation of the unigenes was obtained using Blast2GO pipeline according to the blastx search against nr. From the 66,560 final unigenes set, a total of 10,940 unigenes were assigned various GO terms. In the molecular function category, the genes expressed in the antennae were mostly enriched to molecular binding activity (e.g., nucleotide, ion and odorant binding) and catalytic activity (e.g., hydrolase and oxidoreductase). In the biological process terms, cellular and metabolic processes were the most represented. In the cellular component terms, cell, cell part and organelle were the most abundant (Figure 1B).
Identification of Candidate Chemosensory Receptors
The unigenes related to candidate chemosensory receptors were identified by keyword search of the blastx annotation. The predicted protein sequences of the unigenes were further searched by PSI-blastp with known Lepidopteran chemosensory receptors [4] to indentify more candidate ORs. We identified 47 distinct unigenes that were putative OR genes. Of these, 23 sequences were full-length OR genes because they have intact open reading frames with a general length of 1,200bp and 5-7 transmembrane domains, which are characteristic of typical insect ORs.
The C. suppressalis Orco co-receptor orthologue was easily detected as it has a high degree of identity with the conserved insect co-receptor: this gene was named CsupOrco. Six unigenes were considered to be putative pheromone receptors because they shared considerable similarity with previously characterized Lepidopteran pheromone receptors and were clustered together into one subgroup in the phylogenetic tree (Figure 2). These 6 candidate ORs were named as “CsupPRx” (x=1 through 6), to more clearly indicate function, as previously reported Lepidopteran pheromone receptors were not clearly separated from the general odorant receptors and followed no orderly numbering system. The naming convention followed in this report is also consistent with general naming of odorant binding proteins, where the pheromone binding proteins (PBPs) are distinguished from other odorant binding proteins (OBPs). Other candidate ORs were highly divergent and shared low similarity with other insect ORs, which is common for insect olfactory receptor genes. These genes were named as “CsupOR”, followed by a numeral, in descending order of their coding region lengths.
Phylogenetic tree of candidate CsupORs with known lepidopteran OR sequences. Harm: H. armigera; Hvir: H. virescens; Bmor: B. mori; Pxyl: P. xylostella; Osca: Ostrinia scapulalis; Onub: Ostrinia nubilalis; Ozea: Ostrinia zealis; Ofur: Ostrinia furnacalis; Opal: Ostrinia palustralis; Ozag: Ostrinia zaguliaevi: Oova: Ostrinia ovalipennis; Olat: Ostrinia latipennis. The clade in blue indicates the pheromone receptor gene clade.
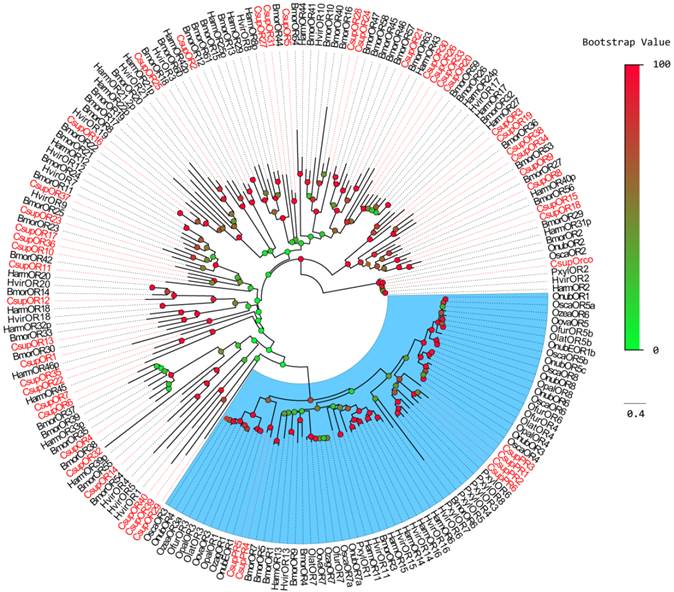
Phylogenetic analysis was performed with ORs from B. mori, H. armigera, H. virescens, and PR sequences from P. xylostella and some Crambidae insects. For the relatively conserved PR genes, the CsupPR4 and CsupPR5 were clustered together with the Crambidae pheromone receptor 1 and 3. CsupPR1, 2, 3 and 6 were not closely grouped with the Crambidae PRs but clustered with the P. xylostella PR clade with high bootstrap support. Almost all CsupOR candidates clustered with at least one Lepidopteran orthologous gene in the phylogenetic tree. No C. suppressalis specific OR family expansion was discovered in our phylogenetic tree.
Information including unigene reference, length, and best blastx hit of all 47 odorant receptors are listed in Table 2. The sequences are listed in Additional File 1: Supplementary Material S1.
Unigenes of candidate olfactory receptors
| Gene name | Length (nt) | ORF (aa) | Unigene reference | Status | TMD (No.) | Evalue | BLASTx best hit |
|---|---|---|---|---|---|---|---|
| ORco | |||||||
| CsupOrco | 3864 | 474 | CL5509.Contig1 | Complete ORF | 7 | 0.0E+00 | gb|AFQ94048.1|olfactory receptor 2 [Chilo suppressalis] |
| Pheromone receptor | |||||||
| CsupPR1 | 1492 | 424 | Unigene14957 | Complete ORF | 6 | 1.0E-94 | gb|ADB89183.1|odorant receptor 6 [Ostrinia nubilalis] |
| CsupPR2 | 2220 | 367 | Unigene18611 | Complete ORF | 4 | 2.0E-95 | gb|ADB89183.1|odorant receptor 6 [Ostrinia nubilalis] |
| CsupPR3 | 2044 | 264 | CL1103.Contig5 | Complete ORF | 6 | 1.0E-56 | gb|AFK30402.1|E-race odorant receptor 6 [Ostrinia nubilalis] |
| CsupPR4 | 483 | 160 | CL812.Contig2 | 5',3' lost | 2 | 2.0E-28 | dbj|BAG71417.1|olfactory receptor-1 [Diaphania indica] |
| CsupPR5 | 297 | 99 | CL4759.Contig1 | 5',3' lost | 1 | 2.0E-21 | dbj|BAH57981.1|olfactory receptor [Ostrinia latipennis] |
| CsupPR6 | 259 | 86 | Unigene43713 | 5',3' lost | 2 | 6.0E-10 | gb|AFK30403.1|odorant receptor 6 [Ostrinia furnacalis] |
| Other odorant receptor | |||||||
| CsupOR1 | 1537 | 457 | Unigene28449 | Complete ORF | 7 | 5.0E-152 | ref|NP_001116817.1|olfactory receptor-like [Bombyx mori] |
| CsupOR2 | 1371 | 446 | Unigene15165 | Complete ORF | 6 | 2.0E-179 | ref|NP_001155301.1|olfactory receptor 60 [Bombyx mori] |
| CsupOR3 | 1612 | 439 | Unigene29790 | Complete ORF | 5 | 5.0E-56 | dbj|BAH66328.1|olfactory receptor [Bombyx mori] |
| CsupOR4 | 1443 | 432 | CL3655.Contig2 | Complete ORF | 6 | 1.0E-140 | gb|AFL70813.1|odorant receptor 50, partial [Manduca sexta] |
| CsupOR5 | 1445 | 429 | Unigene22904 | Complete ORF | 7 | 4.0E-141 | ref|NP_001166607.1|olfactory receptor 44 [Bombyx mori] |
| CsupOR6 | 1381 | 423 | CL5260.Contig2 | Complete ORF | 6 | 1.0E-131 | gb|AFL70813.1|odorant receptor 50, partial [Manduca sexta] |
| CsupOR7 | 2645 | 423 | Unigene26044 | Complete ORF | 4 | 2.0E-27 | gb|EHJ75140.1|olfactory receptor [Danaus plexippus] |
| CsupOR8 | 1423 | 422 | Unigene990 | Complete ORF | 5 | 3.0E-114 | ref|NP_001166893.1|olfactory receptor 27 [Bombyx mori] |
| CsupOR9 | 1429 | 422 | CL3918.Contig4 | 5' lost | 6 | 4.0E-112 | gb|AFC91736.1|putative odorant receptor OR28 [Cydia pomonella] |
| CsupOR10 | 1600 | 414 | Unigene24741 | 5' lost | 7 | 4.0E-116 | ref|NP_001091818.1|olfactory receptor 42 [Bombyx mori] |
| CsupOR11 | 1270 | 406 | CL5145.Contig2 | Complete ORF | 6 | 1.0E-10 | sp|P81922|Odorant receptor 47b [Drosophila melanogaster] |
| CsupOR12 | 1501 | 402 | Unigene11744 | Complete ORF | 5 | 4.0E-116 | gb|ACC63240.1|olfactory receptor 20, partial [Helicoverpa armigera] |
| CsupOR13 | 1318 | 402 | CL2287.Contig1 | Complete ORF | 7 | 7.0E-109 | tpg|DAA05986.1|TPA: TPA_exp: odorant receptor 30 [Bombyx mori] |
| CsupOR14 | 1209 | 400 | Unigene4576 | 5' lost | 6 | 1.0E-119 | ref|NP_001166616.1|olfactory receptor 54 [Bombyx mori] |
| CsupOR15 | 1275 | 400 | Unigene35932 | Complete ORF | 5 | 1.0E-150 | ref|NP_001166617.1|olfactory receptor 56 [Bombyx mori] |
| CsupOR16 | 1344 | 397 | Unigene33520 | 3' lost | 6 | 3.0E-135 | ref|NP_001166613.1|olfactory receptor 22 [Bombyx mori] |
| CsupOR17 | 1365 | 397 | CL458.Contig1 | Complete ORF | 6 | 6.0E-162 | gb|AFC91721.1|putative odorant receptor OR12 [Cydia pomonella] |
| CsupOR18 | 1442 | 397 | CL3235.Contig2 | 3' lost | 6 | 7.0E-134 | ref|NP_001166894.1|olfactory receptor 29 [Bombyx mori] |
| CsupOR19 | 1632 | 395 | CL545.Contig1 | Complete ORF | 5 | 8.0E-54 | gb|EHJ63141.1|olfactory receptor [Danaus plexippus] |
| CsupOR20 | 1382 | 390 | CL1707.Contig3 | Complete ORF | 5 | 6.0E-124 | emb|CAG38118.1|putative chemosensory receptor 17 [Heliothis virescens] |
| CsupOR21 | 1220 | 386 | CL727.Contig1 | Complete ORF | 5 | 2.0E-94 | ref|NP_001166620.1|olfactory receptor 63 [Bombyx mori] |
| CsupOR22 | 1388 | 381 | Unigene18694 | Complete ORF | 6 | 4.0E-118 | gb|AFC91732.1|putative odorant receptor OR24 [Cydia pomonella] |
| CsupOR23 | 1193 | 379 | Unigene18626 | 5' lost | 6 | 9.0E-60 | ref|NP_001166606.1|olfactory receptor 23 [Bombyx mori] |
| CsupOR24 | 1270 | 378 | CL380.Contig1 | Complete ORF | 4 | 3.0E-78 | gb|AFC91719.1|putative odorant receptor OR10 [Cydia pomonella] |
| CsupOR25 | 863 | 377 | Unigene17554 | 5' lost | 2 | 3.0E-86 | ref|NP_001166895.1|olfactory receptor 18 [Bombyx mori] |
| CsupOR26 | 1214 | 375 | Unigene22379 | 5' lost | 4 | 2.0E-54 | ref|NP_001091790.1|candidate olfactory receptor [Bombyx mori] |
| CsupOR27 | 545 | 370 | Unigene24576 | 5' lost | 4 | 1.0E-26 | tpg|DAA05974.1|TPA: TPA_exp: odorant receptor 15 [Bombyx mori] |
| CsupOR28 | 1061 | 353 | Unigene22927 | 5',3' lost | 6 | 3.0E-135 | ref|NP_001104832.2|olfactory receptor 16 [Bombyx mori] |
| CsupOR29 | 1060 | 348 | Unigene33676 | 5' lost | 6 | 4.0E-118 | gb|AEF32141.1|odorant receptor [Spodoptera exigua] |
| CsupOR30 | 1170 | 346 | CL1602.Contig3 | 5' lost | 5 | 2.0E-67 | gb|AFC91739.1|putative odorant receptor OR31 [Cydia pomonella] |
| CsupOR31 | 1022 | 340 | Unigene28661 | 5',3' lost | 6 | 1.0E-26 | gb|EHJ65088.1|olfactory receptor 44 [Danaus plexippus] |
| CsupOR32 | 2686 | 332 | CL4999.Contig1 | Complete ORF | 6 | 3.0E-43 | ref|NP_001091791.1|candidate olfactory receptor [Bombyx mori] |
| CsupOR33 | 971 | 323 | Unigene30218 | 5',3' lost | 4 | 8.0E-87 | ref|NP_001091790.1|candidate olfactory receptor [Bombyx mori] |
| CsupOR34 | 1000 | 295 | Unigene35881 | 5' lost | 4 | 8.0E-82 | ref|NP_001166892.1|olfactory receptor 36 [Bombyx mori] |
| CsupOR35 | 911 | 272 | CL5748.Contig2 | 5' lost | 2 | 6.0E-76 | gb|AFC91725.1|putative odorant receptor OR17 [Cydia pomonella] |
| CsupOR36 | 793 | 264 | Unigene26834 | 5',3' lost | 4 | 5.0E-65 | ref|NP_001091818.1|olfactory receptor 42 [Bombyx mori] |
| CsupOR37 | 834 | 245 | CL296.Contig2 | 5' lost | 2 | 1.0E-66 | dbj|BAH66322.1|olfactory receptor [Bombyx mori] |
| CsupOR38 | 657 | 219 | Unigene35370 | 5',3' lost | 4 | 1.0E-72 | ref|NP_001166892.1|olfactory receptor 36 [Bombyx mori] |
| CsupOR39 | 604 | 163 | CL4235.Contig2 | 3' lost | 3 | 3.0E-16 | dbj|BAH66322.1|olfactory receptor [Bombyx mori] |
| CsupOR40 | 406 | 120 | CL4235.Contig1 | 3' lost | 2 | 1.0E-11 | ref|NP_001104828.1|olfactory receptor 25 [Bombyx mori] |
Identification of Candidate Ionotropic Receptors
The putative IR genes in the C. suppressalis antennal transcriptome were represented according to their similarity to known insect IRs. Bioinformatic analysis led to the identification of 20 candidates IRs, in which 13 sequences contain a full-length ORF, the remaining 7 sequences are marked as incomplete due to lacking a complete 5' or 3' terminus. The insect IRs contained three transmembrane domains. TMHMM2.0 predicted 10 IR candidates with three transmembrane domains (Table 3).
Unigenes of candidate ionotropic receptors
| Gene name | Length (nt) | ORF (aa) | Unigene reference | Status | TMD (No.) | Evalue | BLASTx best hit |
|---|---|---|---|---|---|---|---|
| CsupIR1 | 1934 | 574 | Unigene35421 | 5' lost | 3 | 0.0E+00 | gb|EHJ72198.1|putative ionotropic glutamate receptor-invertebrate [Danaus plexippus] |
| CsupIR1.1 | 2121 | 656 | CL4511.Contig3 | Complete ORF | 3 | 1.0E-113 | gb|EHJ72198.1|putative ionotropic glutamate receptor-invertebrate [Danaus plexippus] |
| CsupIR2 | 2147 | 320 | CL2718.Contig3 | Complete ORF | 4 | 4.0E-163 | gb|AAB62572.1|GABA-gated chloride channel isoform a3 [Heliothis virescens] |
| CsupIR21a | 2972 | 844 | Unigene11518 | Complete ORF | 3 | 0.0E+00 | gb|ADR64678.1|putative chemosensory ionotropic receptor IR21a [Spodoptera littoralis] |
| CsupIR25a | 3304 | 927 | Unigene17452 | Complete ORF | 3 | 2.0E-75 | sp|P39087|Glutamate receptor, ionotropic kainate 2 [Mus musculus] |
| CsupIR3 | 2100 | 474 | Unigene29712 | Complete ORF | 4 | 0.0E+00 | gb|EHJ68597.1|putative glycine receptor beta precursor [Danaus plexippus] |
| CsupIR4 | 931 | 277 | Unigene904 | 5',3' lost | 2 | 5.0E-12 | sp|Q68Y21|Glutamate receptor delta-2 subunit [Danio rerio] |
| CsupIR40a | 2918 | 707 | CL4571.Contig2 | Complete ORF | 3 | 1.0E-15 | sp|Q9ULK0|Glutamate receptor delta-1 subunit [Homo sapiens] |
| CsupIR41a | 1989 | 598 | CL876.Contig2 | 5' lost | 3 | 0.0E+00 | gb|AFC91758.1|putative ionotropic receptor IR41a [Cydia pomonella] |
| CsupIR64a | 1258 | 380 | Unigene22885 | 5' lost | 4 | 8.0E-17 | sp|Q68Y21|Glutamate receptor delta-2 subunit [Danio rerio] |
| CsupIR68a | 2044 | 674 | Unigene14878 | Complete ORF | 6 | 0.0E+00 | gb|ADR64682.1|putative chemosensory ionotropic receptor IR68a [Spodoptera littoralis] |
| CsupIR75d | 1145 | 294 | CL349.Contig2 | 5' lost | 1 | 1.0E-11 | sp|P34299|Glutamate receptor 1 [Caenorhabditis elegans] |
| CsupIR75p | 2200 | 615 | CL46.Contig4 | Complete ORF | 3 | 0.0E+00 | gb|ADR64684.1|putative chemosensory ionotropic receptor IR75p [Spodoptera littoralis] |
| CsupIR75p.1 | 1429 | 441 | CL2655.Contig2 | 5' lost | 4 | 0.0E+00 | gb|EHJ72019.1|putative ionotropic glutamate receptor-invertebrate [Danaus plexippus] |
| CsupIR75q1 | 588 | 196 | Unigene9838 | 5',3' lost | 2 | 1.0E-56 | gb|AFC91752.1|putative ionotropic receptor IR75q2 [Cydia pomonella] |
| CsupIR75q2 | 2011 | 635 | CL1806.Contig1 | Complete ORF | 4 | 0.0E+00 | gb|AFC91752.1|putative ionotropic receptor IR75q2 [Cydia pomonella] |
| CsupIR76b | 2070 | 547 | Unigene33212 | Complete ORF | 3 | 0.0E+00 | gb|AFC91765.1|putative ionotropic receptor IR76b [Cydia pomonella] |
| CsupIR87a | 2067 | 647 | Unigene8213 | Complete ORF | 6 | 4.0E-07 | sp|O43424|Glutamate receptor delta-2 subunit [Homo sapiens] |
| CsupIR8a | 3058 | 912 | Unigene17458 | Complete ORF | 3 | 0.0E+00 | gb|AFC91764.1|putative ionotropic receptor IR8a, partial [Cydia pomonella] |
| CsupIR93a | 2878 | 877 | CL2805.Contig1 | Complete ORF | 3 | 1.0E-33 | sp|Q63226|Glutamate receptor delta-2 subunit [Rattus norvegicus] |
To distinguish putative IRs from iGluRs, putative C. suppressalis IRs were aligned with IR orthologoues from D. melanogaster, B. mori, S. littoralis and some D. melanogaster iGluRs for a phylogenetic analysis. The results revealed a clear segregation between DmeliGluRs and insect IRs (Figure 3). In the phylogenetic tree of IRs, most C. suppressalis IR candidates clustered with their ionotropic receptor orthologues into a separate clade. According to their positions in phylogenetic tree and strong bootstrap support, 15 of 20 candidate C. suppressalis IRs were given names consistent with the number and suffix of the Dmel/Bmor/Slit IR orthologues in the same clade.
Phylogenetic tree of candidate CsupIRs with known lepidopteran IRs and iGluRs. Dmel: D. melanogaster,Bmor: B. mori, Slit: S. littoralis. The clade in blue indicates the iGluR gene clade; the clade in purple indicates the IR8a/IR25a clade.
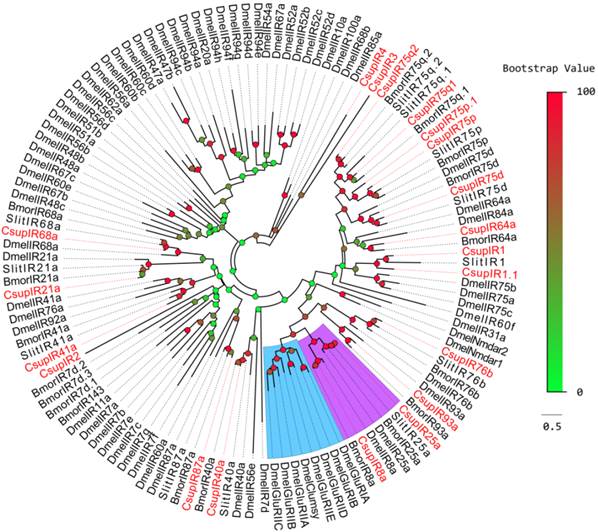
Two of the remaining 5 IR sequences, CL4511.Contig3 and CL2655.Contig2, were clustered into the SlitIR1 and Slit/Bmor IR75p clades, respectively, with reliable bootstrap support, forming small expansions with the CsupIR1 and CsupIR75p genes. Considering that these two sequences contain typical IR characteristics, these two sequences may likely be C. suppressalis specific genes, or their orthologues haven't been detected in other insects. These two sequences were named as “CsupIR1.1” and “CsupIR75p.1”, respectively. The other 3 sequences, CL2718.Contig3, Unigene29712 and Unigene904, had low bootstrap values unable to clearly demonstrate their phylogenetic positions, were named as “CsupIR2”, “CsupIR3” and “CsupIR4”, respectively. The information including unigene reference, length, and best blastx hit of all the 20 IRs are listed in Table 3. The sequences of all 20 IRs were listed in SAdditional File 1: upplementary Material S1.
Identification of Putative Odorant-binding Proteins and Chemosensory Proteins
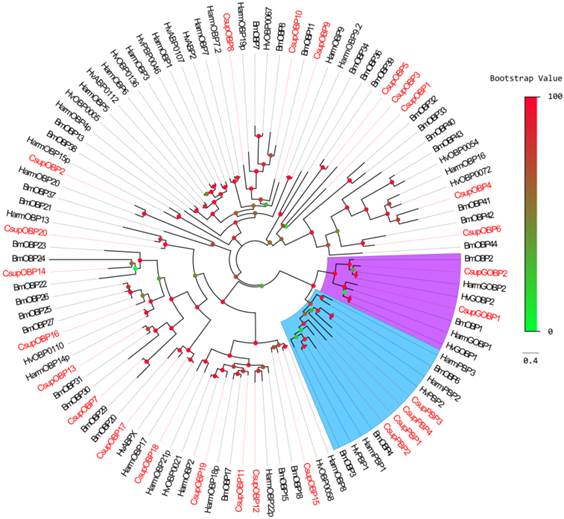
In addition to keyword searching and PSI-Blast, we also used motif scanning to detect the conserved 6 cysteine residues pattern (C1-X5-39-C2-X3-C3-X21-44-C4-X7-12-C5-X8-C6) [22] of the putative odorant-binding proteins. In our transcript set, we identified 26 different sequences encoding odorant binding proteins, including 4 PBPs and 2 GOBPs. In these 26 sequences, 23 had intact ORFs detected; 3 unigenes failed in the signal peptide test which is performed by SignalP. Sequence alignment showed that almost all the putative OBPs shared the classic six-cysteine motif, except 4 sequences (CsupOBP4, 10, 11 and 13), which grouped into the “minus-C” subgroup with their second cysteine residues missing [23]. It was also notable that all 4 “minus-C” OBPs had a lysine residue in place of the C2 cysteine (Figure 4). In the phylogenetic tree, the PBP and GOBP sequences were clustered respectively into the PBP and GOBP clades as expected (Figure 5). All candidate OBP sequences were clustered with at least one lepidopteran orthologue. Comparing our putative OBPs with NCBI records of C. suppressalis, we identified 10 as “discovered genes”, which are GOBP1, 2, PBP1, 2, 3, OBP2, 8, 13, 14 and 17. All of these “discovered genes” have identities over 96% in amino acid to their most similar NCBI records. Therefore, we named these candidate GOBPs and PBPs following the existing NCBI records. We named the candidate OBPs as “CsupOBP” followed by a numeral in descending order of their coding region lengths, as the numbering of existing C. suppressalis OBP records is confusing (Table 4).
Sequences alignment of putative CsupOBPs. The conserved cysteine residues were marked with “*”. Because of the overly long sequence of CsupOBP1, the CsupOBP1 is not included in the multisequence alignment.

Phylogenetic tree of candidate CsupOBPs with known lepidopteran OBPs. Bmor: B. mori, Hv: H. virescens. The clade in blue indicates the PBP gene clade; the clade in purple indicates the GOBP clade.
Unigenes of candidate odorant binding proteins
| Gene name | Length (nt) | ORF (aa) | Unigene reference | Status | Signal Peptide | Evalue | BLASTx best hit |
|---|---|---|---|---|---|---|---|
| CsupGOBP1 | 907 | 173 | CL3430.Contig1 | Complete ORF | Y | 3.0E-97 | gb|ACJ07126.1|general odorant binding protein 1 [Chilo suppressalis] |
| CsupGOBP2 | 1068 | 162 | Unigene17531 | Complete ORF | Y | 1.0E-86 | gb|ACJ07120.1|general odorant binding protein 2 [Chilo suppressalis] |
| CsupPBP1 | 988 | 184 | Unigene28597 | Complete ORF | N | 1.0E-91 | gb|ADK66921.1|pheromone binding protein 1 [Chilo suppressalis] |
| CsupPBP2 | 4508 | 165 | CL470.Contig5 | Complete ORF | Y | 1.0E-89 | gb|ACJ07123.1|pheromone binding protein 2 [Chilo suppressalis] |
| CsupPBP3 | 731 | 170 | Unigene28180 | Complete ORF | Y | 1.0E-91 | gb|ADL09140.1|pheromone binding protein 3 [Chilo suppressalis] |
| CsupPBP4 | 689 | 165 | Unigene31774 | Complete ORF | Y | 1.0E-38 | gb|ADT78499.1|pheromone binding protein 5 [Ostrinia nubilalis] |
| CsupOBP1 | 1224 | 351 | CL5570.Contig1 | Complete ORF | Y | 6.0E-21 | dbj|BAI82446.1|odorant binding protein 6 [Delia antiqua] |
| CsupOBP2 | 1212 | 251 | Unigene24952 | Complete ORF | Y | 1.0E-142 | gb|ADD71058.1|odorant-binding protein [Chilo suppressalis] |
| CsupOBP3 | 917 | 242 | Unigene28587 | Complete ORF | Y | 5.0E-09 | gb|EFA09155.1|odorant binding protein 22 [Tribolium castaneum] |
| CsupOBP4 | 1038 | 194 | CL2795.Contig2 | Complete ORF | Y | 1.0E-44 | gb|EHJ77172.1|odorant binding protein [Danaus plexippus] |
| CsupOBP5 | 753 | 184 | Unigene31885 | Complete ORF | Y | 6.0E-35 | emb|CAX63249.1|odorant-binding protein SaveOBP4 precursor, partial [Sitobion avenae] |
| CsupOBP6 | 1201 | 184 | Unigene22461 | Complete ORF | Y | 1.0E-40 | ref|NP_001159621.1|odorant binding protein LOC100307012 precursor [Bombyx mori] |
| CsupOBP7 | 599 | 156 | Unigene29832 | Complete ORF | Y | 2.0E-14 | gb|AFI57166.1|odorant-binding protein 17 [Helicoverpa armigera] |
| CsupOBP8 | 608 | 153 | Unigene31806 | Complete ORF | Y | 2.0E-79 | gb|AER27567.1|odorant binding protein [Chilo suppressalis] |
| CsupOBP9 | 697 | 150 | Unigene13466 | Complete ORF | Y | 7.0E-27 | gb|AAR28762.1|odorant-binding protein [Spodoptera frugiperda] |
| CsupOBP10 | 620 | 147 | Unigene17320 | Complete ORF | Y | 2.0E-37 | dbj|BAI44701.1|odorant binding protein [Bombyx mori] |
| CsupOBP11 | 1046 | 147 | Unigene542 | Complete ORF | Y | 5.0E-60 | gb|AFG72998.1|odorant-binding protein 1 [Cnaphalocrocis medinalis] |
| CsupOBP12 | 1847 | 145 | Unigene4662 | Complete ORF | N | 4.0E-45 | gb|AFG72998.1|odorant-binding protein 1 [Cnaphalocrocis medinalis] |
| CsupOBP13 | 1151 | 145 | Unigene15206 | 5' lost | Y | 8.0E-60 | gb|AFI57166.1|odorant-binding protein 17 [Helicoverpa armigera] |
| CsupOBP14 | 534 | 140 | Unigene22895 | Complete ORF | Y | 4.0E-15 | gb|ACX53795.1|odorant binding protein [Heliothis virescens] |
| CsupOBP15 | 978 | 139 | Unigene3748 | Complete ORF | Y | 2.0E-63 | gb|AFG73000.1|odorant-binding protein 2 [Cnaphalocrocis medinalis] |
| CsupOBP16 | 636 | 138 | Unigene2333 | Complete ORF | Y | 1.0E-38 | gb|AFI57167.1|odorant-binding protein 18 [Helicoverpa armigera] |
| CsupOBP17 | 1503 | 133 | CL2095.Contig1 | Complete ORF | Y | 2.0E-24 | gb|EFA04687.1|odorant binding protein 08 [Tribolium castaneum] |
| CsupOBP18 | 563 | 129 | CL5651.Contig2 | Complete ORF | N | 2.0E-29 | gb|AFG72998.1|odorant-binding protein 1 [Cnaphalocrocis medinalis] |
| CsupOBP19 | 2038 | 121 | CL5839.Contig1 | 3' lost | Y | 5.0E-44 | gb|EHJ65653.1|odorant-binding protein 1 [Danaus plexippus] |
| CsupOBP20 | 387 | 113 | Unigene36476 | 3' lost | Y | 2.0E-41 | gb|AFD34173.1|odorant binding protein 5 [Argyresthia conjugella] |
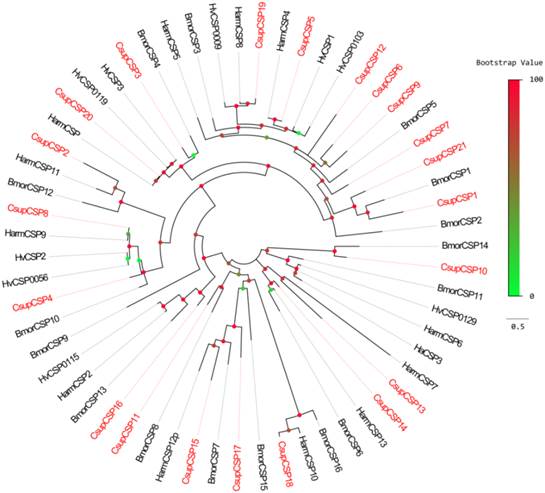
Bioinformatic analysis led to the identification of 21 different sequences encoding candidate CSPs. Among them, 18 sequences have full-length ORFs and signal peptides; Due to incomplete N-termini, the remaining 3 failed in the SignalP test. The conserved cysteine pattern of C1-X6-8-C2-X18-C3-X2-C4 [24] and the six-helix secondary-structure were retained in all 21 candidate CSPs (Figure 6). Neighbor-joining tree analysis showed that all of the 21 sequences clustered with Lepidopteran orthologous genes (Figure 7). These candidate CSPs were named as “CsupCSP” followed by a numeral in descending order of their coding region lengths. The information on the CSPs is listed in Table 5. The sequences are listed in Additional File 1: Supplement Material S1.
Sequences alignment of putative CsupCSPs. The conserved cysteine residues were marked with “*”.

Phylogenetic tree of candidate CsupCSPs with known lepidopteran CSPs. Bmor: B. mori, Hv: H. virescens.
Unigenes of candidate chemosensory proteins
| Gene name | Length (nt) | ORF (aa) | Unigene reference | Status | Signal Peptides | Evalue | BLASTx best hit |
|---|---|---|---|---|---|---|---|
| CsupCSP1 | 727 | 189 | Unigene35502_All | Complete ORF | Y | 2.00E-37 | gb|ACX53806.1|chemosensory protein [Heliothis virescens] |
| CsupCSP2 | 3757 | 151 | CL5573.Contig2_All | Complete ORF | Y | 6.00E-38 | gb|EHJ76401.1|chemosensory protein CSP1 [Danaus plexippus] |
| CsupCSP3 | 1279 | 131 | Unigene21279_All | Complete ORF | Y | 1.00E-52 | gb|ACX53804.1|chemosensory protein [Heliothis virescens] |
| CsupCSP4 | 1782 | 129 | Unigene13693_All | Complete ORF | Y | 1.00E-17 | dbj|BAF91712.1|chemosensory protein [Papilio xuthus] |
| CsupCSP5 | 510 | 128 | CL2398.Contig1_All | Complete ORF | Y | 1.00E-68 | gb|AFR92093.1|chemosensory protein 9 [Helicoverpa armigera] |
| CsupCSP6 | 710 | 126 | Unigene2439_All | Complete ORF | Y | 6.00E-47 | dbj|BAF91714.1|chemosensory protein [Papilio xuthus] |
| CsupCSP7 | 625 | 126 | Unigene31984_All | Complete ORF | Y | 3.00E-43 | dbj|BAG71914.1|chemosensory protein 4a [Papilio xuthus] |
| CsupCSP8 | 503 | 126 | Unigene39861_All | Complete ORF | Y | 5.00E-65 | gb|AAM77040.1|chemosensory protein 2 [Heliothis virescens] |
| CsupCSP9 | 481 | 124 | Unigene39944_All | Complete ORF | Y | 2.00E-64 | gb|ACX53727.1|chemosensory protein [Heliothis virescens] |
| CsupCSP10 | 577 | 123 | CL4176.Contig1_All | Complete ORF | Y | 1.00E-53 | dbj|BAG71921.1|chemosensory protein 13 [Papilio xuthus] |
| CsupCSP11 | 726 | 123 | Unigene13345_All | Complete ORF | Y | 5.00E-46 | dbj|BAF91716.1|chemosensory protein [Papilio xuthus] |
| CsupCSP12 | 456 | 123 | Unigene25770_All | Complete ORF | Y | 9.00E-35 | gb|EHJ78408.1|chemosensory protein [Danaus plexippus] |
| CsupCSP13 | 769 | 121 | CL1604.Contig1_All | Complete ORF | Y | 5.00E-21 | gb|EHJ73331.1|chemosensory protein 12 [Danaus plexippus] |
| CsupCSP14 | 1272 | 121 | CL5801.Contig1_All | Complete ORF | Y | 3.00E-56 | gb|ACX53719.1|chemosensory protein [Heliothis virescens] |
| CsupCSP15 | 503 | 121 | Unigene24547_All | Complete ORF | Y | 9.00E-29 | ref|NP_001037068.1|chemosensory protein 7 precursor [Bombyx mori] |
| CsupCSP16 | 536 | 121 | Unigene28390_All | Complete ORF | Y | 3.00E-41 | dbj|BAF91717.1|chemosensory protein [Papilio xuthus] |
| CsupCSP17 | 2686 | 120 | CL4999.Contig1_All | Complete ORF | Y | 3.00E-27 | dbj|BAG71919.1|chemosensory protein 11b [Papilio xuthus] |
| CsupCSP18 | 1153 | 109 | Unigene15508_All | Complete ORF | Y | 1.00E-48 | dbj|BAF91720.1|chemosensory protein [Papilio xuthus] |
| CsupCSP19 | 367 | 97 | CL2398.Contig2_All | 5' lost | N | 1.00E-51 | gb|AFR92095.1|chemosensory protein 11 [Helicoverpa armigera] |
| CsupCSP20 | 306 | 95 | Unigene38877_All | 5' lost | N | 1.00E-50 | gb|AAK53762.1|chemosensory protein [Helicoverpa armigera] |
| CsupCSP21 | 268 | 89 | Unigene39576_All | 5',3' lost | N | 1.00E-39 | gb|ACX53806.1|chemosensory protein [Heliothis virescens] |
Identification of Candidate Sensory Neuron Membrane Proteins
SNMPs are thought to be involved in the recognition of Lepidopteran pheromone, since they were first identified in Lepidopteran pheromone-sensitive neurons [25, 26]. SNMPs of two families, SNMP1 and 2, were discovered in our C. suppressalis antennal transcriptome. Unigene35775 showed a 99% identity to the CsupSNMP1 published in Genebank. And the CL173.contig15 covered the whole sequence of the CsupSNMP2 (GI: 406668637). Our SNMP unigene sequences are available in Additional File 1: Supplementary Material S1.
Tissue- and Sex- specific Expression of Candidate C. suppressalis OR and IR genes
The expression patterns of the candidate 7 ORs and 20 IRs in male antennae, female antennae and legs were analyzed by semi-quantitative reverse transcription PCR. Results for all of these genes are listed in Figure 8.
Tissue- and sex- specific expressions of candidates CsupORs and CsupIRs. M: male antennae, F: female antennae, L: legs.
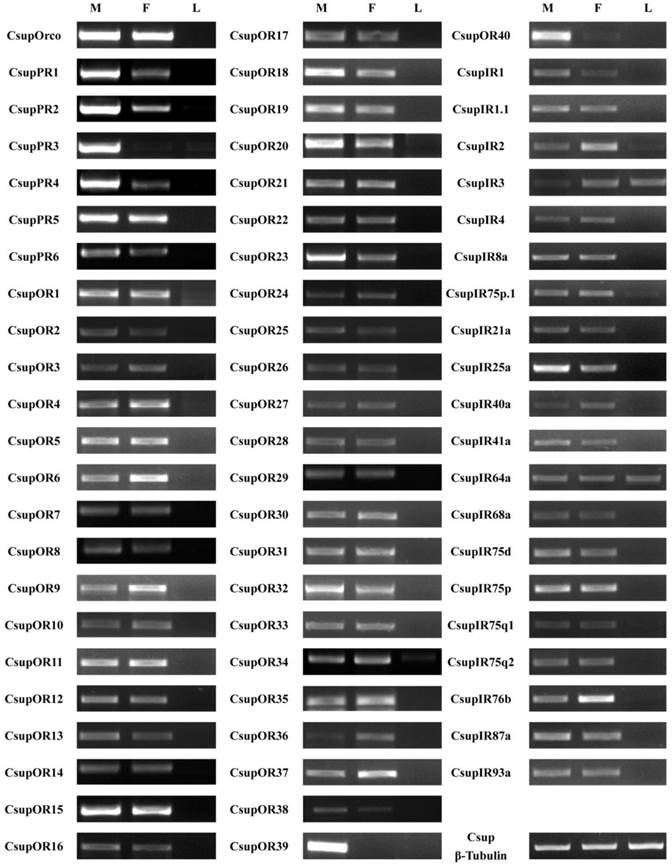

All 47 ORs were successfully detected as expressed in our semi-quantitative RT-PCR analysis. These results indicated that these candidate ORs are expressed in the olfactory organ antennae, but not in non-olfactory organs such as legs. Of the six candidate PRs, only CsupPR3 was found to be exclusive to the male antennae. CsupPR1, 2 and 4 have expression detected in both male and female antennae, but the amounts in male antennae were significantly higher than in female antennae. Of the candidate ORs, two sequences (CsupOR39 and CsupOR40) showed a male-specific expression. The remaining ORs were expressed in both sexes, with some differential expression in male or female antennae. Compared with ORs, the candidate IRs showed no big differences between male and female. Additionally, CsupIR3 and CsupIR64a have considerable expression in leg, at similar levels, based upon luminance of the bands in the agarose gels. CsupIR2 and CsupIR75p.1 are also weakly expressed in legs.
Discussion
Our antennal transcriptome sequencing provides a dataset of 47 ORs, 20 IRs, 26 OBPs and 21 CSPs. All of the previously annotated C. suppressalis chemosensory genes available in NCBI were identified in our dataset. Compared to the antennal transcriptomes of M.sexta with 47 ORs [27], C. pomonella with 43 ORs [28] and Helicoverpa armigera with 47 ORs [4], our OR dataset of 47 sequences is of similar quantity. The neuroanatomical study of C. suppressalis' close relative, the European corn borer (ECB) (Ostrinia nubilalis) suggested that there are 64 glomeruli in the antenna lobe of male and female ECB [29]. If the logic holds true that one olfactory receptor type is expressed in OSN type and axonal projects of different OSNs expressing the same olfactory receptor converge on the same antennal lobe glomeruli, our OR dataset of 47 sequences is quite reasonable, for some glomeruli should also be innervated by OSNs expressing other classes of chemoreceptors such as ionotropic receptors [18].
We identified 6 candidate PR genes by their similarities to PRs in other Lepidopterans and physiologic analysis. But the expression profiles of these six sequences showed that not all of them are exclusive in male antenna. Although PR expression in Bombyx mori and some Lepidoptera have been shown to be restricted to male antennae [30, 31], some recent studies gave examples of PR genes expressed in both sexes. In the antennal transcriptome of Helicoverpa armigera, 6 candidate PR genes were discovered; two of them, HarmOR6 and HarmOR13, display expression in male and female antenna [4]. Two candidate PRs identified in S. littoralis were also found to be expressed in antennae of both sexes [32]. Obviously, this phenomenon cannot be simply ascribed to differences between species. A physiology and morphology study suggested that S. littoralis females detect their own pheromone. The rationale behind female pheromone perception has been proposed to be optimization of pheromone production and spatial dispersion of females over host plants [33, 34].
Ionotropic receptors represent a new member of the chemosensory receptor family, and were first discovered in D. melanogaster [18] through genome analyses. The Ionotropic Receptor family is a variant iGluR subfamily. Animal iGluRs have been best characterized for their essential roles in synaptic transmission as receptors for the excitatory neurotransmitter glutamate [35, 36]. IRs share a considerable degree of commonality with the typical iGluRs: first, they are all located to specialized distal membrane domains of neuronal dendrites (OSN cilia and post-synaptic membranes, respectively); secondly, they responds to binding of extracellular ligands (volatile odorants and neurotransmitter); thirdly, they are comprised of multimeric functional complexes (IR8a/25a co-express with other cell-type specific IRs and iGluRs are formed of heteromeric subunits) [20]. It is easy to conjecture that the IR arose from an iGluR with a change in expression localization from an interneuron to a sensilla neuron [20]. In our study, we found 20 IR candidates in C. suppressalis antennae including two co-receptors, IR8a and IR25a. Compared to ORs, the IR family is relatively conserved both in sequence and expression pattern. Among the 20 CsupIRs we discovered, 15 sequences have orthologs found in Dmel/Bmor/Slit IRs; the expression levels have no significant difference between male and female antenna, which is similar to results in S. littoralis IRs [37] and H. armigera IRs [4]. Considering the relatively high sequence conservation and similarities in expression, the functions of CsupIRs are probably conserved as IRs in other Lepidoptera.
Conclusion
Our goal for this study was to identify genes potentially involved in olfactory signal detection in C. suppressalis, and this is well met by a repertoire of 47 ORs, 20 IRs, 26 OBPs and 21 CSPs. Our approach has been proved to be successful in identifying low-expressing chemosensory receptor genes, especially in a non-model pest species without an available genome sequence. Our findings make it possible for future research of the olfactory system of C. suppressalis at the molecular level, and provide information for comparative and functional genomic analyses of related species.
Materials and Methods
Insects
C. suppressalis were obtained from a laboratory colony maintained at the Institute of Plant Protection, Chinese Academy of Agricultural Sciences, Beijing, China. Larvae were reared on an artificial diet at 28±1°C, 70±5% relative humidity, and under a photoperiod of 14:10 (light: dark). After pupation, male and female pupae were separated for adult eclosion in cages kept at 30±1°C, 80±5% RH and 16: 8 light/dark cycles, and were fed with 10% sugar solution. Antennae of unmated male or female individuals were collected 1-3 days after eclosion and immediately frozen in liquid nitrogen, and stored at -70°C until extraction. Antennae were pulled off with tweezers grasped at the very root of the antennae, in order to reserve the intact structure of antennae.
RNA preparing
Frozen antennae were crushed in a liquid nitrogen cooled vitreous homogenizer and total RNA was extracted using TRIzol reagent (Invitrogen, Carlsbad, CA, USA) following the manufacturer's instructions. Residual DNA in total RNA was removed by DNase I (Promega, Madison, WI, USA). Total RNA was dissolved in RNase-free water and RNA integrity was verified by gel electrophoresis. RNA quantity was determined on a Nanodrop ND-2000 spectrophotometer (NanoDrop products, Wilmington, DE, USA).
cDNA library construction and sequencing
Ten micrograms of total RNA extracted from approximately 500 antennae of 1-3 day old adult male or female moths was used to isolate poly-A RNA using oligo(dT) magnetic beads. Poly-A RNA of each sample was digested into short fragments by fragmentation buffer. Random hexamers were used for first-strand cDNA, followed by second-strand cDNA synthesis using RNase H and DNA polymerase I. These dual-strand DNA samples were treated with T4 DNA Polymerase and T4 Polynucleotide Kinase for end-repairing and dA-tailing, followed by adaptor ligation to the dsDNA's dA tail using T4 DNA ligase. Then bands of insert length around 200bp was collected by 2% agarose gel electrophoresis and purified with QIAquick Gel Extraction Kit (Qiagen, Hilden, Germany), and used as templates for PCR amplification to create the cDNA library.
The library was pair-end sequenced using PE90 strategy (paired-end reads of 90 base pairs per read) on Illumina HiSeq™ 2000 (Illumina, San Diego, CA, USA) at the Beijing Genome Institute (Shenzhen, China). Different libraries were sequenced in one lane and raw-reads were sorted by barcodes in the sequencing adaptor.
Assembly
The raw-reads were treated to generate clean-read datasets by the following procedure. First, reads with adaptors or containing unknown nucleotides (Ns) more than 5% were removed directly. Secondly, low-quality reads containing more than 20% suspect-nucleotides of Phred Quality Score less than 10 were filtered out. Finally, both ends of reads were evaluated to trim unreliable ends containing more than 3 successive suspect-nucleotides. Each clean-read dataset of male and female antenna was feed to Trinity r2012-06-08 [38] separately for De novo assembly using paired reads mode and default parameters. Then the Trinity outputs were clustered by TGICL [39]. The consensus cluster sequences and singletons make up the unigenes dataset.
Functional annotation
The annotation of unigenes were performed by NCBI blastx against a pooled database of non-redundant (nr) and SwissProt protein sequences with e-value < 1e-5. The blast results were then imported into Blast2GO [40] pipeline for GO Annotation. Protein coding region prediction was performed by OrfPredictor [41] according to the blast result. The signal peptide of the protein sequences were predicted using SignalP 4.0 [41] server version (http://www.cbs.dtu.dk/services/SignalP/) with default parameters. The transmembrane-domains of annotated genes were predicted using TMHMM [43] server version2.0 (http://www.cbs.dtu.dk/services/TMHMM) with the new model.
Phylogenetic analyses
The phylogenetic reconstruction of C. suppressalis chemosensory genes was performed according to our previous research [4]. Amino acid sequences were aligned using Clustal Omega [44]. Phylogenetic trees were constructed by the neighbor-joining method, with Jones-Taylor-Thornton (JTT) amino acid substitution model, as implemented in MEGA5.2 software. Node support was assessed using a bootstrap procedure of 1000 replicates.
Expression analysis by semi-quantitative reverse transcription PCR
Semi-quantitative reverse transcription PCR was performed to verify the expression of candidate chemosensory genes. Tissue samples were collected from C. suppressalis adult 1 day after eclosion for 3 biological replicates and total RNA were extracted as mentioned above. The cDNA was synthesized from total RNA using RevertAid First Strand cDNA Synthesis Kit (Thermo Scientific, Waltham, MA, USA) with the gDNA removal procedure performed. Gene-specific primers were designed by Primer3 tool (http://frodo.wi.mit.edu/primer3/) and sequences are available in Additional File 2: supplementary material S2. Taq MasterMix (CWBIO, Beijing, China) was used for PCR reactions under general 3-step amplification of 94 °C for 30s, 55-60 °C for 30s, 72 °C for 30s. The PCR cycle-numbers were adjusted respectively for each gene. For most chemosensory genes, cycle-numbers were range from 30 to 34, but for some high-express-level genes like actin and Orco, cycle-numbers were reduced to 25 to 29. PCR products were run on a 2% agarose gel and verified by DNA sequencing.
Abbreviations
iGluR: ionotropic glutamate receptor; OR: odorant receptor; IR: ionotropic receptor; PR: pheromone receptor; PBP: pheromone binding protein; GOBP: general odorant binding protein; OBP: odorant binding protein; CSP: chemosensory protein; SNMP: sensory neuron membrane protein; GO: gene ontology; FPKM : fragments per kb per million fragments; FDR: false discovery rate; JTT: Jones-Taylor-Thornton amino acid substitution model.
Supplementary Material
Additional File 1Supplementary Material S1 Amino acid sequences of C. suppressalis olfactory genes.
Supplementary Material S2 Primers used in the semi-quantitative RT-PCR analysis.
Acknowledgements
We thank Dr. Yunhe Li (Institute of Plant Protection, Chinese Academy of Agricultural Sciences) for providing the insects. This work was supported by National Basic Research Program of China (973 Program, 2012CB114104), National Natural Science Foundation of China (31230062) and International Science & Technology Cooperation Program of China (2013DFG32230).
Data Deposition
The clean reads of the C. suppressalis antennal transcriptome were stored in the NCBI SRA database, under the accession number of SRX497236 and SRX497239.
Competing Interests
The authors have declared that no competing interest exists.
References
1. Sato K, Touhara K. Insect olfaction: receptors, signal transduction, and behavior. Results and problems in cell differentiation. 2009;47:121-38 doi:10.1007/400_2008_10
2. Gao Q, Chess A. Identification of candidate Drosophila olfactory receptors from genomic DNA sequence. Genomics. 1999;60:31-9 doi:10.1006/geno.1999.5894
3. Zhao XC, Tang QB, Berg BG, Liu Y, Wang YR, Yan FM. et al. Fine structure and primary sensory projections of sensilla located in the labial-palp pit organ of Helicoverpa armigera (Insecta). Cell and tissue research. 2013;353:399-408 doi:10.1007/s00441-013-1657-z
4. Liu Y, Gu S, Zhang Y, Guo Y, Wang G. Candidate olfaction genes identified within the Helicoverpa armigera Antennal Transcriptome. PloS one. 2012;7:e48260. doi:10.1371/journal.pone.0048260
5. Klein U. Sensillum-lymph proteins from antennal olfactory hairs of the moth Antheraea polyphemus (Saturniidae). Insect biochemistry. 1987;17:1193-204 doi: 10.1016/0020-1790(87)90093-X
6. Krieger J, Breer H. Olfactory reception in invertebrates. Science. 1999;286:720-3 doi: 10.1126/science.286.5440.720
7. Campanacci V, Krieger J, Bette S, Sturgis JN, Lartigue A, Cambillau C. et al. Revisiting the specificity of Mamestra brassicae and Antheraea polyphemus pheromone-binding proteins with a fluorescence binding assay. The Journal of biological chemistry. 2001;276:20078-84 doi:10.1074/jbc.M100713200
8. Pelosi P, Maida R. Odorant-binding proteins in insects. Comparative biochemistry and physiology Part B, Biochemistry & molecular biology. 1995;111:503-14 doi: 10.1016/0305-0491(95)00019-5
9. Pelosi P, Zhou JJ, Ban LP, Calvello M. Soluble proteins in insect chemical communication. Cellular and molecular life sciences: CMLS. 2006;63:1658-76 doi:10.1007/s00018-005-5607-0
10. Laughlin JD, Ha TS, Jones DN, Smith DP. Activation of pheromone-sensitive neurons is mediated by conformational activation of pheromone-binding protein. Cell. 2008;133:1255-65 doi:10.1016/j.cell.2008.04.046
11. Kim MS, Repp A, Smith DP. LUSH odorant-binding protein mediates chemosensory responses to alcohols in Drosophila melanogaster. Genetics. 1998;150:711-21
12. Briand L, Swasdipan N, Nespoulous C, Bezirard V, Blon F, Huet JC. et al. Characterization of a chemosensory protein (ASP3c) from honeybee (Apis mellifera L.) as a brood pheromone carrier. European journal of biochemistry / FEBS. 2002;269:4586-96 doi:10.1046/j.1432-1033.2002.03156.x
13. Clyne PJ, Warr CG, Freeman MR, Lessing D, Kim J, Carlson JR. A novel family of divergent seven-transmembrane proteins: candidate odorant receptors in Drosophila. Neuron. 1999;22:327-38 doi:10.1016/S0896-6273(00)81093-4
14. Benton R, Sachse S, Michnick SW, Vosshall LB. Atypical membrane topology and heteromeric function of Drosophila odorant receptors in vivo. PLoS biology. 2006;4:e20. doi:10.1371/journal.pbio.0040020
15. Sato K, Pellegrino M, Nakagawa T, Nakagawa T, Vosshall LB, Touhara K. Insect olfactory receptors are heteromeric ligand-gated ion channels. Nature. 2008;452:1002-6 doi:10.1038/nature06850
16. Wicher D, Schafer R, Bauernfeind R, Stensmyr MC, Heller R, Heinemann SH. et al. Drosophila odorant receptors are both ligand-gated and cyclic-nucleotide-activated cation channels. Nature. 2008;452:1007-11 doi:10.1038/nature06861
17. Larsson MC, Domingos AI, Jones WD, Chiappe ME, Amrein H, Vosshall LB. Or83b encodes a broadly expressed odorant receptor essential for Drosophila olfaction. Neuron. 2004;43:703-14 doi:10.1016/j.neuron.2004.08.019
18. Benton R, Vannice KS, Gomez-Diaz C, Vosshall LB. Variant ionotropic glutamate receptors as chemosensory receptors in Drosophila. Cell. 2009;136:149-62 doi:10.1016/j.cell.2008.12.001
19. Abuin L, Bargeton B, Ulbrich MH, Isacoff EY, Kellenberger S, Benton R. Functional architecture of olfactory ionotropic glutamate receptors. Neuron. 2011;69:44-60 doi:10.1016/j.neuron.2010.11.042
20. Croset V, Rytz R, Cummins SF, Budd A, Brawand D, Kaessmann H. et al. Ancient protostome origin of chemosensory ionotropic glutamate receptors and the evolution of insect taste and olfaction. PLoS genetics. 2010;6:e1001064. doi:10.1371/journal.pgen.1001064
21. Chiu J, DeSalle R, Lam HM, Meisel L, Coruzzi G. Molecular evolution of glutamate receptors: a primitive signaling mechanism that existed before plants and animals diverged. Molecular biology and evolution. 1999;16:826-38
22. Zhou JJ, He XL, Pickett JA, Field LM. Identification of odorant-binding proteins of the yellow fever mosquito Aedes aegypti: genome annotation and comparative analyses. Insect molecular biology. 2008;17:147-63 doi:10.1111/j.1365-2583.2007.00789.x
23. Hekmat-Scafe DS, Scafe CR, McKinney AJ, Tanouye MA. Genome-wide analysis of the odorant-binding protein gene family in Drosophila melanogaster. Genome research. 2002;12:1357-69 doi:10.1101/gr.239402
24. Zhou JJ, Kan Y, Antoniw J, Pickett JA, Field LM. Genome and EST analyses and expression of a gene family with putative functions in insect chemoreception. Chemical senses. 2006;31:453-65 doi:10.1093/chemse/bjj050
25. Rogers ME, Krieger J, Vogt RG. Antennal SNMPs (sensory neuron membrane proteins) of Lepidoptera define a unique family of invertebrate CD36-like proteins. Journal of neurobiology. 2001;49:47-61 doi:10.1002/neu.1065
26. Benton R, Vannice KS, Vosshall LB. An essential role for a CD36-related receptor in pheromone detection in Drosophila. Nature. 2007;450:289-93 doi:10.1038/nature06328
27. Grosse-Wilde E, Kuebler LS, Bucks S, Vogel H, Wicher D, Hansson BS. Antennal transcriptome of Manduca sexta. Proceedings of the National Academy of Sciences of the United States of America. 2011;108:7449-54 doi:10.1073/pnas.1017963108
28. Bengtsson JM, Trona F, Montagne N, Anfora G, Ignell R, Witzgall P. et al. Putative chemosensory receptors of the codling moth, Cydia pomonella, identified by antennal transcriptome analysis. PloS one. 2012;7:e31620. doi:10.1371/journal.pone.0031620
29. Karpati Z, Dekker T, Hansson BS. Reversed functional topology in the antennal lobe of the male European corn borer. The Journal of experimental biology. 2008;211:2841-8 doi:10.1242/jeb.017319
30. Mitsuno H, Sakurai T, Murai M, Yasuda T, Kugimiya S, Ozawa R. et al. Identification of receptors of main sex-pheromone components of three Lepidopteran species. The European journal of neuroscience. 2008;28:893-902 doi:10.1111/j.1460-9568.2008.06429.x
31. Sakurai T, Nakagawa T, Mitsuno H, Mori H, Endo Y, Tanoue S. et al. Identification and functional characterization of a sex pheromone receptor in the silkmoth Bombyx mori. Proceedings of the National Academy of Sciences of the United States of America. 2004;101:16653-8 doi:10.1073/pnas.0407596101
32. Legeai F, Malpel S, Montagné N, Monsempes C, Cousserans F, Merlin C. et al. An Expressed Sequence Tag collection from the male antennae of the Noctuid moth Spodoptera littoralis: a resource for olfactory and pheromone detection research. BMC Genomics. 2011;12:86. doi:10.1186/1471-2164-12-86
33. Palanaswamy P, Seabrook W. Behavioral responses of the female eastern spruce budworm Choristoneura fumiferana (Lepidoptera, Tortricidae) to the sex pheromone of her own species. Journal of Chemical Ecology. 1978;4:649-55
34. Schneider D, Schulz S, Priesner E, Ziesmann J, Francke W. Autodetection and chemistry of female and male pheromone in both sexes of the tiger moth Panaxia quadripunctaria. Journal of Comparative Physiology A. 1998;182:153-61 doi:10.1007/s003590050166
35. Gereau IV RW, Swanson GT. The glutamate receptors. Springer. 2008
36. Madden DR. The structure and function of glutamate receptor ion channels. Nature reviews Neuroscience. 2002;3:91-101 doi:10.1038/nrn725
37. Olivier V, Monsempes C, Francois MC, Poivet E, Jacquin-Joly E. Candidate chemosensory ionotropic receptors in a Lepidoptera. Insect molecular biology. 2011;20:189-99 doi:10.1111/j.1365-2583.2010.01057.x
38. Grabherr MG, Haas BJ, Yassour M, Levin JZ, Thompson DA, Amit I. et al. Full-length transcriptome assembly from RNA-Seq data without a reference genome. Nature biotechnology. 2011;29:644-52 doi:10.1038/nbt.1883
39. Pertea G, Huang X, Liang F, Antonescu V, Sultana R, Karamycheva S. et al. TIGR Gene Indices clustering tools (TGICL): a software system for fast clustering of large EST datasets. Bioinformatics. 2003;19:651-2 doi:10.1093/bioinformatics/btg034
40. Conesa A, Götz S, García-Gómez JM, Terol J, Talón M, Robles M. Blast2GO: a universal tool for annotation, visualization and analysis in functional genomics research. Bioinformatics. 2005;21:3674-6 doi: 10.1093/bioinformatics/bti610
41. Min XJ, Butler G, Storms R, Tsang A. OrfPredictor: predicting protein-coding regions in EST-derived sequences. Nucleic acids research. 2005;33:W677-80 doi:10.1093/nar/gki394
42. Petersen TN, Brunak S, von Heijne G, Nielsen H. SignalP 4.0: discriminating signal peptides from transmembrane regions. Nature methods. 2011;8:785-6 doi:10.1038/nmeth.1701
43. Krogh A, Larsson B, von Heijne G, Sonnhammer EL. Predicting transmembrane protein topology with a hidden Markov model: application to complete genomes. Journal of molecular biology. 2001;305:567-80 doi:10.1006/jmbi.2000.4315
44. Sievers F, Wilm A, Dineen D, Gibson TJ, Karplus K, Li W. et al. Fast, scalable generation of high-quality protein multiple sequence alignments using Clustal Omega. Molecular systems biology. 2011;7:539. doi:10.1038/msb.2011.75
Author contact
Corresponding author: grwangcn (G.R. Wang)

 Global reach, higher impact
Global reach, higher impact