ISSN: 1449-2288International Journal of Biological Sciences
Int J Biol Sci 2015; 11(12):1348-1362. doi:10.7150/ijbs.13132 This issue Cite
Research Paper
Muscle Transcriptional Profile Based on Muscle Fiber, Mitochondrial Respiratory Activity, and Metabolic Enzymes
1. Leibniz Institute for Farm Animal Biology (FBN), Institute for Genome Biology, Wilhelm-Stahl-Allee 2, D-18196 Dummerstorf, Germany
2. 2 Institute of Food Quality and Food Safety, University of Veterinary Medicine Hannover, D-30173 Hannover, Germany.
3. 3 Department of Animal Science, Quality of Food of Animal Origin, Georg-August-University Goettingen, D-37075 Goettingen, Germany.
Abstract
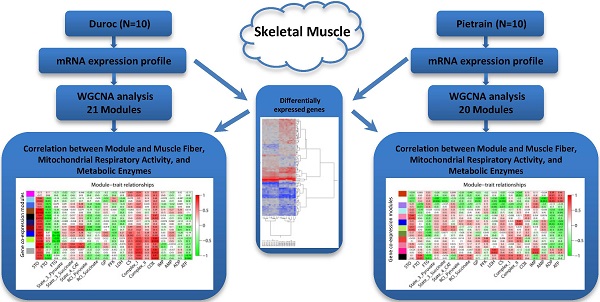
Skeletal muscle is a highly metabolically active tissue that both stores and consumes energy. Important biological pathways that affect energy metabolism and metabolic fiber type in muscle cells may be identified through transcriptomic profiling of the muscle, especially ante mortem. Here, gene expression was investigated in malignant hyperthermia syndrome (MHS)-negative Duroc and Pietrian (PiNN) pigs significantly differing for the muscle fiber types slow-twitch-oxidative fiber (STO) and fast-twitch-oxidative fiber (FTO) as well as mitochondrial activity (succinate-dependent state 3 respiration rate). Longissimus muscle samples were obtained 24 h before slaughter and profiled using cDNA microarrays. Differential gene expression between Duroc and PiNN muscle samples were associated with protein ubiquitination, stem cell pluripotency, amyloid processing, and 3-phosphoinositide biosynthesis and degradation pathways. In addition, weighted gene co-expression network analysis within both breeds identified several co-expression modules that were associated with the proportion of different fiber types, mitochondrial respiratory activity, and ATP metabolism. In particular, Duroc results revealed strong correlations between mitochondrion-associated co-expression modules and STO (r = 0.78), fast-twitch glycolytic fiber (r = -0.98), complex I (r=0.72) and COX activity (r = 0.86). Other pathways in the protein-kinase-activity enriched module were positively correlated with STO (r=0.93), while negatively correlated with FTO (r = -0.72). In contrast to PiNN, co-expression modules enriched in macromolecule catabolic process, actin cytoskeleton, and transcription activator activity were associated with fiber types, mitochondrial respiratory activity, and metabolic enzyme activities. Our results highlight the importance of mitochondria for the oxidative capacity of porcine muscle and for breed-dependent molecular pathways in muscle cell fibers.
Keywords: microarray, muscle, muscle fiber, mitochondrial Respiratory Activity
