Impact Factor ISSN: 1449-2288
Int J Biol Sci 2017; 13(6):748-758. doi:10.7150/ijbs.18472 This issue Cite
Research Paper
Oncogenic Protein Kinase D3 Regulating Networks in Invasive Breast Cancer
1. The Key Laboratory of Developmental Genes and Human Disease, Ministry of Education, Institute of Life Science, Southeast University, Nanjing 210096, PR China;
2. Jiangsu Key Laboratory for Molecular and Medical Biotechnology, College of Life Science, Nanjing Normal University, Nanjing 210046, China;
3. Department of General Surgery, Jinling Hospital, Medical School of Nanjing University, NanJing 210002, P. R. China.
* These authors contributed equally to this work.
Abstract
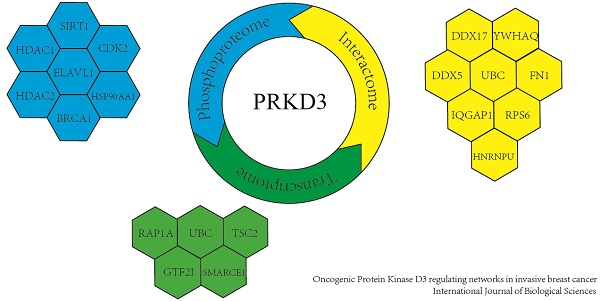
Protein Kinase D3 (PRKD3) functions as an important oncogenic driver in invasive breast cancer, which is the leading cause of women mortality. However, PRKD3 regulating network is largely unknown. In this study, we systematically explored PRKD3 regulating networks via investigating phosphoproteome, interactome and transcriptome to uncover the molecular mechanism of PRKD3 in invasive breast cancer. Using iTRAQ, 270 proteins were identified as PRKD3 regulated phosphoproteins from 4619 phosphosites matching 3666 phosphopeptides from 2016 phosphoproteins with p-value <0.005. Transcriptome analysis using affymetrix microarray identified 45 PRKD3 regulated genes, in which 20 genes were upregulated and 25 genes were downregulated with p-value <0.005 upon silencing PRKD3. Using Co-IP in combination of MS identification, 606 proteins were identified to be PRKD3 interacting proteins from 2659 peptides. Further network analysis of PRKD3 regulated phosphoproteins, interacting proteins and regulated genes, reveals 19 hub nodes, including ELAVL1, UBC and BRCA1. UBC was recognized as the most common hub node in PRKD3 regulating networks. The enriched pathway analysis reveals that PRKD3 regulates pathways contributing to multiple cancer related events, including cell cycle, migration and others. Enrichment of cell cycle and cell mobility related pathways across PRKD3 networks, explained the observations that depletion of oncogenic PRKD3 led to alternation of cell cycle and decrease of cell migration ability. Taken together, our current study provided valuable information on the roles as well as the molecular mechanisms of PRKD3 in invasive breast cancer.
Keywords: Invasive breast cancer, PRKD3, PRKD3 regulating networks.

 Global reach, higher impact
Global reach, higher impact