10
Impact Factor
ISSN: 1449-2288
Int J Biol Sci 2017; 13(11):1438-1449. doi:10.7150/ijbs.20836 This issue Cite
Research Paper
Transcriptome integration analysis in hepatocellular carcinoma reveals discordant intronic miRNA-host gene pairs in expression
1. Life Sciences Institute, Zhejiang University, Hangzhou, 310058, China;
2. University of Hawai'i Cancer Center, Honolulu, HI, 96813, USA;
3. State Key Laboratory of Molecular Oncology, Cancer Institute & Hospital, Chinese Academy of Medical Sciences and Peking Union Medical College, Beijing, 100021, China;
4. Liver Carcinogenesis Section, Laboratory of Human Carcinogenesis, National Cancer Institute, Bethesda, MD, 20892, USA;
5. Sinowell Beijing Tech Ltd, Beijing, 100045, China;
6. Liver Cancer Institute and Zhongshan Hospital, Fudan University, Shanghai, China, 200433;
7. Department of Surgery, John A. Burns School of Medicine, University of Hawai'i, Honolulu, HI, 96813, USA.
* The authors wish it to be known that, in their opinion, the first three authors should be regarded as joint First Authors.
Abstract
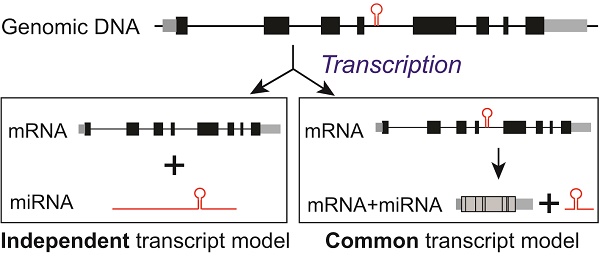
Intronic miRNAs, residing in intronic regions of host genes, are thought to be co-transcribed from their host genes and present consistent expression patterns with host genes. Recent studies reported a few intronic miRNAs with discordant expression with their host genes. We therefore aimed to understand the expression pattern of intronic miRNAs and their host genes in hepatocellular carcinoma (HCC) and reveal possible associated molecular mechanisms. Our genome wide integration analysis of miRNA and mRNA transcriptomes, in three dataset from 550 patients with HCC, found that a large amount of miRNA-host gene pairs were discordantly expressed. Consistent results were also revealed in 775 breast cancer patients. Further, most of HCC-related intronic miRNAs were predicted to have distinct upstream regulators and independent proximal promoter signals from host genes. The discordant expression of representative pairs, miR-26s/CTDSPs, was validated experimentally. We have also identified the independent transcriptional start site, promoter signal, and transcriptional factor of miR-26b from its host gene. Collectively, discordant expression of intronic miRNAs and their host genes was relatively ubiquitous and the intronic miRNA “independent transcription” may partially contribute to such a phenotype.
Keywords: microRNA, Intronic miRNA, Transcriptome, Integration analysis, Hepatocellular carcinoma.

 Global reach, higher impact
Global reach, higher impact