Impact Factor ISSN: 1449-2288
Int J Biol Sci 2018; 14(11):1571-1585. doi:10.7150/ijbs.25328 This issue Cite
Research Paper
Genome-Wide Chromatin Structure Changes During Adipogenesis and Myogenesis
1. Institute of Animal Genetics and Breeding, College of Animal Science and Technology, Sichuan Agricultural University, Chengdu 611130, China;
2. Farm Animal Genetic Resources Exploration and Innovation Key Laboratory of Sichuan Province, Sichuan Agricultural University, Chengdu 611130, China;
3. Novogene Bioinformatics Institute, Beijing 100089, China;
4. Chongqing Academy of Animal Sciences, Chongqing 402460, China;
5. Animal Breeding and Genetics Key Laboratory of Sichuan Province, Pig Science Institute, Sichuan Animal Science Academy, Chengdu 610066, China.
*These authors contributed equally to this work.
Abstract
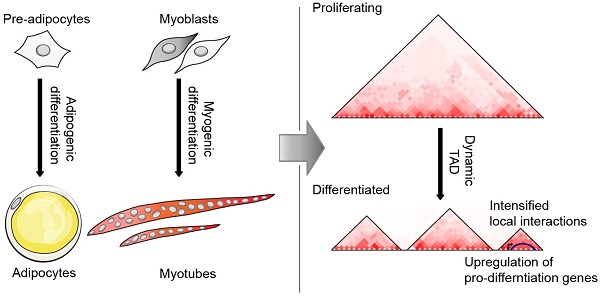
The recently developed high-throughput chromatin conformation capture (Hi-C) technology enables us to explore the spatial architecture of genomes, which is increasingly considered an important regulator of gene expression. To investigate the changes in three-dimensional (3D) chromatin structure and its mediated gene expression during adipogenesis and myogenesis, we comprehensively mapped 3D chromatin organization for four cell types (3T3-L1 pre-adipocytes, 3T3-L1-D adipocytes, C2C12 myoblasts, and C2C12-D myotubes). We demonstrate that the dynamic spatial genome architecture affected gene expression during cell differentiation. A considerable proportion (~22%) of the mouse genome underwent compartment A/B rearrangement during adipogenic and myogenic differentiation, and most (~80%) upregulated marker genes exhibited an active chromatin state with B to A switch or stable A compartment. More than half (65.4%-73.2%) of the topologically associating domains (TADs) are dynamic. The newly formed TAD and intensified local interactions in the Fabp gene cluster indicated more precise structural regulation of the expression of pro-differentiation genes during adipogenesis. About half (32.39%-59.04%) of the differential chromatin interactions (DCIs) during differentiation are promoter interactions, although these DCIs only account for a small proportion of genome-wide interactions (~9.67% in adipogenesis and ~4.24% in myogenesis). These differential promoter interactions were enriched with promoter-enhancer interactions (PEIs), which were mediated by typical adipogenic and myogenic transcription factors. Differential promoter interactions also included more differentially expressed genes than nonpromoter interactions. Our results provide a global view of dynamic chromatin interactions during adipogenesis and myogenesis and are a resource for studying long-range chromatin interactions mediating the expression of pro-differentiation genes.
Keywords: Adipogenesis, Myogenesis, Compartment A/B, Topologically associating domain, Differential chromatin interaction, Gene expression.

 Global reach, higher impact
Global reach, higher impact