Impact Factor ISSN: 1449-2288
Int J Biol Sci 2009; 5(2):205-214. doi:10.7150/ijbs.5.205 This issue Cite
Research Paper
Identifications of SUMO-1 cDNA and Its Expression Patterns in Pacific White Shrimp Litopeanaeus vannamei
1. Dept of Human Nutrition, Food and Animal Sciences, University of Hawaii at Manoa, Honolulu, HI 96822, USA.
2. Dept of Molecular Biosciences and Bioengineering, University of Hawaii at Manoa, Honolulu, HI 96822, USA.
3. Department of Biochemistry, Faculty of Science, Prince of Songkla University, Songkhla, 90112, Thailand.
4. Center for Genomics and Bioinformatics Research, Prince of Songkla University, Songkhla, 90112, Thailand.
Received 2009-1-11; Accepted 2009-2-14; Published 2009-2-19
Abstract
Small ubiquitin-like modifiers (SUMO) work in a similar way as ubiquitin to alter the biological properties of a target protein by conjugation. A shrimp SUMO cDNA named LvSUMO-1 was identified in Litopenaeus vannamei. LvSUMO-1 cDNA contains a coding sequence of 282 nucleotides with untranslated regions of 37 bp at 5'-end and 347 bp at 3'-end, respectively. The deduced 93 amino acids exhibit 83% identity with the Western Honeybee SUMO-1, and more than 65% homologies with human and mouse SUMO-1. LvSUMO-1 mRNA is expressed in most L. vannamei tissues with the highest level in hepatopancrease. The mRNA expression of LvSUMO-1 over development stages in L. Vammamei is distinguished by a low level in nauplius stage and relatively high level in postlarva stage with continuous expression until juvenile stage. The LvSUMO-1 protein and its conjugated proteins are detected in both cytoplasm and nucleus in several tissues. Interestingly, LvSUMO-1 mRNA levels are high in abdominal muscle during the premolt stage, wherein it has significant activities of protein degradation, suggesting its possible role in the regulation of shrimp muscle protein degradation.
Keywords: Invertebrate, Small ubiquitin-like modifiers, Sumoylation, RT-PCR, mRNA, Gene expression
Introduction
Small ubiquitin-like modifiers (SUMO) work in a similar way as ubiquitins to conjugate a target protein (sumoylation) to alter the properties of the modified protein even though it has only 20% identity with ubiquitin in mRNA sequence (1). Sumoylation has been indicated to regulate the stability of protein function, nuclear cytoplasm protein transport and gene transcription. Three members of the SUMO family have been described in vertebrates, SUMO-1, SUMO-2 and SUMO-3 (2). In invertebrates, such as yeast, insects and nematodes, only a single SUMO gene termed SMT3 is expressed (3), whereas in plants up to eight version of SUMO are expressed (4).
SUMO-1 has been reported with various synonyms, such as GMP1, PIC1, sentrin, UBL1 or SMT3 (5-9). The first mammalian protein shown to be sumoylated was RanGAP1, a GTPase-activating protein, which is known to play an important role in transport of proteins across the nuclear pore complex. Sumoylation facilitates the binding of RanGAP1 to nuclear pore, and changes its intracellular localization (6, 9, 10). SUMO-1 modifications of heat shock transcription factors (HSF1 and HSF2) increase their DNA binding activities while blocking the modifications by mutation decreases HSF1 transcriptional activity (11,12). Similarly, SUMO-1 modified p53 tumor suppressor by increasing its transcriptional and apoptotic response. The sumoylation sites in p53 are located in the carboxy-terminal region, which is known to regulate its DNA binding activity (3, 13,14). Another transcription factor, nuclear factor of activated T cell (NFAT), can be sumoylated on the residues K684 and K897 when treated with phorbol ester and calcium ionophore (15). In addition, several other transcription factors were reported to undergo SUMO modifications, including Ets transcription factor (16-18), androgent receptor (19-21), c-Myb(22), AP-2 (22,23) and c-Jun (Muller et al., 2000). SUMO-1 modification can also enhance the transcriptional activity of the transforming growth factor β (TGF-β) signaling proteins (24,25). One of the TGF-β signaling proteins, namely Smad4, was identified as a target protein for SUMO-1 at two evolutionarily conserved lysine residues (Lys-113 and Lys-159) in the MH1 domain. Sumoylation of Smad4 by over-expression of SUMO-1 enhanced transcriptional responses and cellular responses induced by TGF-β (24). Another TGF-β signal protein, SnoN, is also regulated by sumoylation. Loss of sumoylation in the Lys-50 site (via a Lys-to-Arg point mutation) in SnoN potently activates muscle-specific gene expression and enhances myotube formation. These results suggests that SUMO modification plays an important role in regulation of myogenic differentiation (25).
Sumoylation in invertebrate was mainly reported in Drosophila. Transcriptional repressor Tramtrack69 protein (Ttk69), which inhibits neuronal differentiation, was identified as one of the most important substrates for SUMO conjugation. SUMO and Ttk69 proteins localize together on polytene chromosome. SUMO-conjugated Ttk69 proteins are present at the sites of Ttk69 action in vivo (26). Pacific white shrimp (Litopeanaeu Vannamei) is one of the most important seafood products in the United States and Asia. Previously, abdominal muscle cDNA library of L. Vannamei was constructed by our laboratory (27). While we performed phage lift hybridization of the cDNA library by using TGF-β degenerated oligonucleotide DNA probes, a DNA sequence similar to SUMO-1 cDNA, named LvSUMO-1, was identified. We reported here LvSUMO-1 cDNA sequences and its expressions in various tissues during early development stages and different molting stages in L. vannamei. Its expression patterns in abdominal muscle over several molting stages suggest its possible role in the regulation of shrimp muscle protein degradation.
Materials and Methods
Animals, tissue sampling and RNA extraction
Cultured specific pathogens-free (SPF) nauplius, postlarvae and 3-month-old L. vannamei from a local shrimp farm in Honolulu HI were transported alive in plastic bags with 1/3 water and 2/3 compressed air to the lab. Fresh tissue samples were collected for RNA and protein extractions. The tissue samples from 3-month-old shrimp include heart (H), eyestalk (E), intestine (I), hepatopancreas (Hep), haemocyte (Cy), lymphoid (L), brain and nerve ganglion (BN), gill (G), pleopods (P), abdominal muscle (AM), swimming leg muscle (SM) and ovary (O). The tissues samples used for molt stage studies were from a previous report (28). Total RNA was isolated by TRIZOL Reagent (Invitrogen), following the method described by Yang et al (29). Briefly, 100 mg of tissue was homogenized in 1ml TRIZOL reagent using a Polytron homogenizer (Model PT 10/35, Brinkmann Instruments, Westbury, NY). Homogenized sample was incubated for 7 min at room temperature follow by chloroform (0.2 ml per ml of TRIZOL) added. The tube was vigorously shaken by hand and incubated at room temperature for 5 min. Following centrifugation at 12,000 rpm for 15 min at 4°C, the aqueous phase was transferred to a clean tube to which isopropanol was added. After 15 min incubation at room temperature, tubes were centrifuged at 4°C for 15 min at 12,000 rpm, and supernatant was withdrawn and remaining RNA pellets were washed with 70% ethanol, allowed to dry for 10-15 min, dissolved in deionized-distilled water and stored in -80°C. Total RNA concentrations were calculated by spectrophotometer.
Screening of abdominal muscle cDNA library oligonucleotide probe
A phages cDNA library was constructed with the mRNA isolated from Litopenaeus vannamei abdominal muscle of juvenile animals (28) by using the SMART cDNA library construction kit (Clontech). High-density filters corresponding to a shrimp muscle phage cDNA library was represented by two high-density filters with 10,000 plaques spotted on each membrane. A total of six degenerated oligonucleotide DNA probes designed by comparison of the c-terminal conserved region of TGF-β mRNA sequences were labeled with biotin (www.idtdna.com). Hybridization was done as described (30) in hybridization oven. After hybridization, the filters were washed in 2x SSC, 0.1% SDS 2 times for 15 min at room temperature and once for 15 min at 50˚C of 0.1x SSC, 0.1%SDS. Blots were autoradiographed for empirically optimized exposure times by using CDP-Star® Chemiluminescence reagent as substrate for streptavidin alkaline phosphatase conjugates.
Positive phage conversion
Individual positive plaques were excised from plates and incubated overnight at 4˚C in 350 ul 1x phage elution buffer (0.1M NaCl, 0.01M MgSO4, 0.035M Tris-HCl, pH7.5, 0.01% Gelatin). All plaques from single plate were taken, to avoid possible selection bias. Lambda TriplEx2 plaque elutes (150ul) were incubated into 200ul of E.coli BM25.8 overnight cultures in presence of MgCl2 (10mM final concentration) and incubated at 31˚C for 30 min followed by addition of LB medium and another incubation with shaking (225rpm) for 1 hr. Infected cell suspensions were transferred and spread onto LB ampicillin (100mg/ml) plates and incubated overnight at 31ºC.
Sequence analysis
Plasmid DNA was isolated from the conversion clones for plasmid recovery and sequencing. The searches for nucleotide, EST and protein sequence similarities were conducted with BLAST algorithm at the National Center for Biotechnology Information (NCBI) (http://www.ncbi.nlm.nih.gov/BLAST/). The LvSUMO-1 deduced amino acid sequence was analyzed with the NCBI Protein Sequence Analysis software. Multiple alignment of the LvSUMO-1 was performed with the ClustalX Multiple Alignment program (http://www.ncbi.nlm.nih.gov) and Multiple Alignment show by GENEDOC, version 2.6.001 (31). SUMO modification site was determined by SUMOplotTM prediction (http://abgent.com/tools)
Tissue specificity of SUMO-1 gene expressions in L. vannamei
Total RNA samples were treated for possible genomic DNA contamination with DNase I (Invitrogent) in a 10 µl reaction containing 1.5 µg of RNA, 1 µl 10x buffer (200 mM Tris-HCl, pH 8.4, 20 mM MgCl2, 500mM KCl), 1.5 µl DNase I (1.5 U) at 37 ºC for 15 min, followed by inactivation with 25 mM EDTA at 65ºC for 10 min. Reverse transcription (RT) was performed using 1.5 µg RNA, 0.5 µg Oligo (dT) 18 mers, 1.0 mM dNTP, 4 µl of 5x reaction buffer and 40 U of M-MuLV reverse transcriptase (Fermentas) at 37ºC for 60 min. The reaction was stopped by heating at 70ºC for 10 min. For PCR amplification, two oligonucleotide primers, B95_F: 5'-ATGTC TGATAACGCTGACGCCAAGCCA-3'and B95_R: 5'-TCAATGGCCGCCGGTC TGCTCCTGGTA-3'were designed based on the nucleotide sequence corresponding to the nucleotide positions 1-27 and 256-282 of L. vannamei SUMO-1 cDNA, respectively. A set of control, elongation factor (EF-1), EF-1_F: 5'- GAAGATCGACAGGCGTACTGGTAA 3'and EF-1_R: 5'- GGCCTTGGTAGTTTCGCCAGATTGAT -3', served as a control (28) was used during the RT-PCR amplification. PCR reactions were conducted for 40 cycles with denature at 94ºC for 30 sec, annealing at 60ºC for 30 sec, and extension at 72ºC for 30 sec in a reaction buffer consisting of 10 mM Tris (pH 8.3), 50 mM KCl, 200 M dNTPs, 0.2 µM of each primer, and 2.5 U of Taq Polymerase (New England Biolab) in a total volume of 50 µl. To visualize the amplified products, PCR reaction was electrophoresed through 1.5% agarose gel containing 0.025% ethidium bromide.
Cytoplasmic and nuclear protein extractions
150 mg tissue was homogenized in 1 ml buffer A (10 mM HEPES, pH 7.9, 10 mM KCl, 0.1 mM EDTA) contained 1 µl of each protease inhibitor (1mM DTT, 0.5mM PMSF, 10µg/µl leupeptin) and 40µl of 10% NP40, incubated 10 min at room temperature. Cell debases and un-break cell was separated by centrifugation at 4ºC at 1,000 g for 3 min. Follow by centrifugation at 4ºC at 15,000x g for 3 min, nuclear was packed in the bottom of microcentrifuge tube. Supernatant was separated to new tube as the cytosolic protein fraction. 150 µl of buffer B (20mM HEPES, pH 7.9, 0.4 M NaCl, 1mM EDTA, 10% glycerol) was added into nuclear pellet. Pellet was resuspended by pipeting up and down and shacked vigorously at 4 ºC for 2 hr. Supernatant represented nuclear protein fraction was separated by centrifugation at 4ºC at 15,000x g for 5 min. Protein concentration was tested by using BCA method (Pierce).
Western blot analysis
The cytosolic and nuclear protein were subjected to electrophoresis using 15% SDS-PAGE, following by Western blotting according to the Mini Tran-Blot electrophoretic Transfer cell system (BioRad). Proteins were transferred onto PVDF membrane (Bio Rad) in electoblotting buffer (25mM Tris-HCl, 190mM Glycine, 20% methanol) at a constant current of 2.5mA for 2 h. The membrane was immersed in blocking buffer (5% nonfat dry milk, 1X Tris buffer saline (TBS), pH 7.4, 0.1% Tween 20) at room temperature for 1 hr followed by incubation with rabbit polyclonal antibody to human SUMO-1 (c-terminal) (Abgent) in antibody dilution buffer (1% nonfat dry milk, 1x Tris buffer saline(TBS), pH 7.4, 0.1% Tween 20) at a concentration of 1:500 at 4 C overnight. Subsequently, the membrane was incubated in HRP-conjugated anti-rabbit IgG (Santa Cruz) for 2 h at a concentration of 1:1000 and developed with Visualizer EC western blot detection kit (Millipore).
Results and Discussion
L. vannamei SUMO-1 cDNA and amino acid sequences homolog
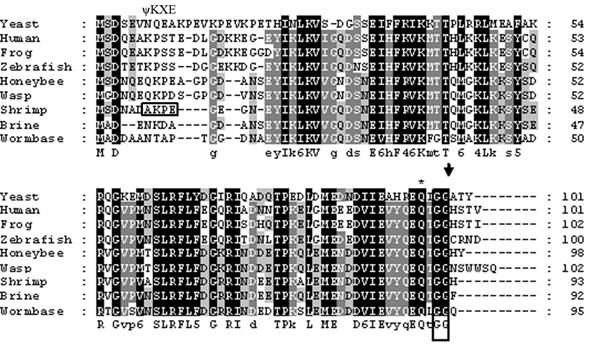
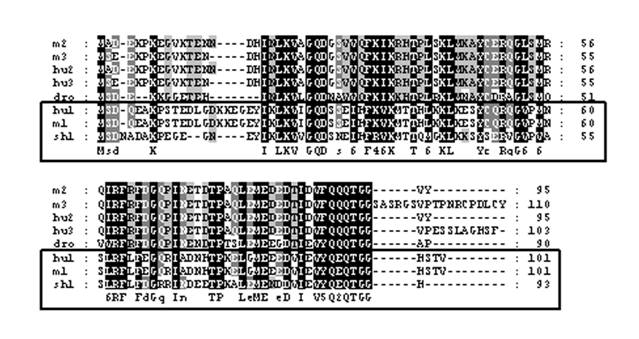
The cDNA library from abdominal muscle of juvenile animals was successfully constructed by a PCR-base method. Library titrations were 5.0 x 106 pfu/ml (27). Phage lift hybridization method was used for screening TGF-β family and other related proteins by a mixture of six biotin-labeled degenerated oligonucleotide DNA probes. A total of 112 positive clones were isolated and sequenced. One of the cDNA sequences (664 nucleotides) showed a high homology to SUMO-1, named as LvSUMO-1. The cDNA sequence contains a protein-coding sequence of 282 nucleotides and 5' (37 nucleotides) and 3' (347 nucleotides) untranslated regions (Fig. 1). The cDNA sequence of LvSUMO-1 was submitted to NCBI Genbank (Accession number: FJ751760). The deduced sequence of 93 amino acids from LvSUMO-1 cDNA has a molecular mass of 10.23 kDa and exhibits 83% identity and 89% similarity with the Western Honeybee (Apis mellifera) SUMO homolog family member Smo-1 (Fig. 2) and 65% identity and 79% similarity with human and mouse SUMO-1, respectively. The deduced amino acid sequences of SUMO proteins from Human, Xenopus, Zebrafish, L. vannamei, Brine shrimp, Honeybee, Nematode and yeast are aligned in Fig. 2. All SUMO proteins from yeast to human share the conserved the C-terminal diglycine cleavage/attachment, which is the critical domain for SUMO protein maturation and conjugation (32). The results from sequence alignment also revealed that SUMO-1 is highly evolutionarily conserved from human SUMO-1 to yeast Smt3 proteins, particularly the double glycine site. A prominent difference between the SUMO proteins and ubiquitin is the presence of a highly variable N-terminal extension in SUMO proteins, wherein it includes several charged amino acids such as glycines and prolines (33). The function of this N-terminal extension in SUMOs is not presently known. The homology of LvSUMO-1 and SMT3 of Drosophila melanogaster is only 45% (Fig. 3). Protein alignment of human SUMO1/2/3, mouse SUMO1/2/3, SMT3 Drosophila and LvSUMO-1 suggests that LvSUMO-1 has a high homology to human and mouse SUMO-1 (Fig. 3). The SUMOplot prediction from http://abgent.com/tools showed the LvSUMO-1 contains the AKPE motive site, which is the sumoylation consensus ψKXE motif where ψ is a large hydrophobic residue and X is any amino acid in the N-terminal (34). Therefore, the name of LvSUMO-1 was assigned to this cDNA sequence.
Nucleotide and deduced amino acid sequences of L. vannamei SUMO-1 (LvSUMO-1) cDNA. Amino acids are indicated as single capital letters under each triplet codon of the nucleotide sequence. The start codon is underlined and an asterisk (*) indicates the stop codon. Sumo modification prediction site on the polypeptide chain was showed in capital letters (AKPE). The polyadenylation signal (ATTAAA) is typed in capital letters underlined.

Amino acid sequence alignment among different SUMO-1 proteins. The amino acid sequences of Human (Accession no. AAH53528), Xenopus (Accession no. Q6DEP7), Zebrafish (Accession no. Q7SZR5), Brine shrimp: (Accession no. ABQ41279), Honeybee (Accession no. XP_392826), Wesp (Accession no. XP_001607301), Nematode (Accession no. CAP33470) and yeast (Accession no. NP_010798) are aligned. C-terminal diglycine motif, a site used for propeptide processing, is boxed, and the predicted cleavage site is marked by an arrowhead. A glutamine residue known to be important for the interaction of human SUMO-1 with protease Senp2 is asterisked.
Alignment of the LvSUMO-1 (sh1) with those of human SUMO-1 (hu1: GenBank Accession No. NP_003343), human SUMO-2(hu2: GenBank Accession No. P61956), human SUMO-3(hu: GenBank Accession No. NP_008867), mouse SUMO-1(m1: GenBank Accession No. NP_033486), mouse SUMO-2(m2: GenBank Accession No. NP_579932), mouse SUMO-3(m3: GenBank Accession No. Q9Z172) and Drosophilar smt3(dro: GenBank Accession No. NP_477411). Identical residues among all sequences are shown in black background and gaps are shown by hyphens. vanSUMO-1 grouped with human and mouse SUMO-1, is boxed.
Specific tissue expression of LvSUMO-1 mRNA in L. vannamei
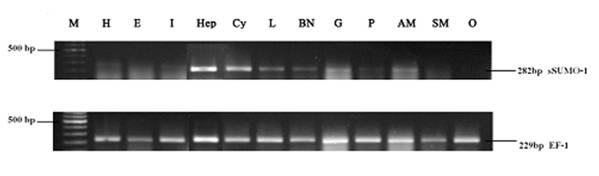
Expressions of LvSUMO-1 in different tissues are shown in Fig. 4. LvSUMO-1 mRNA expressions were detected in hepatopancrease, haemocytes, gill and abdominal muscle with the highest level in hepatopancrease and low levels in heart, intestine, lymphoid, brain and nerve and pleopods. The LvSUMO-1 mRNA was not detected in eyestalk (E), swimming leg muscle (SM) and ovary (O). SUMO-1 mRNA expressions by Northern blot hybridization had been reported in most mouse tissues, including heart, brain, spleen, lung, liver, skeletal muscle, kidney and testis (35). Similarly, the endogenous SUMO-1 mRNA expression was found in human leukocyte, colon, small intestine, ovary, testis, prostate, thymus and spleen and testis with the testis exhibiting the highest level (8). These data reveal that LvSUMO-1 is ubiquitously expressed in many tissues in L Vannamei.
Tissue-specific expressions of LvSUMO-1 proteins
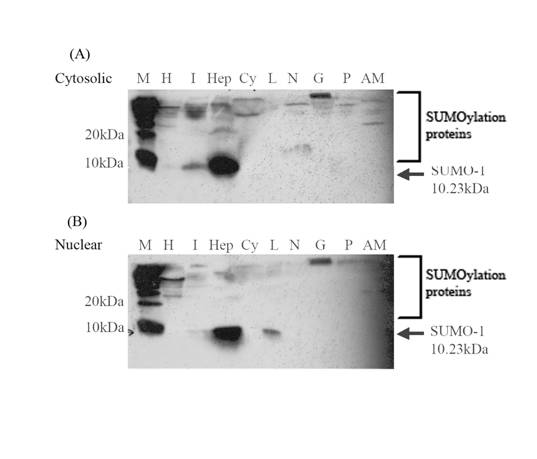
To confirm the mRNA expressions, the LvSUMO-1 proteins expression was detected by using Western blot analysis (Fig. 5). The antibody to SUMO-1 is generated from rabbits immunized with a KLH conjugated synthetic peptide selected from the C-terminal region of human SUMO1. The cytoplasmic and nuclear proteins from different tissues were extracted and tested for LvSUMO-1 protein. In the cytoplasmic protein fraction, LvSUMO-1 was detected at the position of 10 KD in hepatopancreas, heart and intestine tissue (Fig. 5a). The expression of LvSUMO-1 protein in nuclear fraction was also detected in hepatopancreas, intestine and lymphoid (Fig.5b). In both cytoplasmic and nuclear fractions, LvSUMO-1 protein showed a high level of expression in hepatopancreas. The protein blot by this antibody also showed LvSUMO-1 conjugated protein bands since their molecular weight are more 10 KD. The free form of LvSUMO-1 protein was not detected in the abdominal muscle tissue while several conjugated protein signals with higher molecular weights were detected in the tissue (Fig.4).
Tissue specificity of LvSUMO-1 gene expression by RT-PCR. LvSUMO was highly expressed in hepatopancrease(Hep), haemocytes(Cy), Gill(G) and abdominal muscle(AM) but weakly in heart(H), intestine(I), lymphoid(L), brain and nerve(BN) and pleopods(P). Also the expression were undetected in eyestalk (E), swimming leg muscle(SM) and ovary(O) in this amount of RNA template. EF-1 gene was used as the internal control.
Western Blot analysis of shrimp SUMO-1 transcripts from various tissues compost of heart(H), intestine(I), hepatopancrease(Hep), haemocytes(Cy), lymphoid(L), brain and nerve(BN), Gill(G), pleopods(P) and abdominal muscle(AM) in cytosolic and nuclear protein fraction(M=protein marker). Protein expression level of shrimp SUMO-1 was very high expressed in hepatopancreas both in cytosilic and nuclear protein fraction. In addition, the expression in were found in heart and intestine of cytoplasmic protein fraction and in intestine and lymphoid of nuclear protein fraction.
Most of the SUMO-1 protein was previously shown to localized on nuclear membrane (36, 37). Unlike SUMO-2/3 that has an apparent free form, human SUMO-1 has no free form to be detected (33). The LvSUMO-1 protein was detected in free form in intestine and hepatopancreas of cytosolic fraction and in hepatopancrease and lymphoid of nuclear fraction, which are more like the human SUMO-2/3. Many of sumolylated proteins have been found. For example, SUMO-1 is believed to form an isopeptide bond between the carboxyl terminus of SUMO-1 and a lysine side chain(s) of the target protein (2, 9, 32). All SUMO forms are initially processed as inactive precursors, then mature by a carboxy-terminal proteolytic cleavage event, which yields the mature modifiers with exposed carboxy-terminal glycine residues. These residues are required for the formation of and isopeptide bond between the carboxyl terminus of SUMO with an ε-amino group of lysine residue of a target protein. The processing reaction is catalyzed by a group of cysteine proteases, termed ULPs (ubiquitin-like protein-processing enzyme) or SUMO-specific proteases (3). The mechanism of SUMO conjugation has been proposed by Ronald T. Hay (38), which consists of three enzymes cascade involving a Ubl activating enzyme(E1), a Ubl conjugating enzyme(E2), and typically a Ubl protein ligase(E3). To initiate the SUMO modification reaction, activating enzyme catalyzes the formation of adenylated SUMO in which the C-terminal carboxyl group of SUMO is covalently linked to AMP. SUMO was transesterified to conjugating enzyme. The conjugating enzyme called Ubc9 can directly recognize substrate protein that contain the SUMO modification consensus motif, ψKXE motif (39). In the case of LvSUMO-1 conjugation, the polymeric SUMO chain may happen because it contains SUMO modification consensus motif. However several key enzymes such as ligase (E3) in the pathways has been reported in shrimp or crustacean yet.
LvSUMO-1 mRNA expression and shrimp muscle development
The LvSUMO-1 was originally identified from shrimp muscle cDNA library. Its mRNA expression can be easily detected by RT-PCR although only sumoylated proteins were detected in the abdominal muscle tissue by Western blotting. Therefore, we performed RT-PCR to study the function of LvSUMO-1 expression in shrimp muscle tissue instead of western blot analysis. The developmental and life cycle of L. vannamei can be classified to seven stages, including egg, nauplius, protozoea, mysis, postlarva and juvenile and adult (40). Muscle formation in dendrobranchiate shrimp was reported by Kiernan and Hertzler (41). In the dendrobranchiate malacostracan Sicyonia ingentis, F-actin staining was first detected in limb setae at 12 h. Staining of F-actin was detected in the trunk at 15 h and grew into naupliar limbs. Sarcomeres were detected at 19 h, identifying the structures as extrinsic limp muscles. The extrinsic limb muscles enlarged but retain their general pattern during the later nauplius stages. Longitudinal trunk muscle and circumferential visceral muscle developed in the post-naupliar region during nauplius instars 4 and 5, at the time when the gut also formed. Among decapod malacostracan larvae, abdominal muscle development was described in zuea and mysis larvae of the caridean shrimp Crangon vulgaris (42). And the adult muscle of dendrobranchiate shrimp Penaeus (Litopenaeus) setiferus have been carefully examined (43).
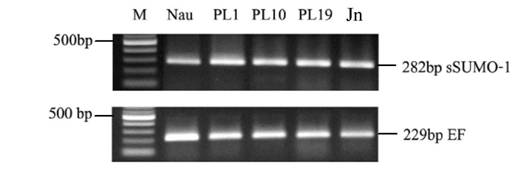
In this study, we sampled shrimp tissue over several developmental stages, mainly nauplius as the larva stage, postlarva and juvenile to study LvSUMO-1 expression. The LvSUMO-1 mRNA expression was detected in a low level in nauplius (Nau) stage and then followed by increased up-regulation in postlarva(PL) stage 1, 10 and 19. The expression was also presented in juvenile (Jn) (Fig.6). These results indicate that LvSUMO-1 was up-regulated during the muscle formation. Other studies have showed that sumoylation of SnoN by SUMO-1 plays a role in TGF-β signaling, which is known to regulate myogenesis. The over-expression of c-Ski/SnoN induces skeletal muscle differentiation (44, 45). Loss of sumoylation activates muscle-specific gene expression and enhanced myotube formation (25). Most regulatory genes related to shrimp muscle formation and development are not known. This study suggests possible target protein by LvSUMO-1 modification may play a role in the regulation of shrimp myogenic differentiation and muscle formation.
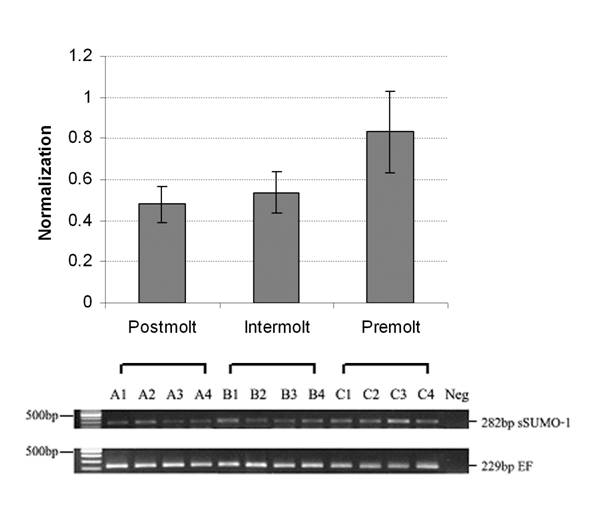
Growth in shrimps, like other crustacean, is intermittent, and muscle mass accumulation occurs through molting-shedding of exoskeleton (46, 47). Muscle morphology and biochemical changes of L. vannamei during molting process have been studied by Yang and his research group (28, 48). Muscle structural α-actin and cytoskeleton β-actin were increased during intermolt and premolt stage, suggesting high muscle growth during these stages. However, the mRNA encoding for ubiquitin and Hsp70 did not increase significantly in premolt stages. In this study, three molting stages, including postmolt, intermolt and premolt, were used for RNA extraction. The LvSUMO-1 transcript was detected in the abdominal muscle tissue in all molting stages tested. The LvSUMO-1 mRNA expression was high at premolt (C) stage, and low during postmolt stage (A) and intermolt stages (B) (Fig. 7). The LvSUMO-1 expression in intermolt stage is much lower than in postmolt stage. As the overall muscle protein synthesis is higher in intermolt than in postmolt (46), the result may indicate that LvSUMO-1 play an important role in the regulation of muscle protein synthesis and degradation during the molt cycle. Further comparisons of LvSUMO-1 and ubiquitin during molting stage are needed to clarify their expression patterns in relation to protein accumulation during shrimp molt cycle.
Differentiation stage of shrimp SUMO-1 gene expression by RT-PCR (Nau=nauplius, PL1= postlarvae 1 day, PL10= postlarvae 10 days, PL19= postlarvae 19 day and Adu= adult). The expression level is up regulated in all postlavae stated and adult.
Shrimp SUMO-1 gene expression in different molting stage by RT-PCR. (A= postmolt, B=intermolt, C=premolt). SUMO-1 mRNA level is low expressed between postmolt and intermolt but high expressed in premolt.
Conclusions
LvSUMO-1 cDNA contains a protein-coding sequence of 282 nucleotides and 37 untranslated nucleotides at 5' and 347 untranslated nucleotides at 3' ends. The deduced 93 amino acids was found to exhibit 83% identity with the Western Honeybee SUMO-1 and 65% identity with human and mouse SUMO-1. The homology of LvSUMO-1 and of Drosophila melanogaster SUMO-1 (SMT3) is only 45%. LvSUMO-1 has a sumoylation motif that is conserved from mouse to human. This is the first report of SUMO-1 in crustacean species. The expression of LvSUMO-1 mRNA was identified in most shrimp tissues. LvSUMO-1 protein was found in both cytoplasm and nucleus while mostly detectable proteins were located in cytoplasm. By using SUMO-1 antibody detection, the free form of LvSUMO-1 protein was detected mostly in hepatopancreas, cytosolic and lymphoid. Several SUMO-1 conjugated proteins were detected in abdominal muscle tissue. LvSUMO-1 level also showed a progressive increase over the early developmental stages. The expression patterns in abdominal muscle during the molt cycle indicate that LvSUMO-1 may be involved in the regulation of muscle protein synthesis and degradation in shrimp.
Acknowledgements
This work was supported by the USDA-CSREES-TSTAR programs (Award #2008-34135-19343) and National Research Council of Thailand. The Royal Golden Jubilee Graduate Program was awarded to Yanisa Laoong-u-thai (PHD/0250/2546).
Conflict of Interests
The authors have declared that they have no conflict of interest.
References
1. Matic I, Macek B, Hilger M, Walther TC, Mann M. Phosphorylation of SUMO-1 Occurs in Vivo and Is Conserved through Evolution. J Proteome Res. 2008;7:4050-57
2. Kamitani T, Nguyen HP, Yeh ET. Preferential modification of nuclear proteins by a novel ubiquitin-like molecule. J Biol Chem. 1997;272:14001-4
3. Muller S, Hoege C, Pyrowolakis G, Jentsch S. Sumo, ubiquitin's mysterious cousin. Nat Rev Mol Cell Bio. 2001;2:202-210
4. Kurepa J, Walker JM, Smalle J, Gosink MM, Davis SJ, Durham TL, Sung DY, Vierstra RD. The small ubiquitin-like modifier (SUMO) protein modification system in Arabidopsis. Accumulation of SUMO1 and -2 conjugates is increased by stress. J Biol Chem. 2003;278:6862-72
5. Boddy MN, Howe K, Etkin LD, Solomon E, Freemont PS. PIC 1, a novel ubiquitin-like protein which interacts with the PML component of a multiprotein complex that is disrupted in acute promyelocytic leukaemia. Oncogene. 1996;13:971-82
6. Matunis MJ, Coutavas E, Blobel G. A novel ubiquitin-like modification modulates the partitioning of the Ran-GTPase-activating protein RanGAP1 between the cytosol and the nuclear pore complex. J Cell Biol. 1996;135:1457-70
7. Okura T, Gong L, Kamitani T, Wada T, Okura I, Wei CF, Chang HM, Yeh ET. Protection against Fas/APO-1- and tumor necrosis factor-mediated cell death by a novel protein, sentrin. J Immunol. 1996;157:4277-81
8. Shen Z, Pardington-Purtymun PE, Comeaux JC, Moyzis RK, Chen DJ. UBL1, a human ubiquitin-like protein associating with human RAD51/RAD52 proteins. Genomics. 1996;36:271-9
9. Mahajan R, Gerace L, Melchior F. Molecular characterization of the SUMO-1 modification of RanGAP1 and its role in nuclear envelope association. J Cell Biol. 1998;140:259-70
10. Bischoff FR, Krebber H, Kempf T, Hermes I, Ponstingl H. Human Rangtpase-Activating Protein Rangap1 Is a Homolog of Yeast Rna1p Involved in Messenger-Rna Processing and Transport. Proc Natl Acad Sci U S A. 1995;92:1749-53
11. Goodson ML, Hong Y, Rogers R, Matunis MJ, Park-Sarge OK, Sarge KD. Sumo-1 modification regulates the DNA binding activity of heat shock transcription factor 2, a promyelocytic leukemia nuclear body associated transcription factor. J Biol Chem. 2001;276:18513-8
12. Hong Y, Rogers R, Matunis MJ, Mayhew CN, Goodson ML, Park-Sarge OK, Sarge KD. Regulation of heat shock transcription factor 1 by stress-induced SUMO-1 modification. J Biol Chem. 2001;276:40263-7
13. Gostissa M, Hengstermann A, Fogal V, Sandy P, Schwarz SE, Scheffner M, Del Sal G. Activation of p53 by conjugation to the ubiquitin-like protein SUMO-1. EMBO J. 1999;18:6462-71
14. Rodriguez MS, Desterro JM, Lain S, Midgley CA, Lane DP, Hay RT. SUMO-1 modification activates the transcriptional response of p53. Embo J. 1999;18:6455-61
15. Terui Y, Saad N, Jia S, McKeon F, Yuan J. Dual role of sumoylation in the nuclear localization and transcriptional activation of NFAT1. J Biol Chem. 2004;279:28257-65
16. Yang SH, Jaffray E, Hay RT, Sharrocks AD. Dynamic interplay of the SUMO and ERK pathways in regulating Elk-1 transcriptional activity. Mol Cell. 2003;12:63-74
17. Yang SH, Jaffray E, Senthinathan B, Hay RT, Sharrocks AD. SUMO and transcriptional repression: dynamic interactions between the MAP kinase and SUMO pathways. Cell Cycle. 2003;2:528-30
18. Degerny C, Monte D, Beaudoin C, Jaffray E, Portois L, Hay RT, de Launoit Y, Baert JL. SUMO modification of the Ets-related transcription factor ERM inhibits its transcriptional activity. J Biol Chem. 2005;280:24330-8
19. Poukka H, Karvonen U, Janne OA, Palvimo JJ. Covalent modification of the androgen receptor by small ubiquitin-like modifier 1 (SUMO-1). Proc Natl Acad Sci U S A. 2000;97:14145-50
20. Callewaert L, Verrijdt G, Haelens A, Claessens F. Differential effect of small ubiquitin-like modifier (SUMO)-ylation of the androgen receptor in the control of cooperativity on selective versus canonical response elements. Mol Endocrinol. 2004;18:1438-49
21. Bies J, Markus J, Wolff L. Covalent attachment of the SUMO-1 protein to the negative regulatory domain of the c-Myb transcription factor modifies its stability and transactivation capacity. J Biol Chem. 2002;277:8999-9009
22. Eloranta JJ, Hurst HC. Transcription factor AP-2 interacts with the SUMO-conjugating enzyme UBC9 and is sumolated in vivo. J Biol Chem. 2002;277:30798-804
23. Muller S, Berger M, Lehembre F, Seeler JS, Haupt Y, Dejean A. c-Jun and p53 activity is modulated by SUMO-1 modification. J Biol Chem. 2000;275:13321-9
24. Liang M, Melchior F, Feng XH, Lin X. Regulation of Smad4 sumoylation and transforming growth factor-beta signaling by protein inhibitor of activated STAT1. J Biol Chem. 2004;279:22857-65
25. Wrighton KH, Liang M, Bryan B, Luo K, Liu M, Feng XH, Lin X. Transforming growth factor-beta-independent regulation of myogenesis by SnoN sumoylation. J Biol Chem. 2007;282:6517-24
26. Lehembre F, Badenhorst P, Muller S, Travers A, Schweisguth F, Dejean A. Covalent modification of the transcriptional repressor tramtrack by the ubiquitin-related protein Smt3 in Drosophila flies. Mol Cell Biol. 2000;20:1072-82
27. Cesar JRO, Zhao B, Yang J. Analysis of expressed sequenced tags from abdominal muscle cDNA library of the pacific white shrimp Litopenaeus vannamei. Animal. 2008;2:1377-83
28. Cesar JRO, Yang J. Expression patterns of ubiquitin, heat shock protein 70, alpha-actin and beta-actin over the molt cycle in the abdominal muscle of marine shrimp Litopenaeus vannamei. Mol Reprod Dev. 2007;74:554-59
29. Yang J, Zhao B, Baracos VE, Kennelly JJ. Effects of bovine somatotropin on beta-casein mRNA levels in mammary tissue of lactating cows. J Dairy Sci. 2005;88:2806-12
30. Deering RE, Arakawa CK, Oshima KH, Ohara PJ, Landolt ML, Winton JR. Development of a Biotinylated DNA Probe for Detection and Identification of Infectious Hematopoietic Necrosis Virus. Dis Aquat Org. 1991;11:57-65
31. Nicholas KB, Nicholas HBJr, Deerfield DWII. GeneDoc: Analysis and Visualization of Genetic Variation. EMBNEW NEWS. 1997;4:14
32. Johnson ES, Blobel G. Ubc9p is the conjugating enzyme for the ubiquitin-like protein Smt3p. J Biol Chem. 1997;272:26799-802
33. Su HL, Li SS. Molecular features of human ubiquitin-like SUMO genes and their encoded proteins. Gene. 2002;296:65-73
34. Matic I, van Hagen M, Schimmel J, Macek B, Ogg SC, Tatham MH, Hay RT, Lamond AI, Mann M, Vertegaal AC. In vivo identification of human small ubiquitin-like modifier polymerization sites by high accuracy mass spectrometry and an in vitro to in vivo strategy. Mol Cell Proteomics. 2008;7:132-44
35. Howe K, Williamson J, Boddy N, Sheer D, Freemont P, Solomon E. The ubiquitin-homology gene PIC1: characterization of mouse (Pic1) and human (UBL1) genes and pseudogenes. Genomics. 1998;47:92-100
36. Kamitani T, Kito K, Nguyen HP, Fukuda-Kamitani T, Yeh ETH. Characterization of a second member of the sentrin family of ubiquitin-like proteins. J Biol Chem. 1998;273:11349-353
37. Saitoh H, Hinchey J. Functional heterogeneity of small ubiquitin-related protein modifiers SUMO-1 versus SUMO-2/3. J Biol Chem. 2000;275:6252-8
38. Hay RT. SUMO: A history of modification. Molecular Cell. 2005;18:1-12
39. Rodriguez MS, Dargemont C, Hay RT. SUMO-1 conjugation in vivo requires both a consensus modification motif and nuclear targeting. J Biol Chem. 2001;276:12654-59
40. Wickens JF. Prawn biology and culture. Oceanogr Mar Biol Ann Rev. 1976;14:435-507
41. Kiernan DA, Hertzler PL. Muscle development in dendrobranchiate shrimp, with comparison with Artemia. Evol & Dev. 2006;8:537-549
42. Daniel RJ. The abdominal muscular systems of the zoea and mysis stages of shrimp (Crangon vulgaris Fabr) and their bearing on phylogeny. Proc Trans Liverpool Biol Soc. 1930;44:95-109
43. Young JH. Morphology of the white shrimp Penaeus setiferus. United states Department of the Interior, Fish and Wildlife Service. Fishery Bull. 1959;59:1-169
44. Colmenares C, Stavnezer E. The ski oncogene induces muscle differentiation in quail embryo cells. Cell. 1989;59:293-303
45. Boyer PL, Colmenares C, Stavnezer E, Hughes SH. Sequence and biological activity of chicken snoN cDNA clones. Oncogene. 1993;8:457-66
46. Mykles DL, Skinner DM. The role of calcium-dependent proteinase in molt-induced claw muscle atrophy. Prog Clin Biol Res. 1985;180:141-50
47. el Haj AJ, Harrison P, Whiteley NM. Regulation of muscle gene expression in Crustacea over the moult cycle. Symp Soc Exp Biol. 1992;46:151-65
48. Cesar JRO, Zhao B, Malcha S, Ako H, Yang J. Morphological and biochemical changes in the muscle of the marine shrimp Litopenaeus vannamei during the molt cycle. Aquaculture. 2005;261:688-94
Author contact
![]() Correspondence to: Jinzeng Yang, Ph.D. Dept of Human Nutrition, Food and Animal Sciences, University of Hawaii at Manoa, Honolulu, HI 96822, USA. E-mail: jinzengedu. Amornrat Phongdara, Ph.D. Center for Genomics and Bioinformatics Research, Faculty of Science, Prince of Songkla University, Hat-Yai, Songkhla 90112, Thailand. E-mail: pamornracom
Correspondence to: Jinzeng Yang, Ph.D. Dept of Human Nutrition, Food and Animal Sciences, University of Hawaii at Manoa, Honolulu, HI 96822, USA. E-mail: jinzengedu. Amornrat Phongdara, Ph.D. Center for Genomics and Bioinformatics Research, Faculty of Science, Prince of Songkla University, Hat-Yai, Songkhla 90112, Thailand. E-mail: pamornracom

 Global reach, higher impact
Global reach, higher impact