Impact Factor ISSN: 1449-2288
Int J Biol Sci 2015; 11(4):448-455. doi:10.7150/ijbs.11164 This issue Cite
Review
Signal Transducer and Activator of Transcription 4 in Liver Diseases
1. Department of Infectious Diseases, Peking University First Hospital, Beijing 100034;
2. Department of Oncology, the First Affiliated Hospital of Anhui Medical University, Hefei 230032;
3. Institute of Hypoxic Disease, School of Basic Medical Sciences, Capital Medical University, Beijing 100069
4. Department of Physiology and Pathophysiology, School of Basic Medical Sciences, Capital Medical University, Beijing 100069, China.
Received 2014-11-24; Accepted 2015-1-24; Published 2015-2-27
Abstract
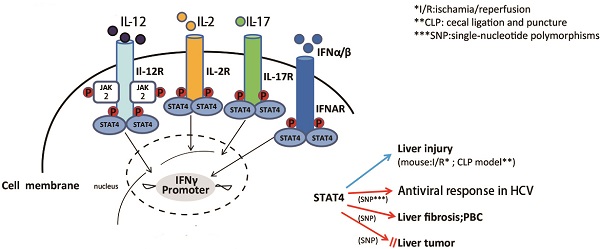
STAT4 is a member of the signal transducer and activator of transcription (STAT) family of molecules that localizes to the cytoplasm. STAT4 regulates various genes expression as a transcription factor after it is phosphorylated, dimerizes and translocates to the nucleus. STAT4 activation is detected virtually in the liver of several mouse models of liver injury, as well as the human liver of chronic liver diseases. STAT4 gene polymorphism has been shown to be associated with the antiviral response in chronic hepatitis C and drug-induced liver injury (DILI), primary biliary cirrhosis (PBC), HCV-associated liver fibrosis and in hepatocellular carcinoma (HCC). However, the roles of STAT4 in the pathogeneses of liver diseases are still not understood entirely. This review summarizes the recent advances on the functional roles of STAT4 and its related cytokines in liver diseases, especially in regulating hepatic anti-viral responses, inflammation, proliferation, apoptosis and tumorigenesis. Targeting STAT4 signaling pathway might be a promising strategy in developing therapeutic approaches for treating hepatitis in order to prevent further injury like cirrhosis and liver cancer.
Keywords: STAT4, cytokines, liver injury, chronic hepatitis, liver fibrosis, hepatocellular carcinoma.
Introduction
The Janus Kinase-Signal Transducers and Activators of Transcription (JAK-STAT) pathways have already been clearly shown to play critical roles in the immune, neuronal, hematopoietic and hepatic systems. Among these complex pathways, STAT4 is an important transcription factor that is crucial for the differentiation of Th1 cells in promoting cellular-mediate immune response (1). Generally, it has been accepted that STAT4 expression is restricted to myeloid cells (especially in activated monocytes, macrophages, and dendritic cells (2)), thymus, and testes. In contrast, the expression of STAT4 in resting human T cells is very low. Diverse cytokines can activate STAT4, including interleukin (IL)-12 (3-5), interferon (IFN)-α/β (6-8), IL-2 (9) and IL-17 (10) produced by T cell, natural killer (NK) cell and several other types of immune cells. In addition, growth arrest and DNA-damage-inducible protein GADD45-β/γ can also activate STAT4 via MKK6/p38 pathway (11). After activation, STAT4 enters the nucleus, binds to the STAT4 element and activates its target genes. Among these target genes, interferon γ(IFN-γ) is the most characterized one. Although it has been extensively investigated in immune system (12-19) and cancer (20), the roles of STAT4 in the pathogenesis of liver diseases are largely unknown. This review summarizes the recent advances on the functions of STAT4 and its related cytokines involved in liver injury, hepatitis, fibrosis and cancer (Figures 1-3).
Functions of IL-12-dependent STAT4. IL-12, synthesized predominantly by dendritic cells, macrophages and human B-lymphoblastoid cells, as the major upstream cytokine of STAT4, that can activate STAT4 via stimulating target cells (NK cell, NKT celland T cell) to release IFN-γ. IFN-γ plays a crucial role on cell proliferation, NK cell cytotoxicity, Th1 differentiation, liver fibrosis, liver tumor, viral replication and anti-viral therapy response as well as inhibiting Fas/FasL mediated apoptosis.
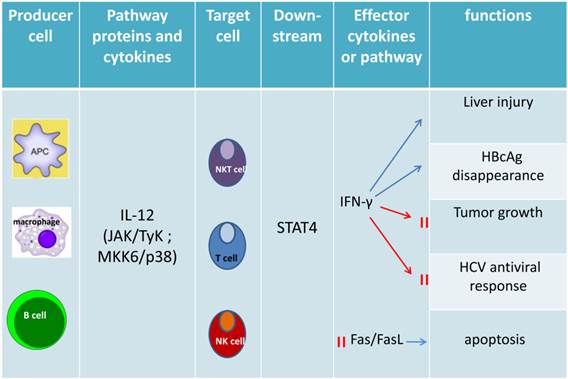
The mainly upstream genes and target genes of STAT4. IL-12, as a major upstream cytokine, stimulate T cell, NK cell and NKT cell to release target cytokines or activated some receptors, transcription factors or proteinsby activated STAT4 to regulate Th1 immune response, autophagic-associated necrotic hepatic cell death, histone acetylation or positive feedback loop. IFN-α and IFN-β both play a crucial role on Th1 cell immunity and cell proliferation by activated STAT4 via different target genes respectively. IL-2 and IL-17 also can be upstream genes that activate STAT4.
A complex role of IL-12, IFN-γ and STAT4 in pathogenesis of liver injury, liver fibrosis and liver tumor. In summary, studies from animal models and human in the recent decades have suggested that STAT4 and related cytokines, such as IL-12, IFN-α/β exhibit complex biological functions, especially in regulating antiviral responses in HCV, liver injury, liver fibrosis and tumorigenesis.
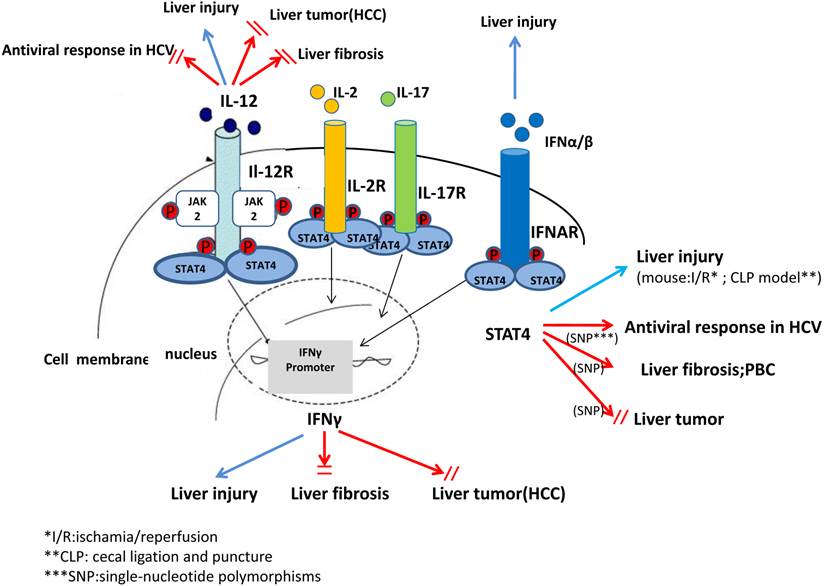
Activation of STAT4 and its downstream target genes
STAT4 is predominately activated by IL-12, which mainly regulates tissue inflammation, fibrogenesis and antiviral defense (21). In CD4+ Th1 cells, the binding of IL-12 and its receptor, results in phosphorylation of tyrosine 693 and serine 721 of STAT4 (22). Then phosphorylated STAT4 translocates into nucleus, binds to specific DNA sequences, leading to the production of inflammatory cytokines, like IFN-γ (22). IFN-γ combined with other inflammatory cytokines may induce liver injury associated with Th1 cellular immunity and autophagy-related necrotic hepatocyte death (23). IFN-α can also activate STAT4, resulting in tyrosine and serine phosphorylation, DNA binding of STAT4, and consequently transcription of its target genes (6). In mast cells, IFN-β may activate STAT4 to induce monocyte chemoattractant protein 1 (MCP-1) expression (7). IL-2 can also induce tyrosine phosphorylation of STAT4 in NK and NKT cells (9). It has also been reported that IL-17 can induce STAT4 activation in human leukemic monocyte lymphoma cell line U937 (10).
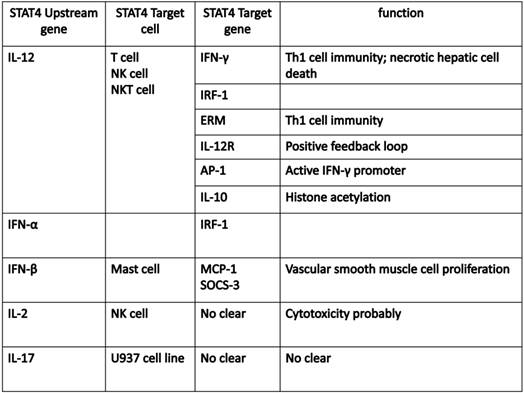
The development of ChIP (chromatin immunoprecipitation)-on-Chip technology greatly expanded our understanding on the genomic occupancies of STAT4. Good SR et al explored that STAT4 target genes could beclustered into several subsets by functions, including Th1 phenotype-related cluster (Furin, Ifng and Il18r1, Myd88, Gadd45g), TCR signaling-related cluster (LCE1B, Lcp2), other receptor (IL-12ra), and cytokine cluster (IL-10, IL-21, IL-24 and IL-17f) (24). Ifng is one of the main target genes for STAT4, which plays a key role in the IL-12-induced differentiation of T cells into the Th1 pathway. Activator Protein 1 (AP-1) is an important target gene,regulated by STAT4. STAT4 can bind with c-Jun, and the phosphorylated c-Jun-STAT4 complex efficiently interacts with the AP-1-relevant promoter sequence. AP-1 binds to the promoter sequence to elicit IFN-γ promoter activation depending on the presence of STAT4 (25). The Ets transcription factor ERM is also one of STAT4 downstream target genes and ERM is Th1-specific which isinduced by IL-12 through the STAT4-dependent pathway (26). IFN regulatory factor-1 (IRF-1) has been well known to be regulated by IFN-α. More interestingly, STAT4 also participates in the regulation of IRF-1 in two different mechanisms. First, STAT4 is required for the IL-12-dependent transactivation of IRF1. Second, STAT4 can directly bind to the IRF-1 promoter (27). In addition, suppressor of cytokine signaling 3 (SOCS-3) and monocyte chemoattractant protein-1(MCP-1) contribute to the human vascular smooth muscle cells (VSMC) proliferation by STAT4-dependent pathway (28). An interesting paper reported STAT4-binding element in the fourth intron of the IL-10 gene. This intronic STAT4 motif is a target for cytokine-induced histone acetylation, which indicates that STAT4 may play a role in regulating histone acetylation via binding downstream gene IL-10 (29).
STAT4 and its related cytokines in liver injury
IL-12 and liver injury
IL-12 is the major cytokine that can activate STAT4, synthesized predominantly by dendritic cells, macrophages and human B-lymphoblastoid cells (30). It comprises of two subunits, including the light chain p35 (IL-12A) and the heavy chain p40 (IL-12B). There are other heterodimeric interleukins, such as IL-23, IL-27, and IL-35, whose subunits consist of either or both the IL-12 p35 or p40 chains (31). The IL-12 receptor belongs to the hematopoietin receptor superfamily and it is composed of two chains that are termed as IL-12Rβ1 and IL-12Rβ2. Ligand binding results in heterodimer formation and activation of the receptor associated with JAK kinases, Jak2 and Tyk2 (32). IL-12 activates the JAK2 and Tyk2, leading to phosphorylation of STAT4 on tyrosine 693 (6). Moreover, IL-12 can also activate the p38/MKK6 signaling pathway that in turn phosphorylates STAT4 on serine 721 (22). Numerous studies have suggested that IL-12 acts as a pro-inflammatory cytokine which induces liver injury and also inhibits liver tumor growth by activating NK and NKT cells to produce IFN-γ (33). IL-12-mediated increased IFN-γ production, cellular proliferation, enhanced NK cell cytotoxicity, and Th1 cell differentiation, were all impaired in STAT4 knockout mice (34, 35). IL-12 therapy enhances type 1 immunity and induces IFN-γ expression. IL-12 could also strongly synergize with other cytokines like IL-18 to enhance both cytolytic activity and IFN-γ production by T, NKT, and NK cells, thereby promoting liver inflammation (36, 37). Overexpression of IL-12 in the liver by hydrodynamic injection of IL-12 cDNA resulted in liver injury (38). However, prolonged IL-12 stimulation of NK cells specifically decreased the level of activated STAT4 protein and induced apoptosis with increasing ROS production in human NK cells (39). Deletion of IL-12 suppressed liver inflammation in dominant negative TGF-β receptor transgenic mice (40), but in concanavalin A(ConA)-induced hepatitis, the role of IL-12 remains as a controversial issue (41, 42). Our recent study showed that disruption of IL-12a or IL-12b augmented Con A-induced hepatocellular damage (43). Other studies have focused on the antiviral role of IL-12. Some data showed that IL-12 treatment results in the disappearance of cytoplasmic hepatitis B core antigen (44). In contrast, other results reveal that IL-12 induces a strong immunosuppressive reaction in the liver of chronic woodchuck hepatitis virus (WHV) carriers that counteracts the antiviral effect of the treatment (45). Moreover, clinical data showed that low serum IL-12p40 levels are associated with a non-virological response in chronic hepatitis C patients whose IL28B polymorphism is TG or GG genotypes (46).
IFN-α/β and liver injury
Type-I IFN α/β, mainly produced by macrophages and fibroblasts, has antiviral, anti-proliferative and immune regulatoryactivities (47). Type-I IFN receptor (IFNAR) is a common cell surface receptor complex, composed of subunits IFNAR1 and IFNAR2. The binding of IFNα/β to its cell surface receptor leads to the phosphorylation of the transcription factor STAT1, STAT4 and activates IFN target genes (48). Previous work has suggested that type-I IFN signaling is required in the innate immunity-mediated ischemia-reperfusion induces liver transplant damage (49). Disruption of type-I IFN signaling decreased serum alanine aminotransferase (ALT) levels, improved histological architecture, and increased 14-day survival in a clinically relevant murine model of extended hepatic cold followed by orthotopic liver transplantation (OLT). The physiological role of type I IFN in controlling IFN-γ production remains to be fully evaluated.
STAT4 and liver injury in mouse models
IFN- γ serves as an important target gene of STAT4 and it has been well documented in liver inflammatory response, liver injury and inhibition of virus replication. Moreover, the activation of STAT4 was also detected in several models of liver injury including Con A-induced T cell hepatitis (1, 50) and hepatic ischemia/reperfusion injury (51, 52). The role of STAT4 in the pathogenesis of animal liver injury models is complicated and summarized below.
In ConA-induced mouse model, peak STAT4 activation in liver was detected at 9 hours (50). Recently, Li et al. reported that suppressive oligodeoxynucleotides (ODNs) inhibit the development of Con A-induced hepatitis through down-regulation of the STAT1/4 and T-bet pathways, suggesting that STAT4 may be involved in the development of liver injury (53). Our studies further found after Con A administration, STAT4 was observed being activated in multiple types of liver immune cells (T cells, NK T cells, macrophages and Kupffer cells). Disruption of IL-12a, IL-12b, or STAT4 significantly augmented ConA-induced liver injury even though the reduced production of Th1 and Th2 cytokines. Higher FasL expression and increased cytotoxicity against hepatocytes in hepatic NKT cells from ConA-treated STAT4-/- mice than those from wild-type mice can explain this phenomenon. In summary, our results show that despite of the up-regulation of pro-inflammatory cytokines, STAT4 protects mice against acute T-cell hepatitis, which is mediated by direct or indirect down-regulation of FasL expression on NKT cells (43).
The function of STAT4 in ischemia/reperfusion (I/R) model remains to be controversial. Shen XD et.al reported that STAT4-deficient mice were resistant to hepatic ischemia/reperfusion injury, suggesting that disruption of STAT4 may be protective in I/R induced liver injury progression (52). However, another study showed that there was no significant difference in the severity of liver injury between STAT4-deficient and wild-type mice after liver ischemia/reperfusion injury (51).
STAT4 and its related cytokines in human liver injury
IL-12 and viral hepatitis
IL12B 3'UTR gene polymorphisms may be associated with HBV susceptibility (54). Serum low IL-12p40 level is relative to non-virological responses in chronic hepatitis C patients who take the treatment with pegylated interferon and ribavirin (46). Modulation of regulatory T-cell activity in combination with IL-12 induces a strong immunosuppressive reaction in the liver of chronic WHV carriers which may indicates that IL-12 involves in counteracting the antiviral effect (45). Recently, in vivo and in vitro experiments have shown that IFN-α-induced regulation of STAT1 and STAT4 phosphorylation, which plays a role in increasing cytotoxicity and decreasing IFN-γ production in HCV infection (55). Another study revealed that type I IFNs can promote IFN-γ production by activating STAT4, but can also inhibit production of IL-12. So the efficacy of IFN-α in treatment of hepatitis C, may depend in part on the balance of IFN-γ-inducing and IL-12-suppressing effect (56).
STAT4 and viral hepatitis
The NK cells from chronic hepatitis C patients treated with IFN-α have higher level of phosphorylated-STAT4 and phosphorylated-STAT1 than those from healthy subjects. It suggested that both STAT4 and STAT1 activation in NK cells may play a crucial role in the antiviral response in chronic hepatitis C (57). There is evidence suggesting that STAT4 variants (rs7574865) might contribute to chronic hepatitis B disease progression, and the variants seemed to be population specific in Tibetans in China (58).
STAT4 and drug induced liver injury (DILI)
The mechanisms of drug-induced liver injury remain unclear. An analysis of 285 DILI patients indicated a trend association between STAT4 variants (rs7574865) and hepatocellular injury (59). Another study which included over 800 patients and over 3000 control subjects showed there is an association between a single-nucleotide polymorphisms (SNP) in the STAT4 gene and a common type of DILI (hepatocellular injury) (60).
STAT4 and its related cytokines in liver fibrosis
Liver fibrosis is an accompanying response and common characteristic of chronic liver disease. The pathologic features of hepatic fibrosis include the accumulation of extracellular matrix (ECM) proteins and the activation of hepatic stellate cells (HSCs). After liver injury and inflammation, HSCs become activated, express alpha-smooth muscle actin (a-SMA), and produce large amounts of collagen (61-63). The JAK-STAT signaling pathway has been proposed to play a central role in liver inflammation and fibrosis development. Kong X et al reported that IL-22 induces HSC senescence through the activation of STAT3, SOCS3, and p53 pathways, thereby inhibiting liver fibrosis (64). It has been well studied that IL-12-mediated activation of STAT4 plays an important role in regulating tissue inflammation, fibrogenesis and antiviral defense (1, 51, 65-67). Cheng et al. reported granuloma size was significantly larger and granuloma fibrosis remarkably intensified in anti-IL-12-treated mice after schistosome infection. These results suggest that IL-12 may play an impeditive role in granuloma formation as well as in liver fibrosis (68). However, the roles of STAT4 in HSC senescence or activation and also in the pathogenesis of liver fibrosis still remain largely unknown.
STAT4 and its related cytokines in human liver fibrosis
Primary biliary cirrhosis
Using the advent of genome-wide association studies, STAT4, IL12A, and IL12RB2 have been identified to be the risk loci that is associated with development of primary biliary cirrhosis (69).
Liver fibrosis
A clinical study genotyped 160 transplant patients with HCV recurrence for STAT4 (rs7574865) and determined their fibrotic stage based on pathologic analysis. This result indicates STAT4-T-allele is associated with development of advanced fibrosis (70).
STAT4 and its related cytokines in liver tumor
IL-12 and liver tumor
Being the STAT4 upstream cytokine, IL-12 is well known to play an important role in the regulation of innate and adaptive immune responses (71). IL-12 can induce prolonged IFN-γ production, NK cell activation and inhibition of liver metastasis of CT-26 colon cancer cells. Upon IL-12 administration, STAT4 activation in NK cells enhanced and IFN-γ production increased (72). It has been shown that IL-12 therapy can induce regression of primary tumors, inhibit the distant metastasis, and prolong the survival of tumor-bearing mice (73-76). Intratumoral gene transfer of murine IL-12 enhances lymphocytic infiltration into the tumor and significantly reduces the number of microvessels and inhibits the growth of HCC (77). A recent study indicates that IL12A rs568408 may contribute to the risk of HCC and modify HCC risk associated with HBV infection in Chinese patients (78). IL-12-mediated anti-tumor effects depend on the production of IFN-γ in vivo in several animal models (74, 79-82). IL-12 can inhibit tumor angiogenesis mainly through IFN-γ-dependent production of the chemokine IP-10 (83). Combined GM-CSF and IL-12 gene therapy induces various types of effectors and high levels of IFN-γ, thereby suppresses the growth of orthotopic liver tumors (84).
IFN-a/β and liver tumor
It is shown that endogenous type I IFN prevents the growth of primary carcinogen-induced and transplantable tumors but does not act directly on tumor cells. Hematopoietic cells, in particular NK cells, are critical IFN-α/β targets involving in the antitumor responses (85, 86). The combination therapy of IFN-α and sorafenib synergistically suppresses HCC growth and apoptosis (87).
STAT4 and human liver tumor
The studies about STAT4 in liver tumor development are very few. Recent studies have shown that STAT4 deficiency may contribute to impaired IFN-γ production in lymphoma patients after autologous peripheral blood stem cell transplantation (PBSCT). In fact, STAT4 deficiency in these patients when acquired may also promote lymphoma development (88). Chemotherapy-induced STAT4 deficiency could be due to the reduced levels of STAT4 mRNA and the protein stability (20).
A genome-wide association study (GWAS) was conducted in 2,514 Chinese patients with chronic HBV carriers (1,161 HCC cases and 1,353 controls) followed by a 2-stage validation among 6 independent populations of chronic HBV carriers (4,319 cases and 4,966 controls). The analysis showed that HCC risk was significantly associated with loci rs7574865 at STAT4. The risk allele G of these loci was strongly associated with lower levels of STAT4 mRNA in HCC tumor tissues (89). These findings indicate that STAT4 is associated with HCC development, although more consolidated studies are still needed.
Conclusions
In summary, in vivo and in vitro studies from recent decades have suggested that STAT4 and its related cytokines exhibit complex biological functions in regulating hepatic anti-viral responses, inflammation, proliferation, apoptosis and tumorigenesis. These findings can help with the characterization of pathophysiology and treatments of liver diseases that may be associated with STAT4 and its cytokines. However, there is still a long way to explore and elucidate STAT4 as a therapeutic approach in treating liver disease (Figure 3).
Acknowledgements
This work was supported by the National Natural Science Foundation of China grant 81300312(Y.W), 81372577(H.W), and TianQing liver diseases research fund of the China Hepatitis Prevention and Treatment of Foundation CFHPC20132028(Y.W). The authors thank Dr. Teresa Ramirez from Laboratory of Liver Diseases, National Institute on Alcohol Abuse and Alcoholism, National Institutes of Health, for critical reading of this manuscript.
Competing Interests
The authors have declared that no competing interest exists.
References
1. Gao B. Cytokines, STATs and liver disease. Cell Mol Immunol. 2005;2:92-100
2. Frucht DM, Aringer M, Galon J, Danning C, Brown M. et al. Stat4 is expressed in activated peripheral blood monocytes, dendritic cells, and macrophages at sites of Th1-mediated inflammation. J Immunol. 2000;164:4659-4664
3. Leonard WJ, O'Shea JJ. Jaks and STATs: biological implications. Annu Rev Immunol. 1998;16:293-322
4. Nishikomori R, Usui T, Wu CY, Morinobu A, O'Shea JJ. et al. Activated STAT4 has an essential role in Th1 differentiation and proliferation that is independent of its role in the maintenance of IL-12R beta 2 chain expression and signaling. J Immunol. 2002;169:4388-4398
5. Bacon CM, Petricoin EF 3rd, Ortaldo JR, Rees RC, Larner AC. et al. Interleukin 12 induces tyrosine phosphorylation and activation of STAT4 in human lymphocytes. Proc Natl Acad Sci U S A. 1995;92:7307-7311
6. Cho SS, Bacon CM, Sudarshan C, Rees RC, Finbloom D. et al. Activation of STAT4 by IL-12 and IFN-alpha: evidence for the involvement of ligand-induced tyrosine and serine phosphorylation. J Immunol. 1996;157:4781-4789
7. Iida K, Suzuki K, Yokota M, Nakagomi D, Wakashin H. et al. STAT4 is required for IFN-beta-induced MCP-1 mRNA expression in murine mast cells. Int Arch Allergy Immunol. 2011;155(Suppl 1):71-76
8. Miyagi T, Gil MP, Wang X, Louten J, Chu WM. et al. High basal STAT4 balanced by STAT1 induction to control type 1 interferon effects in natural killer cells. J Exp Med. 2007;204:2383-2396
9. Wang KS, Ritz J, Frank DA. IL-2 induces STAT4 activation in primary NK cells and NK cell lines, but not in T cells. J Immunol. 1999;162:299-304
10. Subramaniam SV, Cooper RS, Adunyah SE. Evidence for the involvement of JAK/STAT pathway in the signaling mechanism of interleukin-17. Biochem Biophys Res Commun. 1999;262:14-19
11. Morinobu A, Gadina M, Strober W, Visconti R, Fornace A. et al. STAT4 serine phosphorylation is critical for IL-12-induced IFN-gamma production but not for cell proliferation. Proc Natl Acad Sci U S A. 2002;99:12281-12286
12. Barnes J, Agarwal SK. Targeting STAT4 in systemic sclerosis: a promising new direction. Expert Rev Clin Immunol. 2011;7:445-448
13. Wang C, Sandling JK, Hagberg N, Berggren O, Sigurdsson S. et al. Genome-wide profiling of target genes for the systemic lupus erythematosus-associated transcription factors IRF5 and STAT4. Ann Rheum Dis. 2012
14. Piotrowski P, Lianeri M, Wudarski M, Olesinska M, Jagodzinski PP. Contribution of STAT4 gene single-nucleotide polymorphism to systemic lupus erythematosus in the Polish population. Mol Biol Rep. 2012;39:8861-8866
15. Liang YL, Wu H, Shen X, Li PQ, Yang XQ. et al. Association of STAT4 rs7574865 polymorphism with autoimmune diseases: a meta-analysis. Mol Biol Rep. 2012;39:8873-8882
16. Mohamed RH, Pasha HF, El-Shahawy EE. Influence of TRAF1/C5 and STAT4 genes polymorphisms on susceptibility and severity of rheumatoid arthritis in Egyptian population. Cell Immunol. 2012;273:67-72
17. Kim SW, Kim ES, Moon CM, Kim TI, Kim WH. et al. Abnormal Genetic and Epigenetic Changes in Signal Transducer and Activator of Transcription 4 in the Pathogenesis of Inflammatory Bowel Diseases. Dig Dis Sci. 2012
18. Lee HS, Park H, Yang S, Kim D, Park Y. STAT4 polymorphism is associated with early-onset type 1 diabetes, but not with late-onset type 1 diabetes. Ann N Y Acad Sci. 2008;1150:93-98
19. Martinez A, Varade J, Marquez A, Cenit MC, Espino L. et al. Association of the STAT4 gene with increased susceptibility for some immune-mediated diseases. Arthritis Rheum. 2008;58:2598-2602
20. Lupov IP, Voiles L, Han L, Schwartz A, De La Rosa M. et al. Acquired STAT4 deficiency as a consequence of cancer chemotherapy. Blood. 2011;118:6097-6106
21. Wurster AL, Tanaka T, Grusby MJ. The biology of Stat4 and Stat6. Oncogene. 2000;19:2577-2584
22. Visconti R, Gadina M, Chiariello M, Chen EH, Stancato LF. et al. Importance of the MKK6/p38 pathway for interleukin-12-induced STAT4 serine phosphorylation and transcriptional activity. Blood. 2000;96:1844-1852
23. Chang CP, Yang MC, Lei HY. Concanavalin A/IFN-gamma triggers autophagy-related necrotic hepatocyte death through IRGM1-mediated lysosomal membrane disruption. PLoS One. 2011;6:e28323
24. Good SR, Thieu VT, Mathur AN, Yu Q, Stritesky GL. et al. Temporal induction pattern of STAT4 target genes defines potential for Th1 lineage-specific programming. J Immunol. 2009;183:3839-3847
25. Nakahira M, Ahn HJ, Park WR, Gao P, Tomura M. et al. Synergy of IL-12 and IL-18 for IFN-gamma gene expression: IL-12-induced STAT4 contributes to IFN-gamma promoter activation by up-regulating the binding activity of IL-18-induced activator protein 1. J Immunol. 2002;168:1146-1153
26. Ouyang W, Jacobson NG, Bhattacharya D, Gorham JD, Fenoglio D. et al. The Ets transcription factor ERM is Th1-specific and induced by IL-12 through a Stat4-dependent pathway. Proc Natl Acad Sci U S A. 1999;96:3888-3893
27. Galon J, Sudarshan C, Ito S, Finbloom D, O'Shea JJ. IL-12 induces IFN regulating factor-1 (IRF-1) gene expression in human NK and T cells. J Immunol. 1999;162:7256-7262
28. Lv L, Zhang J, Huang X, Zhao Y, Zhou Z. et al. Lentivirus-mediated RNA interference targeting STAT4 inhibits the proliferation of vascular smooth muscle cells. Arch Med Res. 2008;39:582-589
29. Grant LR, Yao ZJ, Hedrich CM, Wang F, Moorthy A. et al. Stat4-dependent, T-bet-independent regulation of IL-10 in NK cells. Genes Immun. 2008;9:316-327
30. Visekruna A, Volkov A, Steinhoff U. A key role for NF-kappaB transcription factor c-Rel in T-lymphocyte-differentiation and effector functions. Clin Dev Immunol. 2012;2012:239368
31. Yuzhalin AE, Kutikhin AG. Interleukin-12: clinical usage and molecular markers of cancer susceptibility. Growth Factors. 2012;30:176-191
32. Sinigaglia F, D'Ambrosio D, Panina-Bordignon P, Rogge L. Regulation of the IL-12/IL-12R axis: a critical step in T-helper cell differentiation and effector function. Immunol Rev. 1999;170:65-72
33. Subleski JJ, Hall VL, Back TC, Ortaldo JR, Wiltrout RH. Enhanced antitumor response by divergent modulation of natural killer and natural killer T cells in the liver. Cancer Res. 2006;66:11005-11012
34. Kaplan MH, Sun YL, Hoey T, Grusby MJ. Impaired IL-12 responses and enhanced development of Th2 cells in Stat4-deficient mice. Nature. 1996;382:174-177
35. Thierfelder WE, van Deursen JM, Yamamoto K, Tripp RA, Sarawar SR. et al. Requirement for Stat4 in interleukin-12-mediated responses of natural killer and T cells. Nature. 1996;382:171-174
36. Trinchieri G. Interleukin-12: a cytokine at the interface of inflammation and immunity. Adv Immunol. 1998;70:83-243
37. Trinchieri G. Interleukin-12 and the regulation of innate resistance and adaptive immunity. Nat Rev Immunol. 2003;3:133-146
38. Rodriguez-Galan MC, Reynolds D, Correa SG, Iribarren P, Watanabe M. et al. Coexpression of IL-18 strongly attenuates IL-12-induced systemic toxicity through a rapid induction of IL-10 without affecting its antitumor capacity. J Immunol. 2009;183:740-748
39. Huang Y, Lei Y, Zhang H, Zhang M, Dayton A. Interleukin-12 treatment down-regulates STAT4 and induces apoptosis with increasing ROS production in human natural killer cells. J Leukoc Biol. 2011;90:87-97
40. Yoshida K, Yang GX, Zhang W, Tsuda M, Tsuneyama K. et al. Deletion of interleukin-12p40 suppresses autoimmune cholangitis in dominant negative transforming growth factor beta receptor type II mice. Hepatology. 2009;50:1494-1500
41. Zhu R, Diem S, Araujo LM, Aumeunier A, Denizeau J. et al. The Pro-Th1 cytokine IL-12 enhances IL-4 production by invariant NKT cells: relevance for T cell-mediated hepatitis. J Immunol. 2007;178:5435-5442
42. Nicoletti F, Di Marco R, Zaccone P, Salvaggio A, Magro G. et al. Murine concanavalin A-induced hepatitis is prevented by interleukin 12 (IL-12) antibody and exacerbated by exogenous IL-12 through an interferon-gamma-dependent mechanism. Hepatology. 2000;32:728-733
43. Wang Y, Feng D, Wang H, Xu MJ, Park O. et al. STAT4 Knockout Mice Are More Susceptible to Concanavalin A-Induced T-Cell Hepatitis. Am J Pathol. 2014;184:1785-1794
44. Cavanaugh VJ, Guidotti LG, Chisari FV. Interleukin-12 inhibits hepatitis B virus replication in transgenic mice. J Virol. 1997;71:3236-3243
45. Otano I, Suarez L, Dotor J, Gonzalez-Aparicio M, Crettaz J. et al. Modulation of regulatory T-cell activity in combination with interleukin-12 increases hepatic tolerogenicity in woodchucks with chronic hepatitis B. Hepatology. 2012;56:474-483
46. Umemura T, Joshita S, Yoneda S, Katsuyama Y, Ichijo T. et al. Serum interleukin (IL)-10 and IL-12 levels and IL28B gene polymorphisms: pretreatment prediction of treatment failure in chronic hepatitis C. Antivir Ther. 2011;16:1073-1080
47. Trinchieri G. Type I interferon: friend or foe? J Exp Med. 2010;207:2053-2063
48. Ronnblom L. The type I interferon system in the etiopathogenesis of autoimmune diseases. Ups J Med Sci. 2011;116:227-237
49. Shen XD, Ke B, Ji H, Gao F, Freitas MC. et al. Disruption of Type-I IFN pathway ameliorates preservation damage in mouse orthotopic liver transplantation via HO-1 dependent mechanism. Am J Transplant. 2012;12:1730-1739
50. Hong F, Jaruga B, Kim WH, Radaeva S, El-Assal ON. et al. Opposing roles of STAT1 and STAT3 in T cell-mediated hepatitis: regulation by SOCS. J Clin Invest. 2002;110:1503-1513
51. Kato A, Graul-Layman A, Edwards MJ, Lentsch AB. Promotion of hepatic ischemia/reperfusion injury by IL-12 is independent of STAT4. Transplantation. 2002;73:1142-1145
52. Shen XD, Ke B, Zhai Y, Gao F, Anselmo D. et al. Stat4 and Stat6 signaling in hepatic ischemia/reperfusion injury in mice: HO-1 dependence of Stat4 disruption-mediated cytoprotection. Hepatology. 2003;37:296-303
53. Li N, Liu YH, Li SL, Fu CY, Zhou RR. et al. Protective role of synthetic oligodeoxynucleotides expressing immunosuppressive TTAGGG motifs in concanavalin A-induced hepatitis. Immunol Lett. 2013;151:54-60
54. Wang C, Zhang X, Zhu B, Hu D, Wu J. et al. Relationships between tumour necrosis factor-alpha, interleukin-12B and interleukin-10 gene polymorphisms and hepatitis B in Chinese Han haemodialysis patients. Nephrology (Carlton). 2012;17:167-174
55. Edlich B, Ahlenstiel G, Zabaleta Azpiroz A, Stoltzfus J, Noureddin M. et al. Early changes in interferon signaling define natural killer cell response and refractoriness to interferon-based therapy of hepatitis C patients. Hepatology. 2012;55:39-48
56. Byrnes AA, Li DY, Park K, Thompson D, Mocilnikar C. et al. Modulation of the IL-12/IFN-gamma axis by IFN-alpha therapy for hepatitis C. J Leukoc Biol. 2007;81:825-834
57. Miyagi T, Takehara T, Nishio K, Shimizu S, Kohga K. et al. Altered interferon-alpha-signaling in natural killer cells from patients with chronic hepatitis C virus infection. J Hepatol. 2010;53:424-430
58. Liao Y, Cai B, Li Y, Chen J, Ying B. et al. Association of HLA-DP/DQ, STAT4 and IL-28B variants with HBV viral clearance in Tibetans and Uygurs in China. Liver Int. 2014
59. Urban TJ, Shen Y, Stolz A, Chalasani N, Fontana RJ. et al. Limited contribution of common genetic variants to risk for liver injury due to a variety of drugs. Pharmacogenet Genomics. 2012;22:784-795
60. Urban TJ, Goldstein DB, Watkins PB. Genetic basis of susceptibility to drug-induced liver injury: what have we learned and where do we go from here? Pharmacogenomics. 2012;13:735-738
61. Bataller R, Brenner DA. Liver fibrosis. J Clin Invest. 2005;115:209-218
62. Friedman SL. Mechanisms of hepatic fibrogenesis. Gastroenterology. 2008;134:1655-1669
63. Friedman SL. Hepatic stellate cells: protean, multifunctional, and enigmatic cells of the liver. Physiol Rev. 2008;88:125-172
64. Kong X, Feng D, Wang H, Hong F, Bertola A. et al. Interleukin-22 induces hepatic stellate cell senescence and restricts liver fibrosis in mice. Hepatology. 2012;56:1150-1159
65. Kong X, Horiguchi N, Mori M, Gao B. Cytokines and STATs in Liver Fibrosis. Front Physiol. 2012;3:69
66. Wang H, Lafdil F, Kong X, Gao B. Signal transducer and activator of transcription 3 in liver diseases: a novel therapeutic target. Int J Biol Sci. 2011;7:536-550
67. Avouac J, Furnrohr BG, Tomcik M, Palumbo K, Zerr P. et al. Inactivation of the transcription factor STAT-4 prevents inflammation-driven fibrosis in animal models of systemic sclerosis. Arthritis Rheum. 2011;63:800-809
68. Cheng YL, Song WJ, Liu WQ, Lei JH, Kong Z. et al. The effects of interleukin (IL)-12 and IL-4 deficiency on worm development and granuloma formation in Schistosoma japonicum-infected mice. Parasitol Res. 2012;110:287-293
69. Hirschfield GM, Invernizzi P. Progress in the genetics of primary biliary cirrhosis. Semin Liver Dis. 2011;31:147-156
70. Eurich D, Boas-Knoop S, Struecker B, Neuhaus R, Neuhaus P. et al. Genetic variants of STAT-4 affect the development of graft fibrosis after liver transplantation for HCV-induced liver disease. Transplantation. 2013;95:203-208
71. Trinchieri G. Interleukin-12: a proinflammatory cytokine with immunoregulatory functions that bridge innate resistance and antigen-specific adaptive immunity. Annu Rev Immunol. 1995;13:251-276
72. Uemura A, Takehara T, Miyagi T, Suzuki T, Tatsumi T. et al. Natural killer cell is a major producer of interferon gamma that is critical for the IL-12-induced anti-tumor effect in mice. Cancer Immunol Immunother. 2010;59:453-463
73. Brunda MJ, Luistro L, Warrier RR, Wright RB, Hubbard BR. et al. Antitumor and antimetastatic activity of interleukin 12 against murine tumors. J Exp Med. 1993;178:1223-1230
74. Nastala CL, Edington HD, McKinney TG, Tahara H, Nalesnik MA. et al. Recombinant IL-12 administration induces tumor regression in association with IFN-gamma production. J Immunol. 1994;153:1697-1706
75. Zou JP, Yamamoto N, Fujii T, Takenaka H, Kobayashi M. et al. Systemic administration of rIL-12 induces complete tumor regression and protective immunity: response is correlated with a striking reversal of suppressed IFN-gamma production by anti-tumor T cells. Int Immunol. 1995;7:1135-1145
76. Mu J, Zou JP, Yamamoto N, Tsutsui T, Tai XG. et al. Administration of recombinant interleukin 12 prevents outgrowth of tumor cells metastasizing spontaneously to lung and lymph nodes. Cancer Res. 1995;55:4404-4408
77. Harada N, Shimada M, Okano S, Suehiro T, Soejima Y. et al. IL-12 gene therapy is an effective therapeutic strategy for hepatocellular carcinoma in immunosuppressed mice. J Immunol. 2004;173:6635-6644
78. Liu L, Xu Y, Liu Z, Chen J, Zhang Y. et al. IL12 polymorphisms, HBV infection and risk of hepatocellular carcinoma in a high-risk Chinese population. Int J Cancer. 2011;128:1692-1696
79. Brunda MJ, Luistro L, Hendrzak JA, Fountoulakis M, Garotta G. et al. Role of interferon-gamma in mediating the antitumor efficacy of interleukin-12. J Immunother Emphasis Tumor Immunol. 1995;17:71-77
80. Hess SD, Egilmez NK, Bailey N, Anderson TM, Mathiowitz E. et al. Human CD4+ T cells present within the microenvironment of human lung tumors are mobilized by the local and sustained release of IL-12 to kill tumors in situ by indirect effects of IFN-gamma. J Immunol. 2003;170:400-412
81. Kozar K, Kaminski R, Switaj T, Oldak T, Machaj E. et al. Interleukin 12-based immunotherapy improves the antitumor effectiveness of a low-dose 5-Aza-2'-deoxycitidine treatment in L1210 leukemia and B16F10 melanoma models in mice. Clin Cancer Res. 2003;9:3124-3133
82. Smyth MJ, Taniguchi M, Street SE. The anti-tumor activity of IL-12: mechanisms of innate immunity that are model and dose dependent. J Immunol. 2000;165:2665-2670
83. Sgadari C, Angiolillo AL, Tosato G. Inhibition of angiogenesis by interleukin-12 is mediated by the interferon-inducible protein 10. Blood. 1996;87:3877-3882
84. Chang CJ, Chen YH, Huang KW, Cheng HW, Chan SF. et al. Combined GM-CSF and IL-12 gene therapy synergistically suppresses the growth of orthotopic liver tumors. Hepatology. 2007;45:746-754
85. Dunn GP, Bruce AT, Sheehan KC, Shankaran V, Uppaluri R. et al. A critical function for type I interferons in cancer immunoediting. Nat Immunol. 2005;6:722-729
86. Swann JB, Hayakawa Y, Zerafa N, Sheehan KC, Scott B. et al. Type I IFN contributes to NK cell homeostasis, activation, and antitumor function. J Immunol. 2007;178:7540-7549
87. Wang L, Jia D, Duan F, Sun Z, Liu X. et al. Combined anti-tumor effects of IFN-alpha and sorafenib on hepatocellular carcinoma in vitro and in vivo. Biochem Biophys Res Commun. 2012;422:687-692
88. Robertson MJ, Chang HC, Pelloso D, Kaplan MH. Impaired interferon-gamma production as a consequence of STAT4 deficiency after autologous hematopoietic stem cell transplantation for lymphoma. Blood. 2005;106:963-970
89. Jiang DK, Sun J, Cao G, Liu Y, Lin D. et al. Genetic variants in STAT4 and HLA-DQ genes confer risk of hepatitis B virus-related hepatocellular carcinoma. Nat Genet. 2013;45:72-75
Author contact
Corresponding author: Hua Wang, M.D., Ph.D., Department of Oncology, the First Affiliated Hospital of Anhui Medical University, Hefei, 230032. Email: wanghuaedu.cn.

 Global reach, higher impact
Global reach, higher impact