Impact Factor ISSN: 1449-2288
Int J Biol Sci 2017; 13(8):1038-1050. doi:10.7150/ijbs.19476 This issue Cite
Research Paper
DNA Methylation Mediated Downregulation of miR-449c Controls Osteosarcoma Cell Cycle Progression by Directly Targeting Oncogene c-Myc
1. Department of Orthopedics, The First Affiliated Hospital of Kunming Medical University, Kunming, Yunnan 650032, China;
2. Department of Dermatology, University of Colorado Anschutz Medical Campus, Aurora, CO 80045, USA.
* These authors contributed equally to this work.
Received 2017-2-3; Accepted 2017-7-1; Published 2017-8-17
Abstract
MicroRNAs (miRNAs) are critical regulators of gene expression, and they have broad roles in the pathogenesis of different diseases including cancer. Limited studies and expression profiles of miRNAs are available in human osteosarcoma cells. By applying a miRNA microarray analysis, we observed a number of miRNAs with abnormal expression in cancerous tissues from osteosarcoma patients. Of particular interest in this study was miR-449c, which was significantly downregulated in osteosarcoma cells and patients, and its expression was negatively correlated with tumor size and tumor MSTS stages. Ectopic expression of miR-449c significantly inhibited osteosarcoma cell proliferation and colony formation ability, and caused cell cycle arrest at the G1 phase. Further analysis identified that miR-449c was able to directly target the oncogene c-Myc and negatively regulated its expression. Overexpression of c-Myc partially reversed miR-449c-mimic-inhibited cell proliferation and colony formation. Moreover, DNA hypermethylation was observed in two CpG islands adjacent to the genomic locus of miR-449c in osteosarcoma cells. Conversely, treatment with the DNA methylation inhibitor AZA caused induction of miR-449c. In conclusion, our results support a model that DNA methylation mediates downregulation of miR-449c, diminishing miR-449c mediated inhibition of c-Myc and thus leading to the activation of downstream targets, eventually contributing to osteosarcoma tumorigenesis.
Keywords: microRNA, miR-449c, DNA methylation, cell cycle, osteosarcoma.
Introduction
MicroRNAs (miRNAs), a class of small non-coding RNAs with variable lengths (∼21-25 nucleotides), have been demonstrated to play important regulatory roles in the pathogenesis of different diseases including cancer, by targeting specific mRNAs for degradation or translation repression [1-6]. In the past several years, investigation of miRNA expression profiling in different diseases has been popular because miRNAs are key regulators in gene expression networks and can affect a broad range of cellular processes such as cell cycle progression, inflammation, cell differentiation and proliferation, cell death, as well as the stress response [1-6]. Owing to these miRNA expression profiles, a great number of unique miRNAs that function in specific cancer types or in specific biological processes have been reported [7-15]. For instance, aberrant expression of miR-191 and miR-193a was associated with poor survival rates in melanoma patients [16]. High miR-155 and low let-7a-2 were correlated with significant reductions in lung cancer survival rate [17]. In chronic lymphocytic leukemia, the miR-15/16 cluster showed tumor-suppressor activity and high levels of this miRNA were associated with good prognosis [18]. Moreover, several studies have also investigated miRNA expression profiles in human osteosarcoma cancerous tissues and cells [19-21]. Aberrant expression of a variety of miRNAs such as miR-1 [13], miR-17-5p [22], miR-26a [23], miR-34a [24], miR-133b [13], miR-150 [25], miR-203 [24], and miR-370 [26], has been reported to be involved in osteosarcoma tumorigenesis. Epigenetic mechanisms including DNA methylation and histone modifications play major roles in the regulation of miRNA expression [27-30]. Hypermethylation of the CpG island adjacent to the genomic locus of tumor suppressive miRNAs is one of the most common mechanisms to attenuate miRNA expression during tumorigenesis [27-32].
c-Myc functions as a transcription factor and plays multiple roles in tumorigenesis, such as cell cycle progression and apoptosis; moreover, c-Myc is usually overexpressed in different cancer types, such as liver, breast, lung and prostate cancers [33-37]. Activation of c-Myc in cancer cells can result in constitutive activation of a pathway, such as Wnt//β-Catenin in tumors with loss of APC, or through direct alterations of the c-Myc gene, such as amplification or chromosomal translocation [33-37]. In addition, several miRNAs such as miR-33b [38], let-7 [39], and miR-145 [40], have also been identified to target the 3′-UTR of c-Myc, thereby affecting its expression. The abnormal expression of c-Myc in cancers presumably causes a sustained increase in c-Myc protein levels, perhaps throughout the entire cell cycle rather than in a restricted manner, because elevated expression of c-Myc activates expression of many cell cycle regulators such as cyclin D1, D2, CDK4, and CDK6 through binding enhancer box sequences (E-boxes) [38-41].
In this study, we subjected mRNAs from three-paired cancerous tissues and their adjacent normal tissues to a miRNA microarray platform. We identified a total number of 28 miRNAs with higher levels and 53 miRNAs with lower levels in cancerous tissues compared to that of normal tissues. Next, we focused our further studies on one of the down-regulated miRNAs, miR-449c, and assessed its role in the pathogenesis of osteosarcoma. Our results demonstrated that miR-449c acted as a tumor suppressor, and it directly targeted c-Myc and regulated the expression of c-Myc downstream targets including Cyclin D1, D2, CDK4, and CDK6. Moreover, we also revealed that DNA hypermethylation of CpG islands adjacent to the miR-449c genomic locus was responsible for the downregulation of miR-449c in osteosarcoma cells, and this process caused activation of c-Myc and its downstream cell cycle regulators.
Materials and Methods
Osteosarcoma patients and cell lines
Forty-eight paired cancerous tissues and their adjacent non-tumor tissues were obtained from 48 patients who underwent surgery at the Department of Orthopedics, Kunming Medical University, Yunnan, China, between January 2010 and December 2014. All patients were diagnosed with osteosarcoma according to histopathological features. The patients' clinicopathological characteristics, including age, gender, MSTS stages, and tumor size are summarized in Supplementary Table-1. The samples were immediately frozen in liquid nitrogen after surgery and stored at -80 °C until RNA extraction. The clinical samples were acquired with written informed consent from all of the participants following protocols approved by the ethical board of Kunming Medical University. All experimental procedures used in this study were carried out in accordance with the approved guidelines of the ethical board of Kunming Medical University.
The four osteosarcoma cell lines including U2OS, Saos-2, MG63 and HOS, as well as the human osteoblast cell line hFOB1.19, were all obtained from the American Type Culture Collection (ATCC, MD, USA). All cell lines were cultured with DMEM medium (Life Technologies, CA, USA) supplemented with 10% heat-inactivated FBS (Invitrogen, CA, USA) and 100 U/ml of penicillin-streptomycin (Invitrogen, CA, USA). Cells were cultured in a humidified incubator (Invitrogen, CA, USA) at 37°C with 5% CO2. Cells were split twice per week with a density of approximately 80% confluent.
miRNA microarray analysis
Microarray analyses of miRNAs were performed as previously described [41]. Briefly, total RNAs were isolated from frozen tissues of osteosarcoma patients by using TRIZOL® Reagent (Invitrogen, CA, USA) and then cleaned with the miRNeasy Mini kit (Qiagen, MD, USA) following the manufacturer's protocols. RNA quality and quantity were determined by spectrophotometry (A260/A280 ratio) and capillary electrophoresis (Agilent 2100 Bioanalyzer, Agilent Technologies, CA, USA). Then, a total of 1 μg of RNA of each sample was labeled with the miRCURY™ Hy3/Hy5 Power kit (Exiqon, Vedbaek, Denmark) following the manufacturer's guidelines. The labeled RNA was subjected to hybridization on the miRCURY™ LNA Array (miRBase.14.0, Exiqon, Vedbaek, Denmark) slides and signals were scanned using the Axon GenePix 4000B microarray machine (Axon Instruments, CA, USA). The average intensities (> 50) were chosen to calculate a normalization factor after background correction. MicroRNA expression data were then normalized using the median values, and differentially expressed miRNAs in cancerous tissues were identified according to the fold changes of intensities compared to normal tissues.
Cell transient transfection
Cells transfection was performed as previously described [42, 43]. Briefly, cells were first seeded onto 24-well plates and incubated for 18 h and then transiently transfected with the required plasmids (100 ng), miR-449c-mimics, anti-miR-449c, or miR-negative control (miR-NC) at a final concentration of 50 nM using Lipofectamine 2000 (Invitrogen, CA, USA) following the manufacturer's instructions. Cells were incubated in 0.5 ml DMEM medium supplemented with 10% FBS at 37°C for 48 h before subjecting to the required experiments.
Cell proliferation and colony formation assays
Cells transfected with miR-NC or miR-449c-mimic were seeded onto 96-well plates for 0, 1, 2, 3, 4, or 5 days. Cell viability was determined at 490 nm using an MTT kit (Roche, IN, USA) following the manufacturer's instructions.
A colony formation assay was performed as previously described [34]. Briefly, cells transfected with miR-449c-mimic or miR-NC were diluted and seeded onto 6-well plates with a density of approximately 1,000 cells per well. Cells were cultured at 37°C for 14 days with a change of media every three days. Cell colonies were then fixed with 3.7% paraformaldehyde (PFA) for 5 min at room temperature, followed by staining with 0.05% crystal violet for 20 min and washed with ddH2O five times to remove excess dye. The colony numbers were counted by a gel documentation system (Bio-Rad, CA, USA).
Western blot analysis
Protein levels were determined by Western blots as previously described [44-46]. Briefly, cells were lysed in 1x RIPA buffer and equal amounts of total cell lysates were separated by SDS-PAGE and then transferred to a PVDF membrane. Blots were subsequently incubated with a primary antibody and a peroxidase-conjugated secondary antibody. The signals were detected by enhanced chemiluminescence reagents (GE Healthcare, NJ, USA) and imaged with the ChemiDoc MP (Bio-Rad, CA, USA). Antibodies to GAPDH, c-Myc, Cyclin D1, Cyclin D2, CDK4, and CDK6 were purchased from Sigma-Aldrich (St. Louis, MO, USA).
Quantitative real-time PCR (qRT-PCR)
Total RNA was extracted from frozen tissues or cultured cells using TRIZOL® Reagent (Invitrogen, CA, USA) according to the manufacturer's instructions. The Verso cDNA Synthesis Kit (Thermo Fisher Scientific, MA, USA) was used to synthesize cDNA. The resulting cDNAs were subjected to qRT-PCR analyses on the Bio-rad CFX96 real-time PCR System (Bio-rad, CA, USA) using the specific primers listed in Supplementary Table-2. The PCR cycling conditions were 95 °C for 5 min and then 45 cycles of 95°C for 15 sec and 60°C for 30 sec. GAPDH was chosen as an internal control to normalize individual gene expression using the 2-ΔCt method.
The expression of miR-449c expression was determined as previously described [24]. Briefly, total RNA was extracted from frozen tissues or cultured cells using the miRNeasy Mini Kit (Qiagen, MD, USA) following the manufacturer's guidelines. After the generation of cDNAs with TaqMan MicroRNA Reverse Transcription kit (Thermo Fisher Scientific, MA, USA), a TaqMan MicroRNA Assay kit (assay ID: 479367, Thermo Fisher Scientific, MA, USA) was used to examine the expression of miR-449c following the manufacturer's protocols. The qRT-PCR program was performed on the Bio-rad CFX96 real-time PCR System (Bio-Rad, CA, USA) at 95°C for 2 min and then 45 cycles of 95°C for 10 sec and 60 for 20 sec. RNU6B was chosen as an internal control to normalize miR-449c expression using the 2-ΔCt method. All reactions were conducted in triplicate.
Flow cytometry analysis
Flow cytometric analyses were performed as previously described [24]. Briefly, cells were washed twice with ice-cold 1×PBS and then treated with 0.25% trypsin-EDTA after transfection with miR-449c-mimic or miR-NC for 48 h. The cell suspension was fixed with 70% ethanol at 4°C for 12 h. Cells were subsequently incubated and stained in a solution containing 50 μg/mL RNase, 50 μg/mL propidium iodide (PI), and 0.1 mM EDTA at 37°C for 30 min. Cells were then subjected to flow cytometry (BD Biosciences, CA, USA) to analyze cell cycle distribution. Cells in different cell cycle stages were counted. All samples were tested in triplicate.
Drug treatment
Cells were seeded onto 6-well plates at a concentration of 1 × 105 cells per well and incubated at 37°C for 18 h. Next, cells were treated with DMSO, 1 μM AZA (Sigma-Aldrich, MO, USA), or 300 nM TSA (Sigma-Aldrich, MO, USA) for three days. The medium was changed every 24 h.
Quantitative methylation-specific PCR (qMSP)
CpG Island identification was performed in a CpG island prediction database (http://www.urogene.org) and two CpG islands around the miR-449c genomic locus were found. Methyl Primer Express v1.0 (Thermo Fisher Scientific, MA, USA) was used to design qMSP primers (Supplementary Table-3). Briefly, the sodium bisulfite modified genomic DNA samples were subjected to PCR to analyze methylated DNA using a KAPA SYBR FAST qPCR Kit (Kapa Biosystems, MA, USA) with the following cycling conditions: 95 °C for 5 min, then 45 cycles of 95 °C for 15 sec, 60°C for 60 sec. GAPDH was used as an internal control to normalize expression of CpG islands. The experiments were replicated three times.
Statistical analysis
All experiments were independently performed in triplicate. Experimental data were applied to analyze using student's t-test. Data were represented as the mean ± standard deviation (SD). Statistical significance was set at a P < 0.05.
Results
miR-449c was significantly downregulated in cancerous osteosarcoma tissues
To investigate miRNA expression profile and miRNA roles in the pathogenesis of osteosarcoma, we compared miRNA expression in cancerous tissues and their corresponding adjacent normal tissues from three osteosarcoma patients (MSTS stage II) by miRNA microarray analysis. The hierarchical clustering analysis revealed distinctive miRNA expression patterns between cancerous tissues and normal tissues (Figure 1A). Overall, a total number of 81 miRNAs were identified to have aberrant expression in cancerous tissues compared to normal tissues. Of them, 28 miRNAs were upregulated and the others were downregulated. As shown in Figure 1A, we selected 15 upregulated miRNAs and 15 downregulated miRNAs that showed consistent expression patterns in three cancerous tissues.
miR-449c was decreased in human osteosarcoma cancerous tissues. (A) Heat maps of the most significantly (P < 0.01) altered 30 miRNAs in osteosarcoma tissues were shown. RNA from three paired cancerous tissues and adjacent normal tissues were subjected to miRNA microarray analysis. Each column represented a tissue sample, and each row represented a probe set. The heat maps indicate high (red) or low (green) levels of miRNA expression. (B) Expression of miR-449c in osteosarcoma cancerous tissues was shown. Relative expression of miR-449c in osteosarcoma tumors (n = 48) was normalized to corresponding adjacent normal tissues (n = 48), P < 0.001. (C-D) Expression of miR-449c is negatively correlated with osteosarcoma tumor size and MSTS stages. The expression of miR-449c was significantly lower in larger tumors (tumor maximal diameter ≥ 12 cm) (C) and was significantly lower in the osteosarcoma patients with an advanced MSTS stages (II/III/IV) than those with an early MSTS stage (I) (D). ** P < 0.001.
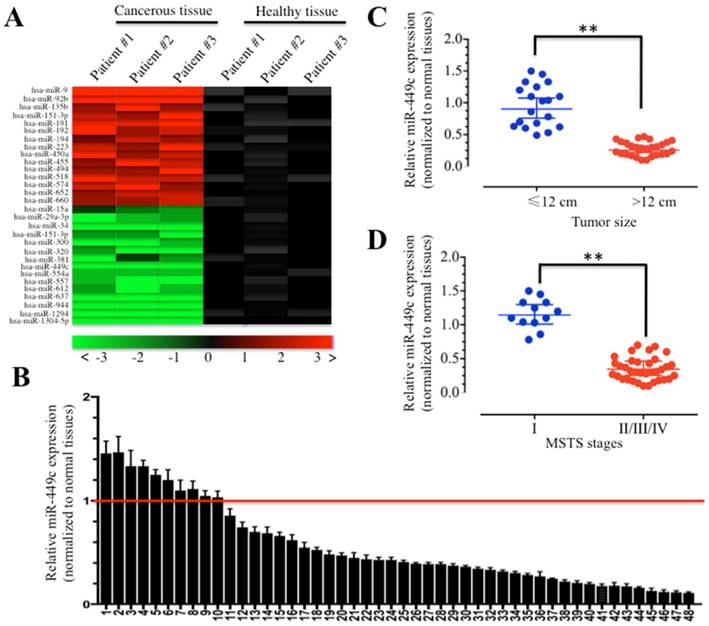
Among these downregulated miRNAs, we first focused on miR-449c, which exhibited consistent decrease in expression (more than 3-fold) in all of these three cancerous tissues. We next examined the expression of miR-449c in 48-paired cancerous tissues and normal tissues obtained from osteosarcoma patients who underwent surgery. Compared to levels in healthy tissues, we found 10 cancerous tissues that showed increased miR-449c levels (≥ 1-fold) and 38 cancerous tissues that had decreased miR-449c levels (< 1-fold) (Figure 1B). Moreover, the reduced expression of miR-449c in osteosarcoma patients was associated with larger tumor size (≥ 12 cm, P = 0.0003) (Figure 1C), and higher MSTS stages (P = 0.0001) (Figure 1D). However, the expression of miR-449c had no significant correlation with other parameters including age and gender (Supplementary Table-1). Thus, significant downregulation of miR-449c was frequently observed in the majority of the osteosarcoma tumors.
Ectopic expression of miR-449c suppressed osteosarcoma cell proliferation and colony formation
To further examine whether miR-449c expression was also decreased in human osteosarcoma cell lines, we detected its level by qRT-PCR in U2OS, Saos-2, MG63, and HOS cells. Our results showed that miR-449c expression was also markedly downregulated in all four cell lines compared to hFOB1.19 control cells (Figure 2A). Of these four malignant cell lines, U2OS and MG63 were selected to further analyze the role of miR-449c. With the observation that miR-449c was dramatically suppressed in cancerous osteosarcoma tissues and cell lines, we considered that miR-449c functioned as a tumor suppressor. To investigate whether higher level of miR-449c had effects on osteosarcoma cell growth, we re-introduced miR-449c-mimics into U2OS and MG63 cells. As shown in Figure 2B, the qRT-PCR results indicated that miR-449c was successfully overexpressed in these two cell lines. The transfected cells were then used for cell proliferation assays and the results indicated that the overexpression of miR-449c was able to significantly inhibit cell proliferation compared to cells transfected with miR-NC (Figure 2C). In addition, a significant decrease of colony formation rates was also observed in U2OS and MG63 cells overexpressing miR-449c (Figures 2D and Supplementary Figure 1). These results further demonstrate that miR-449c expression is important for osteosarcoma cell growth and the increase of miR-449c expression is a promising strategy for osteosarcoma cancer therapy.
Ectopic expression of miR-449c inhibited osteosarcoma cell proliferation and colony formation ability. (A) Expression of miR-449c was decreased in osteosarcoma cell lines. The miR-449c levels in hFOB1.19, U2OS, Saos-2, MG63, and HOS cells were determined by qRT-PCR. Expression of miR-449c was normalized to RNU6B levels in each cell line, and the resulting ratios in hFOB1.19 cells were arbitrarily defined as 1-fold. (B) Overexpression of miR-449c was found in hFOB1.19, U2OS, and MG63 cells; miR-449c-mimic and its negative control miR-NC were transfected into hFOB1.19, U2OS, and MG63 cells for 24 h. Expression of miR-449c was determined by qRT-PCR. (C) Overexpression of miR-449c inhibited osteosarcoma cell proliferation. Cell used in (B) were subject to the MTT assay to evaluate degree of cell proliferation, and the cell viability was determined at 490 nm. (D) Overexpression of miR-449c inhibited osteosarcoma cell colony formation ability. Cells used in (B) were seeded onto 6-well plates with a density of 1 x 103 cells per well and cultured with 0.1 ml DMEM medium for two weeks. Then, cells were stained with 0.5% crystal violet and the number of colonies was counted. Representative data from three independent experiments are shown. **P < 0.001.

miR-449c negatively regulated c-Myc expression by directly targeting its 3′-UTR
Given the important role of miR-449c in osteosarcoma cells, we sought to study its mechanism in osteosarcoma carcinogenesis. To this aim, we searched for mRNA targets of miR-449c in miRDB, an online database for miRNA target prediction (http://www.mirdb.org/miRDB). Surprisingly, a total number of 394 predicted targets of miR-449c were identified (data not shown). Among these putative targets, we found c-Myc, a critical oncogene involved in tumorigenesis. After sequence alignment, we identified the binding site of miR-449c in the 3′-UTR of c-Myc mRNA (Figure 3A). To validate whether miR-449c targeted c-Myc in osteosarcoma and affected c-Myc expression, qRT-PCR was carried out to analyze c-Myc expression in the 48-paired cancerous and non-tumor tissues. Interestingly, c-Myc was significantly overexpressed in the majority of cancerous tissues (38/48) after normalization with non-tumor tissues, opposite in direction from what was observed with miR-449c expression (Figure 3B). In addition, we also examined the correlation between miR-449c and c-Myc expression. As shown in Supplementary Figure 2, a two-dimensional plot depicting average expression values demonstrated a significant (P < 0.001) linear correlation (R2 = 0.81) with a Pearson correlation coefficient (r) of 0.904 between miR-449c and c-Myc expression in osteosarcoma tissue. These results indicated that miR-449c levels were negatively correlated expression levels of c-Myc in osteosarcoma cancerous tissues. To further confirm this observation, we examined mRNA and protein levels of c-Myc in U2OS, Saos-2, MG63, and HOS cells. As expected, c-Myc was overexpressed at both transcriptional and protein levels in these four cell lines (Figures 3C and 3D). Next, we further tested if c-Myc was a direct target of miR-449c by using a luciferase reporter assay. Accordingly, wild-type (WT) and mutant 3′-UTRs of c-Myc were cloned into luciferase reporter vectors (pMIR-report vectors) and then co-transfected with miR-449c-mimic or miR-NC. The luciferase assay results showed that overexpression of miR-449c significantly decreased luciferase activity in the 3′-UTR WT, but failed to repress luciferase activity in the 3′-UTR mutant. These results suggested that mutation of the miR-449c-binding site in the c-Myc 3′-UTR abolished inhibition of miR-449c and that c-Myc was directly and negatively regulated by miR-449c.
miR-449c directly targeted oncogene c-Myc. (A) Schematic representation of c-Myc 3′-UTRs showed a putative miR-449c binding site. The seed location for WT 3′-UTR of c-Myc was indicated with red, while it is indicated with blue for mutant 3′-UTR. (B) Expression of c-Myc in 48 paired cancerous tissues and non-tumor tissues was shown. (C-D) c-Myc was upregulated in osteosarcoma cells at both mRNA (C) and protein levels (D). (E) The mutant of c-Myc 3′-UTR failed to bind miR-449c. miR-449c-mimic or miR-NC was co-transfected with pMIR-c-Myc-3′-UTR-WT or pMIR-c-Myc-3′-UTR-mutant into U2OS cells for 24 h. Luciferase activity was quantified using the dual-luciferase assay reporter system by normalizing to Renilla activity. (F) Cell cycle regulators were activated in osteosarcoma cells. Protein levels of c-Myc, Cyclin D1, D2, CDK4, CDK6, and GAPDH in hFOB1.19, U2OS, MG63, Saos-2, and HOS cells were determined by Western blot. Representative data from three independent experiments are shown. **P < 0.001.
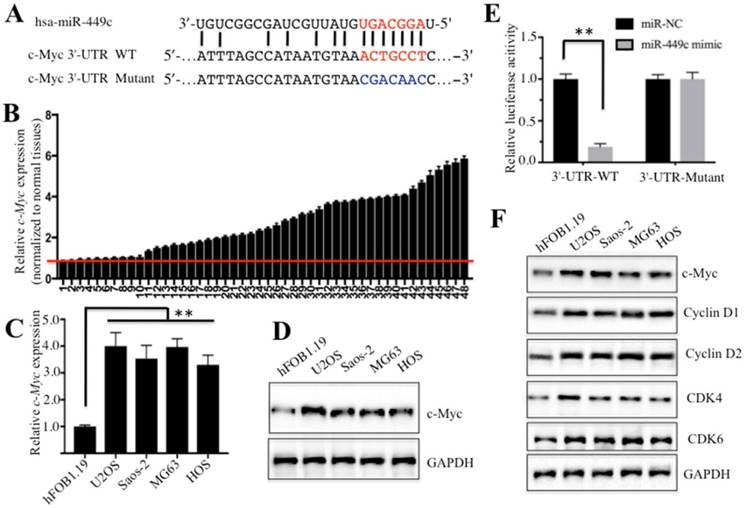
With the observation that miR-449c overexpression was able to inhibit osteosarcoma cell proliferation, one possibility that might explain this phenomenon is that miR-449c overexpression caused the downregulation of c-Myc, which further repressed the expression of some critical genes involved in cell cycle progression. To confirm this hypothesis, we primarily measured expression of some cell cycle regulators including Cyclin D1, D2, CDK4, and CDK6, which are downstream targets of c-Myc. Our results indicated that these four proteins and their mRNAs were upregulated in all four osteosarcoma cell lines compared to hFOB1.19 cells (Figure 3E and Supplementary Figures 3A and 3B), suggesting the activation of c-Myc downstream events in osteosarcoma cells.
Ectopic expression of miR-449c suppressed c-Myc expression and caused osteosarcoma cell cycle arrest at G1 phase
We further investigated whether ectopic expression of miR-449c could regulate osteosarcoma cell cycle progression. Accordingly, miR-449c-mimic and miR-NC were transfected into U2OS and MG63 cells, respectively. Nearly a 9-fold induction of miR-449c was observed in both U2OS and MG63 cells by qRT-PCR (Figures 4A and 4B). Meanwhile, expression of c-Myc and its downstream cell cycle regulators were also examined. Our results indicated that ectopic expression of miR-449c significantly inhibited levels of c-Myc and its downstream cell cycle regulators including Cyclin D1, D2, CDK4 and CDK6 (Figure 4C). In addition, we also transfected miR-NC or anti-miR-449c into hFOB1.19 cells to see if downregulation of miR-449c was able to increase the expression of these cell cycle regulators. As expected, our results indicated that expression of all of these cell cycle regulators was significantly upregulated with the downregulation of miR-449c in hFOB1.19 cells (Supplementary Figures 4A and 4B).
Ectopic expression of miR-449c caused cell cycle arrest at G1 phase. (A-B) Expression of miR-449c in U2OS and MG63 cells. miR-449c-mimic and miR-NC were transfected into U2OS and MG63 cells, respectively, for 24 h. Expression levels of miR-449c were determined by qRT-PCR. (C) Protein levels of cell cycle regulators were significantly decreased with the overexpression of miR-449c. Protein levels of c-Myc, Cyclin D1, D2, CDK4, and GAPDH in U2OS and MG63 cells transfected with miR-449c-mimic or miR-NC were determined by Western blot. (D) Overexpression of miR-449c induced cell cycle arrest at the G1 phase. Cells used in A and B were subject to flow cytometry analysis of cell cycle distribution. Representative data from three independent experiments are shown. **P < 0.001.
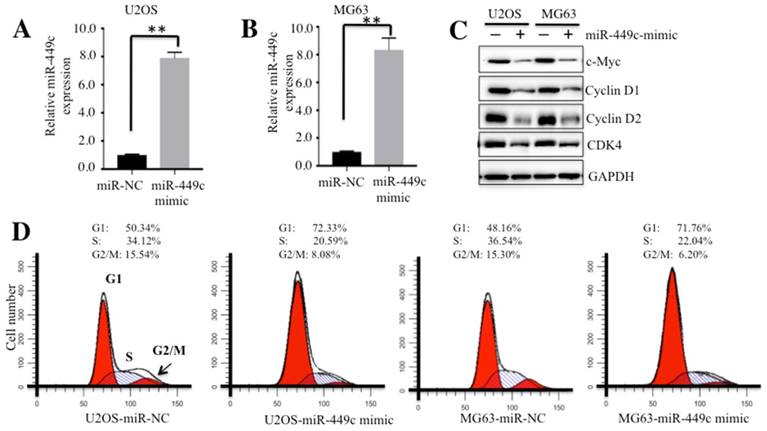
Given that Cyclin D1, D2, CDK4, and CDK6 are critical regulators of the G1-to-S transition, our results suggested that overexpression of miR-449c might affect osteosarcoma cell cycle progression. To confirm this hypothesis, we subjected cells overexpressing miR-449c to flow cytometry analysis. The results revealed that ectopic expression of miR-449c significantly increased cell population in G1 phase, but markedly decreased the percentage of cells in S phase and G2/M phase (Figure 4D).
Overexpression of c-Myc partially reversed miR-449c's effects on osteosarcoma cells
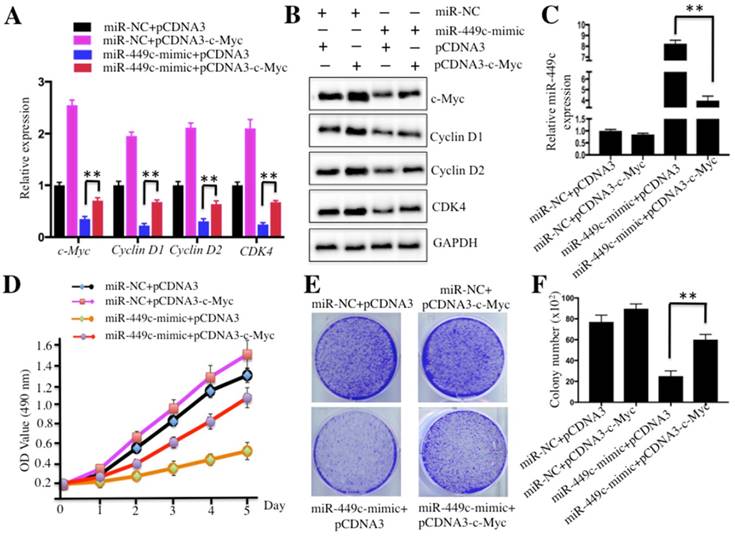
Thus far, our data demonstrated that miR-449c direct targeted the 3′-UTR of c-Myc and repressed its expression. To further investigate whether expression of c-Myc was able to affect miR-449c level, we co-transfected c-Myc and miR-370-mimic into U2OS cells and then investigated the related effects. Accordingly, the combination plasmids including miR-NC+pCDNA3, miR-NC+pCDNA3-c-Myc, miR-449c-mimic+pCDNA3, and miR-449c-mimic+pCDNA3-c-Myc were co-transfected into U2OS cells, respectively. As shown in Figure 5A, cells co-transfected with miR-449c-mimic and pCDNA3-c-Myc had ∼1.5-fold higher c-Myc levels compared to cells transfected miR-449c-mimic+pCDNA3. Similar inductions were also observed in other c-Myc targets including Cyclin D1, D2, CDK4, and CDK6 (Figures 5A and 5B). However, expression of these genes was not comparable to that in cells co-transfected with miR-NC and pCDNA3-c-Myc (Figures 5A and 5B). Additionally, miR-449c expression was also measured by qRT-PCR in these cells. The results indicated that the reintroduction of c-Myc significantly decreased the expression of miR-449c, though not to the level of the negative controls (Figure 5C). Cell proliferation assay results showed that cells co-transfected with miR-449c-mimic and pCDNA3-c-Myc enhanced osteosarcoma cell viability compared to cells transfected with the miR-449c-mimic and pCDNA3 (Figure 5D). In addition, increased colony forming ability was also observed in cells co-transfected with miR-449c-mimic and pCDNA3-c-Myc compared to those co-transfected with the miR-449c-mimic and pCDNA3 (Figures 5E and 5F). Taken together, these results demonstrate that overexpression of c-Myc partially reverses effects of miR-449c on osteosarcoma cells.
Overexpression of c-Myc partially rescued miR-449c-inhibited cell growth. U2OS cells were transfected with the following combination plasmids: miR-NC+pCDNA3, miR-NC+pCDNA3-c-Myc, miR-449c-mimic+pCDNA3, and miR-449c-mimic+pCDNA3-c-Myc. After 48 h, multiple studies were performed: mRNA levels of cell cycle regulators including Cyclin D1, D2 and CDK4 (A), protein levels of these same regulators (B), miR-449c levels (C), cell proliferation assay assessed by MTT assay (D), and colony formation assay (E and F). Representative data from three independent experiments are shown. **P < 0.001.
DNA hypermethylation caused downregulation of miR-449c in osteosarcoma cells
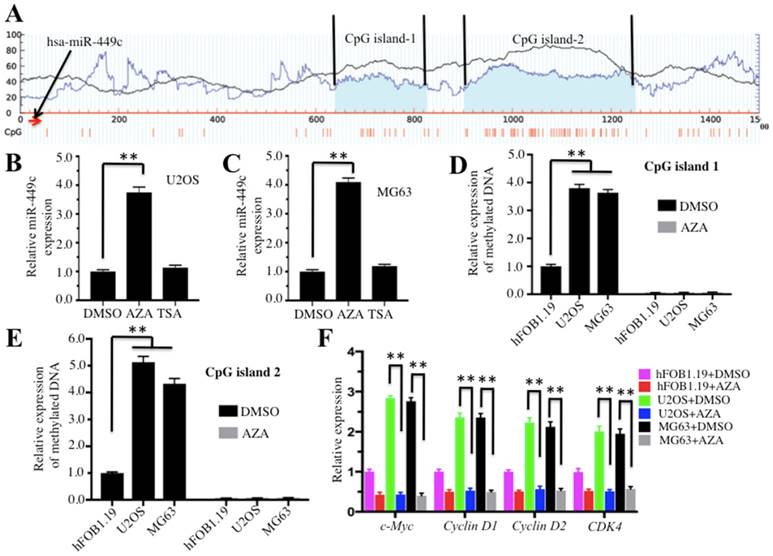
Given that miR-449c is an important regulator in osteosarcoma cells, we sought to analyze the regulatory mechanism of miR-449c expression. Because DNA methylation is a general mechanism for regulating aberrant expression of miRNAs, we initially analyzed CpG islands upstream (-1500) and downstream (+1500) of the miR-449c genome locus. Unfortunately, we did not identify any CpG island in the upstream of the miR-449c genome locus (data not shown). However, two CpG islands located downstream of the miR-449c genomic locus were identified (Figure 6A). To determine if DNA methylation had any effect on miR-449c expression, we treated U2OS and MG63 cells with the DNA methylation inhibitor AZA or the acetylation inhibitor TSA, respectively, and then evaluated miR-449c expression by qRT-PCR analysis. The results showed that there was a ∼4-fold increase in miR-449c levels in both U2OS and MG63 cells following AZA treatment, but no obvious increases were observed after TSA treatment (Figures 6B and 6C), suggesting that miR-449c expression in osteosarcoma cells might be regulated by DNA methylation. To confirm this hypothesis, we detected the DNA methylation statuses of CpG islands 1 and 2 using qMSP in hFOB1.19, U2OS, and MG63 cells with or without AZA treatment. Our results revealed that both CpG islands were significantly methylated in U2OS and MG63 cells without AZA treatment compared to hFOB1.19 cells (Figures 6D and 6E). However, DNA methylation levels of both CpG islands were dramatically decreased when cells were treated with AZA, but no differences were identified among these cell lines after AZA treatment (Figures 6D and 6E). In addition, we also examined expression of c-Myc and its targets in cells treated with or without AZA. As shown in Figure 6F, AZA treatment significantly suppressed expression of these genes. These results indicated that decreased methylation of CpG islands correlated with the induction of miR-449c in osteosarcoma cells.
DNA hypermethylation was responsible for the downregulation of miR-449c in osteosarcoma cells. (A) The miR-449c genomic locus has two adjacent CpG islands. CpG island predictions in the adjacent miR-449c genomic locus were performed in a database (http://www.urogene.org). The two predicted CpG islnds were indicated as island-1 and island-2. The genomic locus of miR-449c was indicated with a red arrow. (B-C) Effects of AZA and TSA on miR-449c expression in U2OS and MG63 cell lines are shown. The U2OS and MG63 cells were treated with DMSO, AZA (1 μM), or TSA (300 nM). Next, expression of miR-449c was determined by qRT-PCR. (D-E) Expression levels of methylated CpG islands with or without AZA treatment are shown. Three cell lines (hFOB1.19, U2OS, and MG63) were treated with or without 1 μM AZA and then expression levels of methylated CpG islands 1 and 2 were determined by qMSP analysis. (F) Expression levels of cell cycle regulators in the conditions with or without AZA treatment are shown. Messenger RNAs from cells used in D were used in qRT-PCR analyses. Expression levels of cell cycle regulators were normalized to GAPDH in each cell line, and the resulting ratios in hFOB1.19 cells with DMSO treatment were arbitrarily defined as 1-fold. Representative data from three independent experiments are shown. **P < 0.001.
Given that AZA treatment caused the upregulation of miR-449c, we next sought to investigate whether DNA hymermethyaltion in hFOB1.19 cells would have opposite effect on miR-449c and c-Myc expression. DNA Methyltransferase 1 (DNMT1) is the major enzyme responsible for maintaining methylation patterns in cells, thus, we intended to constructed its overexpression vector pCDNA3-DNMT1-Flag, and evaluated its overexpression effect on miR-449c and c-Myc expression. Accordingly, we transfected the pCDNA3-DNMT1-Flag vector into hFOB1.19 cells, followed by examination of DNMT1 level to verify its overexpression. As shown in Supplementary Figure 5A, we observed DNMT1-overexpression hFOB1.19 cells (hFOB1.19+pCDNA3-DNMT1-Flag) had higher DNMT1 protein and mRNA levels than the non-overexpression cells (hFOB1.19+pCDNA3-Flag). We also examined the DNA methylation statuses of CpG islands 1 and 2 using qMSP in these cells. Our results indicated that both CpG islands were significantly methylated in DNMT1-overexpression cells compared to non-overexpression cells (Supplementary Figures 5B and 5C). In contrast to AZA treatment results, we found expression of miR-449c was dramatically inhibited in DNMT1-overexpression cells (Supplementary Figures 5D), while expression of c-Myc and its targets was significantly upregulated (Supplementary Figures 5E). These results suggested that the enhanced methylation of CpG islands correlated with the decrease of miR-449c level in osteosarcoma cells, suggesting that the downregulation of miR-449c may be regulated, at least in part, by DNA methylation in osteosarcoma cells.
Discussion
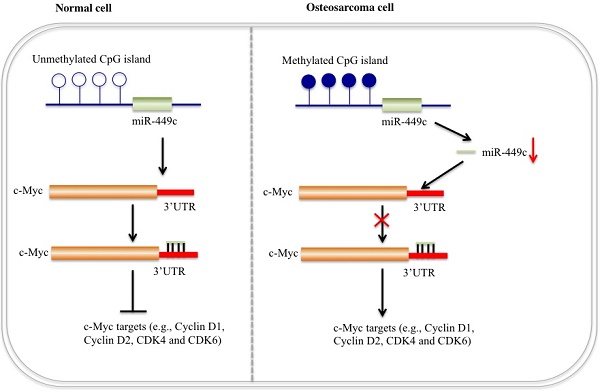
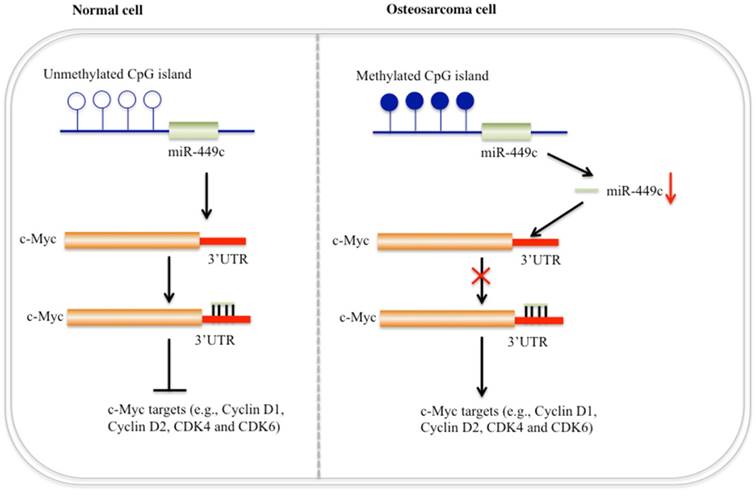
Accumulating evidence demonstrates that miRNAs are critical regulators acting as oncogenes or tumor suppressors in different processes of tumorigenesis [1-4]. Although a few miRNAs have been reported to play important roles in the pathogenesis of human osteosarcoma [21-26], the molecular mechanisms of the vast majority of miRNAs in human osteosarcoma are far from being fully elucidated. In this study, we obtained a miRNA expression profile from human osteosarcoma tissues by applying an miRNA microarray platform. A variety of miRNAs with aberrant expression were identified, but we focused our study on revealing role of miR-449c. The expression of miR-449c was analyzed in 48 cases and showed downregulation tendencies in more than 79% (38/48) patients. Moreover, expression of miR-449c was gradually decreased in stage II/III/IV of osteosarcoma patients and was negatively correlated with tumor size. In addition, we identified that miR-449c directly targeted the oncogene c-Myc, and DNA hypermethylation caused down-regulation of miR-449c in osteosarcoma cells, thereby resulting in c-Myc overexpression and activation of its downstream targets (Figure 7). These findings indicate that miR-449c may function as a tumor suppressor in the modulation of osteosarcoma cell growth.
Schematic diagram of miR-449c function in human osteosarcoma cells. In normal cells (left), CpG islands adjacent to the genomic locus of miR-449c are not methylated or are in a very low DNA methylation state. Expression of miR-449c is very high, which represses c-Myc expression and eventually results in decreased expression of cell cycle regulators. In osteosarcoma cells (right), DNA hypermethylation in the adjacent genomic region of miR-449c results in miR-449c downregulation, which diminishes its inhibition to c-Myc, leading to activation of the c-Myc gene. Activation of c-Myc further induces expression of cell cycle regulators and eventually contributes to osteosarcoma tumorigeneis.
Decreased expression of miRNAs is observed in almost all types of human malignancies and is correlated with cell proliferation, invasion, metastasis, and poor prognosis. Recent studies show that a number of miRNAs, including miR-34a and miR-203, play critical roles in controlling osteosarcoma cell growth [24]. However, it is still largely unknown what the roles of many other miRNAs in osteosarcoma are. By applying a miRNA microarray analysis, a great number of miRNAs with aberrant expression were identified. Of the identified miRNAs, we focused on a new miRNA, miR-449c, which has not been reported to play roles in tumorigenesis. Our data indicated that miR-449c was able to directly target the 3′-UTR of c-Myc. The oncogene c-Myc is an important transcription factor that is responsible for the regulation of a great number of human genes (up to one-third) [47], and plays critical roles in many cellular processes, such as cell cycle progression, cell differentiation, and cell proliferation [27, 28]. Although overexpression of c-Myc has been observed in human osteosarcoma cells for a long time [48], the underlying regulatory mechanism of this overexpression is still obscure. Our findings provide a regulatory mechanism for c-Myc overexpression in osteosarcoma cells. Further studies are required to investigate whether miR-449c is involved in signaling pathways that c-Myc participates in, which may help us more fully understand the roles of miR-449c in the pathogenesis of human osteosarcoma.
In addition, we also investigated the underlying mechanism of miR-449c downregulation in human osteosarcoma. DNA methylation and acetylation have long been recognized as key players in the regulation of miRNA expression [27-30]. Thus, we treated osteosarcoma cells with either AZA or TSA and found that miR-449c was significantly increased with the treatment of AZA but not with TSA, suggesting DNA methylation was involved in the regulatory mechanisms of miR-449c expression. In addition, overexpression of DNMT1 in hFOB1.19 cells resulted in the downregulation of miR-449c, as well as the upregulation of c-Myc and its downstream targets. These results clearly demonstrated that DNA hypermethylation is a major mechanism for the downregulation of miR-449c in osteosarcoma cells. Recently, miR-33b was also identified to target c-Myc in gastric cancer, and AZA treatment increased miR-33b expression [38]. However, we did not find alteration of miR-33b expression in human osteosarcoma cells (data not shown), suggesting that miRNA expression profiles might be different in different cancer types. DNA hypermethylation within the CpG island located in the promoter regions of tumor-suppressive miRNAs is known as an epigenetic mechanism to regulate miRNA alteration in cancer cells [27-30]. Here, we found two CpG islands adjacent to the genomic locus of miR-449c. Both of them were hypermethylated in osteosarcoma cells, and AZA treatment was able to diminish this methylation, suggesting that the hypermethylation of CpG islands might regulate the expression of miR-449c in human osteosarcoma cells.
In summary, we identified a total number of 81 miRNAs with aberrant expression in osteosarcoma cancerous tissues by applying a miRNA microarray analysis. We focused on a tumor suppressive miRNA, miR-449c, which was decreased in the majority of osteosarcoma cancerous tissues and was negatively correlated with osteosarcoma tumor size and MSTS stages. We applied both in vitro and in vivo experimental and clinical findings to investigate the roles of miR-449c in the pathogenesis of osteosarcoma. Our results demonstrated that miR-449c direct targets the oncogene c-Myc and downregulation of miR-449c in osteosarcoma cancerous tissues and osteosarcoma cells resulted in activation of c-Myc and its downstream targets, including Cyclin D1, D2, CDK4, and CDK6. Abnormal expression of these cell cycle regulators led to osteosarcoma tumorigenesis. In addition, we also found that osteosarcoma cells had higher DNA methylation level in the two CpG islands adjacent to the miR-449c genomic locus, which suggested that DNA methylation might be important for the downregulation of miR-449c in osteosarcoma cells. Our results provide promising new insights for osteosarcoma therapy in the future.
Supplementary Material
Supplementary figures and tables.
Abbreviations
APC, adenomatous polyposis coli gene; AZA, 5-AZA-2'-deoxycytidine; CDK, cyclin-dependent kinase; miRNA-seq, microRNA sequencing; MSP, methylation-specific PCR; MSTS, musculoskeletal tumor stages; TSA, trichostatin A.
Acknowledgements
This study was supported by a grant from the National Natural Science Foundation of China (No.81601939).
Authors' contributions
C. Z and F. Z designed the research. Q. L and H. L performed most of the experiments. X. L, B. W and L. Z performed some parts of the experiments. Q. L and H. L analyzed data and performed statistical analyses. C. Z organized the figures and wrote the manuscript. All authors reviewed and approved the manuscript.
Ethical approval and informed consent
The clinical samples were acquired with written informed consent from all of the participants following protocols approved by the ethical board of Kunming Medical University. All experimental procedures used in this study were carried out in accordance with the approved guidelines of the ethical board of Kunming Medical University.
Competing Interests
The authors have declared that no competing interest exists.
References
1. Zhang B, Pan X, Cobb GP, Anderson TA. microRNAs as oncogenes and tumor suppressors. Dev Biol. 2007;302:1-12
2. Drakaki A, Iliopoulos D. MicroRNA Gene Networks in Oncogenesis. Curr Genomics. 2009;10:35-41
3. Hayes J, Peruzzi PP, Lawler S. MicroRNAs in cancer: biomarkers, functions and therapy. Trends Mol Med. 2014;20:460-69
4. Jansson MD, Lund AH. MicroRNA and cancer. Mol Oncol. 2012;6:590-610
5. Chen X, Duan N, Zhang C, Zhang W. Survivin and tumorigenesis: molecular mechanism and therapeutic strategies. J Cancer. 2016;7(3):314-23
6. Zhang F, Zhang L, Zhang C. Long noncoding RNAs and tumorigenesis: genetic associations, molecular mechanisms, and therapeutic strategies. Tumour Biol. 2016;37:163-75
7. Lu J, Getz G, Miska EA, Alvarez-Saavedra E, Lamb J, Peck D, Sweet-Cordero A. et al. MicroRNA expression profiles classify human cancers. Nature. 2005;435:834-38
8. Zhu J, Zheng Z, Wang J, Sun J, Wang P, Cheng X. et al. Different miRNA expression profiles between human breast cancer tumors and serum. Front Genet. 2014;5:149
9. Song C, Chen H, Wang T, Zhang W, Ru G, Lang J. Expression profile analysis of microRNAs in prostate cancer by next-generation sequencing. Prostate. 2015;75:500-16
10. Yang J, Han S, Huang W, Chen T, Liu Y, Pan S, Li S. A meta-analysis of microRNA expression in liver cancer. PLoS One. 2014;9:e114533
11. Murakami Y, Tanahashi T, Okada R, Toyoda H, Kumada T, Enomoto M. et al. Comparison of hepatocellular carcinoma miRNA expression profiling as evaluated by next generation sequencing and microarray. PLoS One. 2014;9:e106314
12. Zhang C, Zhang F. The multifunctions of WD40 proteins in genome integrity and cell cycle progression. J Genomics. 2015;3:40-50
13. Zhang C, Liu Y. Targeting cancer with sesterterpenoids: the new potential antitumor drugs. J Nat Med. 2015;69:255-66
14. Yang C, Sun C, Liang X, Xie S, Huang J, Li D. Integrative analysis of microRNA and mRNA expression profiles in non-small-cell lung cancer. Cancer Gene Ther. 2016;23:90-97
15. Novello C, Pazzaglia L, Cingolani C, Conti A, Quattrini I, Manara MC. et al. miRNA expression profile in human osteosarcoma: role of miR-1 and miR-133b in proliferation and cell cycle control. Int J Oncol. 2013;42:667-75
16. Caramuta S, Egyhazi S, Rodolfo M, Witten D, Hansson J, Larsson C, Lui WO. MicroRNA expression profiles associated with mutational status and survival in malignant melanoma. J Invest Dermatol. 2010;130:2062-70
17. Yanaihara N, Caplen N, Bowman E, Seike M, Kumamoto K, Yi M. et al. Unique microRNA molecular profiles in lung cancer diagnosis and prognosis. Cancer Cell. 2006;9:189-98
18. Pekarsky Y, Croce CM. Role of miR-15/16 in CLL. Cell Death Differ. 2015;22:6-11
19. Gougelet A, Pissaloux D, Besse A, Perez J, Duc A, Dutour A. et al. Micro-RNA profiles in osteosarcoma as a predictive tool for ifosfamide response. Int J Cancer. 2011;129:680-690
20. Zhao F, Lv J, Gan H, Li Y, Wang R, Zhang H. et al. MiRNA profile of osteosarcoma with CD117 and stro-1 expression: miR-1247 functions as an onco-miRNA by targeting MAP3K9. Int J Clin Exp Pathol. 2015;8:1451-58
21. Lulla RR, Costa FF, Bischof JM, Chou PM, de FBM, Vanin EF, Soares MB. Identification of Differentially Expressed MicroRNAs in Osteosarcoma. Sarcoma. 2011;2011:732690
22. Wang W, Zhang L, Zheng K, Zhang X. miR-17-5p promotes the growth of osteosarcoma in a BRCC2-dependent mechanism. Oncol Rep. 2016;35:1473-82
23. Tan X, Fan S, Wu W, Zhang Y. MicroRNA-26a inhibits osteosarcoma cell proliferation by targeting IGF-1. Bone Res. 2015;3:15033
24. Chen X, Chen XG, Hu X, Song T, Ou X, Zhang C, Zhang W. MiR-34a and miR-203 Inhibit Survivin Expression to Control Cell Proliferation and Survival in Human Osteosarcoma Cells. J Cancer. 2016;7:1057-65
25. Qu Y, Pan S, Kang M, Dong R, Zhao J. MicroRNA-150 functions as a tumor suppressor in osteosarcoma by targeting IGF2BP1. Tumour Biol. 2016;37:5275-84
26. Duan N, Hu X, Yang X, Cheng H, Zhang W. MicroRNA-370 directly targets FOXM1 to inhibit cell growth and metastasis in osteosarcoma cells. Int J Clin Exp Pathol. 2015;8:10250-60
27. Barter MJ, Bui C, Young DA. Epigenetic mechanisms in cartilage and osteoarthritis: DNA methylation, histone modifications and microRNAs. Osteoarthritis Cartilage. 2012;20:339-49
28. Chuang JC, Jones PA. Epigenetics and microRNAs. Pediatr Res. 2007;61:24R-29R
29. Chen Z, Sui J, Zhang F, Zhang C. Cullin family proteins and tumorigenesis: genetic association and molecular mechanisms. J Cancer. 2015;6:233-42
30. Zhang F, Zhao X, Shen H, Zhang C. Molecular mechanisms of cell death in intervertebral disc degeneration. Int J Mol Med. 2016;37:1439-48
31. Lopez-Serra P, Esteller M. DNA methylation-associated silencing of tumor-suppressor microRNAs in cancer. Oncogene. 2012;31:1609-22
32. Braconi C, Huang N, Patel T. MicroRNA-dependent regulation of DNA methyltransferase-1 and tumor suppressor gene expression by interleukin-6 in human malignant cholangiocytes. Hepatology. 2010;51:881-90
33. Nilsson JA, Cleveland JL. Myc pathways provoking cell suicide and cancer. Oncogene. 2003;22:9007-21
34. Dang CV. MYC on the path to cancer. Cell. 2012;149:22-35
35. Zhang C. Essential functions of iron-requiring proteins in DNA replication, repair and cell cycle control. Protein Cell. 2014;5:750-60
36. Zhang C, Zhang F. Iron homeostasis and tumorigenesis: molecular mechanisms and therapeutic opportunities. Protein Cell. 2015;6:88-100
37. Zhang C, Liu G, Huang M. Ribonucleotide reductase metallocofactor: assembly, maintenance and inhibition. Front Bio. 2014;9:104-13
38. Yin H, Song P, Su R, Yang G, Dong L, Luo M. et al. DNA Methylation mediated down-regulating of MicroRNA-33b and its role in gastric cancer. Sci Rep. 2016;6:18824
39. Sampson VB, Rong NH, Han J, Yang Q, Aris V, Soteropoulos P. et al. MicroRNA let-7a down-regulates MYC and reverts MYC-induced growth in Burkitt lymphoma cells. Cancer Res. 2007;67:9762-70
40. Sachdeva M, Zhu S, Wu F, Wu H, Walia V, Kumar S. et al. p53 represses c-Myc through induction of the tumor suppressor miR-145. Proc Natl Acad Sci U S A. 2009;106:3207-3212
41. Liu CG, Calin GA, Volinia S, Croce CM. MicroRNA expression profiling using microarrays. Nat Protoc. 2008;3:563-578
42. Zhang W, Duan N, Zhang Q, Song T, Li Z, Zhang C. et al. DNA methylation mediated down-regulation of miR-370 regulates cell growth through activation of Wnt/b-Catenin signaling pathway in human osteosarcoma cells. Int J Biol Sci. 2017;13:561-73
43. Chen Z, Wang K, Hou C, Jiang K, Chen B, Chen J. et al. CRL4BDCAF11 E3 ligase targets p21 for degradation to control cell cycle progression in human osteosarcoma cells. Sci Rep. 2017;7:1175
44. Zhang C, Guo H, Zhang J, Guo G, Schumaker KS, Guo Y. Arabidopsis cockayne syndrome A-like proteins 1A and 1B form a complex with CULLIN4 and damage DNA binding protein 1A and regulate the response to UV irradiation. Plant Cell. 2010;22:2353-69
45. Zhang Y, Li H, Zhang C, An X, Liu L, Stubbe J. et al. Conserved electron donor complex Dre2-Tah18 is required for ribonucleotide reductase metallocofactor assembly and DNA synthesis. Proc Natl Acad Sci U S A. 2014;111:E1695-704
46. Li H, Stumpfig M, Zhang C, An X, Stubbe J, Lill R, Huamg M. The diferric-tyrosyl radical cluster of ribonucleotide reductase and cytosolic iron-sulfur clusters have distinct and similar biogenesis requirements. J Biol Chem. 2017 doi: 10.1074/jbc.M117.786178
47. Stine ZE, Walton ZE, Altman BJ, Hsieh AL, Dang CV. MYC, Metabolism, and Cancer. Cancer Discov. 2015;5:1024-39
48. Han G, Wang Y, Bi W. C-Myc overexpression promotes osteosarcoma cell invasion via activation of MEK-ERK pathway. Oncol Res. 2012;20:149-56
Author contact
Corresponding authors: caiguo.zhangedu and spinezhangfcom

 Global reach, higher impact
Global reach, higher impact