Impact Factor ISSN: 1449-2288
Int J Biol Sci 2018; 14(12):1669-1677. doi:10.7150/ijbs.27819 This issue Cite
Research Paper
BERMP: a cross-species classifier for predicting m6A sites by integrating a deep learning algorithm and a random forest approach
1. School of Data Science and Software Engineering, Qingdao University, 266021, Qingdao, China
2. School of Basic Medicine, Qingdao University, 266021, Qingdao, China
3. Cancer institute, the Affiliated Hospital of Qingdao University, Qingdao, Shandong, 266061, China
4. Qingdao Cancer Institute, Qingdao, Shandong 266061, China
# Contributed equally.
Abstract
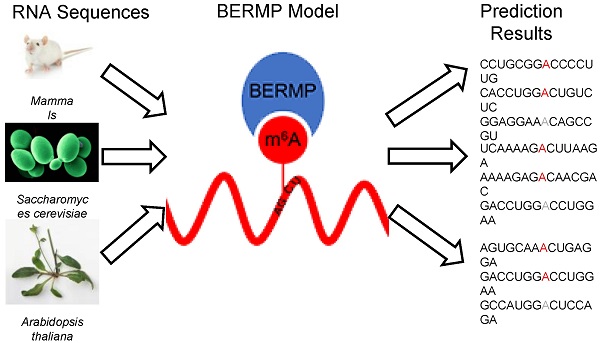
N6-methyladenosine (m6A) is a prevalent RNA methylation modification involved in several biological processes. Hundreds or thousands of m6A sites identified from different species using high-throughput experiments provides a rich resource to construct in-silico approaches for identifying m6A sites. The existing m6A predictors are developed using conventional machine-learning (ML) algorithms and most are species-centric. In this paper, we develop a novel cross-species deep-learning classifier based on bidirectional Gated Recurrent Unit (BGRU) for the prediction of m6A sites. In comparison with conventional ML approaches, BGRU achieves outstanding performance for the Mammalia dataset that contains over fifty thousand m6A sites but inferior for the Saccharomyces cerevisiae dataset that covers around a thousand positives. The accuracy of BGRU is sensitive to the data size and the sensitivity is compensated by the integration of a random forest classifier with a novel encoding of enhanced nucleic acid content. The integrated approach dubbed as BGRU-based Ensemble RNA Methylation site Predictor (BERMP) has competitive performance in both cross-validation test and independent test. BERMP also outperforms existing m6A predictors for different species. Therefore, BERMP is a novel multi-species tool for identifying m6A sites with high confidence. This classifier is freely available at
Keywords: Deep learning, Recurrent neural network, bidirectional Gated Recurrent Unit, N6-methyladenosine, Random forest

 Global reach, higher impact
Global reach, higher impact