Impact Factor ISSN: 1449-2288
Int J Biol Sci 2021; 17(6):1555-1564. doi:10.7150/ijbs.59191 This issue Cite
Research Paper
In silico structure-based discovery of a SARS-CoV-2 main protease inhibitor
1. State Key Laboratory of Emerging Infectious Diseases, Carol Yu Centre for Infection, Department of Microbiology, Li Ka Shing Faculty of Medicine, The University of Hong Kong, Pokfulam, Hong Kong Special Administrative Region, China.
2. Key Laboratory of Tropical Translational Medicine of Ministry of Education, Hainan Medical University, Haikou, Hainan, China.
3. Hainan-Medical University-The University of Hong Kong Joint Laboratory of Tropical Infectious Diseases, Hainan Medical University, Haikou, Hainan, and The University of Hong Kong, Pokfulam, Hong Kong Special Administrative Region, China.
4. Department of Pathogen Biology, Hainan Medical University, Haikou, Hainan, China.
5. School of Biomedical Sciences, Li Ka Shing Faculty of Medicine, The University of Hong Kong, Pokfulam, Hong Kong Special Administrative Region, China.
6. Department of Clinical Microbiology and Infection Control, The University of Hong Kong-Shenzhen Hospital, Shenzhen, Guangdong Province, China.
*Equal contribution.
Abstract
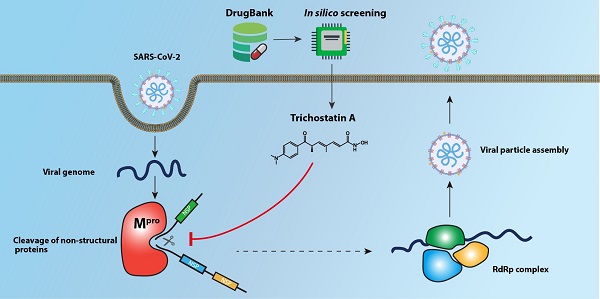
The Coronavirus Disease 2019 (COVID-19) pandemic caused by the novel lineage B betacoroanvirus severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) has resulted in significant mortality, morbidity, and socioeconomic disruptions worldwide. Effective antivirals are urgently needed for COVID-19. The main protease (Mpro) of SARS-CoV-2 is an attractive antiviral target because of its essential role in the cleavage of the viral polypeptide. In this study, we performed an in silico structure-based screening of a large chemical library to identify potential SARS-CoV-2 Mpro inhibitors. Among 8,820 compounds in the library, our screening identified trichostatin A, a histone deacetylase inhibitor and an antifungal compound, as an inhibitor of SARS-CoV-2 Mpro activity and replication. The half maximal effective concentration of trichostatin A against SARS-CoV-2 replication was 1.5 to 2.7µM, which was markedly below its 50% effective cytotoxic concentration (75.7µM) and peak serum concentration (132µM). Further drug compound optimization to develop more stable analogues with longer half-lives should be performed. This structure-based drug discovery platform should facilitate the identification of additional enzyme inhibitors of SARS-CoV-2.

 Global reach, higher impact
Global reach, higher impact