Impact Factor
ISSN: 1449-2288
Int J Biol Sci 2022; 18(9):3762-3776. doi:10.7150/ijbs.71385 This issue Cite
Research Paper
Establishment and Validation of an Interferon-Stimulated Genes (ISGs) Prognostic Signature in Pan-cancer Patients: A Multicenter, Real-world Study
1. Department of Thoracic Surgery, National Cancer Center/National Clinical Research Center for Cancer/Cancer Hospital, Chinese Academy of Medical Sciences and Peking Union Medical College, Beijing, China.
2. Department of Colorectal Surgery, Fudan University Shanghai Cancer Center, Shanghai 200032, China.
3. Department of Colorectal Surgery, National Cancer Center/National Clinical Research Center for Cancer/Cancer Hospital, Chinese Academy of Medical Sciences and Peking Union Medical College, Beijing 100021, China.
4. Department of Pharmacology, Nanjing University of Chinese Medicine, Nanjing, Jiangsu, 210023, China.
#These authors contributed equally to this work.
Abstract
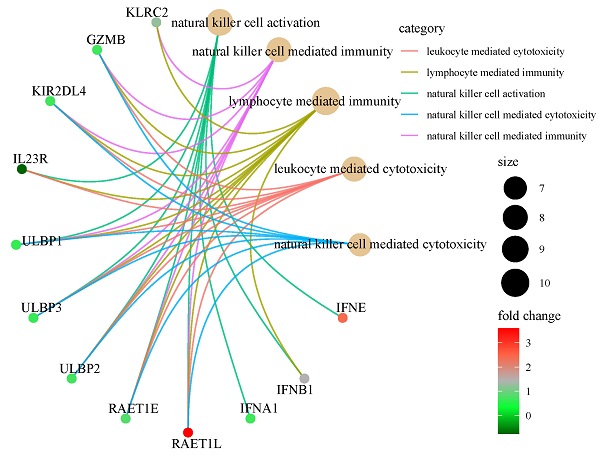
Our study aims at developing an interferon-stimulated genes (ISGs) signature that could predict overall survival (OS) in cancer patients, which enrolled a total of 5643 pan-cancer patients. Linear models for microarray data method analysis were conducted to identify the differentially expressed prognostic genes in the global ISGs family. Time-dependent receiver operating characteristic (ROC) and Kaplan-Meier survival analysis were used to test the efficiency of a multi-gene signature in predicting the prognosis of pan-cancer patients. The prognostic performance and potential biological function of gene signature were verified by quantitative real-time PCR in a pan-cancer independent cohort. Three ISGs genes were finally identified to build a classifier, a specific risk score formula, with which patients were classified into the low- or high-risk groups. Time-dependent ROC analyses proved prognostic accuracy. Then, its prognostic value was validated in seven external validation series. A nomogram was constructed to guide the individualized treatment of patients with lung adenocarcinoma. Biological pathway and tumor immune infiltration analysis showed that the signature might cause poor prognosis by blocking NK cell activation. Finally, the signature in our centers was confirmed by real-time quantitative PCR. A robust ISGs-related feature was discovered to effectively classify pan-cancer patients into subgroups with different OS.
Keywords: Natural Killer Cells, Tumor Biomarkers, Interferon-stimulated Genes, Pan-cancer Study

 Global reach, higher impact
Global reach, higher impact