Impact Factor ISSN: 1449-2288
Int J Biol Sci 2026; 22(7):3564-3579. doi:10.7150/ijbs.129521 This issue Cite
Review
The Quartet of Core Oncogenic Drivers in Neuroendocrine Prostate Cancer: Multi-Omics Dataset Integration to Forge a Translational Link Between Biology and Precision Therapy
1. Department of Urology, Peking University Third Hospital, Beijing, 100191, China.
2. State Key Laboratory of Female Fertility Promotion, Center for Reproductive Medicine, Department of Obstetrics and Gynecology, Center for Reproductive Medicine, Peking University Third Hospital, Beijing, 100191, China.
3. Department of Urology, Tianjin Institute of Urology, The Second Hospital of Tianjin Medical University, Tianjin, 300211, China.
4. Department of Urology, The Affiliated Hospital of Xuzhou Medical University, Xuzhou, 221004, China.
# These authors contributed equally to this work.
Received 2025-12-5; Accepted 2026-3-2; Published 2026-3-25
Abstract
Prostate cancer (PC) is the most common cancer among American men and the second leading cause of cancer-related deaths. For advanced or metastatic PC, anti-androgen therapies, including androgen deprivation therapy (ADT), are considered standard treatment options. However, these therapies often result in the development of castration-resistant prostate cancer (CRPC) or neuroendocrine prostate cancer (NEPC), both of which present significant treatment challenge. The molecular mechanisms driving the progression from androgen - sensitive PC to castration-resistant and neuroendocrine phenotypes are still being actively investigated. This review aims to comprehensively evaluate the cellular and molecular mechanisms underlying the development of NEPC. Specifically, it will focus on the roles of cancer stem cells (CSCs), epithelial - mesenchymal transition (EMT), and autophagy in the pathogenesis and progression of NEPC. Furthermore, the review will explore the potential of targeting these processes for therapeutic intervention in advanced P. This review will integrate current findings from clinical trials, pre-clinical models, and molecular research to clarify the promising approaches for improving treatment outcomes for patients with advanced PC.
Keywords: PC, NEPC, EMT, CSCs, autophagy, lineage plasticity
Introduction
Prostate cancer (PC) is one of the most frequently diagnosed malignancies among men over the age of 50 in the United States and represents a growing global health burden [1-3]. Despite substantial advances in therapeutic strategies, PC remains a major cause of cancer-related morbidity and mortality in urology [4-6]. ADT is the cornerstone of treatment for advanced or metastatic PC. Although most patients initially respond to ADT, durable disease control is uncommon, and the majority eventually experience disease progression due to the development of therapeutic resistance [7-9]. This stage of disease progression is referred to as CRPC. Resistance to androgen receptor (AR)-targeted therapy can arise through multiple mechanisms, including AR amplification or mutation, the emergence of constitutively active AR splice variants such as AR variant 7 (AR-V7), and activation of bypass signaling pathways, including glucocorticoid receptor (GR)-dependent programs [10-12]. Prolonged suppression of AR signaling has also been associated with phenotypic alterations in PC cells, including lineage switching and histological dedifferentiation toward neuroendocrine differentiation (NED) and/or EMT [13-15]. This adaptive process is commonly referred to as treatment-induced lineage plasticity or lineage crisis [16-18]. Clinical and experimental observations indicate that prostate tumors with low AR activity often display increased cellular heterogeneity and enrichment of stem-like states, accompanied by activation of cell cycle and survival programs that contribute to aggressive clinical behavior [19, 20]. Although the precise molecular mechanisms through which androgen receptor pathway inhibitors (ARPIs) promote lineage plasticity remain incompletely understood, accumulating evidence supports cellular plasticity as a key adaptive resistance strategy under sustained therapeutic pressure [21, 22].
Autophagy, a conserved cellular stress - response and recycling pathway, has emerged as an additional process that is potentially relevant to the biology of advanced PC and NEPC [23, 24]. Significantly, the current evidence does not support autophagy as a unidirectional promoter or suppressor of NEPC [25, 26]. Instead, it seems to function as a context-dependent, stress-adaptive process that may permissively support lineage plasticity and therapy resistance [27-29]. Remarkably, autophagy has been reported to have context - dependent effects during cancer progression, acting as either a tumor-suppressive or tumor-supportive mechanism depending on the disease stage and microenvironmental conditions [30-32]. In advanced tumors exposed to metabolic, oxidative, or therapeutic stress, autophagy may facilitate cellular adaptation by supporting survival, metabolic flexibility, and persistence. Nevertheless, its precise functional role in NEPC remains an area of active research [33-35].
Importantly, emerging evidence suggests that neuroendocrine prostate tumors represent a spectrum of differentiation states characterized by variable expression of neuroendocrine markers, EMT - associated features, and CSC - related properties [36, 37]. Rather than functioning as independent or deterministic causal drivers, EMT programs, CSC - associated states, and autophagy - related processes are increasingly recognized as context-dependent regulatory and permissive mechanisms that may facilitate lineage plasticity, neuroendocrine differentiation, and therapy resistance during NEPC evolution [38-40].
Accordingly, this review aims to synthesize current clinical, preclinical, and molecular evidence to clarify how EMT-, CSC-, and autophagy-associated regulatory programs intersect with NEPC development and progression [41, 42]. By focusing on their functional involvement within a broader network of lineage plasticity rather than proposing linear causal models, we seek to provide a balanced framework for understanding NEPC biology and for exploring potential therapeutic vulnerabilities in advanced PC [43-45].
Mechanisms of Therapy Resistance and NE Phenotype in PC
Hormonal therapy demonstrates initial efficacy in nearly all PC patients, with chemical castration typically achieved using agents that inhibit testicular androgen secretion, such as gonadotropin-releasing hormone (GnRH) agonists or antagonists [46, 47]. Nevertheless, this treatment is palliative, and castration resistance develops over time [48, 49]. Despite low androgen levels, this progression primarily occurs because PC cells evade ADT by restoring AR signaling through genetic alterations [50, 51]. The AR locus is associated with various AR-dependent resistance mechanisms, such as genomic amplification, AR mutations that transform antiandrogens into agonists, and AR transcription factor variants missing the carboxy-terminal ligand-binding domain, enabling AR signaling without ligand binding [52-54].
With the development of more potent ARPIs (e.g., abiraterone and enzalutamide) for CRPC treatment, additional resistance mechanisms have emerged [55, 56]. For example, increased levels and/or activity of the GR enable bypass of the AR-blocking effect of enzalutamide without the need to restore AR function [57, 58]. By binding to canonical AR binding sites, GR activates transcriptional programs similar to those regulated by AR [59, 60]. A study reported that GR protein expression was detected in 30% (8 out of 22) of tumors following enzalutamide treatment, compared to only 10% (3 out of 22) before treatment [61, 62]. Elevated GR expression following enzalutamide treatment correlated with unfavorable clinical outcomes. These data suggest that GR-mediated bypass acts as an adaptive resistance mechanism when the AR pathway is blocked [63-65].
Lineage switching is recognized as a third mechanism for PC cells to evade AR pathway inhibition. In prostate cancer, 20-25% of metastatic castration-resistant prostate cancer (mCRPC) patients treated with ARPIs relapse with tumor cells exhibiting neuroendocrine characteristics, suggesting that lineage plasticity plays a role in antiandrogen resistance [66-69]. Two primary hypotheses explain the origin of NEPC: one proposes that genetic or epigenetic dysregulation drives the transdifferentiation of adenocarcinomas into the NE lineage, while the other suggests that oncogenic mutations arise in normal NE cells [70-72]. Regardless of its origin, NEPC's metastatic and highly aggressive nature often requires platinum-based chemotherapy, administered concurrently with small-cell carcinoma treatments [73-75]. Even with chemotherapy, these tumors typically lead to patient death within two years of diagnosis (Figure 1).
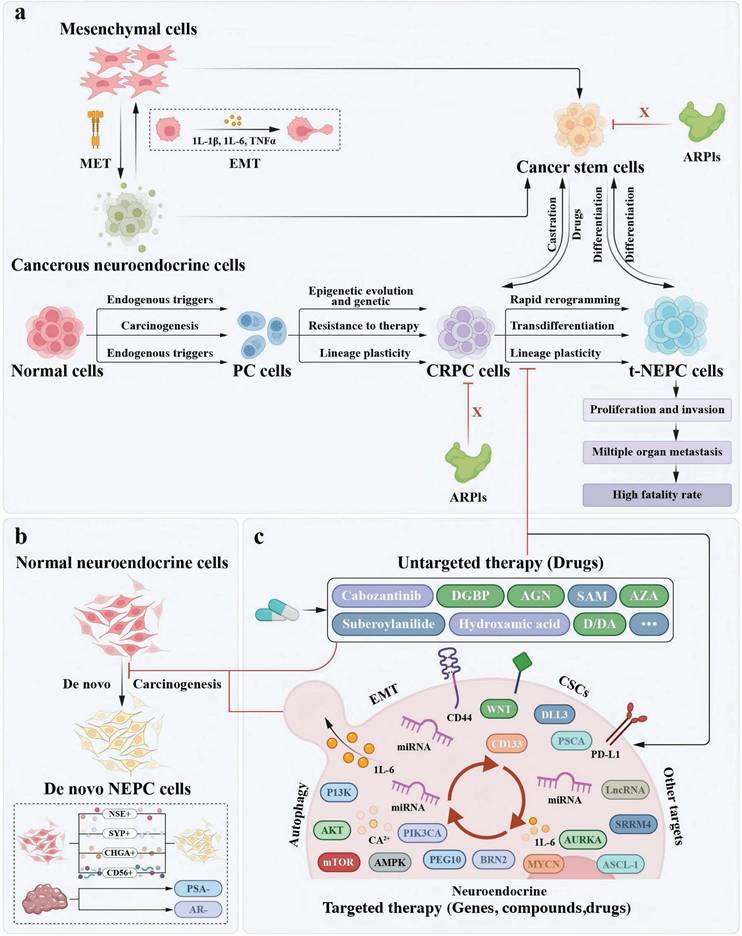
The origin of two models of NEPC development and the diagram of integrated treatment for NE progression. a. In the epigenetic and/or genetic dysregulation that causes PC/CRPC to transdifferentiate into the NE lineage and NEPC hierarchical model, tumours are heterogeneous, that is, they contain a mix of proliferative-type and stem-like cells. CSCs and/or NE cells cannot be eliminated by ARPIs to reduce tumour burden. These cell types can differentiate into NEPC through their plasticity. A partial EMT induces a plastic phenotype, which enables transdifferentiation to proceed via a transient pluripotent and stem-like state to generate CSCs. CRPC cells and CSCs can be interconverted by drugs and castration. Lethal NEPC induced by the interaction of cells further accelerates the proliferation and invasive activity of cancer cells and metastasis to multiple internal organs, ultimately leading to high mortality in patients. b. The de novo transformation of normal NE cells into NEPC. c. Mitigating the progression of NEPC by the agents that target EMT, NED, CSCs, and autophagy as well as other untargeted therapy drugs. Mitigating the progression of NEPC by other targets such lncRNA, miRNA, SRRM4, etc.
EMT, CSCs, and NED
EMT program is associated with PC progression
Accumulating evidence indicates that PC cells frequently engage EMT-associated transcriptional and phenotypic programs that are associated with malignant traits, including enhanced invasiveness, therapeutic tolerance, and migratory capacity [76, 77]. Importantly, current evidence does not support a simple linear model in which EMT-related molecules either directly induce NED or merely emerge as passive markers after NED is established [78, 79]. In clinical settings, EMT-related features have been correlated with bone and lymph node metastasis, higher Gleason scores, and shortened time to biochemical recurrence, highlighting their relevance to aggressive disease phenotypes [80, 81].
Consistent with this observation, elevated expression of N-cadherin (CDH2) has been reported in CRPC cell lines and in tumors exhibiting low AR pathway activity, a feature commonly associated with AR-independent cellular states [82, 83]. Functional studies further demonstrate that monoclonal antibodies targeting N-cadherin can suppress invasion, metastasis, and castration-resistant growth in CRPC xenograft models, supporting a contributory role for EMT-associated molecules in advanced disease progression [84, 85].
Therapeutic suppression of AR signaling has also been linked to changes in cellular state associated with EMT and stem-like properties [86, 87]. ADT has been shown to correlate with increased expression of AR splice variants, such as AR-V7, alongside enrichment of stemness-associated markers and EMT-related transcripts [88, 89]. In AR3Tg transgenic mouse models, an increased population of prostate progenitor cells within the prostate epithelium (defined as LIN⁻SCA-1⁺CD49f^high cells) has been observed, suggesting that altered AR signaling may create a permissive context for disrupted differentiation programs rather than directly inducing EMT [90, 91].
In clinical practice, circulating tumor cells (CTCs) detected in patients with CRPC undergoing treatment with enzalutamide and/or abiraterone frequently display phenotypic heterogeneity and have been associated with resistance to ARPIs [92, 93]. These observations are consistent with a model in which sustained AR suppression is accompanied by the emergence or stabilization of EMT- and stemness-associated cellular states [94-96].
From a therapeutic perspective, strategies targeting AR signaling have been explored for their potential to limit these adaptive phenotypes [97, 98]. Preclinical studies demonstrate that galeterone, an AR degrader, can reduce the expression of EMT- and stem cell-associated markers and exhibit antitumor activity [99, 100]. However, clinical studies indicate that in patients with AR-V7-positive CRPC, galeterone did not confer a progression-free survival benefit compared with enzalutamide, underscoring the complexity of targeting EMT- and stemness-associated programs through AR-directed approaches alone [17, 101, 102].
CSC program is associated with PC progression
Prostate stem cells (PSCs), which reside in both the luminal and basal compartments of the prostate epithelium, have been proposed as potential cells of origin for PC due to their long-lived nature and susceptibility to oncogenic alterations [103-105]. In advanced disease, particularly metastatic CRPC, prostate tumors frequently harbor genetic alterations in key tumor suppressors such as TP53, RB1, and PTEN, highlighting their central role in disease progression [106-108].
Experimental studies indicate that combined genetic perturbations, rather than single events, are required to create cellular states permissive for aggressive behavior. For example, the concurrent loss of PTEN and activation of RAS signaling has been shown to induce partial EMT-like features in PSCs and CSC-enriched populations, endowing these cells with increased plasticity and metastatic potential [109-111]. In contrast, PTEN inactivation alone is generally insufficient to drive metastatic progression, underscoring the importance of cooperative genetic contexts [112-114].
Importantly, the combined inactivation of PTEN, TP53, and RB1 has been consistently associated with enhanced lineage plasticity in PC cells. Rather than directly initiating NEPC, these alterations appear to establish a permissive cellular and epigenetic context in which differentiation programs are destabilized. In this setting, impaired differentiation of PSCs and CSC-enriched populations facilitates phenotypic switching, increases metastatic competence, and supports the emergence of therapy-resistant states [115, 116].
Mechanistically, TP53 and RB1 have been shown to restrain lineage plasticity, at least in part, through suppression of pluripotency-associated transcriptional programs such as SOX2. Loss of these regulatory constraints promotes stabilization of stem-like and plastic cellular states, which can subsequently cooperate with additional signaling pathways and environmental pressures to shape advanced disease phenotypes, including NEPC [117, 118].
EMT- and CSC-associated transcriptomic programs overlap with NEPC
Multiple classical signaling pathways, including IL-6 and WNT, have been implicated in the regulation of cellular plasticity and NED in prostate cancer. Rather than operating through isolated linear cascades, these pathways converge on EMT- and CSC-associated transcriptional programs, shaping dynamic gene expression states that are permissive for lineage plasticity [119, 120]. In this context, EMT- and CSC-related processes are increasingly viewed as intersecting regulatory modules that coexist with, and interact alongside, neuroendocrine transcriptional programs [121-123].
Transcriptomic analyses have revealed that EMT-associated genes, such as TWIST, SNAIL, and VIM, as well as CSC-related markers including NOTCH1, CD133, and KIT, are frequently enriched in prostate tumors exhibiting neuroendocrine features [124, 125]. At the same time, canonical neuroendocrine markers—such as CHGA, SYP, and NSE, widely used in pathological and clinical settings—are elevated in NEPC. These observations support the concept that EMT, CSC, and neuroendocrine transcriptional signatures substantially overlap, rather than representing mutually exclusive or sequential states [126-128].
Several signaling mediators appear to bridge these transcriptional programs. For example, IL-6 and CD44 have been reported to correlate with EMT-associated, CSC-associated, and neuroendocrine gene expression patterns, and are consistently linked to increased cellular plasticity in PC models. Importantly, these factors are best interpreted as shared regulatory nodes within a broader plasticity network, rather than as singular determinants of neuroendocrine fate [129, 130].
To further examine the molecular intersections among EMT, CSC, and neuroendocrine transcriptional programs, we performed a systematic literature survey of PubMed-indexed studies published between 2013 and 2023 and identified 38 relevant reports (Supplementary Figure 1). Across these studies, overlapping signaling pathways, transcription factors, and cell surface receptors were repeatedly observed in models of epithelial plasticity and neuroendocrine differentiation [131-133]. For instance, CD44 has been associated with enhanced NED in PC cell lines (PC3 and LNCaP) and in patient-derived NEPC tissues. Similarly, suppression of GSN was shown to reduce IL-6 expression and attenuate NED in LNCaP cells, while IL-6-associated activation of MAPK signaling facilitates formation of the TGF-β/SMAD2 complex and is linked to neuroendocrine differentiation in NCI-H660 cells [134-136].
In cell-derived xenograft models of neuroendocrine transdifferentiation, upregulation of EMT-associated transcription factors encoded by PEG10 has been observed during the transition from adenocarcinoma to neuroendocrine-like states in LNCaP-derived and CRPC tissues [137, 138]. Collectively, these findings do not support a single linear causal pathway but instead point to redundant, cooperative, and context-dependent mechanisms through which EMT-, CSC-, and neuroendocrine-associated transcriptional programs intersect to shape lineage plasticity and phenotypic diversity in advanced PC (Figure 2). In this framework, EMT-associated programs may precede, accompany, or stabilize neuroendocrine differentiation depending on genetic background, therapeutic pressure, and microenvironmental context, rather than functioning as a unidirectional trigger or a late-stage byproduct of NED [139, 140].
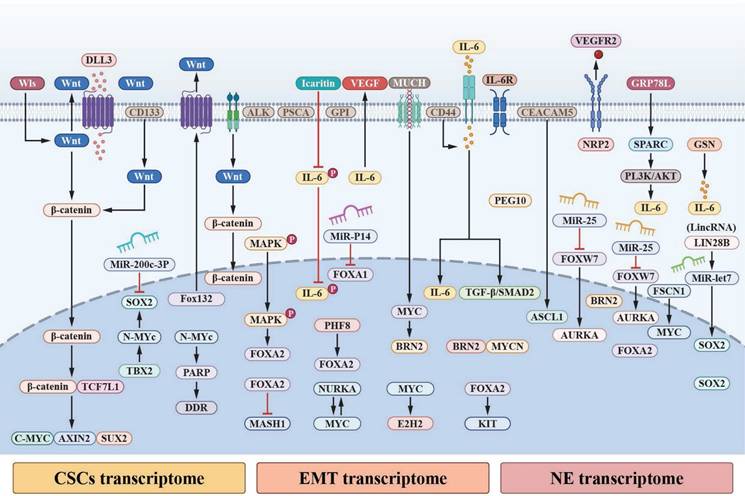
Pivotal signalling nodes and pathways of mediating cellular plasticity and NED aggregation. a. A convergence of WNT-β-catenin signalling pathway and IL-6 signal transducer and activator of STAT3 pathway occurs during EMT, CSCs, and NED to sustain plasticity and transcriptome dynamics. Through the interaction of IL-6 with its receptor, IL-6R, Janus Kinases (JAKs) are activated, such as JAK1 and JAK2, which in turn phosphorylate STAT3 at Tyr705; as a result, activated STAT3 dimers are translocated to the nucleus. The transcription of genes associated with NED, CSCs, and EMT are mediated by STAT3 in the nucleus by interacting with γ-activated sequences (GASs) and interferon-stimulated response elements (ISREs). Canonical WNT signalling further activate STAT3; Dishevelled (DVL) associates with the Frizzled (FZD) receptor to sequester glycogen synthase kinase-3β (GSK3β), thereby enabling the stabilization of β-catenin and its translocation to the nucleus. In the nucleus, β-catenin associates with members of the T cell factor (TCF)-lymphoid enhancer factor (LEF) transcription factor family. Activation of WNT-responsive genes is mediated by β-catenin's association with TCF-LEF transcription factors, such as STAT3. Additionally, activated FZD can activate the tyrosine-protein kinase FYN during non-canonical WNT signalling; activated FYN recruits and phosphorylates STAT3 at Tyr705 to accelerate STAT3 translocation into the nucleus and transcription. b. Collected signaling pathways converge and single or multiple regulate the three transcriptomes to influence NED and intertwine with IL-6 and WNT signaling pathways.
Autophagy as a Stress-Adaptive Program in NEPC
miRNA-mediated regulation of autophagy and NED
Autophagy is a conserved cellular stress-adaptation process that enables cells to maintain metabolic and proteostatic homeostasis under adverse conditions and has been reported to exert context-dependent, and sometimes opposing, effects during cancer progression [141-143]. In the context of advanced PC and NEPC, autophagy has attracted increasing attention because tumor cells are frequently exposed to androgen deprivation, metabolic stress, hypoxia, and therapeutic pressure. Rather than serving solely as a housekeeping mechanism, autophagy in these settings is increasingly viewed as a context-dependent adaptive program that may support cellular persistence and phenotypic flexibility [144-146].
MicroRNAs (miRNAs) have emerged as important post-transcriptional regulators that fine-tune autophagy-related signaling networks, particularly under stress conditions relevant to lineage plasticity [147-149]. Autophagy-associated genes are regulated at multiple stages—from initiation to lysosomal degradation—and numerous components of this machinery are subject to miRNA-mediated control [150, 151]. Stress-induced alterations in miRNA expression can therefore modulate autophagic activity and influence downstream cellular states [152-154].
In PC models, environmental stressors such as nutrient deprivation, hypoxia, and oxidative stress—conditions frequently encountered during androgen receptor pathway inhibition—have been shown to reshape miRNA expression profiles [155, 156]. These changes are closely linked to activation of autophagy-related pathways and are increasingly associated with NED and therapy-adaptive phenotypes, rather than with autophagy in isolation [157, 158].
Although thousands of miRNAs have been identified, only a subset has been implicated in autophagy regulation with relevance to prostate cancer progression [159, 160]. Several miRNAs, including miR-199a-3p, miR-100, miR-128, miR-338-3p, miR-7, and miR-96, have been reported to influence autophagic activity by targeting mTOR, a central metabolic checkpoint frequently dysregulated in advanced PC [161-163]. In addition, miRNA-mediated modulation of upstream regulators such as AKT signaling components, the calcium sensor CaMKKβ, and energy sensors including LKB1 and AMPK provides a mechanistic link between metabolic stress, autophagy regulation, and lineage plasticity [164, 165]. Taken together, these findings suggest that miRNAs function as critical integrative nodes that connect stress-responsive autophagy programs with transcriptional and phenotypic changes associated with NEPC, rather than acting as generic regulators of autophagy alone [166, 167].
Accumulating evidence suggests that miRNAs are involved in NED-associated transcriptional reprogramming in PC, primarily by modulating stress-responsive signaling networks rather than acting as isolated initiating factors. Several miRNAs have been reported to associate with NEPC-related phenotypes and lineage plasticity in experimental and clinical contexts [168].
For example, miR-421-mediated regulation of ATM has been linked to NEPC progression, while the LIN28B/let-7/SOX2 axis has been implicated in shaping transcriptional networks associated with stem-like and plastic cellular states in NEPC [169, 170]. Inhibition of miR-194 has been reported to attenuate epithelial-to-neuroendocrine-like differentiation in PC models, suggesting that specific miRNAs may modulate, rather than directly determine, neuroendocrine phenotypes. In NE-like LNCaP cells, altered expression of miR-20b, miR-106b, and miR-17 has been associated with changes in AKT3 biosynthesis, further linking miRNA dynamics to signaling adaptations observed during NED [171, 172].
Lineage switching toward NE-associated states has also been correlated with coordinated changes in miRNA expression, including upregulation of miR-301a and miR-375 and downregulation of the miR-106a~363 cluster. Importantly, these miRNA alterations are best interpreted as part of a broader transcriptional reprogramming process that accompanies NED, rather than as singular causal triggers [173].
Notably, several miRNAs implicated in autophagy regulation—such as let-7a, miR-17, miR-20a, miR-106a, miR-106b, and miR-143—have also been reported in association with NED in PC, highlighting a potential regulatory intersection between autophagy, metabolic adaptation, and lineage plasticity [174, 175]. In advanced-stage TRAMP prostate tumors, AKT1 has been shown to influence the expression of multiple miRNAs (including let-7a, miR-10a, miR-143, and miR-145a), which are linked to metastatic and EMT-associated phenotypes (Table 1).
MiRNA known to regulate autophagy and are associated with NEPC
| miRNA | Effect on autophagy | Autophagy-related target | Autophagy-related stimuli | Cell lines | Ref |
|---|---|---|---|---|---|
| MIRLET7A | Induction | RICTOR | N.D. | MGC-803 and SGC-7901 | [169] |
| MIR17 | Suppression | BECLIN1, BCL-2 | Paclitaxel | A549, A549-T24, H596-TxR, H1734, H1734-T5, H1299, H1299-Tx | [170, 171] |
| MIR20A | Suppression | ATG16L1 | Hypoxia | RAW264.7 | [172] |
| MIR106A | Suppression | ATG7 | N.D. | HCT116, SW620 | [173] |
| MIR106B | Suppression | ATG16L1 | Escherichia coli | HCT116, SW480, HeLa, U2OS | [174] |
| MIR143 | Suppression | ATG7 | Hydrogen peroxide | Mice cardiac progenitor cells | [175] |
In models of neuroendocrine transdifferentiation, particularly LNCaP-derived systems, integrated analyses reveal that coordinated changes in both miRNAs and mRNAs contribute to phenotypic plasticity. Members of the miR-17 family (miR-106b, miR-106a, miR-20b, miR-17, and miR-20a) have been repeatedly highlighted in this context. Consistent with the heterogeneous nature of treatment-emergent NEPC, clinical data further indicate that CRPC-NE more frequently exhibits alterations in miR-143 compared with adenocarcinoma, while t-NEPC samples display substantial intertumoral heterogeneity, including differences in extracellular vesicle-derived miRNA profiles (Figure 3; Supplementary Figure 2).
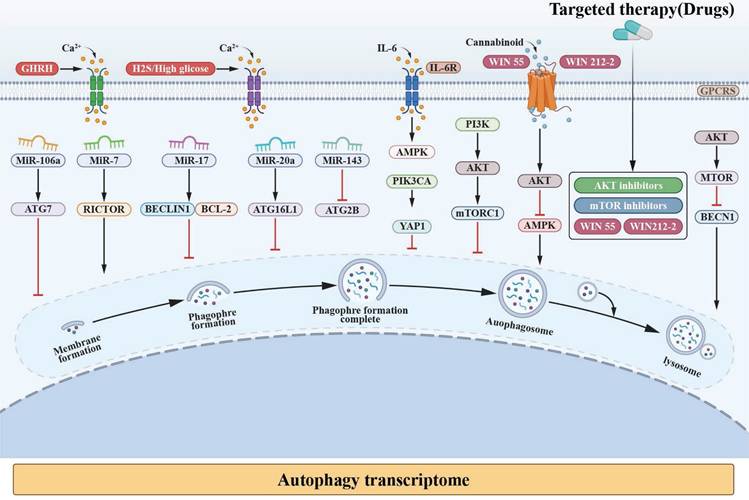
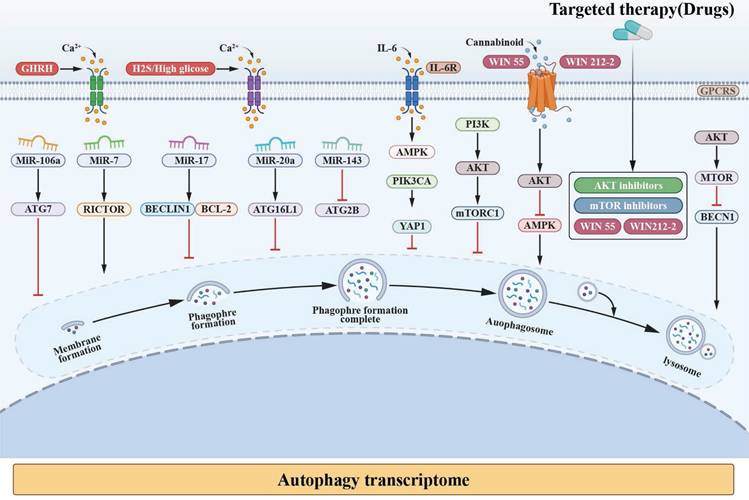
Potential relationship between autophagy and NED in PC. In the formation of autolysosome, an endless stream of mechanisms are involved. The key regulators, such as miRNA, PI3K-AKT-mTOR, Ca²⁺, AMPK, and PIK3CA, which rolling in the progress of autophagy and NED in PC are interviewed.
PI3K/AKT/mTOR-modulated autophagy under therapeutic stress
Autophagy is tightly regulated by nutrient- and energy-sensing pathways, among which the PI3K/AKT/mTOR signaling axis functions as a central metabolic checkpoint [176, 177]. In advanced PC, particularly under conditions of androgen deprivation and therapeutic stress, dysregulation of this pathway has been repeatedly implicated in adaptive survival responses and lineage plasticity [178-180].
In PC models, suppression of mTORC1 activity—whether through metabolic stress, androgen receptor pathway inhibition, or pharmacologic intervention—has been consistently associated with activation of autophagy-related programs [181, 182]. This response is increasingly interpreted as a stress-adaptive mechanism rather than a constitutive oncogenic driver [183, 184]. In this context, autophagy may support cellular persistence by maintaining metabolic homeostasis and buffering proteotoxic and oxidative stress, conditions frequently encountered during treatment-emergent NED [185, 186].
Importantly, PI3K/AKT/mTOR signaling intersects with multiple regulatory layers relevant to NEPC biology, including EMT-associated programs, cancer stem-like states, and miRNA-mediated post-transcriptional control [187, 188]. Aberrant activation of PI3K/AKT signaling—often arising from PTEN loss—has been linked to therapy resistance and lineage plasticity, while downstream modulation of mTOR activity influences autophagic flux and cellular stress tolerance [189, 190]. These interactions suggest that PI3K/AKT/mTOR-regulated autophagy participates in a broader plasticity-supporting network, rather than acting as an isolated pathway [185, 186].
Although numerous signaling cascades have been reported to modulate autophagy in cancer cells, their relevance to NEPC is highly context dependent [191, 192]. In this review, we focus specifically on the PI3K/AKT/mTOR axis because of its frequent alteration in advanced PC and its documented interaction with therapeutic pressure-induced phenotypic adaptation [193, 194]. Pharmacological agents targeting this pathway can either enhance or suppress autophagy depending on treatment context, dosage, and cellular state, underscoring the complexity of autophagy modulation in NEPC-relevant settings (Table 2).
Compounds and genes known to regulate autophagy and are associated with NEPC
| Compounds and genetic methods | Effect | Ref |
|---|---|---|
| PI3K/AKT/mTOR signaling pathway | ||
| 3-methyladenine | A PtdIns3K inhibitor that effectively blocks an early stage of autophagy by inhibiting the class III PtdIns3K, but not a specifific autophagy inhibitor. 3-MA also inhibits the class I PI3K and can thus, at suboptimal concentrations in long-term experiments, promote autophagy in some systems, as well as affect cell survival through AKT and other kinases. 3-MA does not inhibit BECN1-independent autophagy. | [226, 227] |
| 10-NCP | 10-(40-N-diethylamino)butyl)-2-chlorophenoxazine; an AKT inhibitor that induces autophagy in neurons. | [228, 229] |
| Akti-1/2 | An allosteric inhibitor of AKT1 and AKT2 that promotes autophagy in B-cell lymphoma. | [230] |
| AZD8055 | A catalytic MTOR inhibitor that acts as a potent autophagy inducer. | [231] |
| Clonidine | Activates the imidazoline receptor, which decrease cAMP in cells. An MTOR-independent inducer of autophagy. | [232] |
| Eriocalyxin B | An autophagy inducer that exerts anti-tumor activity in breast cancer by inhibition of the AKT-MTOR-RPS6KB signaling pathway. | [233] |
| ESC8 | A cationic estradiol derivative that induces autophagy and apoptosis simultaneously by downregulating the MTOR kinase pathway in breast cancer cells. | [234, 235] |
| Everolimus | An inhibitor of MTORC1 that induces both autophagy and apoptosis in B-cell lymphoma primary cultures. | [236] |
| Ezetimibe | A cholesterol absorption inhibitor that acts by binding to NPC1L1, which induces autophagy via MTORC1-dependent and independent pathways. Ezetimibe also activates TFEB and could potentially exert therapeutic effects on steatohepatitis and fibrosis. | [237] |
| Fasudil | An inhibitor of ROCK (Rho associated coiled-coil containing protein kinase) enhancing autophagy via phosphorylation of MAPK8/JNK1 and BCL2, and promoting BECN1-PIK3C3/VPS34 complex formation; shRNA-mediated approaches to inhibiting ROCK have similar results. | [238] |
| KU-0063794 | An MTOR inhibitor that binds the catalytic site and activates autophagy. | [239] |
| MLN4924 | A small molecule inhibitor of NAE (NEDD8 activating enzyme); induces autophagy by blockage of MTOR activity via both DEPTOR and the HIF1A-DDIT4/REDD1-TSC1/2 axis as a result of inactivation of cullin-RING ligases. | [240] |
| NVP-BEZ235 | A dual inhibitor of PIK3CA/p110 and the MTOR catalytic site that activates autophagy. | [241] |
| Rapamycin | Inhibits MTOR by binding to RPTOR, thus inducing autophagy, but only provides partial inhibition. | [242] |
| SMER28 | An MTOR-independent inducer of autophagy. | [243] |
| TMS | Trans-3,5,4-trimethoxystilbene upregulates the expression of TRPC4, resulting in MTOR inhibition. | [244] |
| Torin1 | A catalytic MTOR inhibitor that induces autophagy and provides more complete inhibition than rapamycin (it inhibits all forms of MTOR). | [245] |
| Wortmannin | An inhibitor of PtdIns 3-kinase that blocks autophagy, but is not a specific inhibitor. | [246] |
| miRNA signaling pathway | ||
| Knockdown | This method (including miRNA, RNAi, shRNA and siRNA) can be used to inhibit gene expression and provides relatively direct evidence for the role of an autophagic component. However, the efficiency of knockdown varies, as does the stability of the targeted protein. In addition, more than one gene involved in autophagy should be targeted to avoid misinterpreting indirect effects. | [247] |
| microRNA | Can be used to reduce the levels of target mRNA(s) or block translation. | [248] |
| Other signaling pathways (calcium, AMPK, PIK3CA, etc) | ||
| Calcium | An autophagy activator that can be released from ER or lysosomal stores under stress conditions; however, calcium can also inhibit autophagy. | [249, 250] |
| Thapsigargin | An inhibitor of the sarcoplasmic/endoplasmic reticulum Ca2+ ATPase (SERCA) that inhibits autophagic sequestration through the depletion of intracellular Ca2+ stores; however, thapsigargin may also block fusion of autophagosomes with endosomes by interfering with recruitment of RAB7, resulting in autophagosome accumulation. Long-term thapsigargin treatment may induce ER stress and a secondary stimulation of autophagy. | [251, 252] |
| Resveratrol | A natural polyphenol that induces autophagy via activation of AMPK. | [253] |
| RSVAs | Synthetic small-molecule analogs of resveratrol that potently activate AMPK and induce autophagy. | [254] |
| SBI-0206965 | A highly selective ULK1 kinase inhibitor in vitro that suppresses ULK1-mediated phosphorylation events in cells, regulating autophagy and cell survival. This compound is also an inhibitor of AMPK, competitively inhibiting ATP binding, and also inhibiting the binding of AMPK to its substrates. | [255] |
| Butein | A plant-derived natural molecule that induces autophagy through the activation of AMPK. | [256] |
| CCCP | Carbonyl cyanide m-chlorophenylhydrazone is a prototype protonophore, uncoupler of oxidative phosphorylation that stimulates autophagy via the AMPK-ULK1 pathway or alternative pathways and mitophagy, but inhibits autophagosomelysosome fusion due to the increase of intralysosomal pH. | [257] |
| Cinacalcet HCl | A calcimimetic that increases the sensitivity of CASR (calcium sensing receptor) to extracellular calcium. In some models, cinacalcet induces the formation of GFP-LC3 puncta during starvation, whereas in others it causes an increase in LC3-II accumulation in basal and CQ conditions. In a diabetic nephropathy model, the proposed pathway through cinacalcet-induced autophagy is CAMKK2/CaMKKβ-STK11/LKB1-AMPK-PPARGC1A/PGC1α to decrease oxidative stress, which results in a decrease of apoptosis (increased BCL2:BAX ratio) and increased autophagy (increase of BECN1 and LC3-I to LC3-II conversion). Cinacalcet may have a dual effect inducing autophagosome formation and inhibiting the late steps of autophagy. | [258, 259] |
| NVP-BEZ235 | A dual inhibitor of PIK3CA/p110 and the MTOR catalytic site that activates autophagy. | [260, 261] |
Supplementary: this table is not meant to be complete, as there are many compounds and genetic methods that regulate autophagy, and new ones are being discovered routinely.
Collectively, current evidence supports a model in which PI3K/AKT/mTOR-regulated autophagy contributes to stress adaptation and phenotypic flexibility in advanced PC, including NEPC, rather than serving as a unidirectional driver of NED [195, 196].
Recent literature underscores the critical importance of the PI3K/AKT/mTOR signaling pathway in PC NED (Supplementary Figure 2). For instance, the lack of PKCλ/ι promotes serine production through the activation of mTORC1/ATF4, resulting in increased levels of S-adenosylmethionine, which in turn supports the development of NEPC. The cannabinoid WIN suppressed the PI3K/AKT/mTOR signaling pathway in order to modulate NEPC [197-199]. Research indicates MAOA/mTOR/HIF-1α pathway that inhibiting MAOA can delay progression of NED, while hyperactive mTOR promotes NED in PC cells and is associated with increased IRF1 expression. Of the protein kinases (PKs) that showed notable alterations during NEPC transdifferentiation, 54 potentially druggable and targetable PKs were identified, primarily associated with major signaling pathways such as PI3K-AKT, MAPK, and mTOR. As previously highlighted, miRNAs and AKT are involved in a regulatory interaction during NE transdifferentiation [200-202]. Research indicates that members of the miR-17 family suppress the expression of AKT3, and leading to apoptotic proteins' activation. AKT inhibitor (Triciribine), suppresses NEPC development in mice by increasing the levels of miR-669h-3p and miR-3104-3p while decreasing miR-375-3p, miR-let7a-5p, miR-10a-5p, and miR-143-3p (Figures 3) (Supplementary Figure 2).
In addition to miRNA- and PI3K/AKT/mTOR-mediated regulation, calcium (Ca²⁺)-dependent signaling has emerged as another modulatory layer linking autophagy to neuroendocrine-associated plasticity. Ca²⁺ functions as a second messenger, crucial for regulating physiological processes [203-205]. Precise spatial regulation of cytosolic Ca²⁺ levels is essential for maintaining cellular homeostasis, which depends on the coordinated function of Ca²⁺ storage within organelles and the activity of multiple calcium channels, proteins, and transporters. The store-operated Ca²⁺ influx pathway, which is activated by the depletion of Ca²⁺ stores in the endoplasmic reticulum (ER), plays a crucial role in preserving Ca²⁺ balance and has been extensively studied for its regulatory effects in various types of cells, including both excitable and non-excitable ones. Ca²⁺ is also known to modulate opposing processes such as autophagy, which enhances cell survival. The lysosome functions as a signaling hub, with Ca²⁺-mediated regulation of autophagy occurring through calcineurin and TFEB activation [206-208].
NED is associated with the advancement of PC to an androgen-resistant phenotype [209, 210]. Disruption of AR signaling in PC cells induces functional T-type Ca²⁺ channels, leading to significant biochemical and morphological changes (Supplementary Figure 2). In the NCI-H660 NEPC cell line, inhibiting the calcium-sensing receptor (CaSR) decreased the expression of neuroendocrine markers [211]. In PC3 and 22RV1 cells transfected with SiCaSR, NE and neuronal marker expression was reduced; conversely, CaSR activation upregulated synaptophysin and NSE expression in PC3 cells [212, 213]. Increased levels of exogenous agents, such as hydrogen sulfide and glucose, have been shown to enhance the expression of functional T-type Ca²⁺ channels and Cav3.2 proteins, thereby facilitating the progression of NEPC (Figure 3).
Taken together, miRNAs, PI3K/AKT/mTOR signaling, and Ca²⁺-dependent pathways converge on autophagy as a stress-adaptive regulatory program in NEPC. Importantly, most supporting evidence is derived from experimental and preclinical models, and collectively supports a context-dependent and permissive role in neuroendocrine differentiation and therapy resistance rather than a deterministic causal driver [214-216].
Additional molecular events related to autophagy in NEPC development
In addition to classical autophagy regulators such as Ca²⁺ channels, microRNAs, and the PI3K/AKT/mTOR signaling pathway, several other molecular factors that impact NEPC are discussed below (Supplementary Figure 2). Research has shown that under conditions of androgen depletion, the AMPK-SIRT1-NO signaling axis promotes NED in LNCaP cells, a process mediated by p38MAPK-induced IL-6 secretion [217-220]. Furthermore, immunohistochemical studies have identified the presence of PIK3CA mutations in patients with NEPC (Figure 3).
Regulating the NEPC process by focusing on the “quartet”
Targeting cellular plasticity represents a promising therapeutic approach, as it plays a key role in the emergence of aggressive NE phenotypes and the development of resistance to next-generation ARPIs [115, 221, 222]. Researchers are currently exploring multiple strategies to address this challenge: promoting the re-differentiation of NE carcinoma cells into AR-dependent epithelial cells to reverse NED; selectively targeting NEPC cells; and inhibiting stem cell-related and EMT pathways to prevent the initiation of NEPC (Table 3). Moreover, ongoing studies continue to identify and validate autophagy-related modulators [156, 223-225]. The involvement of signaling components such as miRNAs, PI3K/AKT/mTOR pathway, Ca²⁺ signaling, and other regulators of autophagy in NEPC is being examined using a range of genetic models and pharmacological agents capable of influencing autophagic processes via these mechanisms (Table 2).
Treatment for directing against drivers of cellular plasticity and neuroendocrine differentiation in prostate cancer
| Inhibitor | Mechanism | Clinical status | Clinical outcome | Ref |
|---|---|---|---|---|
| CSC inhibitors | ||||
| Disulfiram | Inhibitor of ALDH activity, which is highly expressed in prostate CSCs. | Phase Ib trial in mCRPC and NEPC ongoing. | NA | [262-264] |
| Rovalpituzumab tesirine (Rova-T) | Antibody-drug conjugate targeting the NOTCH ligand DLL3, which is expressed on CSCs and neuroendocrine cells. | Phase I trial in DLL3-expressing solid tumours, including a dedicated NEPC arm ongoing. | NA | [265, 266] |
| IL-6-STAT3 inhibitors | ||||
| Galiellalactone | Directly inhibits STAT3 transcriptional activity and induces apoptosis in ALDHhigh prostate CSCs. | Preclinical | NA | [267, 268] |
| Lycopene | Inhibits IL-6 signalling and decreases STAT3 expression and phosphorylation. | Phase I trial in combination with docetaxel in CRPC completed. | Well tolerated with docetaxel; correlative analysis ongoing. | [132, 269, 270] |
| Siltuximab (CNTO 328) | Chimeric monoclonal antibody that binds to and neutralizes IL-6 bioactivity and inhibits STAT3 activity. | Phase I trial in combination with Docetaxel completed. | Efficacy in CRPC | [12, 271, 272] |
| MYCN and AURKA inhibitors | ||||
| Alisertib (MLN8237) | Inhibits AURKA and disrupts the AURKA- MYCN complex to inhibit MYCN-dependent Transcription. | Phase II trial in confirmed or suspected NEPC completed. | Moderate clinical response (18 of 60 patients); clinical progression-free survival of 2.3 months in NEPC arm. | [273-275] |
| GS-5829 | Inhibits BET proteins. | Phase I and II trials as a single agent and in combination with enzalutamide in mCRPC ongoing. | NA | [276-278] |
| ZEN003694 | Inhibits BET proteins and induces apoptosis in NEPC cell lines. | Phase I trial as a single agent and in combination with enzalutamide in mCRPC ongoing. | NA | [279-281] |
Conclusions
Advancements in both basic and translational PC biology have enabled an unprecedentedly detailed understanding of the mechanisms and effects of antiandrogen therapy. In terms of lineage identity, the remarkable plasticity of PC cells has provided new insights into clinically relevant challenges.
In recent years, numerous studies on NEPC regulation have emerged in the field; however, the difficulty of clinical treatment and diagnosis underscores the urgency of adjusting therapeutic strategies. Currently, research on the EMT, CSC, and NED transcriptomes that regulate the NEPC process is ongoing. We continue to expand our understanding of the mechanisms by which these three regulatory modules promote NED. Moreover, novel mechanisms involved in the regulation of NEPC—such as autophagy—have gained considerable research interest. As a well-established pathway in cell death regulation, autophagy is closely linked to the progression of NEPC, offering potential avenues for innovative diagnostic and treatment strategies. Recent evidence strongly indicates that autophagy can also crosstalk with the canonical IL-6 signaling pathway to influence the development of NEPC. Observations of this process have been made not only in experimental systems but also in clinical settings.
This study demonstrates that lineage plasticity is governed not only by highly redundant and complex EMT, CSC, and NED mechanisms but also by dysregulation and crosstalk within the autophagy network. In-depth research on lineage plasticity and autophagy, along with thorough clinical trials, is crucial for enhancing our understanding of these processes and effectively regulating cell plasticity and autophagy in PC. Importantly, each of these regulatory programs is discussed in this review as a distinct yet interconnected parameter, with clearly delineated molecular determinants and context-dependent associations with NEPC rather than as linear causal drivers.
Supplementary Material
Supplementary figures.
Acknowledgements
Funding
This work was supported by the grant from the National Nature Science Foundation of China [No. 82473287], the National Nature Science Foundation of China [No. 82273389], the Peking University Third Hospital Clinical Cohort Construction Project [BYSYDL2023016], and the Peking University Third Hospital Clinical Key Project [BYSYZD2023035].
Author contributions
All authors reviewed the manuscript. WYZ contributed to the conception and design of the review. WYZ, WN, LJB wrote the initial draft. WYZ, CQ, YJF, and ZSD reviewed the manuscript and provided suggestions for revision. All authors read and approved the final manuscript.
Competing Interests
The authors have declared that no competing interest exists.
References
1. Guo H, Ci X, Ahmed M, Hua JT, Soares F, Lin D. et al. ONECUT2 is a driver of neuroendocrine prostate cancer. Nat Commun. 2019;10:278
2. Haffner MC, Morris MJ, Ding C-KC, Sayar E, Mehra R, Robinson B. et al. Framework for the Pathology Workup of Metastatic Castration-Resistant Prostate Cancer Biopsies. Clin Cancer Res. 2025;31:466-78
3. Han M, Li F, Zhang Y, Dai P, He J, Li Y. et al. FOXA2 drives lineage plasticity and KIT pathway activation in neuroendocrine prostate cancer. Cancer Cell. 2022 40
4. Liu C, Chen J, Cong Y, Chen K, Li H, He Q. et al. PROX1 drives neuroendocrine plasticity and liver metastases in prostate cancer. Cancer Lett. 2024;597:217068
5. Lu X, Keo V, Cheng I, Xie W, Gritsina G, Wang J. et al. NKX2-1 drives neuroendocrine transdifferentiation of prostate cancer via epigenetic and 3D chromatin remodeling. Nat Genet. 2025;57:1966-80
6. McCartney M. Prostate cancer screening: what work should GPs stop doing so they can do this work instead? BMJ. 2023;380:284
7. McQuillen CN, Brady NJ. ASCL1 Drives the Development of Neuroendocrine Prostate Cancer. Cancer Res. 2024;84:3499-501
8. Mizuno K, Ku S-Y, Venkadakrishnan VB, Bakht MK, Sigouros M, Chan J. et al. Intraindividual epigenetic heterogeneity underlying phenotypic subtypes of advanced prostate cancer. Nat Commun. 2025;16:5543
9. Rawat K, Punia V, Mathews P, McCoy S, Song W, Saeed MA. et al. Synergistic potential of sipuleucel-T in enhancing immunotherapy for metastatic castration-resistant prostate cancer. J Immunother Cancer. 2025 13
10. Rodarte KE, Nir Heyman S, Guo L, Flores L, Savage TK, Villarreal J. et al. Neuroendocrine Differentiation in Prostate Cancer Requires ASCL1. Cancer Res. 2024;84:3522-37
11. Romero R, Chu T, González Robles TJ, Smith P, Xie Y, Kaur H. et al. The neuroendocrine transition in prostate cancer is dynamic and dependent on ASCL1. Nat Cancer. 2024;5:1641-59
12. Sooreshjani MA, Kamra M, Zoubeidi A, Shah K. Reciprocal deregulation of NKX3.1 and AURKA axis in castration-resistant prostate cancer and NEPC models. J Biomed Sci. 2021;28:68
13. Venkadakrishnan VB, Presser A, Voss NCE, Neiswender J, Brenan L, Sosa KP. et al. Epigenetic Derepression of PROX1 Promotes Neuroendocrine Prostate Cancer Progression. Cancer Res. 2025;85:3842-54
14. Venkadakrishnan VB, Presser AG, Singh R, Booker MA, Traphagen NA, Weng K. et al. Lineage-specific canonical and non-canonical activity of EZH2 in advanced prostate cancer subtypes. Nat Commun. 2024;15:6779
15. Wang D, Du G, Chen X, Wang J, Liu K, Zhao H. et al. Zeb1-controlled metabolic plasticity enables remodeling of chromatin accessibility in the development of neuroendocrine prostate cancer. Cell Death Differ. 2024;31:779-91
16. Wang H, Zhang S, Pan Q, Guo J, Li N, Chen L. et al. Targeting the histone reader ZMYND8 inhibits antiandrogen-induced neuroendocrine tumor transdifferentiation of prostate cancer. Nat Cancer. 2025;6:629-46
17. Wang HT, Yao YH, Li BG, Tang Y, Chang JW, Zhang J. Neuroendocrine Prostate Cancer (NEPC) progressing from conventional prostatic adenocarcinoma: factors associated with time to development of NEPC and survival from NEPC diagnosis-a systematic review and pooled analysis. J Clin Oncol. 2014;32:3383-90
18. Wang L-Y, Hung C-L, Wang T-C, Hsu H-C, Kung H-J, Lin K-H. PROTACs as Therapeutic Modalities for Drug Discovery in Castration-Resistant Prostate Cancer. Annu Rev Pharmacol Toxicol. 2024;65:375-96
19. Wang Y, Wang Y, Ci X, Choi SYC, Crea F, Lin D. et al. Molecular events in neuroendocrine prostate cancer development. Nat Rev Urol. 2021;18:581-96
20. Xin L. Autocrine amphiregulin signaling sustains castration-resistant Ly6d+ prostate cancer cells. Trends Cell Biol. 2023;33:619-20
21. Xiong S, Li S, Li Z, Song Y, Yang L, Yang H. et al. A noncanonical E3 ubiquitin ligase RNF41-mediated MYO1C stability promotes prostate cancer metastasis by inducing actin remodeling. Oncogene. 2024;43:2696-707
22. Yamada Y, Venkadakrishnan VB, Mizuno K, Bakht M, Ku S-Y, Garcia MM. et al. Targeting DNA methylation and B7-H3 in RB1-deficient and neuroendocrine prostate cancer. Sci Transl Med. 2023;15:eadf6732
23. Zhang D, Ma B, Liu D, Wu W, Zhou T, Gao Y. et al. Discovery of a peptide proteolysis-targeting chimera (PROTAC) drug of p300 for prostate cancer therapy. EBioMedicine. 2024;105:105212
24. Zhang T, Zhao F, Lin Y, Liu M, Zhou H, Cui F. et al. Integrated analysis of single-cell and bulk transcriptomics develops a robust neuroendocrine cell-intrinsic signature to predict prostate cancer progression. Theranostics. 2024;14:1065-80
25. Aggarwal R, Rottey S, Bernard-Tessier A, Mellado B, Kosaka T, Stadler WM. et al. Safety and Efficacy of Tarlatamab in Patients with Neuroendocrine Prostate Cancer: Results from the Phase 1b DeLLpro-300 Study. Clin Cancer Res. 2025;31:3854-63
26. Aggarwal R, Zhang T, Small EJ, Armstrong AJ. Neuroendocrine prostate cancer: subtypes, biology, and clinical outcomes. J Natl Compr Canc Netw. 2014;12:719-26
27. Baca SC, Takeda DY, Seo J-H, Hwang J, Ku SY, Arafeh R. et al. Reprogramming of the FOXA1 cistrome in treatment-emergent neuroendocrine prostate cancer. Nat Commun. 2021;12:1979
28. Belge Bilgin G, Lucien-Matteoni F, Chaudhuri AA, Orme JJ, Childs DS, Muniz M. et al. Current and future directions in theranostics for neuroendocrine prostate cancer. Cancer Treat Rev. 2025;136:102941
29. Beltran H, Hruszkewycz A, Scher HI, Hildesheim J, Isaacs J, Yu EY. et al. The Role of Lineage Plasticity in Prostate Cancer Therapy Resistance. Clin Cancer Res. 2019;25:6916-24
30. Beltran H, Jendrisak A, Landers M, Mosquera JM, Kossai M, Louw J. et al. The Initial Detection and Partial Characterization of Circulating Tumor Cells in Neuroendocrine Prostate Cancer. Clin Cancer Res. 2015;22:1510-9
31. Berchuck JE, Baca SC, McClure HM, Korthauer K, Tsai HK, Nuzzo PV. et al. Detecting Neuroendocrine Prostate Cancer Through Tissue-Informed Cell-Free DNA Methylation Analysis. Clin Cancer Res. 2022;28:928-38
32. Berger A, Brady NJ, Bareja R, Robinson B, Conteduca V, Augello MA. et al. N-Myc-mediated epigenetic reprogramming drives lineage plasticity in advanced prostate cancer. J Clin Invest. 2019;129:3924-40
33. Bergmann L, Greimeier S, Riethdorf S, Rohlfing T, Kaune M, Busenbender T. et al. Transcriptional profiles of circulating tumor cells reflect heterogeneity and treatment resistance in advanced prostate cancer. J Exp Clin Cancer Res. 2025;44:111
34. Bhagirath D, Liston M, Patel N, Akoto T, Lui B, Yang TL. et al. MicroRNA determinants of neuroendocrine differentiation in metastatic castration-resistant prostate cancer. Oncogene. 2020;39:7209-23
35. Bhattacharya S, Harris HL, Islam R, Bodas S, Polavaram N, Mishra J. et al. Understanding the function of Pax5 in development of docetaxel-resistant neuroendocrine-like prostate cancers. Cell Death Dis. 2024;15:617
36. Bigot L, Sabio J, Poiraudeau L, Annereau M, Menssouri N, Helissey C. et al. Development of Novel Models of Aggressive Variants of Castration-resistant Prostate Cancer. Eur Urol Oncol. 2023;7:527-36
37. Brady NJ, Bagadion AM, Singh R, Conteduca V, Van Emmenis L, Arceci E. et al. Temporal evolution of cellular heterogeneity during the progression to advanced AR-negative prostate cancer. Nat Commun. 2021;12:3372
38. Cai M, Zheng F, Ren Y-Z, Zhen C, Fu D, Song X-L. et al. New Insights into Potential Therapeutic Targets for Neuroendocrine Prostate Cancer: From Bench to Clinic. Research (Wash D C). 2025;8:0791
39. Cejas P, Xie Y, Font-Tello A, Lim K, Syamala S, Qiu X. et al. Subtype heterogeneity and epigenetic convergence in neuroendocrine prostate cancer. Nat Commun. 2021;12:5775
40. Chakraborty G, Gupta K, Kyprianou N. Epigenetic mechanisms underlying subtype heterogeneity and tumor recurrence in prostate cancer. Nat Commun. 2023;14:567
41. Chen B, Xu P, Yang JC, Nip C, Wang L, Shen Y. et al. Plexin D1 emerges as a novel target in the development of neural lineage plasticity in treatment-resistant prostate cancer. Oncogene. 2024;43:2325-37
42. Chen W-Y, Thuy Dung PV, Yeh H-L, Chen W-H, Jiang K-C, Li H-R. et al. Targeting PKLR/MYCN/ROMO1 signaling suppresses neuroendocrine differentiation of castration-resistant prostate cancer. Redox Biol. 2023;62:102686
43. Chen W-Y, Zeng T, Wen Y-C, Yeh H-L, Jiang K-C, Chen W-H. et al. Androgen deprivation-induced ZBTB46-PTGS1 signaling promotes neuroendocrine differentiation of prostate cancer. Cancer Lett. 2018;440-441:35-46
44. Chen Y, Zhou Q, Hankey W, Fang X, Yuan F. Second generation androgen receptor antagonists and challenges in prostate cancer treatment. Cell Death Dis. 2022;13:632
45. Ci X, Hao J, Dong X, Choi SY, Xue H, Wu R. et al. Heterochromatin Protein 1α Mediates Development and Aggressiveness of Neuroendocrine Prostate Cancer. Cancer Res. 2018;78:2691-704
46. Cyrta J, Augspach A, De Filippo MR, Prandi D, Thienger P, Benelli M. et al. Role of specialized composition of SWI/SNF complexes in prostate cancer lineage plasticity. Nat Commun. 2020;11:5549
47. Dardenne E, Beltran H, Benelli M, Gayvert K, Berger A, Puca L. et al. N-Myc Induces an EZH2-Mediated Transcriptional Program Driving Neuroendocrine Prostate Cancer. Cancer Cell. 2016;30:563-77
48. Davies AH, Beltran H, Zoubeidi A. Cellular plasticity and the neuroendocrine phenotype in prostate cancer. Nat Rev Urol. 2018;15:271-86
49. De Sarkar N, Patton RD, Doebley A-L, Hanratty B, Adil M, Kreitzman AJ. et al. Nucleosome Patterns in Circulating Tumor DNA Reveal Transcriptional Regulation of Advanced Prostate Cancer Phenotypes. Cancer Discov. 2023;13:632-53
50. DeLucia DC, Cardillo TM, Ang L, Labrecque MP, Zhang A, Hopkins JE. et al. Regulation of CEACAM5 and Therapeutic Efficacy of an Anti-CEACAM5-SN38 Antibody-drug Conjugate in Neuroendocrine Prostate Cancer. Clin Cancer Res. 2020;27:759-74
51. Dong B, Jaeger AM, Hughes PF, Loiselle DR, Hauck JS, Fu Y. et al. Targeting therapy-resistant prostate cancer via a direct inhibitor of the human heat shock transcription factor 1. Sci Transl Med. 2020 12
52. Dwivedi S, Bautista M, Shrestha S, Elhasasna H, Chaphekar T, Vizeacoumar FS. et al. Sympathetic signaling facilitates progression of neuroendocrine prostate cancer. Cell Death Discov. 2021;7:364
53. Fenner A. Surfaceome profiling for NEPC target antigens. Nat Rev Urol. 2018;15:396-7
54. Flores-Morales A, Bergmann TB, Lavallee C, Batth TS, Lin D, Lerdrup M. et al. Proteogenomic Characterization of Patient-Derived Xenografts Highlights the Role of REST in Neuroendocrine Differentiation of Castration-Resistant Prostate Cancer. Clin Cancer Res. 2018;25:595-608
55. Gao D, Shen Y, Xu L, Sun Y, Hu H, Xu B. et al. Acetate utilization promotes hormone therapy resistance in prostate cancer through neuroendocrine differentiation. Drug Resist Updat. 2024;77:101158
56. Ge R, Wang Z, Montironi R, Jiang Z, Cheng M, Santoni M. et al. Epigenetic modulations and lineage plasticity in advanced prostate cancer. Ann Oncol. 2020;31:470-9
57. Gritsina G, Fong K-W, Lu X, Lin Z, Xie W, Agarwal S. et al. Chemokine receptor CXCR7 activates Aurora Kinase A and promotes neuroendocrine prostate cancer growth. J Clin Invest. 2023 133
58. Guedes LB, Antonarakis ES, Schweizer MT, Mirkheshti N, Almutairi F, Park JC. et al. MSH2 Loss in Primary Prostate Cancer. Clin Cancer Res. 2017;23:6863-74
59. Hagiwara M, Yasumizu Y, Yamashita N, Rajabi H, Fushimi A, Long MD. et al. MUC1-C Activates the BAF (mSWI/SNF) Complex in Prostate Cancer Stem Cells. Cancer Res. 2020;81:1111-22
60. Hou Z, Huang S, Li Z. Androgens in prostate cancer: A tale that never ends. Cancer Lett. 2021 516
61. Hsu E-C, Rice MA, Bermudez A, Marques FJG, Aslan M, Liu S. et al. Trop2 is a driver of metastatic prostate cancer with neuroendocrine phenotype via PARP1. Proc Natl Acad Sci U S A. 2020;117:2032-42
62. Jiang J, Liu S, Xu C, He L, Li H, Zhou Y. et al. LHX2 Rewires the Metabolic and Epigenetic Landscape to Drive Tumor Progression in Prostate Cancer. Cancer Res. 2025;85:4751-68
63. Jing N, Du X, Liang Y, Tao Z, Bao S, Xiao H. et al. PAX6 promotes neuroendocrine phenotypes of prostate cancer via enhancing MET/STAT5A-mediated chromatin accessibility. J Exp Clin Cancer Res. 2024;43:144
64. Kelsey R. Prostate cancer: SRRM4 drives NEPC progression. Nat Rev Urol. 2016;13:371
65. Kim D-H, Sun D, Storck WK, Welker Leng K, Jenkins C, Coleman DJ. et al. BET Bromodomain Inhibition Blocks an AR-Repressed, E2F1-Activated Treatment-Emergent Neuroendocrine Prostate Cancer Lineage Plasticity Program. Clin Cancer Res. 2021;27:4923-36
66. Kim J, Jin H, Zhao JC, Yang YA, Li Y, Yang X. et al. FOXA1 inhibits prostate cancer neuroendocrine differentiation. Oncogene. 2017;36:4072-80
67. Korsen JA, Kalidindi TM, Khitrov S, Samuels ZV, Chakraborty G, Gutierrez JA. et al. Molecular Imaging of Neuroendocrine Prostate Cancer by Targeting Delta-Like Ligand 3. J Nucl Med. 2022;63:1401-7
68. Lee J, Park J, Hur Y, Um D, Choi H-S, Park J. et al. ETV5 reduces androgen receptor expression and induces neural stem-like properties during neuroendocrine prostate cancer development. Proc Natl Acad Sci U S A. 2025;122:e2420313122
69. Lee JK, Bangayan NJ, Chai T, Smith BA, Pariva TE, Yun S. et al. Systemic surfaceome profiling identifies target antigens for immune-based therapy in subtypes of advanced prostate cancer. Proc Natl Acad Sci U S A. 2018;115:E4473-E82
70. Lee JK, Phillips JW, Smith BA, Park JW, Stoyanova T, McCaffrey EF. et al. N-Myc Drives Neuroendocrine Prostate Cancer Initiated from Human Prostate Epithelial Cells. Cancer Cell. 2016;29:536-47
71. Li Y, Donmez N, Sahinalp C, Xie N, Wang Y, Xue H. et al. SRRM4 Drives Neuroendocrine Transdifferentiation of Prostate Adenocarcinoma Under Androgen Receptor Pathway Inhibition. Eur Urol. 2016;71:68-78
72. Li Y, Xie N, Chen R, Lee AR, Lovnicki J, Morrison EA. et al. RNA Splicing of the BHC80 Gene Contributes to Neuroendocrine Prostate Cancer Progression. Eur Urol. 2019;76:157-66
73. Liang L, Shen Z, Zhang Y, Cheng Y, Yao B, Feng N. et al. OPRK1 drives SLC9A3R1 progression to neuroendocrine prostate cancer. Cell Death Dis. 2025
74. Liu B, Li L, Yang G, Geng C, Luo Y, Wu W. et al. PARP Inhibition Suppresses GR-MYCN-CDK5-RB1-E2F1 Signaling and Neuroendocrine Differentiation in Castration-Resistant Prostate Cancer. Clin Cancer Res. 2019;25:6839-51
75. Liu B, Zhang W, Ji Y, Wu J, Su R, Liu X. et al. Targeting the CCL5/CCR5 axis in tumor-stromal crosstalk to overcome cisplatin resistance in neuroendocrine prostate cancer. J Exp Clin Cancer Res. 2025;44:296
76. Kim Y-B, Ahn Y-H, Jung J-H, Lee Y-J, Lee J-H, Kang JL. Programming of macrophages by UV-irradiated apoptotic cancer cells inhibits cancer progression and lung metastasis. Cell Mol Immunol. 2019;16:851-67
77. Odero-Marah VA, Wang R, Chu G, Zayzafoon M, Xu J, Shi C. et al. Receptor activator of NF-kappaB Ligand (RANKL) expression is associated with epithelial to mesenchymal transition in human prostate cancer cells. Cell Res. 2008;18:858-70
78. Liu L, Rascón IA, Lin D, Ni Y, Dong X, Xue H. et al. CXCR4-LASP1-G9a-SNAIL axis drives NEPC transdifferentiation via induction of EMT and downregulation of REST. Cell Genom. 2025;5:100916
79. Liu S, Alabi BR, Yin Q, Stoyanova T. Molecular mechanisms underlying the development of neuroendocrine prostate cancer. Semin Cancer Biol. 2022;86:57-68
80. Xie Z, Fang Y, Huang J, Shen J, Li J, Li H. et al. Integrative multi-omics and radiogenomic profiling decodes NNK-related tumor remodeling and prognostic stratification in pancreatic cancer. Int J Surg. 2026
81. Zheng X, Lin J, Wu H, Mo Z, Lian Y, Wang P. et al. Forkhead box (FOX) G1 promotes hepatocellular carcinoma epithelial-Mesenchymal transition by activating Wnt signal through forming T-cell factor-4/Beta-catenin/FOXG1 complex. J Exp Clin Cancer Res. 2019;38:475
82. Long Z, Deng L, Li C, He Q, He Y, Hu X. et al. Loss of EHF facilitates the development of treatment-induced neuroendocrine prostate cancer. Cell Death Dis. 2021;12:46
83. Lovnicki J, Gan Y, Feng T, Li Y, Xie N, Ho C-H. et al. LIN28B promotes the development of neuroendocrine prostate cancer. J Clin Invest. 2020;130:5338-48
84. Macedo-Silva C, Albuquerque-Castro Â, Carriço I, Lencart J, Carneiro I, Altucci L. et al. Decoding MUC1 and AR axis in a radiation-induced neuroendocrine prostate cancer cell-subpopulation unveils novel therapeutic targets. Cell Death Discov. 2025;11:306
85. Mak CSL, Zhu M, Fu J, Liang X, Wang X, Yuan F. et al. KDM4A promotes NEPC progression through regulation of MYC expression. Cancer Lett. 2025;638:218184
86. Masone MC. A non-invasive approach for NEPC diagnosis. Nat Rev Urol. 2022;19:67
87. Masone MC. FOXA2-KIT-driven lineage plasticity in NEPC. Nat Rev Urol. 2023;20:8
88. Mosquera MJ, Kim S, Bareja R, Fang Z, Cai S, Pan H. et al. Extracellular Matrix in Synthetic Hydrogel-Based Prostate Cancer Organoids Regulate Therapeutic Response to EZH2 and DRD2 Inhibitors. Adv Mater. 2021;34:e2100096
89. Pan X, Chen K, Gao W, Xu M, Meng F, Wu M. et al. Circular RNA circBNC2 inhibits tumorigenesis by modulating ferroptosis and acts as a nanotherapeutic target in prostate cancer. Mol Cancer. 2025;24:29
90. Qian C, Yang Q, Rotinen M, Huang R, Kim H, Gallent B. et al. ONECUT2 acts as a lineage plasticity driver in adenocarcinoma as well as neuroendocrine variants of prostate cancer. Nucleic Acids Res. 2024;52:7740-60
91. Rawat C, Heemers HV. Alternative splicing in prostate cancer progression and therapeutic resistance. Oncogene. 2024;43:1655-68
92. Reina-Campos M, Linares JF, Duran A, Cordes T, L'Hermitte A, Badur MG. et al. Increased Serine and One-Carbon Pathway Metabolism by PKCλ/ι Deficiency Promotes Neuroendocrine Prostate Cancer. Cancer Cell. 2019 35
93. Séguier D, Parent P, Duterque-Coquillaud M, Labreuche J, Fromont-Hankard G, Dariane C. et al. Emergence of Neuroendocrine Tumors in Patients Treated with Androgen Receptor Pathway Inhibitors for Metastatic Prostate Cancer: A Systematic Review and Meta-analysis. Eur Urol Oncol. 2025;8:581-90
94. Shan J, Al-Muftah MA, Al-Kowari MK, Abuaqel SWJ, Al-Rumaihi K, Al-Bozom I. et al. Targeting Wnt/EZH2/microRNA-708 signaling pathway inhibits neuroendocrine differentiation in prostate cancer. Cell Death Discov. 2019;5:139
95. Shi M, Wang Y, Lin D, Wang Y. Patient-derived xenograft models of neuroendocrine prostate cancer. Cancer Lett. 2021;525:160-9
96. Singh N, Ramnarine VR, Song JH, Pandey R, Padi SKR, Nouri M. et al. The long noncoding RNA H19 regulates tumor plasticity in neuroendocrine prostate cancer. Nat Commun. 2021;12:7349
97. Sulsenti R, Scialpi GB, Frossi B, Botti L, Ferri R, Tripodi I. et al. Intracellular Osteopontin Promotes the Release of TNFα by Mast Cells to Restrain Neuroendocrine Prostate Cancer. Cancer Immunol Res. 2024;12:1147-69
98. Sun Y, Ren S, Wen W, Jing J, Luo X, Shao S. et al. Targeting CXCR2 in prostate cancer cells can block CD47-SIRPα interaction and reverse M2 macrophage polarization in the TME. Mol Cancer. 2025;24:273
99. Tang DG. Serotonin sets up neutrophil extracellular traps to promote neuroendocrine prostate cancer metastasis in the liver. J Clin Invest. 2025 135
100. Teng M, Guo J, Xu X, Ci X, Mo Y, Kohen Y. et al. Circular RMST cooperates with lineage-driving transcription factors to govern neuroendocrine transdifferentiation. Cancer Cell. 2025 43
101. Unno K, Chalmers ZR, Pamarthy S, Vatapalli R, Rodriguez Y, Lysy B. et al. Activated ALK Cooperates with N-Myc via Wnt/β-Catenin Signaling to Induce Neuroendocrine Prostate Cancer. Cancer Res. 2021;81:2157-70
102. Walke P, Price JDW, Vizeacoumar FS, Joseph N, Maranda V, Chowdhury B. et al. A novel role for Neurog2 in MYCN driven neuroendocrine plasticity of prostate cancer. Oncogene. 2025;44:2460-73
103. Ali A, Baena E. The Needle in the Haystack: The Presence of Castrate-resistant Prostate Cancer Cells in Hormone-naïve Prostate Cancer. Eur Urol. 2022;81:456-7
104. Chaudagar K, Hieromnimon HM, Khurana R, Labadie B, Hirz T, Mei S. et al. Reversal of Lactate and PD-1-mediated Macrophage Immunosuppression Controls Growth of PTEN/p53-deficient Prostate Cancer. Clin Cancer Res. 2023;29:1952-68
105. Chen C-C, Tran W, Song K, Sugimoto T, Obusan MB, Wang L. et al. Temporal evolution reveals bifurcated lineages in aggressive neuroendocrine small cell prostate cancer trans-differentiation. Cancer Cell. 2023 41
106. Feng W, Ladewig E, Lange M, Salsabeel N, Zhao H, Lee YS. et al. ERG-driven prostate cancer initiation is cell-context dependent and requires KMT2A and DOT1L. Nat Genet. 2025;57:2177-91
107. Freeland J, Crowell PD, Giafaglione JM, Boutros PC, Goldstein AS. Aging of the progenitor cells that initiate prostate cancer. Cancer Lett. 2021;515:28-35
108. Guo W, Zhang X, Li L, Shao P, Liang C, Zhang H. et al. JAK/STAT signaling maintains an intermediate cell population during prostate basal cell fate determination. Nat Genet. 2024;56:2776-89
109. Jamroze A, Chatta G, Tang DG. Androgen receptor (AR) heterogeneity in prostate cancer and therapy resistance. Cancer Lett. 2021;518:1-9
110. Jiang C, Song Y, Rorive S, Allard J, Tika E, Zahedi Z. et al. Innate immunity and the NF-κB pathway control prostate stem cell plasticity, reprogramming and tumor initiation. Nat Cancer. 2025;6:1537-58
111. Jin T, Yang L, Chang C, Luo H, Wang R, Gan Y. et al. HnRNPA2B1 ISGylation Regulates m6A-Tagged mRNA Selective Export via ALYREF/NXF1 Complex to Foster Breast Cancer Development. Adv Sci (Weinh). 2024;11:e2307639
112. Karthaus WR, Hofree M, Choi D, Linton EL, Turkekul M, Bejnood A. et al. Regenerative potential of prostate luminal cells revealed by single-cell analysis. Science. 2020;368:497-505
113. Lee Y, Yoon J, Ko D, Yu M, Lee S, Kim S. TMPRSS4 promotes cancer stem-like properties in prostate cancer cells through upregulation of SOX2 by SLUG and TWIST1. J Exp Clin Cancer Res. 2021;40:372
114. Li W, Shen MM. Prostate cancer cell heterogeneity and plasticity: Insights from studies of genetically-engineered mouse models. Semin Cancer Biol. 2021;82:60-7
115. Mukha A, Kahya U, Linge A, Chen O, Löck S, Lukiyanchuk V. et al. GLS-driven glutamine catabolism contributes to prostate cancer radiosensitivity by regulating the redox state, stemness and ATG5-mediated autophagy. Theranostics. 2021;11:7844-68
116. Peitzsch C, Gorodetska I, Klusa D, Shi Q, Alves TC, Pantel K. et al. Metabolic regulation of prostate cancer heterogeneity and plasticity. Semin Cancer Biol. 2020 82
117. Qu Y, Guo B, Zhao S, Sun J, Cao J, Xia M. et al. Photothermal treatment of prostate tumor with micellar indocyanine green and napabucasin to co-ablate cancer cells and cancer stem cells. J Control Release. 2025;382:113704
118. Tang DG. Understanding and targeting prostate cancer cell heterogeneity and plasticity. Semin Cancer Biol. 2021;82:68-93
119. Wang H, Cui C, Li W, Wu H, Sha J, Pan J. et al. AGD1/USP10/METTL13 complexes enhance cancer stem cells proliferation and diminish the therapeutic effect of docetaxel via CD44 m6A modification in castration resistant prostate cancer. J Exp Clin Cancer Res. 2025;44:12
120. Wang Z, Townley SL, Zhang S, Liu M, Li M, Labaf M. et al. FOXA2 rewires AP-1 for transcriptional reprogramming and lineage plasticity in prostate cancer. Nat Commun. 2024;15:4914
121. Tang F, Xu D, Wang S, Wong CK, Martinez-Fundichely A, Lee CJ. et al. Chromatin profiles classify castration-resistant prostate cancers suggesting therapeutic targets. Science. 2022;376:eabe1505
122. Tang G, Liu J, Gao X, Yuan K, Chen M, Wang L. et al. Luminal Stem-like Cells in High-Risk/Locally Advanced Prostate Cancer Promote Resistance to Hormonal Therapy. Theranostics. 2025;15:9558-79
123. Wang C, Zhang Y, Gao W-Q. The evolving role of immune cells in prostate cancer. Cancer Lett. 2021 525
124. White JM, Ramos N, Saliganan A-D, Chung J-Y, Bell M, Lindquist J. et al. Selective ablation of TRA-1-60+ pluripotent stem cells suppresses tumor growth of prostate cancer. Theranostics. 2023;13:2057-71
125. Zhang W, Bado IL, Hu J, Wan Y-W, Wu L, Wang H. et al. The bone microenvironment invigorates metastatic seeds for further dissemination. Cell. 2021 184
126. Wang Y, Gao W, Li Y, Chow ST, Xie W, Zhang X. et al. Interplay between orphan nuclear receptors and androgen receptor-dependent or-independent growth signalings in prostate cancer. Mol Aspects Med. 2020;78:100921
127. Wang Y, Wu N, Li J, Liang J, Zhou D, Cao Q. et al. The interplay between autophagy and ferroptosis presents a novel conceptual therapeutic framework for neuroendocrine prostate cancer. Pharmacol Res. 2024;203:107162
128. Wang Y, Xue H, Zhu X, Lin D, Chen Z, Dong X. et al. Deciphering the Transcription Factor Landscape in Prostate Cancer Progression: A Novel Approach to Understand NE Transdifferentiation. Adv Sci (Weinh). 2025;12:e2404938
129. Wen Y-C, Tram VTN, Chen W-H, Li C-H, Yeh H-L, Thuy Dung PV. et al. CHRM4/AKT/MYCN upregulates interferon alpha-17 in the tumor microenvironment to promote neuroendocrine differentiation of prostate cancer. Cell Death Dis. 2023;14:304
130. Xiao Z, Ma J, Cen J, Xie T, Zheng L, Shi G. et al. Rapid diagnostic imaging and targeted immunotoxin delivery in aggressive prostate cancer using CEACAM5-specific nanobodies. J Nanobiotechnology. 2025;23:525
131. Xie W, Agarwal S, Yu J. Ferroptosis: the vulnerability within a cancer monster. J Clin Invest. 2023 133
132. Xu P, Yang JC, Chen B, Ning S, Zhang X, Wang L. et al. Proteostasis perturbation of N-Myc leveraging HSP70 mediated protein turnover improves treatment of neuroendocrine prostate cancer. Nat Commun. 2024;15:6626
133. Yin L, Ye Y, Zou L, Lin J, Dai Y, Fu Y. et al. AR antagonists develop drug resistance through TOMM20 autophagic degradation-promoted transformation to neuroendocrine prostate cancer. J Exp Clin Cancer Res. 2023;42:204
134. Yasumizu Y, Rajabi H, Jin C, Hata T, Pitroda S, Long MD. et al. MUC1-C regulates lineage plasticity driving progression to neuroendocrine prostate cancer. Nat Commun. 2020;11:338
135. Yuan S, He S-H, Li L-Y, Xi S, Weng H, Zhang J-H. et al. A potassium-chloride co-transporter promotes tumor progression and castration resistance of prostate cancer through m6A reader YTHDC1. Cell Death Dis. 2023;14:7
136. Zhang X, Yang Y, Zou H, Yang Y, Zheng X, Corey E. et al. Effective therapeutic targeting of tumor lineage plasticity in neuroendocrine prostate cancer by BRD4 inhibitors. Acta Pharm Sin B. 2025;15:1415-29
137. Zhang X-W, Li J-Y, Li L, Hu W-Q, Tao Y, Gao W-Y. et al. Neurokinin-1 receptor drives PKCɑ-AURKA/N-Myc signaling to facilitate the neuroendocrine progression of prostate cancer. Cell Death Dis. 2023;14:384
138. Zhang Y, Song X-W, Zhang N, Li X-H, Wu F-C, Wei Y-A. et al. Ezetimibe Engineered L14-8 Suppresses Advanced Prostate Cancer by Activating PLK1/TP53-SAT1-Induced Ferroptosis. Adv Sci (Weinh). 2025;12:e04192
139. Zhao SG, Sperger JM, Schehr JL, McKay RR, Emamekhoo H, Singh A. et al. A clinical-grade liquid biomarker detects neuroendocrine differentiation in prostate cancer. J Clin Invest. 2022 132
140. Zhou Q, Yang M, Fu J, Sun X, Wang J, Zhang H. et al. KIF1A promotes neuroendocrine differentiation in prostate cancer by regulating the OGT-mediated O-GlcNAcylation. Cell Death Dis. 2024;15:796
141. Anzules JM, Sayegh M, Li YR. Exploiting AR-Synergistic Metabolic Vulnerabilities in Prostate Cancer. Cancer Res. 2025;85:4045-6
142. Ashrafizadeh M, Paskeh MDA, Mirzaei S, Gholami MH, Zarrabi A, Hashemi F. et al. Targeting autophagy in prostate cancer: preclinical and clinical evidence for therapeutic response. J Exp Clin Cancer Res. 2022;41:105
143. Bae TH, Sung KW, Pham TM, Najy AJ, Zamiri A, Jang H. et al. An Autophagy-Targeting Chimera Induces Degradation of Androgen Receptor Mutants and AR-v7 in Castration-Resistant Prostate Cancer. Cancer Res. 2025;85:342-59
144. Bi J, Witt E, McGovern MK, Cafi AB, Rosenstock LL, Pearson AB. et al. Oral Carbon Monoxide Enhances Autophagy Modulation in Prostate, Pancreatic, and Lung Cancers. Adv Sci (Weinh). 2023;11:e2308346
145. Chen S, Lin J, Yang Z, Wang Y, Wang Q, Wang D. et al. TRIM24-mediated K27-linked ubiquitination of ULK1 alleviates energy stress-induced autophagy and promote prostate cancer growth in the context of SPOP mutation. Cell Death Differ. 2025
146. Clark A, Villarreal MR, Huang S-B, Jayamohan S, Rivas P, Hussain SS. et al. Targeting S6K/NFκB/SQSTM1/Polθ signaling to suppress radiation resistance in prostate cancer. Cancer Lett. 2024;597:217063
147. Cui H, Wang Y, Zhou T, Qu L, Zhang X, Wang Y. et al. Targeting DGAT1 inhibits prostate cancer cells growth by inducing autophagy flux blockage via oxidative stress. Oncogene. 2023;43:136-50
148. Don Wai Luu L, Kaakoush NO, Castaño-Rodríguez N. The role of ATG16L2 in autophagy and disease. Autophagy. 2022;18:2537-46
149. Gao K, Shi Q, Liu Y, Wang C. Enhanced autophagy and NFE2L2/NRF2 pathway activation in SPOP mutation-driven prostate cancer. Autophagy. 2022;18:2013-5
150. Ivarsdottir EV, Gudmundsson J, Tragante V, Sveinbjornsson G, Kristmundsdottir S, Stacey SN. et al. Gene-based burden tests of rare germline variants identify six cancer susceptibility genes. Nat Genet. 2024;56:2422-33
151. Ji F, Dai E, Kang R, Klionsky DJ, Liu T, Hu Y. et al. Mammalian nucleophagy: process and function. Autophagy. 2025;21:1396-412
152. Ke Z-B, Chen J-Y, Xue Y-T, Lin B, Huang Q, Huang X-Y. et al. Mechanical signal modulates prostate cancer immune escape by USP8-mediated ubiquitination-dependent degradation of PD-L1 and MHC-1. Cell Death Dis. 2025;16:413
153. Liang D, Feng Y, Zandkarimi F, Wang H, Zhang Z, Kim J. et al. Ferroptosis surveillance independent of GPX4 and differentially regulated by sex hormones. Cell. 2023 186
154. Mei W, Wei M, Tang C, Li W, Ye B, Xin S. et al. BCAT2 binding to PCBP1 regulates the PI3K/AKT signaling pathway to inhibit autophagy-related apoptosis and ferroptosis in prostate cancer. Cell Death Dis. 2025;16:337
155. Portale F, Carriero R, Iovino M, Kunderfranco P, Pandini M, Marelli G. et al. C/EBPβ-dependent autophagy inhibition hinders NK cell function in cancer. Nat Commun. 2024;15:10343
156. Shi Q, Jin X, Zhang P, Li Q, Lv Z, Ding Y. et al. SPOP mutations promote p62/SQSTM1-dependent autophagy and Nrf2 activation in prostate cancer. Cell Death Differ. 2022;29:1228-39
157. Vitale I, Pietrocola F, Guilbaud E, Aaronson SA, Abrams JM, Adam D. et al. Apoptotic cell death in disease-Current understanding of the NCCD 2023. Cell Death Differ. 2023;30:1097-154
158. Wang Y, Lei H, Yan B, Zhang S, Xu B, Lin M. et al. Tumor acidity-activatable macromolecule autophagy inhibitor and immune checkpoint blockade for robust treatment of prostate cancer. Acta Biomater. 2023;168:593-605
159. Wei X, Liang M, Deng M, Zheng J, Luo F, Ma Q. A switch from lysosomal degradation to secretory autophagy initiates osteogenic bone metastasis in prostate cancer. J Extracell Vesicles. 2024;13:e70002
160. Wei Y, Hankey W, Xu D, Yuan F. Programmed Cell Death in Cancer. MedComm (2020). 2025;6:e70357
161. Wu N, Zheng W, Zhou Y, Tian Y, Tang M, Feng X. et al. Autophagy in aging-related diseases and cancer: Principles, regulatory mechanisms and therapeutic potential. Ageing Res Rev. 2024;100:102428
162. Xu J, Yang KC, Go NE, Colborne S, Ho CJ, Hosseini-Beheshti E. et al. Chloroquine treatment induces secretion of autophagy-related proteins and inclusion of Atg8-family proteins in distinct extracellular vesicle populations. Autophagy. 2022;18:2547-60
163. Xu W, Li Y, Liu L, Xie J, Hu Z, Kuang S. et al. Icaritin-curcumol activates CD8+ T cells through regulation of gut microbiota and the DNMT1/IGFBP2 axis to suppress the development of prostate cancer. J Exp Clin Cancer Res. 2024;43:149
164. Yang J, Lu X, Hao J-L, Li L, Ruan Y-T, An X-N. et al. VSTM2L protects prostate cancer cells against ferroptosis via inhibiting VDAC1 oligomerization and maintaining mitochondria homeostasis. Nat Commun. 2025;16:1160
165. Yu H, Zhuang J, Zhou Z, Song Q, Lv J, Yang X. et al. METTL16 suppressed the proliferation and cisplatin-chemoresistance of bladder cancer by degrading PMEPA1 mRNA in a m6A manner through autophagy pathway. Int J Biol Sci. 2024;20:1471-91
166. Yu Y, Liu B, Li X, Lu D, Yang L, Chen L. et al. ATF4/CEMIP/PKCα promotes anoikis resistance by enhancing protective autophagy in prostate cancer cells. Cell Death Dis. 2022;13:46
167. Yu Y, Song Y, Cheng L, Chen L, Liu B, Lu D. et al. CircCEMIP promotes anoikis-resistance by enhancing protective autophagy in prostate cancer cells. J Exp Clin Cancer Res. 2022;41:188
168. Zhong B, Zhao Z, Jiang X. RP1-59D14.5 triggers autophagy and represses tumorigenesis and progression of prostate cancer via activation of the Hippo signaling pathway. Cell Death Dis. 2022;13:458
169. Brunner E, Damisch E, Groninger ME, Baschieri F, Tyckaert F, Nommensen L. et al. Unraveling the YAP1-TGFβ1 axis: a key driver of androgen receptor loss in prostate cancer-associated fibroblasts. J Exp Clin Cancer Res. 2025;45:11
170. Hashemi M, Mirdamadi MSA, Talebi Y, Khaniabad N, Banaei G, Daneii P. et al. Pre-clinical and clinical importance of miR-21 in human cancers: Tumorigenesis, therapy response, delivery approaches and targeting agents. Pharmacol Res. 2022;187:106568
171. Hashemi M, Sabouni E, Rahmanian P, Entezari M, Mojtabavi M, Raei B. et al. Deciphering STAT3 signaling potential in hepatocellular carcinoma: tumorigenesis, treatment resistance, and pharmacological significance. Cell Mol Biol Lett. 2023;28:33
172. Li C, Qiao Y, Jiang X, Liu L, Zheng Y, Qiu Y. et al. Discovery of a First-in-Class Degrader for the Lipid Kinase PIKfyve. J Med Chem. 2023;66:12432-45
173. Liu C-J, Fan X-D, Jiang J-G, Chen Q-X, Zhu W. Potential anticancer activities of securinine and its molecular targets. Phytomedicine. 2022;106:154417
174. Liu Z, Fan M, Hou J, Pan S, Xu Y, Zhang H. et al. Serine hydroxymethyltransferase 2 knockdown induces apoptosis in ccRCC by causing lysosomal membrane permeabilization via metabolic reprogramming. Cell Death Dis. 2023;14:144
175. Zetrini AE, Lip H, Abbasi AZ, Alradwan I, Ahmed T, He C. et al. Remodeling Tumor Immune Microenvironment by Using Polymer-Lipid-Manganese Dioxide Nanoparticles with Radiation Therapy to Boost Immune Response of Castration-Resistant Prostate Cancer. Research (Wash D C). 2023;6:0247
176. Acosta Ingram D, Turkes E, Kim TY, Vo S, Sweeney N, Bonte M-A. et al. GRAMD1B is a regulator of lipid homeostasis, autophagic flux and phosphorylated tau. Nat Commun. 2025;16:3312
177. Al-Kuraishy HM, Jabir MS, Al-Gareeb AI, Albuhadily AK, Klionsky DJ, Rafeeq MF. Epilepsy and autophagy modulators: a therapeutic split. Autophagy. 2025;21:1863-87
178. Baltgalvis KA, Lamb KN, Symons KT, Wu C-C, Hoffman MA, Snead AN. et al. Chemoproteomic discovery of a covalent allosteric inhibitor of WRN helicase. Nature. 2024;629:435-42
179. Bansaccal N, Vieugue P, Sarate R, Song Y, Minguijon E, Miroshnikova YA. et al. The extracellular matrix dictates regional competence for tumour initiation. Nature. 2023;623:828-35
180. Bethlehem L, Estevinho MM, Grinspan A, Magro F, Faith JJ, Colombel J-F. Microbiota therapeutics for inflammatory bowel disease: the way forward. Lancet Gastroenterol Hepatol. 2024;9:476-86
181. Bock C, Datlinger P, Chardon F, Coelho MA, Dong MB, Lawson KA. et al. High-content CRISPR screening. Nat Rev Methods Primers. 2022 2
182. Boike L, Henning NJ, Nomura DK. Advances in covalent drug discovery. Nat Rev Drug Discov. 2022;21:881-98
183. Chataway J, Williams T, Li V, Marrie RA, Ontaneda D, Fox RJ. Clinical trials for progressive multiple sclerosis: progress, new lessons learned, and remaining challenges. Lancet Neurol. 2024;23:277-301
184. De Santis MC, Bockorny B, Hirsch E, Cappello P, Martini M. Exploiting pancreatic cancer metabolism: challenges and opportunities. Trends Mol Med. 2024;30:592-604
185. Eggers N, Becker PB. Cell-free genomics reveal intrinsic, cooperative and competitive determinants of chromatin interactions. Nucleic Acids Res. 2021;49:7602-17
186. Endicott SJ, Miller RA. PTEN activates chaperone-mediated autophagy to regulate metabolism. Autophagy. 2023;20:216-7
187. Fang C, Liu S, Zhang S, Zheng H, Fang G, Chen C. et al. Jianpi Huayu decoction enhances the antitumor effect of doxorubicin via piezo1-mediated autophagy in hepatocellular carcinoma. Phytomedicine. 2025;143:156908
188. Fujimoto S, Leiwe MN, Aihara S, Sakaguchi R, Muroyama Y, Kobayakawa R. et al. Activity-dependent local protection and lateral inhibition control synaptic competition in developing mitral cells in mice. Dev Cell. 2023 58
189. Fennell KA, Vassiliadis D, Lam EYN, Martelotto LG, Balic JJ, Hollizeck S. et al. Non-genetic determinants of malignant clonal fitness at single-cell resolution. Nature. 2021;601:125-31
190. Gao Y, Xu X, Zhang X. Targeting different phenotypes of macrophages: A potential strategy for natural products to treat inflammatory bone and joint diseases. Phytomedicine. 2023;118:154952
191. Guzelsoy G, Elorza SD, Ros M, Schachtner LT, Hayashi M, Hobson-Gutierrez S. et al. Cooperative nutrient scavenging is an evolutionary advantage in cancer. Nature. 2025;640:534-42
192. Han Q-F, Li W-J, Hu K-S, Gao J, Zhai W-L, Yang J-H. et al. Exosome biogenesis: machinery, regulation, and therapeutic implications in cancer. Mol Cancer. 2022;21:207
193. He C, Wang D, Shukla SK, Hu T, Thakur R, Fu X. et al. Vitamin B6 Competition in the Tumor Microenvironment Hampers Antitumor Functions of NK Cells. Cancer Discov. 2024;14:176-93
194. Hesse L, Oude Elberink JNG, van Oosterhout AJM, Nawijn MC. Allergen immunotherapy for allergic airway diseases: Use lessons from the past to design a brighter future. Pharmacol Ther. 2022;237:108115
195. van Neerven SM, Vermeulen L. Cell competition in development, homeostasis and cancer. Nat Rev Mol Cell Biol. 2022;24:221-36
196. Wang J, Zhang L, Liu Y, Liu Y, Xiong A, Ran Q. et al. Epithelial Atg5 Deficiency Intensifies Caspase-11 Activation, Fueling Extracellular mtDNA Release to Activate cGAS-STING-NLRP3 Axis in Macrophages During Pseudomonas Infection. MedComm (2020). 2025;6:e70239
197. Wang S, Guo S, Guo J, Du Q, Wu C, Wu Y. et al. Cell death pathways: molecular mechanisms and therapeutic targets for cancer. MedComm (2020). 2024;5:e693
198. Wang Y, Mu N, Wang X, Liu J, Qu Y, Liu X. et al. BEX2 regulates autophagy by inhibiting PIK3CA-p85 interaction in non-small-cell lung cancer cells. Cell Commun Signal. 2025;23:528
199. Wen X, Tang S, Wan F, Zhong R, Chen L, Zhang H. The PI3K/Akt-Nrf2 Signaling Pathway and Mitophagy Synergistically Mediate Hydroxytyrosol to Alleviate Intestinal Oxidative Damage. Int J Biol Sci. 2024;20:4258-76
200. Weng Y, He B, Zhou J, Luo P, Xu Z, Yan H. et al. Potential saviour of pulmonary fibrosis: multi-pathway treatment of natural products. Phytomedicine. 2025;147:157174
201. Wu Y, Li Z, Ding T, Yang Y, Wei C, Zhang S. et al. Bidirectional regulation of neuronal autophagy in ischemic stroke: Mechanisms and therapeutic potential. Ageing Res Rev. 2025;111:102842
202. Xie Y, Han L, Jin Y, Li J, Tao Q, Wu Y. et al. Exploring the role of GLS1 in colorectal cancer progression through the glutamine metabolism-PI3K signaling-autophagy axis. Int J Surg. 2025
203. Yan J-J, Wang Y-Y, Shi Z-Y, Ding Y-Y, Wen H-Q, Wu M-P. et al. SIRT5 modulates mitochondria function via mitophagy and antioxidant mechanisms to facilitate oocyte maturation in mice. Int J Biol Macromol. 2025;306:141488
204. Yan S, Bao S, Chen T, Chen J, Zhang J, Hu X. et al. Cinnamaldehyde alleviates aspirin-induced gastric mucosal injury by regulating pi3k/akt pathway-mediated apoptosis, autophagy and ferroptosis. Phytomedicine. 2024;132:155791
205. Yang J, Wang J, Cheng R, Liao T, Pan S, Du M. et al. Microseparation of Lipophilic and Hydrophilic Metabolites for Single Oocyte Mass Spectrometry Analysis. Anal Chem. 2025;97:1704-10
206. Zhang X, Li S, Malik I, Do MH, Ji L, Chou C. et al. Reprogramming tumour-associated macrophages to outcompete cancer cells. Nature. 2023;619:616-23
207. Zhou X-H, Luo Y-X, Yao X-Q. Exercise-driven cellular autophagy: A bridge to systematic wellness. J Adv Res. 2025;76:271-91
208. Zhuo Y, Li W-S, Lu W, Li X, Ge L-T, Huang Y. et al. TGF-β1 mediates hypoxia-preconditioned olfactory mucosa mesenchymal stem cells improved neural functional recovery in Parkinson's disease models and patients. Mil Med Res. 2024;11:48
209. Ruth JH, Gurrea-Rubio M, Athukorala KS, Rasmussen SM, Weber DP, Randon PM. et al. CD6 is a target for cancer immunotherapy. JCI Insight. 2021 6
210. Shiri I, Salimi Y, Maghsudi M, Jenabi E, Harsini S, Razeghi B. et al. Differential privacy preserved federated transfer learning for multi-institutional 68Ga-PET image artefact detection and disentanglement. Eur J Nucl Med Mol Imaging. 2023;51:40-53
211. Sugier P-E, Lucotte EA, Domenighetti C, Law MH, Iles MM, Brown K. et al. Investigation of Shared Genetic Risk Factors Between Parkinson's Disease and Cancers. Mov Disord. 2023;38:604-15
212. Albert TK, Interlandi M, Sill M, Graf M, Moreno N, Menck K. et al. An extracellular vesicle-related gene expression signature identifies high-risk patients in medulloblastoma. Neuro Oncol. 2021;23:586-98
213. Bakr A, Hey J, Sigismondo G, Liu C-S, Sadik A, Goyal A. et al. ID3 promotes homologous recombination via non-transcriptional and transcriptional mechanisms and its loss confers sensitivity to PARP inhibition. Nucleic Acids Res. 2021;49:11666-89
214. Burns D, Anokian E, Saunders EJ, Bristow RG, Fraser M, Reimand J. et al. Rare Germline Variants Are Associated with Rapid Biochemical Recurrence After Radical Prostate Cancer Treatment: A Pan Prostate Cancer Group Study. Eur Urol. 2022;82:201-11
215. Niu Y, Yang H, Yu Z, Gao C, Ji S, Yan J. et al. Intervention with the Bone-Associated Tumor Vicious Cycle through Dual-Protein Therapeutics for Treatment of Skeletal-Related Events and Bone Metastases. ACS Nano. 2022;16:2209-23
216. Praharaj M, Shen F, Lee AJ, Zhao L, Nirschl TR, Theodros D. et al. Metabolic Reprogramming of Tumor-Associated Macrophages Using Glutamine Antagonist JHU083 Drives Tumor Immunity in Myeloid-Rich Prostate and Bladder Cancers. Cancer Immunol Res. 2024;12:854-75
217. Ariotti N, Wu Y, Okano S, Gambin Y, Follett J, Rae J. et al. An inverted CAV1 (caveolin 1) topology defines novel autophagy-dependent exosome secretion from prostate cancer cells. Autophagy. 2020;17:2200-16
218. Kudo Y, Endo S, Fujita M, Ota A, Kamatari YO, Tanaka Y. et al. Discovery and Structure-Based Optimization of Novel Atg4B Inhibitors for the Treatment of Castration-Resistant Prostate Cancer. J Med Chem. 2022;65:4878-92
219. Lin C, Blessing AM, Pulliam TL, Shi Y, Wilkenfeld SR, Han JJ. et al. Inhibition of CAMKK2 impairs autophagy and castration-resistant prostate cancer via suppression of AMPK-ULK1 signaling. Oncogene. 2021;40:1690-705
220. Zhang Z, Cheng L, Zhang Q, Kong Y, He D, Li K. et al. Co-Targeting Plk1 and DNMT3a in Advanced Prostate Cancer. Adv Sci (Weinh). 2021;8:e2101458
221. Lin Y-X, Wang Y, Ding J, Jiang A, Wang J, Yu M. et al. Reactivation of the tumor suppressor PTEN by mRNA nanoparticles enhances antitumor immunity in preclinical models. Sci Transl Med. 2021 13
222. Mukha A, Kahya U, Dubrovska A. Targeting glutamine metabolism and autophagy: the combination for prostate cancer radiosensitization. Autophagy. 2021;17:3879-81
223. Qin L, Berk M, Chung Y-M, Cui D, Zhu Z, Chakraborty AA. et al. Chronic hypoxia stabilizes 3βHSD1 via autophagy suppression. Cell Rep. 2024;43:113575
224. Shen M, Li X, Qian B, Wang Q, Lin S, Wu W. et al. Crucial Roles of microRNA-Mediated Autophagy in Urologic Malignancies. Int J Biol Sci. 2021;17:3356-68
225. Wu S, Wu X, Wang Q, Chen Z, Li L, Chen H. et al. Bufalin induces ferroptosis by modulating the 2,4-dienoyl-CoA reductase (DECR1)-SLC7A11 axis in breast cancer. Phytomedicine. 2024;135:156130
226. Ho P, Melms JC, Rogava M, Frangieh CJ, Poźniak J, Shah SB. et al. The CD58-CD2 axis is co-regulated with PD-L1 via CMTM6 and shapes anti-tumor immunity. Cancer Cell. 2023 41
227. Hu T, Allam M, Cai S, Henderson W, Yueh B, Garipcan A. et al. Single-cell spatial metabolomics with cell-type specific protein profiling for tissue systems biology. Nat Commun. 2023;14:8260
228. Huang G, Cai Y, Ren M, Zhang X, Fu Y, Cheng R. et al. Salidroside sensitizes Triple-negative breast cancer to ferroptosis by SCD1-mediated lipogenesis and NCOA4-mediated ferritinophagy. J Adv Res. 2024;74:589-607
229. Hundsdorfer L, Muenkel M, Aparicio-Yuste R, Sanchez-Rendon JC, Gomez-Benito MJ, Balmes A. et al. ERK activation waves coordinate mechanical cell competition leading to collective elimination via extrusion of bacterially infected cells. Cell Rep. 2025;44:115193
230. Jiang C, Zhou Q, Yi K, Yuan Y, Xie X. Colorectal cancer initiation: Understanding early-stage disease for intervention. Cancer Lett. 2024;589:216831
231. Jiang M, Zhang K, Zhang Z, Zeng X, Huang Z, Qin P. et al. PI3K/AKT/mTOR Axis in Cancer: From Pathogenesis to Treatment. MedComm (2020). 2025;6:e70295
232. Kim K, Huang H, Parida PK, He L, Marquez-Palencia M, Reese TC. et al. Cell Competition Shapes Metastatic Latency and Relapse. Cancer Discov. 2023;13:85-97
233. Krammer T, Stuart HT, Gromberg E, Ishihara K, Cislo D, Melchionda M. et al. Mouse neural tube organoids self-organize floorplate through BMP-mediated cluster competition. Dev Cell. 2024 59
234. Kuo H-Y, Khan KA, Kerbel RS. Antiangiogenic-immune-checkpoint inhibitor combinations: lessons from phase III clinical trials. Nat Rev Clin Oncol. 2024;21:468-82
235. Li S, Ye T, Hou Z, Wang Y, Hao Z, Chen J. FOXO6: A unique transcription factor in disease regulation and therapeutic potential. Pharmacol Res. 2025;214:107691
236. Li Y, Ma T, Liang X, Jin T, Zhao X, Huang J. et al. Non-coding RNAs regulate autophagy in kidney disease: friend or foe? Autophagy. 2025;21:2537-60
237. Li-Bao L, Díaz-Díaz C, Raiola M, Sierra R, Temiño S, Moya FJ. et al. Regulation of Myc transcription by an enhancer cluster dedicated to pluripotency and early embryonic expression. Nat Commun. 2024;15:3931
238. Liang H, Yang S, Huang Y, Zhu Y, Wu Q, Wu Z. et al. PTPN22 as a therapeutic target in intervertebral disc degeneration: Modulating mitophagy and pyroptosis through the PI3K/AKT/mTOR axis. J Adv Res. 2025
239. Lin F, Wang T, Ai J, Wang J, Huang C, Tian W. et al. Topical application of Tea leaf-derived nanovesicles reduce melanogenesis by modulating the miR-828b/MYB4 axis: better permeability and therapeutic efficacy than conventional tea extracts. Mater Today Bio. 2025;34:102108
240. Liu C, Liu K, Zhang D, Liu Y, Yu Y, Kang H. et al. Dual-layer microneedles with NO/O2 releasing for diabetic wound healing via neurogenesis, angiogenesis, and immune modulation. Bioact Mater. 2024;46:213-28
241. Liu S, Wu J, Lin Y, Liang H, Cai Y, Wang L. et al. Neuropeptide Y1 receptor antagonist alleviated osteoarthritis by restoring chondrocyte autophagy through PI3K/AKT/mTOR signaling pathway. J Orthop Translat. 2025;55:146-58
242. Liu X, Gao X, Yang Y, Yang D, Guo Q, Li L. et al. EVA1A reverses lenvatinib resistance in hepatocellular carcinoma through regulating PI3K/AKT/p53 signaling axis. Apoptosis. 2024;29:1161-84
243. Liu Y, Su Z, Tavana O, Gu W. Understanding the complexity of p53 in a new era of tumor suppression. Cancer Cell. 2024;42:946-67
244. López de Sá A, Díaz-Tejeiro C, Poyatos-Racionero E, Nieto-Jiménez C, Paniagua-Herranz L, Sanvicente A. et al. Considerations for the design of antibody drug conjugates (ADCs) for clinical development: lessons learned. J Hematol Oncol. 2023;16:118
245. Lou L, Zhu Z, Xu L, Wu Z, Wang X, Yuan L. et al. Glabridin mitigates osteoarthritis progression through modulation of the PI3K/AKT/FOXO3A autophagy axis. Phytomedicine. 2025;148:157359
246. Lu J, Wang H, Zhang H, Li J, Li H, Chen Q. et al. Gut Metabolite Indole-3-Propionic Acid Regulates Macrophage Autophagy Through PPT1 Inhibiting Aging-Related Myocardial Fibrosis. Adv Sci (Weinh). 2025;12:e01070
247. Lv W, Yuan J, Ruan Z, Dou Y-N, Zhang H, Fei X. et al. Single-Cell Transcriptome Analysis Reveals That Hmga2 Regulates Neuroinflammation and Retinal Function by Modulating Müller Cell Autophagy Through PI3K/AKT Signaling Following MCAO-Induced Retinal Ischemia. Adv Sci (Weinh). 2025;12:e02534
248. Mannan AA, Darlington APS, Tanaka RJ, Bates DG. Design principles for engineering bacteria to maximise chemical production from batch cultures. Nat Commun. 2025;16:279
249. Nachmias N, Dotan N, Rocha MC, Fraenkel R, Detert K, Kluzek M. et al. Systematic discovery of antibacterial and antifungal bacterial toxins. Nat Microbiol. 2024;9:3041-58
250. Ni Y, Yang W, Wang S, Pan Y, Du H, Zheng L. et al. Modified huangfeng decoction alleviates diabetic nephropathy by activating autophagy and regulating the gut microbiota. Phytomedicine. 2025;141:156677
251. Nishikawa S, Sato G, Takada S, Kohyama S, Honda G, Yanagisawa M. et al. Multimolecular Competition Effect as a Modulator of Protein Localization and Biochemical Networks in Cell-Size Space. Adv Sci (Weinh). 2023;11:e2308030
252. Niu X, You Q, Hou K, Tian Y, Wei P, Zhu Y. et al. Autophagy in cancer development, immune evasion, and drug resistance. Drug Resist Updat. 2024;78:101170
253. O'Dwyer PJ, Gray RJ, Flaherty KT, Chen AP, Li S, Wang V. et al. The NCI-MATCH trial: lessons for precision oncology. Nat Med. 2023;29:1349-57
254. Pan Y, Wei J, Qin D, Zhou X, Gao F, Yu L. et al. Prosapogenin CP4 exacerbates mitophagy to induce apoptosis via AMPK-mTOR and PINK1/Parkin pathways in A549 cells. Phytomedicine. 2025;148:157333
255. Pourmorady AD, Bashkirova EV, Chiariello AM, Belagzhal H, Kodra A, Duffié R. et al. RNA-mediated symmetry breaking enables singular olfactory receptor choice. Nature. 2023;625:181-8
256. Qi Z, Yang W, Xue B, Chen T, Lu X, Zhang R. et al. ROS-mediated lysosomal membrane permeabilization and autophagy inhibition regulate bleomycin-induced cellular senescence. Autophagy. 2024;20:2000-16
257. Rihal V, Khan H, Kaur A, Singh TG. Vitamin D as therapeutic modulator in cerebrovascular diseases: a mechanistic perspectives. Crit Rev Food Sci Nutr. 2022;63:7772-94
258. Shi L, Duan R, Sun Z, Jia Q, Wu W, Wang F. et al. LncRNA GLTC targets LDHA for succinylation and enzymatic activity to promote progression and radioiodine resistance in papillary thyroid cancer. Cell Death Differ. 2023;30:1517-32
259. Song N-J, Xie J, Jung KJ, Wang Y, Pozniak J, Roda N. et al. Tumor-associated NK Cells Regulate Distinct CD8+ T-cell Differentiation Program in Cancer and Contribute to Resistance against Immune Checkpoint Blockers. Cancer Discov. 2025;15:1835-57
260. Song W, Liu W, Li S-Y, Yu Y, Xu H, Shi T. et al. Remodeling the Senescent Microenvironment for Promoting Osteoporotic Tendon-to-Bone Healing via Synergizing Senolytic Quercetin and Aligned Nanowire-Structured Hydrogels. ACS Nano. 2025;19:18364-85
261. Sun G, Xu D, Jiang Y, Su K, Lu J, Zhou C. PPARγ-Axl axis ameliorates intervertebral disc degeneration by activating PI3K/AKT-mediated autophagy to suppress ferroptosis. Cell Mol Biol Lett. 2025;30:128
262. Chen X, Li Q, Liu X, Liu C, Liu R, Rycaj K. et al. Defining a Population of Stem-like Human Prostate Cancer Cells That Can Generate and Propagate Castration-Resistant Prostate Cancer. Clin Cancer Res. 2016;22:4505-16
263. Pai VC, Hsu C-C, Chan T-S, Liao W-Y, Chuu C-P, Chen W-Y. et al. ASPM promotes prostate cancer stemness and progression by augmenting Wnt-Dvl-3-β-catenin signaling. Oncogene. 2018;38:1340-53
264. Bradley CA. Correlates of alisertib response. Nat Rev Urol. 2018;15:726
265. Ahmed I, Mohamed A, Savari O, Xue Y, Asa SL. Expression of DLL3 and SEZ6 in the Spectrum of Neuroendocrine Neoplasia. Endocr Pathol. 2025;36:27
266. Canalis E. Notch in skeletal physiology and disease. Osteoporos Int. 2018;29:2611-21
267. Kim S, Park JM, Park S, Jung E, Ko D, Park M. et al. Suppression of TNBC metastasis by doxazosin, a novel dual inhibitor of c-MET/EGFR. J Exp Clin Cancer Res. 2023;42:292
268. Zhou Q, Chen X, He H, Peng S, Zhang Y, Zhang J. et al. WD repeat domain 5 promotes chemoresistance and Programmed Death-Ligand 1 expression in prostate cancer. Theranostics. 2021;11:4809-24
269. Schroeder A, Herrmann A, Cherryholmes G, Kowolik C, Buettner R, Pal S. et al. Loss of androgen receptor expression promotes a stem-like cell phenotype in prostate cancer through STAT3 signaling. Cancer Res. 2013;74:1227-37
270. Zhang W, Liu B, Wu W, Li L, Broom BM, Basourakos SP. et al. Targeting the MYCN-PARP-DNA Damage Response Pathway in Neuroendocrine Prostate Cancer. Clin Cancer Res. 2017;24:696-707
271. Viswanadhapalli S, Dileep KV, Zhang KYJ, Nair HB, Vadlamudi RK. Targeting LIF/LIFR signaling in cancer. Genes Dis. 2021;9:973-80
272. Sooreshjani MA, Kamra M, Zoubeidi A, Shah K. Correction: Reciprocal deregulation of NKX3.1 and AURKA axis in castration-resistant prostate cancer and NEPC models. J Biomed Sci. 2025;32:100
273. Abida W, Cyrta J, Heller G, Prandi D, Armenia J, Coleman I. et al. Genomic correlates of clinical outcome in advanced prostate cancer. Proc Natl Acad Sci U S A. 2019;116:11428-36
274. Beltran H, Oromendia C, Danila DC, Montgomery B, Hoimes C, Szmulewitz RZ. et al. A Phase II Trial of the Aurora Kinase A Inhibitor Alisertib for Patients with Castration-resistant and Neuroendocrine Prostate Cancer: Efficacy and Biomarkers. Clin Cancer Res. 2018;25:43-51
275. Deng B, Ke B, Tian Q, Gao Y, Zhai Q, Zhang W. Targeting AURKA with multifunctional nanoparticles in CRPC therapy. J Nanobiotechnology. 2024;22:803
276. Carver BS. Defining and Targeting the Oncogenic Drivers of Neuroendocrine Prostate Cancer. Cancer Cell. 2016;29:431-2
277. Dong Q, Qiu H, Piao C, Li Z, Cui X. LncRNA SNHG4 promotes prostate cancer cell survival and resistance to enzalutamide through a let-7a/RREB1 positive feedback loop and a ceRNA network. J Exp Clin Cancer Res. 2023;42:209
278. Formaggio N, Rubin MA, Theurillat J-P. Loss and revival of androgen receptor signaling in advanced prostate cancer. Oncogene. 2021;40:1205-16
279. Mosquera JM, Beltran H, Park K, MacDonald TY, Robinson BD, Tagawa ST. et al. Concurrent AURKA and MYCN gene amplifications are harbingers of lethal treatment-related neuroendocrine prostate cancer. Neoplasia. 2013 15
280. Nieto-Jiménez C, Morafraile EC, Alonso-Moreno C, Ocaña A. Clinical considerations for the design of PROTACs in cancer. Mol Cancer. 2022;21:67
281. Shah K, Cook M. LIMK2: A Multifaceted kinase with pleiotropic roles in human physiology and pathologies. Cancer Lett. 2023;565:216207
Author contact
Corresponding authors: Youzhi Wang. Department of Urology, Peking University Third Hospital, Beijing, 100191, P.R. China. 2516394211edu.cn. Shudong Zhang. Department of Urology, Peking University Third Hospital, Beijing, 100191, P.R. China. zhangshudongedu.cn.

 Global reach, higher impact
Global reach, higher impact