Impact Factor
ISSN: 1449-2288
Int J Biol Sci 2017; 13(1):32-45. doi:10.7150/ijbs.16287 This issue Cite
Research Paper
Deformed Skull Morphology Is Caused by the Combined Effects of the Maldevelopment of Calvarias, Cranial Base and Brain in FGFR2-P253R Mice Mimicking Human Apert Syndrome
Department of Rehabilitation Medicine, Center of Bone Metabolism and Repair, State Key Laboratory of Trauma, Burns and Combined Injury, Trauma Center, Research Institute of Surgery, Daping Hospital, Third Military Medical University, Chongqing 400042, China.
† The first two authors contributed equally to this work.
Abstract
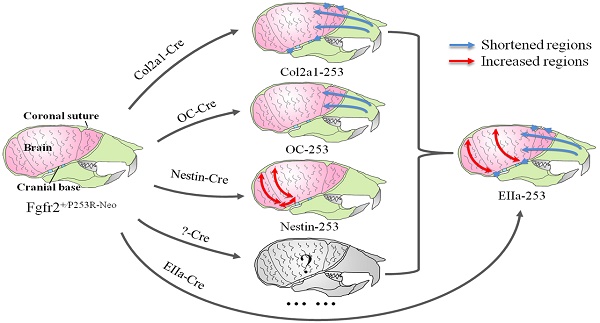
Apert syndrome (AS) is a common genetic syndrome in humans characterized with craniosynostosis. Apert patients and mouse models showed abnormalities in sutures, cranial base and brain, that may all be involved in the pathogenesis of skull malformation of Apert syndrome. To distinguish the differential roles of these components of head in the pathogenesis of the abnormal skull morphology of AS, we generated mouse strains specifically expressing mutant FGFR2 in chondrocytes, osteoblasts, and progenitor cells of central nervous system (CNS) by crossing Fgfr2+/P253R-Neo mice with Col2a1-Cre, Osteocalcin-Cre (OC-Cre), and Nestin-Cre mice, respectively. We then quantitatively analyzed the skull and brain morphology of these mutant mice by micro-CT and micro-MRI using Euclidean distance matrix analysis (EDMA). Skulls of Col2a1-Fgfr2+/P253R mice showed Apert syndrome-like dysmorphology, such as shortened skull dimensions along the rostrocaudal axis, shortened nasal bone, and evidently advanced ossification of cranial base synchondroses. The OC-Fgfr2+/P253R mice showed malformation in face at 8-week stage. Nestin-Fgfr2+/P253R mice exhibited increased dorsoventral height and rostrocaudal length on the caudal skull and brain at 8 weeks. Our study indicates that the abnormal skull morphology of AS is caused by the combined effects of the maldevelopment in calvarias, cranial base, and brain tissue. These findings further deepen our knowledge about the pathogenesis of the abnormal skull morphology of AS, and provide new clues for the further analyses of skull phenotypes and clinical management of AS.
Keywords: Apert syndrome, Fgfr2+/P253R mouse, Tissue specific activation, Skull morphology, Euclidean distance matrix analysis.

 Global reach, higher impact
Global reach, higher impact