ISSN: 1449-2288
Int J Biol Sci 2019; 15(10):2119-2127. doi:10.7150/ijbs.32860 This issue Cite
Research Paper
The Grass Carp Genomic Visualization Database (GCGVD): an informational platform for genome biology of grass carp
1. College of Information Technology, Shanghai Ocean University, Shanghai, 201306, China.
2. Key Laboratory of Fisheries Information Ministry of Agriculture, Shanghai, 201306, China.
3. College of Fisheries and Life Science, Shanghai Ocean University, Shanghai, 201306, China.
Abstract
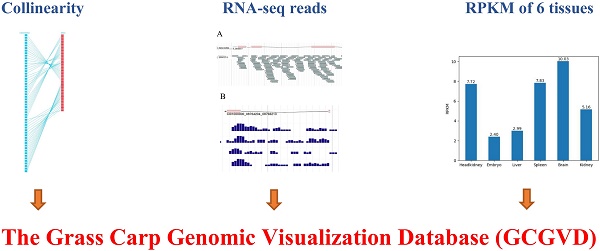
With the release of the draft genome of the grass carp, researches on the grass carp from the genetic level and the further molecular mechanisms of economically valuable physiological behaviors have gained great attention. In this paper, we integrated a large number of genomic, genetic and some other data resources and established a web-based grass carp genomic visualization database (GCGVD). To view these data more effectively, we visualized grass carp and zebrafish gene collinearity and genetic linkage map using Scalable Vector Graphics (SVG) format in the browser, and genomic annotations by JBrowse. Furthermore, we carried out some preliminary study on a whole-genome alternative splicing (AS)of the grass carp. The RNA-seq reads of 15 samples were aligned to the reference genome of the grass carp by Bowtie2 software. RNA-seq reads of each sample and density map of reads were also exhibited in JBrowse. Additionally, we designed a universal grass carp genome annotation data model to improve the retrieval speed and scalability. Compared with the published database GCGD previously, we newly added the visualization of some more genomic annotations, conserved domain and RNA-seq reads aligned to the reference genome. GCGVD can be accessed at
Keywords: Grass carp, Database, Conserved domain, RNA-seq

 Global reach, higher impact
Global reach, higher impact