Impact Factor ISSN: 1449-2288
Int J Biol Sci 2020; 16(9):1575-1585. doi:10.7150/ijbs.41852 This issue Cite
Review
STAT4: an immunoregulator contributing to diverse human diseases
State Key Laboratory of Organ Failure Research, Guangdong Key Laboratory of Viral Hepatitis Research, Department of Infectious Diseases and Hepatology Unit, Nanfang Hospital, Southern Medical University, GuangZhou, China.
Received 2019-11-6; Accepted 2020-2-13; Published 2020-3-5
Abstract
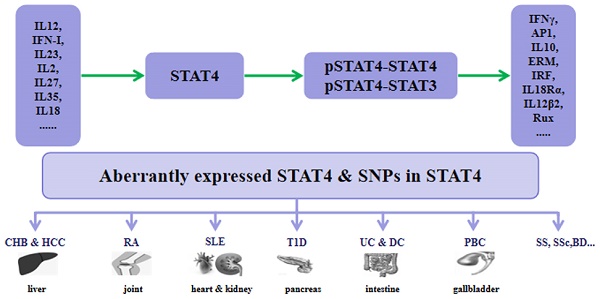
Signal transducer and activator of transcription 4 (STAT4) is a member of the STAT family and localizes to the cytoplasm. STAT4 is phosphorylated after a variety of cytokines bind to the membrane, and then dimerized STAT4 translocates to the nucleus to regulate gene expression. We reviewed the essential role played by STAT4 in a wide variety of cells and the pathogenesis of diverse human diseases, especially many kinds of autoimmune and inflammatory diseases, via activation by different cytokines through the Janus kinase (JAK)-STAT signaling pathway.
Introduction
Signal transducer and activator of transcription (STAT) proteins were identified as the major components of DNA-binding proteins that activate gene transcription in response to a variety of cytokines [1]. Seven different members in the STAT family (STAT1, STAT2, STAT3, STAT4, STAT5A, STAT5B, and STAT6) have been found to regulate many physiological and pathological processes, from pathogen response to cytokine secretion [2]. The Janus kinase (JAK)-STAT pathway is a common pathway in the signaling activated by many cytokines. In JAK-STAT pathway, STAT4 was first discovered to be crucial for promoting cellular-mediated immune responses via the differentiation of Th1 cells [3].
STAT4 protein contains six domains that have different functions in the JAK-STAT pathway: 1. N-terminal domain: dimerizes inactivated STATs and promotes nuclear translocation; 2. helical coiled coil: provides a carbonized hydrophilic surface and binds to regulatory factors; 3. DNA-binding domain: binds to an enhancer of the GAS family; 4. linker domain: involves in the DNA binding process; 5. Src homology (SH2) domain: binds specifically to the cytokine receptor after tyrosine phosphorylation; and 6. C-terminal transactivation domain: activates transcriptional process [4]. Additionally, with or without C-terminal transactivation, there are two spliced STAT4 transcripts, including STAT4α and STAT4β. STAT4α induces more IFN-γ production than that by STAT4β, whereas STAT4β proliferates more vigorously in response to IL12 stimulation [5].
In this review, we discuss the cytokines activating STAT4 in different cells, as well as the role of STAT4 in inflammation and complex diseases, especially autoimmune diseases.
Cytokines that activate STAT4
As shown in figure 1, different combinations of STAT4 are activated by a variety of cytokines, including interleukin (IL)12, type I interferon (IFN-I), IL23, IL2, IL27, and IL35, etc.
Cytokines that activate STAT4. Cytokines and receptor complexes involved in STAT4 signaling. JAK: Janus kinase; STAT: Signaling transducer and activator of transcription; TYK: Tyrosine kinase; STAT4 is phosphorylated after a variety of cytokines (IL12/ IFN-I/ IL23/ IL2/ IL27/ IL35) bind to the membrane, and then dimerized STAT4 translocates to the nucleus to regulate gene expression. Additionally, IL2 enhances the response of cells through up-regulation of the IL12 receptor.
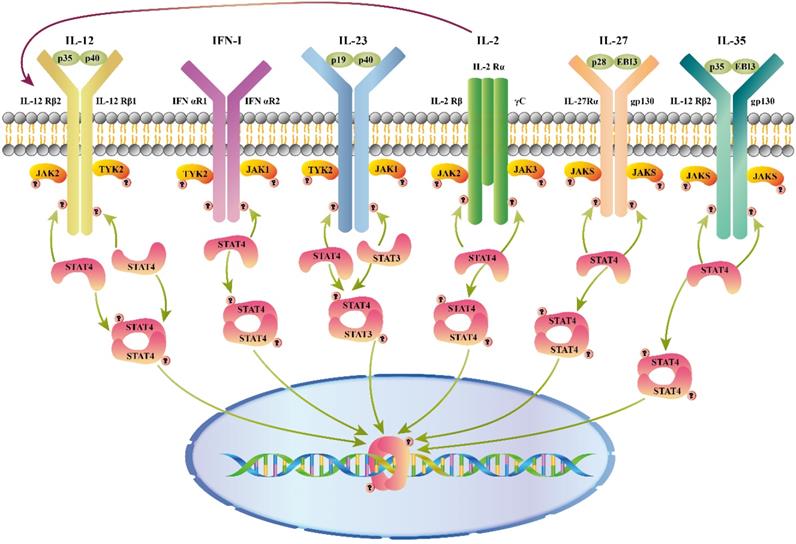
IL12
IL12 is produced by B cells and antigen- presenting cells and is secreted as a pro-inflammatory cytokine in the form of a heterodimer [6]. IL12 receptor (IL12R) is composed of two different subunits, including IL12Rβ1 and IL12Rβ2 [7]. Upon binding to IL12R, the JAK2 and TYK2 are linked to IL12Rβ2 and IL12Rβ1, and then STAT4 is phosphorylated on tyrosine 693 [8, 9]. Moreover, STAT4 is phosphorylated on serine 721 during activation of the p38/MKK6 signaling pathway [10].
The IL12-JAK-STAT4 pathway increases IFNγ production and Th1 cell differentiation [11, 12]. Several other genes that require STAT4 for transcriptional activation have been identified, including activator protein 1 (AP1), IL10, ERM, IFN regulating factor (IRF)-1/4/8, IL18Rα, IL12β2, and Rux. STAT4 binds c-Jun, and then interacts with AP1-relevant promoter [13]. A conserved STAT4- binding element was found in the fourth intron of the IL10 gene [14]. The ETS transcription factor, EMR, is selectively expressed in Th1 cells. ERM can modulate IFNγ gene transcription with STAT4 or some STAT4 inducible factors [15]. IRF1 gene is induced via IL12-dependent transactivation of IRF1 in human natural killer (NK) and T cells [16]. Additionally, it suggests that IL12 may further strengthen innate immune responses by inducing the expression of IRF4 and IRF8 genes [17]. IL12 induces the binding of STAT4 to the IL12Rβ2 enhancer to form a positive feedback loop of IL12/STAT4 axis during T cell receptor (TCR) stimulation [18]. STAT4 binds directly to the IL18Rα locus and alters its acetylation, reducing metastatic binding and DNA methylation transiently and resulting in high expression of IL18Rα in Th1 cells [19]. The promoter regions of Runx1 and Runx3 are targets of STAT4 to promote the antiviral activity of NK cells [20]. Therefore, the IL12/STAT4 axis is vital for inflammatory cytokines secretion that participates in many diseases and anti-tumor responses. Interestingly, Gao B et al. found that Stat4 -/- mice are more susceptible to concanavalin A-induced T-cell hepatitis, which is inconsistent with the traditional view. Compared with wild-type mice, higher FasL expression in NK T cells in ConA-treated Stat4-/- mice results in stronger cytotoxicity against hepatocytes. STAT4 may directly or indirectly inhibit the expression of FasL in NK T cells in case of the up-regulation of pro-inflammatory cytokines [21].
IL12 synergizes with IL18 to enhance both cytotoxicity and IFNγ production [22-24]. MicroRNA (miRNA)-21 inhibits the expression of IL12 by targeting its mRNA [25]. Wu et al. showed that miRNA-21 inhibits the expression of IL12 and STAT4 by regulating IL12 and STAT4 translation post- transcriptionally [26]. Moreover, it has recently been recognized that T follicular helper cells (Tfh cells), a different subset of Th cells, have specialized functions on promoting germinal center formation and regulating B cell function. STAT4 is required for the induction of multiple genes in the development of Tfh cells [27]. IL12Rβ1-deficient subjects do not completely lack Tfh cells, because they are able to show nearly normal levels of CXCR5+ CD4+ T cells during aging. Blocking the IL12/STAT4 signaling pathway might be beneficial for the treatment of these diseases by promoting the Tfh response [28]. Stat4 -/- mice with allergic lung inflammation have increased numbers of regulatory T (Treg) cells in the lung, which correlates with decreased inflammation. Stat4 may not only promote inflammation caused by T cell subsets but also limit the development of Treg cells via promoting a repressive chromatin configuration at the Foxp3 locus [29]. A recent study showed that Stat4-deficient mice have up-regulated numbers of CD4+Foxp3+ Tregs in mesenteric lymph nodes. Similarly, Foxp3 expression was found to be significantly increased. These findings suggest that impaired STAT4 contributes to Foxp3+ Treg response [30].
IFN-I (IFNα/β)
IFN-I is divided into two categories: IFNα, produced by human fibroblasts, and IFNβ, produced by human leukocytes, which function as regulators of antiviral, cell proliferation inhibitory, immunity, and anti-tumor effects. Type I IFN receptor (IFNAR) contains IFNAR1 and IFNAR2. The binding of IFNα/β to IFNAR leads to the phosphorylation of STAT1 and STAT4 and activation of IFN target genes [31]. During the initial phase of a viral infection, IFNα/β mainly activate STAT4 instead of STAT1 in NK cells [32]. IFNα/β also have been reported to induce IFNγ in activated T cells and macrophages via the IFNα/β-STAT4 pathway [33, 34]. In response to IFNβ, STAT4 is activated in bone marrow-derived mast cells (BMMCs), which participate in immune regulation (T and B cells, APC cell activation) [34]. IL5 and IL13 induce eosinophil activation and enhance mucus production as highly inflammatory cytokine. IFNα/β inhibit expression of IL5 and IL13 in PBMCs [35,36]. IFNα/β induce STAT4 binding within the IL5 promoter, repressing nascent transcription in human memory Th2 cells [37-39]. Moreover, Zhao et al. reported that STAT4 promotes retinoic acid inducible gene-1 (RIG1)-triggered type I IFN production when exposed to viral double-stranded (ds) RNA. IFNβ production in macrophages decreases after RNA virus infection in the absence of STAT4 [40]. Unconventionally, during RNA virus infection, STAT4 was found unphosphorylated in the cytoplasm of macrophages. Mechanistically, STAT4 interacts with E3 ligase CHIP, which mediates the K48-linked ubiquitination of RIG1, preventing proteasomal degradation of RIG1. These findings indicate a novel role of STAT4 [40].
IL23
When exposed to the pathogen-derived molecules such as Gram-positive/negative bacterial and viral products, IL23 is produced by monocytes, activated DC, and macrophages [41-43]. IL23 consists of IL23p19 and IL12/IL23p40 subunits. IL23 receptor (IL23R) is composed of IL12β1 and IL23R [44,45]. IL23 can exert biological effects similar to IL12 via IL23R-dependent STAT4 phosphorylation, but its effects are significantly weaker than those of IL12. Unlike IL12, which only activates STAT4-STAT4 DNA-binding complexes, IL23 induces several STAT-containing DNA-binding complexes, including STAT4-STAT4 homo-dimers and STAT3-STAT4 heterodimers [46]. Notably, IL23 is involved in chronic inflammatory by inducing the differentiation and expansion of Th17 through the IL23/STAT4 signaling pathway [47]. In addition to Th cells, memory Th cells and NK T cells also display decreased IL17 production induced by IL23 [48]. The IL23/STAT4 pathway has been implicated in a variety of human autoimmune and inflammatory diseases [49,50]. Significantly, although IL12 and IL23 have overlapping effects in disease, it is unclear how IL23 negatively regulates IFNγ production induced by IL12 [50-51].
IL2
IL2 is a multifunctional cytokine produced mainly by the subset of activated CD4+ T helper cells that act on a large variety of cells, including T and B cells, NK cells, NK T cells, neutrophils, macrophages, and monocytes after antigen activation [52]. IL2 does not affect the phosphorylation of STAT4 in T lymphocytes, although it does activate STAT4 in NK cells [53]. After stimulation with IL2, NK cells that have continuous STAT4 expression exhibit hyper-responsiveness to IL12. IL2 up-regulates the expression of IL12β1 and IL12Rβ2, and contributes to enhance and maintain the expression of STAT4 in NK cells. Through over-expression of IL12R and STAT4, NK cells show increased functional responses to IL12 [54]. IL2 receptor (IL2R) has been shown to be induced via STAT4 signaling [55]. In addition, IL2 plus IL18 suppress allergic inflammation through up-regulation of IL12/STAT4 signaling, increasing IFNγ produced by NK cells [56].
IL27
IL27 is composed of the Epstein - Barr virus induced 3 (EBI3, IL12B-associated protein) and IL30 (IL12A-related protein), mainly secreted by antigen-presenting cells. It is reported to drive rapid expansion of CD4+ T cells [57]. IL27 receptor (IL27R) consists of the IL27R alpha (IL27Rα) subunit and gp130, which is expressed in various immune and non-immune cells. IL27 signaling induces phosphorylation of STAT4 through TCCR/WSX-1 [58]. Additionally, IL27 induces the expression of IL12Rβ2 in CD4+ T cells by developing Th1 cells via inducing T-bet. Therefore, IL27 can establish IL12 responsiveness to STAT4 phosphorylation, contributing to the differentiation of naïve Th cells into Th1 cells during T cell immune response caused by antigen-presenting cells in vivo [59].
IL35
IL35 is an immunosuppressive cytokine produced by Treg cells and plays an important role in many disease models. IL35 receptor (IL35R) consists of the IL12Rβ2 and gp130 domains. Naive T cells are converted into regulatory T cells (iTr35 cells) via IL35R signaling with STAT1-STAT4 heterodimer [60]. Phosphorylated STAT1 and STAT4 expand the number of Tregs in the spleen after stimulation by IL35 [61]. IL35 plays critical roles in the pathogenesis of concanavalin A-induced fulminant hepatitis, reducing the FASL expression in liver mononuclear cells through the STAT1/STAT4 pathway [62]. In addition to affecting inflammation, IL35 reduces cardiac rupture, improves wound healing, and attenuates cardiac remodeling in mice with myocardial infarction by activating CX3C chemokine receptor 1 (Cx3cr1) and transforming growth factor beta-1 (Tgfβ1) in macrophages via phosphorylating Stat1 and Stat4 [63].
IL18
IL18 belongs to the IL1 family and can be produced by a variety of tissue cells. It can induce Th1 cells to produce cytokines, enhance cytotoxic activity of NK cells, promote T cell proliferation, and synergize with IL12 [64]. Unlike IL12, IL18 functions as a cofactor for Th1 cell development and produces substantial amounts of IFNγ [65]. IL12 and STAT4 affect IL18 signaling by inducing expression of MyD88 and directly affecting cellular responses to the expression of IL8 receptor (IL18R). Regulation of IFNγ may also include interactions between STAT4 and AP1 at the IFNγ promoter [66]. When exposed to IL12 and IL18, it enhances co-binding of the c-Jun-STAT4 complex to the AP1-relevant promoter sequence, increasing IFNγ production mediated by AP1 [67].
IL21
IL21 is produced by Tfh cells, and regulates the functions of NK and T cells. IL21 enhances the expression of the IFNγ gene via activating STAT4 and subsequently recruiting pSTAT4 to IFNγ transcriptional sites [68]. Interestingly, IL12 also regulates the production of IL21 in CD4+ T cells via STAT4 signaling [69]. Details of this feedback loop remain to be determined.
Negative regulators of STAT4 signaling pathways
A great deal has been discovered about STAT4 activation in response to cytokines mentioned above, whereas some studies elucidated the mechanism by which the activity of STAT4 is turned off. As shown in figure 2, suppressor of cytokine signaling 3 (SOCS3), protein inhibitor of activated STAT (PAIS), protein tyrosine phosphatases (PTPs), and STAT-interacting LIM protein (SLIM) are believed to regulate the IL12/STAT4 signaling pathway. In addition, some miRNAs were reported to regulate STAT4 expression.
Negative regulators of STAT4 signaling pathways. Negative regulators of STAT protein suppressors of cytokine signaling (SOCS3) blocks STAT activation in the cytoplasm by binding to phosphorylated sites of receptor, turning off the initial signal at the source in cytoplasm. Protein tyrosine phosphatase (PTP) inhibits activation by direct dephosphorylation of activated proteins in cytoplasm and nucleus. SLIM can impair the tyrosine phosphorylation of STAT4 to recruit a nuclear tyrosine phosphatase. PIASx forms a complex with activated Stat4 binding to DNA and prevents Stat4 from binding to DNA.
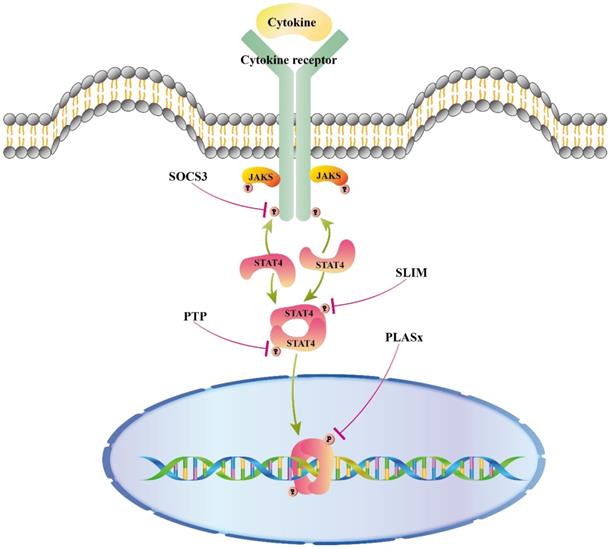
SOCS3
SOCSs are released by cytokine induction, such as IL12 and IL6, and suppress the signaling of cytokines in a negative feedback loop. Phosphorylation of JAK and STAT is down-regulated in cells with high expression of SOCSs [70]. Previous studies showed that SOCS3 inhibits IL6 signaling via blocking activation of STAT3 [71]. More recently, SOCS3 was shown to inhibit IL12 signaling in a negative-feedback loop. SOCS3 is recruited to IL12Rβ2, Tyr-800, the pSTAT4-recruitment site, via its SH2 domain, resulting in decreased levels of activated STAT4 [72].
PAIS
PIAS functions as a regulator of transcriptional activity in the nucleus [73]. PIASx was found in the STAT4-DNA binding complex, inhibiting STAT4- induced gene activation in T cells instead of inhibiting the combination of STAT4 and DNA. By suppressing histone deacetylase (HDAC) activity, the inhibitory activity of PIASx can be diminished. These results suggest that PIASx may be a transcriptional co-repressor of STAT4 [74].
PTPs
PTPs can remove phosphate groups from phosphorylated tyrosine in proteins. As a requirement of JAK-STAT signaling for tyrosine phosphorylation, PTPs can dephosphorylate phosphorylated proteins such as STAT4 and PTP non-receptor type 13 (PTPN13) through its PTPase domain [75].
SLIM
SLIM is a STAT ubiquitin E3 ligase. It recruits a nuclear tyrosine phosphatase through its unique domains and then blocks tyrosine phosphorylation of STAT4 [76]. IFNγ production is decreased in CD4+ T cells in EAE mice via up-regulating SLIM by berbamine [77].
MiRNA
MiRNA functions by degrading mRNA and regulating the post-transcriptional modifications of gene expression that leads to inhibition of protein translation [78-79]. Some miRNAs have been found to interfere with IL12/STAT4 signaling (Table 1). MiRNA-146a, a key player for regulating inflammation and immune response, targets protein kinase C epsilon (PRKCε). PRKCε binds to STAT4 and then drives Th1 cell differentiation in human CD4+ T cells. Therefore, miRNA-146a can weaken the Th1 cell immune response by targeting PRKCε [80-81]. MiRNA-132, miRNA-212, and miRNA-200a contribute to the down-regulation of STAT4 in NK cells via targeting the STAT4 3′-UTR [82]. SiRNA-mediated knockdown of miRNA-155 leads to up-regulation in STAT4 expression in MyLa cells, and up-regulation of oncogenic miRNA-155 results in loss of STAT4 expression via targeting the 3'-UTR of STAT4 [83]. MiR-141 have been found to target the 3′-UTR of STAT4, inhibiting the gastric cancer cells invasion and migration [84]. STAT4 expression is down-regulated in macrophages after overexpression of miRNA-320a, which has binding sites in the 3′-UTR of the STAT4 gene [85].
MicroRNAs had been found to interfere with the expression of STAT4 and its pathway.
| MicroRNAs | Targets |
|---|---|
| MicroRNA-146a [80] | PRKCε |
| MicroRNA-132 [81] | 3'-UTR of STAT4 |
| MicroRNA-212 [81] | 3'-UTR of STAT4 |
| MicroRNA-200a [81] | 3'-UTR of STAT4 |
| MicroRNA-155 [82] | 3'-UTR of STAT4 |
| MicroRNA-141 [83] | 3'-UTR of STAT4 |
| MicroRNA-320a [84] | 3'-UTR of STAT4 |
STAT4 and human diseases
STAT4 is a pivotal mediator in inflammation and tumor development. Understanding the molecular mechanisms of STAT4 in immune responses and immune-mediated diseases would allow the development of novel therapeutic options for human diseases such as chronic hepatitis B (CHB), hepatocellular carcinoma (HCC), rheumatoid arthritis (RA), systemic lupus erythematosus (SLE), type 1 diabetes (T1D), psoriasis, inflammatory bowel diseases (IBDs), Behçet's disease (BD), Sjögren's syndrome (SS), systemic sclerosis (SSc), primary biliary cirrhosis (PBC), and other diseases.
CHB and HCC
CHB is the most critical risk factor for HCC and is caused by hepatitis B virus (HBV) infection [86]. In the 1990s, a complex segregation analysis provided evidence for the association of host genetics with the occurrence of HCC and suggested an interaction between HBV infection and susceptibility loci [87]. Jiang DK et al. showed that a single nucleotide polymorphism (SNP), rs7574865, at STAT4 with lower mRNA levels of STAT4 is significantly associated with the HBV-related HCC risk [88]. Clark A et al. confirmed that rs7574865 at STAT4 is associated with the susceptibility of CHB induced HCC [89]. In addition, STAT4 rs7574865 was reported to be associated with CHB susceptibility as well as HBV natural clearance [90-92]. Recently, Jiang DK et al. reported that STAT4 rs7574865 is a reliable predictor of response to IFNα therapy, a first-line therapy for CHB, in hepatitis B e antigen (HBeAg)-positive CHB patients [93, 94]. Hepatectomy is a main therapy for HCC, and lower STAT4 expression showed higher incidence of recurrence after this procedure. STAT4 may regulate the IFNγ production by CD8+ T-cell infiltration of tumor tissue [95]. STAT4 was reported to be expressed at lower levels in HCC than in normal liver tissues and to be associated with serum hepatitis B surface antigen (HBsAg) level, tumor number, tumor size, and severity of HCC. STAT4 may modulate the pathological progression of HCC by inhibiting HCC cells proliferation, growth, and apoptosis, via acting as a tumor suppressor [96].
RA
RA is a systemic autoimmune disorder that leads to increased risk of progressive joint degeneration, disability, and cardiovascular complications [97]. Both genetic and environmental factors have been reported to affect the pathogenesis of RA. STAT4 contributes to the differentiation and proliferation of both Th1 and Th17 cells, which are crucial effectors in chronic inflammatory disorders [98]. Increased expression of STAT4 protein in dendritic cells in the synovial membrane is associated with the rheumatoid factor in the serum, which is a risk factor for RA [99, 100].
Dysregulation of STAT4 in fibroblasts also promotes the development and progression of RA [101]. IL6 is mainly secreted by fibroblasts in arthritis, regulating the inflammation process. During inflammation, fibroblasts up-regulate the expression of leukemia inhibitory factor (LIF, IL6 family cytokine) and STAT4, and the LIF/STAT4 signaling pathways contribute to IL6 transcription [102]. It is reported that Gαq in mice leads to autoimmune arthritis [103]. The level of phospho-STAT4 is higher in Gαq-/- CD4+ T cells than in controls, showing that Gαq modulates T-bet and STAT4 and then regulates Th1 cell differentiation in Gαq-/- mice [104]. Studies indicated that inappropriate activation of STAT4 in T cells drives the inflammation in RA after tumor necrosis factor alpha (TNFα) therapy, and is unlikely to contribute to RA disease activity against a pro-inflammatory effect of STAT4 [105]. Further studies on the activation of STAT4 are needed to clarify this contradiction.
SLE
SLE affects several organs (kidney, bone, muscle, and heart) and is caused by both environmental factors and genetic susceptibility variants [106]. STAT4 deficiency is associated with deterioration of clinical nephritis with the absence of anti-dsDNA Ab [107]. STAT4-specific antisense oligonucleotide improves advanced nephritis caused by SLE [108]. Serum IFNα levels are frequently elevated in SLE patients. The SNP rs7574865 of STAT4 affects the sensitivity of IFNα to PBMCs, resulting in greater IFNα-induced gene expression [109].
T1D
T1D is caused by multiple environmental and genetic factors. A variety of autoimmune antibodies that damage insulin-producing B cells in human islets can be detected in patients with T1D, leading to hyperglycemia [110]. The response of islets and β-cells to the inflammatory cytokines weakens the cell viability and function and increases the induction of apoptosis. The IL12/STAT4 axis was found to induce β-cell apoptosis [111]. Serum levels of IFNγ and IL12 are reduced in Stat4-/-NOD (non-obese diabetic) mice, and the development of diabetes in NOD mice is prevented with the absence of Stat4. Importantly, Stat4-deficient pancreatic islets restore insulin levels. Moreover, Stat4-/- DIO (diet-induced obese) mice have increased insulin sensitivity and better glucose tolerance compared with controls [112]. In addition, Stat4 ameliorates adipose tissue inflammation that induces pancreatic β-cell dysfunction and atherosclerosis insulin resistance. Reducing the costimulatory effects of CD40 on CD8+ T cells protects against Stat4-deficiency in a diet-induced obesity model [113, 114].
Psoriasis
Psoriasis is characterized by the hyper- proliferation and aberrant differentiation of keratinocytes [115]. The migration of Th1 cells produces IFNγ, which leads to keratinocyte hyper-proliferation, small vessel proliferation, and inflammatory infiltration in the skin, events that play a key role in the pathogenesis of psoriasis [116]. Higher levels of pSTAT4 have been described in psoriatic T cells compared with the skin of non-psoriatic donors, which increases responses to IFNα and leads to up-regulated IFNγ production [117].
IBDs
IBDs, including ulcerative colitis (UC) and Crohn's disease (CD), are chronic inflammatory disorders characterized by aberrant mucosal Th1 cell activation and production of cytokines promoted by IL12/ STAT4 signaling [118]. IL12 is up-regulated in active CD and UC with high amounts of IFNγ produced by lamina propria lymphocytes, and STAT4 is activated in patients with increased expression of IL12Rβ2 [119]. As an alternatively spliced isoform of STAT4, STAT4β activates neutrophils present in the lamina propria by enhancing TNFα and GM-CSF secretion and promoting colitis in vivo. Regulation of STAT4 mRNA splicing is important in the progression of human disease [120]. Moreover, STAT4 activated by LIF inhibits Th17 accumulation and promotes repair of damaged intestinal epithelium blocking STAT3-dependent IL17α/IL17 promoter activation [121]. Interestingly, STAT4 variants affect DNA methylation status at position -172 of the STAT4 gene in IBD patients, and the T allele of STAT4 rs7574865 shows significantly higher methylation levels [122].
BD
BD is a systemic immune system disease that invades many organs of the human body, including the mouth, skin, joint muscles, eyes, blood vessels, heart, lungs, and the nervous system [123]. It has been reported that STAT4 polymorphisms confer susceptibility to BD. The risk alleles A of rs7574070 and A of rs897200 are associated with up-regulation of STAT4 along with increased expression of IL17 in patients with more serious BD, suggesting that these risk alleles contribute to BD through the Th17 pathway instead of the Th1 pathway [124, 125].
SS
SS is an autoimmune disorder caused by lymphocytic infiltration of exocrine glands that results in dysfunction of the salivary and lacrimal glands [124]. Patients with primary SS (pSS) have an increased response of IFN-I, leading to the etiopathogenesis and clinical outcome of the disease [126]. IFN-I/STAT4 signaling pathways play a key role in autoimmune diseases [127].Nordmark G et al. showed that IRF5 and STAT4 are components of the IFN-I system and emphasized the importance of this system in the etiopathogenesis of pSS [128].
SSc
SSc is a systemic connective-tissue disease characterized by excessive deposition of collagen in many tissues, causing fibrosis of skin, internal organs, and microvascular diseases [129]. STAT4 may play an essential role in fibrosis. Avouac J et al. found that fibrotic skin lesions induced by subcutaneous injections of bleomycin or saline in WT mice were almost totally absent in the Stat4-/- mice and had lower numbers of infiltrating leukocytes. STAT4 may increase the release of cytokines involved in fibrotic processes to regulate the activation of fibroblasts [130]. SSc includes two clinical phenotypes, limited cutaneous SSc (lcSSc) and diffuse cutaneous SSc (dcSSc). Rueda et al. found that STAT4 variants were associated with susceptibility to lcSSc instead of dcSSc [131].
PBC
PBC is characterized by a progressive T cell-predominant lymphocytic cholangitis, and positive anti-mitochondrial antibodies (AMA). Genes involved in IL12/STAT4 signaling in CD4+ T cells confer susceptibility risks for PBC [132]. Moreover, antinuclear antibodies (ANA) are associated with more rapid progression and a poorer prognosis in patients with PBC, and variants of STAT4 are reported to be associated with ANA [133]. Apart from IL12/Th1 signaling, bile duct cells increase IL23/Th17 signaling as expression of IL12Rβ2 and IL23R is increased in cholangiocytes in the initial phases of PBC [134]. Liaskou E et al. found increased sensitivity of PBC Treg cells to low doses of IL12 through the IL12-STAT4 pathway, driving their differentiation into IFN-γ secreting cells [135].
Other diseases
Endometriosis, ovarian cancer (OC), and chronic lymphocytic leukemia (CLL) B cells have also been found to be associated with STAT4 in recent years. Although there are only a few studies till present, they are important for understanding the development of these diseases.
Endometriosis is a common gynecological disorder that is histologically similar to the endometrial and/or stromal tissue grown in the uterine cavity and can lead to pelvic pain, dysmenorrhea, and infertility [136]. IFNγ has been found to be reduced in peripheral lymphocytes in endometriosis, and its concentration in PF is decreased [137]. Zamani MR et al. found a significant association between STAT4 rs7582694 and susceptibility to endometriosis [138].
OC is usually diagnosed at a late stage in gynecological cancers. Overexpression of STAT4 has been found in epithelial cells of OC and with poor prognosis of patients. STAT4 mediates the EMT process via cancer-stroma interactions, and promotes metastasis via cancer-associated fibroblasts induced by Wnt7a [139].
CLL is a B-cell neoplasm that accumulates monoclonal CD5+ B cells [140]. P66Shc attenuates BCR-dependent survival signals and modulates Bcl-2 expression as a regulator of B-cell apoptosis. The lack of p66Shc in CLL B cells causes an imbalance toward antiapoptotic Bcl-2 family members [141-143]. Recently, Guo et al. showed that silencing or over-expression of STAT4 resulted in co-modulation of transcriptional regulator of p66Shc in B cells. STAT4 activation increases the coordination of STAT4 and p66Shc expression, enhancing B cell apoptosis. P66Shc has been shown to promote STAT4 expression in a positive feedback loop [144].
Conclusion
Distinct types of cytokines can activate STAT4 in multiple cells such as tumor or immune cells via the JAK-STAT pathway. In vivo and in vitro studies in recent decades have suggested that STAT4 may induce inflammation and autoimmune diseases, inhibit tumor growth or promote tumors via regulating many facets of the innate and adaptive immune responses.
STAT4 increases the Th1 cells differentiation, cytotoxicity and IFNγ production of immune cells. In addition, STAT4 regulates the migration and proliferation of tumor cells. As it can be activated in both tumor cells and immune cells, we suspect that STAT4 may modulate the interaction between tumor cells and host immunity. More researches need to be carried out to understand whether STAT4 affects anti-cancer immune responses and immunologic microenvironment in tumors.
While STAT4 exerts functions in many tumor cells or immune cells such as T cell and NK cell, we lack insight into whether and how it regulates other kinds of cells, including Tfh cell [27-28], Treg cell [29-30], fibroblast [102], mast cell [145] and endothelial precursor cell [146]. If further researches are performed, additional molecular activities of STAT4 in these cells may be discovered.
Additionally, as we have mentioned, SNPs in STAT4 are associated with a variety of diseases. There have been limited researches about the molecular mechanisms by which the variants of STAT4 affect these diseases. Further studies in vivo and in vitro should focus on how the SNPs in STAT4 contribute to immune dysregulation and autoimmunity.
In summary, the cytokines activating STAT4, the responsive genes of STAT4 and the molecular mechanisms of how STAT4 is involved in human diseases require more studies to accelerate the breakthrough process, and treatment strategies for human diseases are also needed to be established based on breakthroughs regarding STAT4.
Acknowledgements
This study was supported by the General Programs from the National Natural Science Foundation of China (No. 81472618, 81670535), the National Science and Technology Major Project (No. 2017ZX10202202 and 2018ZX10301202), the Local Innovative and Research Teams Project of Guangdong Pearl River Talents Program (No. 2017BT01S131), the Research and Development Project in Key Areas of Guangdong Province (No. 2019B020227004), the Innovative Research Team Project of Guangxi Province (no. 2017GXNSFGA198002), the Grant for Recruited Talents to Start Scientific Research from Nanfang Hospital, and the Outstanding Youths Development Scheme of Nanfang Hospital, Southern Medical University (No. 2017J001).
Author Contributions
Chou Yang wrote the review article. Haoming Mai, Bin Zhou, Jinxin Peng and Deke Jiang reviewed the manuscript. All authors read and approved the final manuscript.
Competing Interests
The authors have declared that no competing interest exists.
References
1. Darnell JE Jr, Kerr IM, Stark GR. Stark. Jak-STAT pathways and transcriptional activation in response to IFNs and other extracellular signaling proteins. Science. 1994;264:1415-1421
2. Turkson J. STAT proteins as novel targets for cancer drug discovery. Expert Opin Ther Targets. 2004;8:409-422
3. Gao B. Cytokines, STATs and liver disease. Cell Mol Immunol. 2005;2:92-100
4. Levy DE, Darnell JJ. Stats: transcriptional control and biological impact. Nat Rev Mol Cell Biol. 2002;3:651-662
5. Hoey T, Zhang S, Schmidt N, Yu Q, Ramchandani S, Xu X. et al. Distinct requirements for the naturally occurring splice forms Stat4alpha and Stat4beta in IL-12 responses. EMBO J. 2003;22:4237-4248
6. Visekruna A, Volkov A, Steinhoff U. A key role for NF-kappaB transcription factor c-Rel in T-lymphocyte-differentiation and effector functions. Clin Dev Immunol. 2012;2012:239368
7. Teng MW, Bowman EP, McElwee JJ, Smyth MJ, Casanova JL, Cooper AM. et al. IL-12 and IL-23 cytokines: from discovery to targeted therapies for immune-mediated inflammatory diseases. Nat Med. 1996;21:719-729
8. Cho SS, Bacon CM, Sudarshan C, Rees RC, Finbloom D, Pine R. et al. Activation of STAT4 by IL-12 and IFN-alpha: evidence for the involvement of ligand-induced tyrosine and serine phosphorylation. J Immunol. 1996;157:4781-4789
9. Naeger LK, McKinney J, Salvekar A, Hoey T. Identification of a STAT4 binding site in the interleukin-12 receptor required for signaling. J Biol Chem. 1999;274:1875-1878
10. Visconti R, Gadina M, Chiariello M, Chen EH, Stancato LF, Gutkind JS. et al. Importance of the MKK6/p38 pathway for interleukin-12-induced STAT4 serine phosphorylation and transcriptional activity. Blood. 2000;96:1844-1852
11. Kaplan MH, Sun YL, Hoey T, Grusby MJ. Impaired IL-12 responses and enhanced development of Th2 cells in Stat4-deficient mice. Nature. 1996;382:174-177
12. Thierfelder WE, van Deursen JM, Yamamoto K, Tripp RA, Sarawar SR, Carson RT. et al. Requirement for Stat4 in interleukin-12-mediated responses of natural killer and T cells. Nature. 1996;382:171-174
13. Park WR, Nakahira M, Sugimoto N, Bian Y, Yashiro-Ohtani Y. et al. SA mechanism underlying STAT4-mediated up-regulation of IFN-gamma induction inTCR-triggered T cells. Int Immunol. 2004;16:295-302
14. Grant LR, Yao ZJ, Hedrich CM, Wang F, Moorthy A, Wilson K. et al. Stat4-dependent, T-bet-independent regulation of IL-10 in NK cells. Genes Immun. 2008;9:316-327
15. Ouyang W, Jacobson NG, Bhattacharya D, Gorham JD, Fenoglio D, Sha WC. et al. The Ets transcription factor ERM is Th1-specific and induced by IL-12 through a Stat4-dependent pathway. Proc Natl Acad Sci U S A. 1999;96:3888-3893
16. Galon J, Sudarshan C, Ito S, Finbloom D, O'Shea JJ. IL-12 induces IFN regulating factor-1 (IRF-1) gene expression in human NK and T cells. J Immunol. 1999;162:7256-7262
17. Lehtonen A, Lund R, Lahesmaa R, Julkunen I, Sareneva T, Matikainen S. IFN-alpha and IL-12 activate IFN regulatory factor 1 (IRF-1), IRF-4, and IRF-8 gene expression in human NK and T cells. Cytokine. 2003;24:81-90
18. Letimier FA, Passini N, Gasparian S, Bianchi E, Rogge L. Chromatin remodeling by the SWI/SNF-like BAF complex and STAT4 activation synergistically induce IL-12Rbeta2 expression during human Th1 cell differentiation. EMBO J. 2007;26:1292-1302
19. Yu Q, Thieu VT, Kaplan MH. Stat4 limits DNA methyltransferase recruitment and DNA methylation of the IL-18Ralpha gene during Th1 differentiation. EMBO J. 2007;26:2052-2060
20. Rapp M, Lau CM, Adams NM, Weizman OE, O'Sullivan TE, Geary CD. et al. Core-binding factor beta and Runx transcription factors promote adaptive natural killer cell responses. Sci Immunol. 2017;2:eaan3796
21. Wang Y, Feng D, Wang H, Xu MJ, Park O, Li Y. et al. STAT4 knockout mice are more susceptible to concanavalin A-induced T-cell hepatitis. Am J Pathol. 2014;184:1785-1794
22. Zheng Y, Wang M, Tian T, Liu K, Liu X, Zhai Y. et al. Role of interleukin-12 gene polymorphisms in the onset risk of cancer: a meta-analysis. Oncotarget. 2017;8:29795-29807
23. Trinchieri G. Interleukin-12: a cytokine at the interface of inflammation and immunity. Adv Immunol. 1998;70:83-243
24. Trinchieri G. Interleukin-12 and the regulation of innate resistance and adaptive immunity. Nat Rev Immunol. 2003;3:133-146
25. Lu TX, Munitz A, Rothenberg ME. MicroRNA-21 is up-regulated in allergic airway inflammation and regulates IL-12p35 expression. J Immunol. 2009;182:4994-5002
26. Wu SQ, Wang GL, Li LY, Ji J. Effects of microRNA-21 on the interleukin 12/signal transducer and activator of transcription 4 signaling pathway in asthmatic mice. Cent Eur J Immunol. 2014;39:40-45
27. Nakayamada S, Kanno Y, Takahashi H, Jankovic D, Lu KT, Johnson TA. et al. Early Th1 cell differentiation is marked by a Tfh cell-like transition. Immunity. 2011;35:919-931
28. Schmitt N, Bustamante J, Bourdery L, Bentebibel SE, Boisson-Dupuis S, Hamlin F. et al. IL-12 receptor beta1 deficiency alters in vivo T follicular helper cell response in humans. Blood. 2013;121:.3375-3385
29. O'Malley JT, Sehra S, Thieu VT, Yu Q, Chang HC, Stritesky GL. et al. Signal transducer and activator of transcription 4 limits the development of adaptive regulatory T cells. Immunology. 2009;127:587-595
30. Xu J, Yang Y, Qiu G, Lal G, Yin N, Wu Z. et al. Stat4 is critical for the balance between Th17 cells and regulatory T cells in colitis. J Immunol. 2011;186:6597-6606
31. Sieve AN, Meeks KD, Lee S, Berg RE. A novel immunoregulatory function for IL-23: Inhibition of IL-12-dependent IFN-gamma production. Eur J Immunol. 2010;40:2236-2247
32. Ronnblom L. The type I interferon system in the etiopathogenesis of autoimmune diseases. Ups J Med Sci. 2011;116:227-237
33. Nguyen KB, Watford WT, Salomon R, Hofmann SR, Pien GC, Morinobu A. et al. Critical role for STAT4 activation by type 1 interferons in the interferon-gamma response to viral infection. Science. 2002;297:2063-2066
34. Hunter CA, Gabriel KE, Radzanowski T, Neyer LE, Remington JS. et al. Type I interferons enhance production of IFN-gamma by NK cells. Immunol Lett. 1997;59:1-5
35. Farina GA, York MR, Di Marzio M, Collins CA, Meller S, Homey B. et al. Poly(I:C) drives type I IFN- and TGFbeta-mediated inflammation and dermal fibrosis simulating altered gene expression in systemic sclerosis. J Invest Dermatol. 2010;130:2583-2593
36. Lewis CC, Aronow B, Hutton J, Santeliz J, Dienger K, Herman N. et al. Unique and overlapping gene expression patterns driven by IL-4 and IL-13 in the mouse lung. J Allergy Clin Immunol. 2009;123:795-804
37. Finkelman FD, Hogan SP, Hershey GK, Rothenberg ME, Wills-Karp M. Importance of cytokines in murine allergic airway disease and human asthma. J Immunol. 2010;184:1663-1674
38. Kaser A, C. Molnar, H. Tilg. Differential regulation of interleukin 4 and interleukin 13 production by interferon alpha. Cytokine. 1998;10:75-81
39. Shibuya H, Hirohata S. Differential effects of IFN-alpha on the expression of various TH2 cytokines in human CD4+ T cells. J Allergy Clin Immunol. 2005;116:205-212
40. Gonzales-van Horn SR, Estrada LD, van Oers NS, Farrar JD. et al. STAT4-mediated transcriptional repression of the IL5 gene in human memory Th2 cells. Eur J Immunol. 2016;46:1504-1510
41. Zhao K, Zhang Q, Li X, Zhao D, Liu Y, Shen Q. et al. Cytoplasmic STAT4 Promotes Antiviral Type I IFN Production by Blocking CHIP-Mediated Degradation of RIG-I. J Immunol. 2016;96:1209-1217
42. Oppmann B, Lesley R, Blom B, Timans JC, Xu Y, Hunte B. et al. Novel p19 protein engages IL-12p40 to form a cytokine, IL-23, with biological activities similar as well as distinct from IL-12. Immunity. 2000;13:715-725
43. Lankford CS, Frucht DM. A unique role for IL-23 in promoting cellular immunity. J Leukoc Biol. 2003;73:49-56
44. Langrish CL, McKenzie BS, Wilson NJ, de Waal Malefyt R, Kastelein RA. et al. IL-12 and IL-23: master regulators of innate and adaptive immunity. Immunol Rev. 2004;202:96-105
45. Lupardus PJ, Garcia KC. The structure of interleukin-23 reveals the molecular basis of p40 subunit sharing with interleukin-12. J Mol Biol. 2008;382:931-941
46. Aggarwal S, Ghilardi N, Xie MH, de Sauvage FJ, Gurney AL. Interleukin-23 promotes a distinct CD4 T cell activation state characterized by the production of interleukin-17. J Biol Chem. 2003;278:1910-1914
47. Parham C, Chirica M, Timans J, Vaisberg E, Travis M, Cheung J. et al. A receptor for the heterodimeric cytokine IL-23 is composed of IL-12Rbeta1 and a novel cytokine receptor subunit, IL-23R. J Immunol. 2002;168:5699-5708
48. Toussirot E. The IL23/Th17 pathway as a therapeutic target in chronic inflammatory diseases. Inflamm Allergy Drug Targets. 2012;11:159-168
49. Mathur AN, Chang HC, Zisoulis DG, Stritesky GL, Yu Q, O'Malley JT. et al. Stat3 and Stat4 direct development of IL-17-secreting Th cells. J Immunol. 2007;178:4901-4907
50. Kaplan MH. STAT4: a critical regulator of inflammation in vivo. Immunol Res. 2005;31:231-342
51. Maddur MS, Miossec P, Kaveri SV, Bayry J. Th17 cells: biology, pathogenesis of autoimmune and inflammatory diseases, and therapeutic strategies. Am J Pathol. 2012;181:8-18
52. Malek TR. The biology of interleukin-2. Annu Rev Immunol. 2008;26:453-479
53. Wang KS, Ritz J, Frank DA. IL-2 induces STAT4 activation in primary NK cells and NK cell lines, but not in T cells. J Immunol. 1999;162:299-304
54. Wang KS, Frank DA, Ritz J. Interleukin-2 enhances the response of natural killer cells to interleukin-12 through up-regulation of the interleukin-12 receptor and STAT4. Blood. 2000;95:p3183-3190
55. O'Sullivan A, Chang HC, Yu Q, Kaplan MH. STAT4 is required for interleukin-12-induced chromatin remodeling of the CD25 locus. J Biol Chem. 2004;279:7339-7345
56. Matsubara S, Takeda K, Kodama T, Joetham A, Miyahara N, Koya T. et al. IL-2 and IL-18 attenuation of airway hyperresponsiveness requires STAT4, IFN-gamma, and natural killer cells. Am J Respir Cell Mol Biol. 2007;36:324-332
57. Hunter CA, Kastelein R. Interleukin-27: balancing protective and pathological immunity. Immunity. 2012;37:960-769
58. Pflanz S, Hibbert L, Mattson J, Rosales R, Vaisberg E, Bazan JF. et al. WSX-1 and glycoprotein 130 constitute a signal-transducing receptor for IL-27. J Immunol. 2004;172:2225-2231
59. Lucas S, Ghilardi N, Li J, de Sauvage FJ. IL-27 regulates IL-12 responsiveness of naive CD4+ T cells through Stat1-dependent and -independent mechanisms. Proc Natl Acad Sci U S A. 2003;100:15047-15052
60. Collison LW, Delgoffe GM, Guy CS, Vignali KM, Chaturvedi V, Fairweather D. et al. The composition and signaling of the IL-35 receptor are unconventional. Nat Immunol. 2012;13:290-299
61. Zhang XH, Zhou Y, Zhang JM, Zhou SY, Wang M, Feng R. et al. IL-35 inhibits acute graft-versus-host disease in a mouse model. Int Immunopharmacol. 2015;29:383-392
62. Wang W, Guo H, Li H, Yan Y, Wu C, Wang X. et al. Interleukin-35 Gene-Modified Mesenchymal Stem Cells Protect Concanavalin A-Induced Fulminant Hepatitis by Decreasing the Interferon Gamma Level. Hum Gene Ther. 2018;29:234-241
63. Jia D, Jiang H, Weng X, Wu J, Bai P, Yang W. et al. Interleukin-35 Promotes Macrophage Survival and Improves Wound Healing After Myocardial Infarction in Mice. Circ Res. 2019;124:1323-1336
64. Dinarello CA, Novick D, Puren AJ, Fantuzzi G, Shapiro L, Mühl H. et al. Overview of interleukin-18: more than an interferon-gamma inducing factor. J Leukoc Biol. 1998;63:658-664
65. Takeda K, Tsutsui H, Yoshimoto T, Adachi O, Yoshida N, Kishimoto T. et al. Defective NK cell activity and Th1 response in IL-18-deficient mice. Immunity. 1998;8:383-390
66. Lawless VA, Zhang S, Ozes ON, Bruns HA, Oldham I, Hoey T. et al. Stat4 regulates multiple components of IFN-gamma-inducing signaling pathways. J Immunol. 2000;165:6803-6808
67. Nakahira M, Ahn HJ, Park WR, Gao P, Tomura M, Park CS. et al. Synergy of IL-12 and IL-18 for IFN-gamma gene expression: IL-12-induced STAT4 contributes to IFN-gamma promoter activation by up-regulating the binding activity of IL-18-induced activator protein 1. J Immunol. 2002;168:1146-53
68. Strengell M, Sareneva T, Foster D, Julkunen I, Matikainen S. IL-21 up-regulates the expression of genes associated with innate immunity and Th1 response. J Immunol. 2002;169:3600-5
69. Monteleone G, Monteleone I, Fina D, Vavassori P, Del Vecchio Blanco G, Caruso R. et al. Interleukin-21 enhances T-helper cell type I signaling and interferon-gamma production in Crohn's disease. Gastroenterology. 2005;128:687-694
70. Alexander WS. Suppressors of cytokine signalling (SOCS) in the immune system. Nat Rev Immunol. 2002;2:410-416
71. Croker BA, Krebs DL, Zhang JG, Wormald S, Willson TA, Stanley EG. et al. SOCS3 negatively regulates IL-6 signaling in vivo. Nat Immunol. 2003;4:540-554
72. Yamamoto K, Yamaguchi M, Miyasaka N, Miura O. SOCS-3 inhibits IL-12-induced STAT4 activation by binding through its SH2 domain to the STAT4 docking site in the IL-12 receptor beta2 subunit. Biochem Biophys Res Commun. 2003;310:1188-1193
73. Shuai K. Modulation of STAT signaling by STAT-interacting proteins. Oncogene. 2000;19:2638-2644
74. Arora T, Liu B, He H, Kim J, Murphy TL, Murphy KM. et al. PIASx is a transcriptional co-repressor of signal transducer and activator of transcription 4. J Biol Chem. 2003;278:21327-21330
75. Nakahira M, Tanaka T, Robson BE, Mizgerd JP, Grusby MJ. Regulation of signal transducer and activator of transcription signaling by the tyrosine phosphatase PTP-BL. Immunity. 2007;26:163-176
76. Tanaka T, Soriano MA, Grusby MJ. SLIM is a nuclear ubiquitin E3 ligase that negatively regulates STAT signaling. Immunity. 2005;22:729-736
77. Ren Y, Lu L, Guo TB, Qiu J, Yang Y, Liu A. et al. Novel immunomodulatory properties of berbamine through selective down-regulation of STAT4 and action of IFN-gamma in experimental autoimmune encephalomyelitis. J Immunol. 2008;18:1491-1498
78. Ambros V. The functions of animal microRNAs. Nature. 2004;431:350-355
79. Wong CF, Tellam RL. MicroRNA-26a targets the histone methyltransferase Enhancer of Zeste homolog 2 during myogenesis. J Biol Chem. 2008;283:9836-9843
80. Lu LF, Boldin MP, Chaudhry A, Lin LL, Taganov KD, Hanada T. Function of miR-146a in controlling Treg cell-mediated regulation of Th1 responses. Cell. 2010;142:914-929
81. Möhnle P, Schütz SV, van der Heide V, Hübner M, Luchting B, Sedlbauer J. et al. MicroRNA-146a controls Th1-cell differentiation of human CD4+ T lymphocytes by targeting PRKC. Eur J Immunol. 2015;45:260-272
82. Huang Y, Lei Y, Zhang H, Hou L, Zhang M, Dayton AI. MicroRNA regulation of STAT4 protein expression: rapid and sensitive modulation of IL-12 signaling in human natural killer cells. Blood. 2011;118:6793-6802
83. Litvinov IV, Cordeiro B, Fredholm S, Ødum N, Zargham H, Huang Y. et al. Analysis of STAT4 expression in cutaneous T-cell lymphoma (CTCL) patients and patient-derived cell lines. Cell Cycle. 2014;13:2975-2982
84. Zhou X, Xia Y, Su J, Zhang G. Down-regulation of miR-141 induced by helicobacter pylori promotes the invasion of gastric cancer by targeting STAT4. Cell Physiol Biochem. 2014;33:1003-1012
85. Fortunato O, Borzi C, Milione M, Centonze G, Conte D, Boeri M. et al. Circulating mir-320a promotes immunosuppressive macrophages M2 phenotype associated with lung cancer risk. Int J Cancer. 2018;144:2746-2761
86. Yang JD, Roberts LR. Roberts. Hepatocellular carcinoma: A global view. Nat Rev Gastroenterol Hepatol. 2010;7:448-458
87. Shen FM, Lee MK, Gong HM, Cai XQ, King MC. Complex segregation analysis of primary hepatocellular carcinoma in Chinese families: interaction of inherited susceptibility and hepatitis B viral infection. Am J Hum Genet. 1991;49:88-93
88. Jiang DK, Sun J, Cao G, Liu Y, Lin D, Gao YZ. et al. Genetic variants in STAT4 and HLA-DQ genes confer risk of hepatitis B virus-related hepatocellular carcinoma. Nat Genet. 2013;45:72-75
89. Clark A, Gerlach F, Tong Hv, Hoan NX, Song le H, Toan NL. et al. A trivial role of STAT4 variant in chronic hepatitis B induced hepatocellular carcinoma. Infect Genet Evol. 2013;18:257-261
90. Liao Y, Cai B, Li Y, Chen J, Tao C, Huang H. et al. Association of HLA-DP/DQ and STAT4 polymorphisms with HBV infection outcomes and a mini meta-analysis. PLoS One. 2014;9:e111677
91. Jiang DK, Ma XP, Wu X, Peng L, Yin J, Dan Y. et al. Genetic variations in STAT4, C2, HLA-DRB1 and HLA-DQ associated with risk of hepatitis B virus-related liver cirrhosis. Sci Rep. 2015;5:16278
92. Lu Y, Zhu Y, Peng J, Wang X, Wang F, Sun Z. STAT4 genetic polymorphisms association with spontaneous clearance of hepatitis B virus infection. Immunol Res. 2015;62:146-152
93. Jiang DK, Wu X, Qian J, Ma XP, Yang J, Li Z. et al. Genetic variation in STAT4 predicts response to interferon-alpha therapy for hepatitis B e antigen-positive chronic hepatitis B. Hepatology. 2016;63:1102-1111
94. Chen H, Sun J, Zhou B, Xie Q, Liang X, Fan R. et al. Variants in STAT4 associated with cure of chronic HBV infection in HBeAg-positive patients treated with pegylated interferon-alpha. Clin Gastroenterol Hepatol. 2019:19 pii: S1542-3565
95. Wubetu GY, Utsunomiya T, Ishikawa D, Yamada S, Ikemoto T, Morine Y. et al. High STAT4 expression is a better prognostic indicator in patients with hepatocellular carcinoma after hepatectomy. Ann Surg Oncol. 2014;21( Suppl 4 ):S721-728
96. Wang G, Chen JH, Qiang Y, Wang DZ, Chen Z. Decreased STAT4 indicates poor prognosis and enhanced cell proliferation in hepatocellular carcinoma. World J Gastroenterol. 2015;21:3983-3993
97. Totoson P, Maguin-Gaté K, Prati C, Wendling D, Demougeot C. Mechanisms of endothelial dysfunction in rheumatoid arthritis: lessons from animal studies. Arthritis Res Ther. 2014;16:202
98. Watford WT, Hissong BD, Bream JH, Kanno Y, Muul L, O'Shea JJ. et al. Signaling by IL-12 and IL-23 and the immunoregulatory roles of STAT4. Immunol Rev. 2004;202:139-156
99. Walker JG, Ahern MJ, Coleman M, Weedon H, Papangelis V, Beroukas D. et al. Characterisation of a dendritic cell subset in synovial tissue which strongly expresses Jak/STAT transcription factors from patients with rheumatoid arthritis. Ann Rheum Dis. 2007;66:992-999
100. Gu E, Lu J, Xing D, Chen X, Xie H, Liang J. et al. Rs7574865 polymorphism in signal transducers and activators of transcription 4 gene and rheumatoid arthritis: an updated meta-analysis of 28 case-control comparisons. Int J Rheum Dis. 2015;18:3-16
101. Lefevre S, Meier FM, Neumann E, Muller-Ladner U. et al. Role of synovial fibroblasts in rheumatoid arthritis. Curr Pharm Des. 2015;21:130-141
102. Nguyen HN, Noss EH, Mizoguchi F, Huppertz C, Wei KS, Watts GFM. et al. Autocrine Loop Involving IL-6 Family Member LIF, LIF Receptor, and STAT4 Drives Sustained Fibroblast Production of Inflammatory Mediators. Immunity. 2017;46:220-232
103. Wang Y, Li Y, He Y, Sun Y, Sun W, Xie Q. et al. Expression of G protein alphaq Subunit is Decreased in Lymphocytes from Patients with Rheumatoid Arthritis and is Correlated with Disease Activity. Scand J Immunol. 2012;75:203-209
104. Wang D, Liu Y, Li Y, He Y, Zhang J, Shi G. Galphaq Regulates the Development of Rheumatoid Arthritis by Modulating Th1 Differentiation. Mediators Inflamm. 2017;2017:4639081
105. Aerts NE, Ebo DG, Bridts CH, Stevens WJ, De Clerck LS. T cell signal transducer and activator of transcription (STAT) 4 and 6 are affected by adalimumab therapy in rheumatoid arthritis. Clin Exp Rheumatol. 2010;28:208-214
106. Goropevšek A, Holcar M, Avčin T. The Role of STAT Signaling Pathways in the Pathogenesis of Systemic Lupus Erythematosus. Clin Rev Allergy Immunol. 2017;52:164-181
107. Singh RR, Saxena V, Zang S, Li L, Finkelman FD, Witte DP. et al. Differential contribution of IL-4 and STAT6 vs STAT4 to the development of lupus nephritis. J Immunol. 2003;170:4818-4825
108. Menke J, Bork T, Kutska B, Byrne KT, Blanfeld M, Relle M. et al. Targeting transcription factor Stat4 uncovers a role for interleukin-18 in the pathogenesis of severe lupus nephritis in mice. Kidney Int. 2011;79:452-463
109. Kariuki SN, Kirou KA, MacDermott EJ, Barillas-Arias L, Crow MK, Niewold TB. Cutting edge: autoimmune disease risk variant of STAT4 confers increased sensitivity to IFN-alpha in lupus patients in vivo. J Immunol. 2009;182:34-38
110. Zullo A, Sommese L, Nicoletti G, Donatelli F, Mancini FP, Napoli C. Epigenetics and type 1 diabetes: mechanisms and translational applications. Transl Res. 2017;185:85-93
111. Weaver JR, Nadler JL, Taylor-Fishwick DA. Interleukin-12 (IL-12)/STAT4 Axis Is an Important Element for beta-Cell Dysfunction Induced by Inflammatory Cytokines. PLoS One. 2015;10:e0142735
112. Trembleau S, Penna G, Bosi E, Mortara A, Gately MK, Adorini L. Interleukin 12 administration induces T helper type 1 cells and accelerates autoimmune diabetes in NOD mice. J Exp Med. 1995;181:817-821
113. Dobrian AD, Galkina EV, Ma Q, Hatcher M, Aye SM, Butcher MJ. et al. STAT4 deficiency reduces obesity-induced insulin resistance and adipose tissue inflammation. Diabetes. 2013;62:4109-4121
114. Dobrian AD, Ma K, Glenn LM, Hatcher MA, Haynes BA, Lehrer EJ. et al. Key Role of STAT4 Deficiency in the Hematopoietic Compartment in Insulin Resistance and Adipose Tissue Inflammation. Mediators Inflamm. 2017;2017:5420718
115. Bowcock AM, Krueger JG. Getting under the skin: the immunogenetics of psoriasis. Nat Rev Immunol. 2005;5:699-711
116. Ghoreschi K, Mrowietz U, Röcken M. A molecule solves psoriasis? Systemic therapies for psoriasis inducing interleukin 4 and Th2 responses. J Mol Med (Berl). 2003;81(8):471-480
117. Eriksen KW, Lovato P, Skov L, Krejsgaard T, Kaltoft K, Geisler C. et al. Increased sensitivity to interferon-alpha in psoriatic T cells. J Invest Dermatol. 2005;125:936-944
118. Baumgart DC, Carding SR. Inflammatory bowel disease: cause and immunobiology. Lancet. 2007;369:1627-1640
119. Parrello T, Monteleone G, Cucchiara S, Monteleone I, Sebkova L, Doldo P. et al. Up-regulation of the IL-12 receptor beta 2 chain in Crohn's disease. J Immunol. 2000;165:7234-7239
120. O'Malley JT, Eri RD, Stritesky GL, Mathur AN, Chang HC, Hogenesch H. et al. STAT4 isoforms differentially regulate Th1 cytokine production and the severity of inflammatory bowel disease. J Immunol. 2008;181:5062-5070
121. Zhang YS, Xin DE, Wang Z, Song X, Sun Y, Zou QC. et al. STAT4 activation by leukemia inhibitory factor confers a therapeutic effect on intestinal inflammation. EMBO J. 2019:38 pii: e99595
122. Kim SW, Kim ES, Moon CM, Kim TI, Kim WH, Cheon JH. et al. Abnormal genetic and epigenetic changes in signal transducer and activator of transcription 4 in the pathogenesis of inflammatory bowel diseases. Dig Dis Sci. 2012;57:2600-2607
123. Takeuchi M, Kastner DL, Remmers EF. The immunogenetics of Behcet's disease: A comprehensive review. J Autoimmun. 2015;64:137-148
124. Kirino Y, Bertsias G, Ishigatsubo Y, Mizuki N, Tugal-Tutkun I, Seyahi E. et al. Genome-wide association analysis identifies new susceptibility loci for Behcet's disease and epistasis between HLA-B*51 and ERAP1. Nat Genet. 2013;45:202-207
125. Hou S, Yang Z, Du L, Jiang Z, Shu Q, Chen Y. et al. Identification of a susceptibility locus in STAT4 for Behcet's disease in Han Chinese in a genome-wide association study. Arthritis Rheum. 2012;64:4104-4113
126. Nordmark G, Eloranta ML, Ronnblom L. Primary Sjogren's syndrome and the type I interferon system. Curr Pharm Biotechnol. 2012;13:2054-2062
127. Frucht DM, Aringer M, Galon J, Danning C, Brown M, Fan S. et al. Stat4 is expressed in activated peripheral blood monocytes, dendritic cells, and macrophages at sites of Th1-mediated inflammation. J Immunol. 2000;164:4659-4664
128. Nordmark G, Kristjansdottir G, Theander E, Eriksson P, Brun JG, Wang C. et al. Additive effects of the major risk alleles of IRF5 and STAT4 in primary Sjogren's syndrome. Genes Immun. 2009;10:68-76
129. Chairta P, Nicolaou P, Christodoulou K. Genomic and genetic studies of systemic sclerosis: A systematic review. Hum Immunol. 2017;78:153-165
130. Avouac J, Fürnrohr BG, Tomcik M, Palumbo K, Zerr P, Horn A. et al. Inactivation of the transcription factor STAT-4 prevents inflammation-driven fibrosis in animal models of systemic sclerosis. Arthritis Rheum. 2011;63:800-809
131. Rueda B, Broen J, Simeon C, Hesselstrand R, Diaz B, Suárez H. et al. The STAT4 gene influences the genetic predisposition to systemic sclerosis phenotype. Hum Mol Genet. 2009;18:2071-2077
132. Webb GJ, Siminovitch KA, Hirschfield GM. Siminovitch, G.M. Hirschfield. The immunogenetics of primary biliary cirrhosis: A comprehensive review. J Autoimmun. 2015;64:42-52
133. Joshita S, Umemura T, Nakamura M, Katsuyama Y, Shibata S, Kimura T. et al. STAT4 gene polymorphisms are associated with susceptibility and ANA status in primary biliary cirrhosis. Dis Markers. 2014;2014:727393
134. Yang CY, Ma X, Tsuneyama K, Huang S, Takahashi T, Chalasani NP. et al. IL-12/Th1 and IL-23/Th17 biliary microenvironment in primary biliary cirrhosis: implications for therapy. Hepatology. 2014;59:1944-1953
135. Liaskou E, Patel SR, Webb G, Bagkou Dimakou D, Akiror S, Krishna M. et al. Increased sensitivity of Treg cells from patients with PBC to low dose IL-12 drives their differentiation into IFN-gamma secreting cells. J Autoimmun. 2018;94:143-155
136. Parente Barbosa C, Bentes De Souza AM, Bianco B, Christofolini DM. The effect of hormones on endometriosis development. Minerva Ginecol. 2011;63:375-386
137. Antsiferova YS, Sotnikova NY, Posiseeva LV, Shor AL. Changes in the T-helper cytokine profile and in lymphocyte activation at the systemic and local levels in women with endometriosis. Fertil Steril. 2005;84:1705-1711
138. Zamani MR, Salmaninejad A, Akbari Asbagh F, Masoud A, Rezaei N. STAT4 single nucleotide gene polymorphisms and susceptibility to endometriosis-related infertility. Eur J Obstet Gynecol Reprod Biol. 2016;203:20-24
139. Zhao L, Ji G, Le X, Luo Z, Wang C, Feng M. et al. An integrated analysis identifies STAT4 as a key regulator of ovarian cancer metastasis. Oncogene. 2017;36:3384-3396
140. Caligaris-Cappio F, Ghia P. Novel insights in chronic lymphocytic leukemia: are we getting closer to understanding the pathogenesis of the disease? J Clin Oncol. 2008;26:p. 4497-4503
141. Migliaccio E, Giorgio M, Mele S, Pelicci G, Reboldi P, Pandolfi PP. et al. The p66shc adaptor protein controls oxidative stress response and life span in mammals. Nature. 1999;402:309-313
142. Ventura A, Luzi L, Pacini S, Baldari CT, Pelicci PG. The p66Shc longevity gene is silenced through epigenetic modifications of an alternative promoter. J Biol Chem. 2002;277(25):22370-22376
143. Capitani N, Lucherini OM, Sozzi E, Ferro M, Giommoni N, Finetti F. et al. Impaired expression of p66Shc, a novel regulator of B-cell survival, in chronic lymphocytic leukemia. Blood. 2010;115(18):3726-36
144. Cattaneo F, Patrussi L, Capitani N, Frezzato F, D'Elios MM, Trentin L. et al. Expression of the p66Shc protein adaptor is regulated by the activator of transcription STAT4 in normal and chronic lymphocytic leukemia B cells. Oncotarget. 2016;7:57086-57098
145. Iida K, Suzuki K, Yokota M, Nakagomi D, Wakashin H, Iwata A. et al. STAT4 Is Required for IFN-β-induced MCP-1 mRNA Expression in Murine Mast Cells. Int Arch Allergy Immunol. 2011;155(Suppl 1):71-6
146. Meng ZZ, Liu W, Xia Y, Yin HM, Zhang CY, Su D. et al. The pro-inflammatory signalling regulator Stat4 promotes vasculogenesis of great vessels derived from endothelial precursors. Nat Commun. 2017;8:14640
Author contact
Corresponding author: Deke Jiang, PhD, Department of Infectious Diseases and Hepatology Unit, Nanfang Hospital, Southern Medical University, GuangZhou 510515, China. Phone: (86) 20-62786533; Email: dekejiang17edu.cn

 Global reach, higher impact
Global reach, higher impact