Impact Factor ISSN: 1449-2288
Int J Biol Sci 2020; 16(11):1846-1860. doi:10.7150/ijbs.45018 This issue Cite
Review
Initial success in the identification and management of the coronavirus disease 2019 (COVID-19) indicates human-to-human transmission in Wuhan, China
1. Lab of Chemical Biology and Molecular Drug Design, College of Pharmaceutical Science, Zhejiang University of Technology, Hangzhou, 310014, China
2. Lab of Molecular Immunology, Virus Inspection Department, Zhejiang Provincial Center for Disease Control and Prevention, Hangzhou, 310051, China
3. Center for Natural Products Research, Chengdu Institute of Biology, Chinese Academy of Sciences, Chengdu 610041, PR China.
4. Center for Medical Innovation, Nagasaki University, 1-7-1 Sakamoto, Nagasaki 852-8588, Japan.
Received 2020-2-17; Accepted 2020-3-19; Published 2020-4-6
Abstract
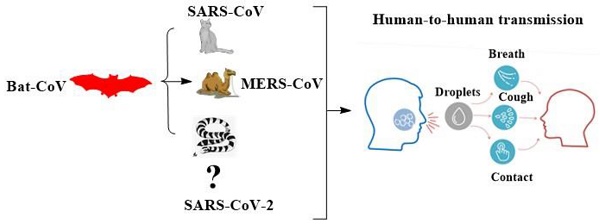
Coronavirus (CoV) has been one of the major pandemic threats to human health in the last two decades. The human coronavirus was first identified in 1960s. CoVs 229E, NL63, OC43, HKU1, SARS-CoV, and MERS-CoV have caused numerous disasters or human deaths worldwide. Recently, an outbreak of the previously unknown deadly CoV disease 2019 (COVID-19) caused by Severe Acute Respiratory Syndrome CoV 2 (SARS-CoV-2, early named 2019-nCoV) occurred in Wuhan, China, and it had caused 81238 cases of confirmed infection, including 3250 deaths until March 19, 2020. Its risks and pandemic potential have brought global consideration. We summarized epidemiology, virological characteristics, clinical symptoms, diagnostic methods, clinical treatments, and prevention methods for COVID-19 to present a reference for the future wave of probable CoV outbreaks.
Keywords: COVID-19, SARS-CoV-2, infection, treatment, prevention
Background
On December 31, 2019, the Health Commission of Hubei Province, China, first declared a cluster of unexplainable cases of pneumonia [1]. Subsequently, on January 10, 2020, the World Health Organization (WHO) announced that there was an outbreak of pneumonia of undiscovered cause in the city of Wuhan, Hubei Province, China. With an epidemiological association to the Huanan Seafood Wholesale Market where birds, bats, snakes, and other wildlife animals are sold [2]. A previously unknown coronavirus (CoV) was immediately discovered in the specimens from the patients. Which WHO provisionally named as the 2019 novel coronavirus (2019-nCoV). Later on February 11, 2020, WHO officially announced that the novel coronavirus was renamed as Severe Acute Respiratory Syndrome CoV 2 (SARS-CoV-2) and named the disease caused by SARS-CoV-2 as COVID-19 that stands for the CoV disease 2019.
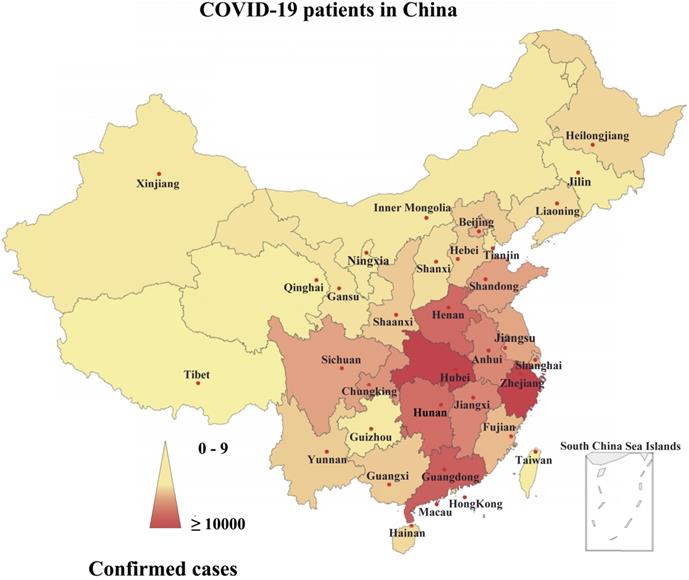
Since December 31, 2019 and as of March 19, 2020-6.00 PM GMT, overall, 81238 laboratory-confirmed cases of COVID-19, and 3250 deaths were announced in all over China, especially in Wuhan, Hubei Province (Fig. 1) [3]. Most of the fatal cases had typical symptoms such as fever, dry cough, dyspnoea, and radiological findings of bilateral lung infiltrates several days after the start of the initial symptoms [4]. Subsequent etiological studies showed that the patients were infected with SARS-CoV-2 [4]. This is the first report that SARS-CoV-2 can infect humans. In addition, SARS-CoV-2 displays strong ability of human-to-human transmission worldwide; although it currently has low pathogenicity, COVID-19 may lead to human death” [1]. As a result, it raises global public health concerns about its risks and pandemic potential.
Geographical distribution of the confirmed cases of COVID-19 in China. Note: the statistical data as of March 19, 2020.
Epidemiology
In December 2019, Wuhan, in China's Hubei Province, became the epicenter of an unexplained outbreak of pneumonia, drawing widespread attention not only in China but also in other countries [5]. As of March 19, 2020, 81238 laboratory-confirmed human cases in China, 131320 confirmed cases in 153 countries, and other 696 confirmed cases on an international conveyance in Japan have been reported to WHO (Table 1) [3]. Confirmed cases of Wuhan passengers were announced in Thailand on January 13, in Japan on January 15 and in South Korea on January 19, and so on [6]. It was soon found that SARS-CoV-2 caused COVID-19 and was epidemiologically related to the Huanan Seafood market in Wuhan [7]. Where restaurants supply wild animals, suggesting that the main route of SARS-CoV-2 transmission may be the point-source zoonotic (animal-to-human) route [8]. According to the "diagnosis and treatment of pneumonia" program of COVID-19, the main source of infection after the initial transmission to patients was SARS-CoV-2 infection in patients [8]. Recent studies suggest that SARS-CoV-2 may be transmitted from person to person primarily through respiratory droplets and through contact [9-11]. Compared to SARS-CoV and MERS-CoV, SARS-CoV-2 in Wuhan currently appears to be less pathogenic and safe if we assumed that the virus spreads effectively worldwide [12]. Still, the possibility of SARS-CoV-2 mutation cannot be ruled out [13]. The incubation period of SARS-CoV-2 is about 7 days on average, the short one is 2-3 days, and the long one could be over 21 days [4, 14]. Referring to the incubation period for other CoVs, the SARS-CoV-2 case was placed under close contact medical observation for 14 days, and close contacts were placed under medical observation in the hospital. After 14 days, if there is no evidence of the disease, the person can be determined as not infected [4, 14]. Everybody is susceptible to SARS-CoV-2, and disease in immunocompromised individuals progress relatively faster and more severe [2, 14]. Children have less exposure and a lower risk of infection; in the same way, the elderly, patients with chronic diseases, and people with weak immunity are more likely to be infected [4, 14].
COVID-19 cases-worldwide. Note: the statistical data as of March 19, 2020.
| Continent | Country / Territory / Area | Confirmed cases | Deaths |
|---|---|---|---|
| Africa | Algeria | 73 | 6 |
| Africa | Cameroon | 10 | 0 |
| Africa | Congo | 3 | 0 |
| Africa | Egypt | 196 | 6 |
| Africa | Ethiopia | 6 | 0 |
| Africa | Gabon | 3 | 0 |
| Africa | Ghana | 7 | 0 |
| Africa | Guinea | 1 | 0 |
| Africa | Kenya | 7 | 0 |
| Africa | Mauritania | 1 | 0 |
| Africa | Morocco | 54 | 2 |
| Africa | Namibia | 2 | 0 |
| Africa | Nigeria | 8 | 0 |
| Africa | Rwanda | 11 | 0 |
| Africa | Senegal | 36 | 0 |
| Africa | Seychelles | 6 | 0 |
| Africa | Sudan | 2 | 1 |
| Africa | Togo | 1 | 0 |
| Africa | Tunisia | 29 | 0 |
| Africa | Eswatini | 1 | 0 |
| Africa | Benin | 1 | 0 |
| Africa | Liberia | 2 | 0 |
| Africa | Somalia | 1 | 0 |
| Africa | Burkina Faso | 26 | 1 |
| Africa | Central African Republic | 1 | 0 |
| Africa | Cote dIvoire | 6 | 0 |
| Africa | Democratic Republic of the Congo | 14 | 0 |
| Africa | Equatorial Guinea | 3 | 0 |
| Africa | South Africa | 116 | 0 |
| Africa | United Republic of Tanzania | 3 | 0 |
| Africa | Gambia | 1 | 0 |
| Africa | Djibouti | 1 | 0 |
| Africa | Zambia | 2 | 0 |
| America | Argentina | 97 | 2 |
| America | Bahamas | 1 | 0 |
| America | Bolivia | 12 | 0 |
| America | Brazil | 428 | 4 |
| America | Canada | 690 | 9 |
| America | Chile | 238 | 0 |
| America | Colombia | 102 | 0 |
| America | Cuba | 11 | 1 |
| America | Ecuador | 168 | 3 |
| America | Guatemala | 8 | 1 |
| America | Guyana | 5 | 1 |
| America | Honduras | 12 | 0 |
| America | Jamaica | 15 | 0 |
| America | Mexico | 118 | 0 |
| America | Panama | 109 | 1 |
| America | Paraguay | 11 | 0 |
| America | Peru | 145 | 0 |
| America | Suriname | 1 | 0 |
| America | Uruguay | 79 | 0 |
| America | Venezuela | 33 | 0 |
| America | Antigua and Barbuda | 1 | 0 |
| America | Costa Rica | 69 | 1 |
| America | Dominican Republic | 21 | 0 |
| America | Saint Lucia | 2 | 0 |
| America | Saint Vincent and the Grenadines | 1 | 0 |
| America | Trinidad and Tobago | 9 | 0 |
| America | United States of America | 9415 | 150 |
| America | Barbados | 1 | 0 |
| America | El Salvador | 1 | 0 |
| America | Nicaragua | 1 | 0 |
| Asia | Afghanistan | 22 | 0 |
| Asia | Bahrain | 256 | 1 |
| Asia | Bangladesh | 10 | 0 |
| Asia | Bhutan | 1 | 0 |
| Asia | Cambodia | 24 | 0 |
| Asia | China | 81238 | 3250 |
| Asia | India | 165 | 3 |
| Asia | Indonesia | 172 | 5 |
| Asia | Iran | 17361 | 1135 |
| Asia | Iraq | 164 | 12 |
| Asia | Israel | 433 | 0 |
| Asia | Japan | 873 | 29 |
| Asia | Jordan | 52 | 0 |
| Asia | Kazakhstan | 37 | 0 |
| Asia | Kuwait | 142 | 0 |
| Asia | Lebanon | 133 | 4 |
| Asia | Malaysia | 790 | 2 |
| Asia | Maldives | 13 | 0 |
| Asia | Mongolia | 5 | 0 |
| Asia | Nepal | 1 | 0 |
| Asia | Oman | 39 | 0 |
| Asia | Pakistan | 302 | 0 |
| Asia | Palestine | 44 | 0 |
| Asia | Philippines | 202 | 17 |
| Asia | Qatar | 452 | 0 |
| Asia | Singapore | 313 | 0 |
| Asia | Taiwan | 108 | 1 |
| Asia | Thailand | 177 | 1 |
| Asia | Uzbekistan | 23 | 0 |
| Asia | Vietnam | 76 | 0 |
| Asia | Myanmar | 0 | 0 |
| Asia | Brunei Darussalam | 68 | 0 |
| Asia | Saudi Arabia | 171 | 0 |
| Asia | South Korea | 8565 | 91 |
| Asia | Sri Lanka | 42 | 0 |
| Asia | United Arab Emirates | 113 | 0 |
| Asia | Kyrgyzstan | 3 | 0 |
| Europe | Albania | 59 | 2 |
| Europe | Andorra | 53 | 0 |
| Europe | Armenia | 115 | 0 |
| Europe | Austria | 1646 | 4 |
| Europe | Azerbaijan | 34 | 0 |
| Europe | Belarus | 46 | 0 |
| Europe | Belgium | 1486 | 14 |
| Europe | Bulgaria | 92 | 2 |
| Europe | Croatia | 81 | 0 |
| Europe | Cyprus | 58 | 0 |
| Europe | Denmark | 1115 | 4 |
| Europe | Estonia | 258 | 0 |
| Europe | Finland | 369 | 0 |
| Europe | France | 9134 | 244 |
| Europe | Georgia | 34 | 0 |
| Europe | Germany | 8198 | 13 |
| Europe | Greece | 418 | 5 |
| Europe | Hungary | 73 | 1 |
| Europe | Iceland | 250 | 0 |
| Europe | Ireland | 366 | 2 |
| Europe | Italy | 35713 | 2978 |
| Europe | Kosovo | 19 | 0 |
| Europe | Latvia | 71 | 0 |
| Europe | Liechtenstein | 25 | 0 |
| Europe | Lithuania | 33 | 0 |
| Europe | Luxembourg | 210 | 2 |
| Europe | Malta | 48 | 0 |
| Europe | Moldova | 36 | 1 |
| Europe | Monaco | 9 | 0 |
| Europe | Netherlands | 2051 | 58 |
| Europe | Norway | 1423 | 3 |
| Europe | Poland | 287 | 5 |
| Europe | Portugal | 642 | 2 |
| Europe | Romania | 260 | 0 |
| Europe | Russia | 147 | 0 |
| Europe | Serbia | 94 | 0 |
| Europe | Slovakia | 107 | 0 |
| Europe | Slovenia | 286 | 1 |
| Europe | Spain | 13716 | 598 |
| Europe | Sweden | 1301 | 10 |
| Europe | Switzerland | 3010 | 21 |
| Europe | Turkey | 191 | 1 |
| Europe | Ukraine | 19 | 2 |
| Europe | Bosnia and Herzegovina | 36 | 0 |
| Europe | Czech Republic | 522 | 0 |
| Europe | Holy See | 1 | 0 |
| Europe | North Macedonia | 42 | 0 |
| Europe | San Marino | 109 | 14 |
| Europe | United Kingdom | 2630 | 103 |
| Europe | Montenegro | 8 | 0 |
| Oceania | Australia | 565 | 6 |
| Oceania | New Zealand | 28 | 0 |
| Oceania | French Polynesia | 3 | 0 |
| Oceania | Guam | 3 | 0 |
| Other | Cases on an international conveyance Japan | 696 | 7 |
| Total | 213254 | 8843 |
SARS-CoV-2 belongs to the new coronavirus species in the genus containing SARS-CoV and MERS-CoV [15]. It is enveloped, and the particles are round or elliptic, usually pleomorphic, with a diameter of 60~140 nm [16, 17]. Its genetic characteristics are different from those of SARS-CoV and MERS-CoV [18]. Current studies have shown that SARS-like coronavirus in bat (bat-SL-CoVZC45, MG772933.1) has over 85% homology to SARS-CoV-2 [16]. SARS-CoV-2 can be found in human respiratory epithelial cells at around 96 hours after infection [14, 16]. In contrast, it takes about 6 days to isolate the viruses when Vero E6 and Huh-7 cell lines are used in the in-vitro culture system [14, 16]. CoVs are single-stranded RNA viruses that mutate readily, making it challenging to develop sustained immunity to them [19]. Flu viruses, for example, circulate every year and require the latest vaccine because the virus type often changes [20].
Virology
Phylogenetic analysis
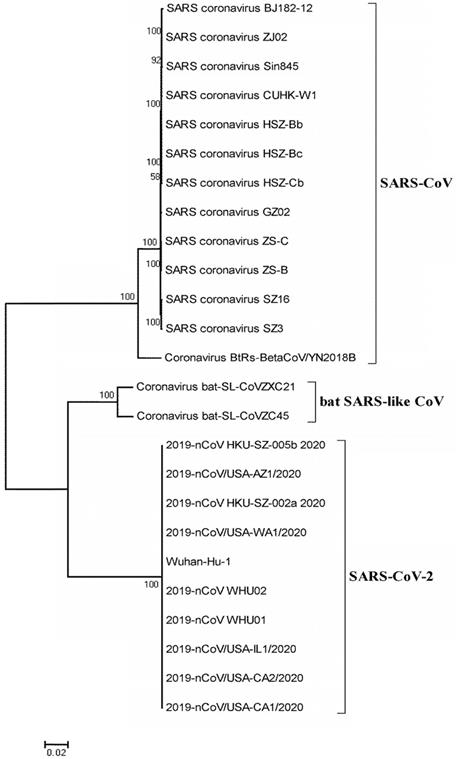
The circulating CoV is a newly discovered CoV, named SARS-CoV-2 by WHO. People are susceptible to the virus because they lack immunity to new strains of CoV [2, 14]. SARS-CoV-2 is different from any of the six known CoVs that can infect humans, including HCoV-229E, HCoV-OC43, HCoV-HKU1, HCoV-NL63, SARS-CoV, and MERS-CoV. It is the seventh CoV that can infect humans and cause severe pneumonia. Genomic analysis of SARS-CoV-2 showed that it was markedly different from both SARS-CoV and MERS-CoV, with the genomic difference being 21% and 48.2%, respectively [21]. Despite being a β-CoV, SARS-CoV-2 is not the same as the SARS-CoV and MERS-CoV, and is considered to be a distant relative of SARS-CoV, rather than a variant of SARS-CoV or a resurgence of SARS-CoV [21]. Previous studies showed that two complete virus genomes (HKU-SZ-002a and HKU-SZ-005b) were sequenced using Nanopore technology and revealed a SARS-CoV-2 is most closely related to bat SARS-like CoVs, bat-SL-CoVZXC21 (NCBI registration number MG772934) and bat-SL-CoVZC45 (NCBI registration number MG772933) [1], which belong to the Betacoronavirus genus (Fig. 2). The virus genome sizes of HKU-SZ-002a and HKU-SZ-005b were about 29.8 kbp, and the GC contents were 38%. The difference between HKU-SZ-002a and HKU-SZ-005b was only two bases [1]. One of them is a non-synonymous mutation at the amino acid position of the unstructured protein at 336 (Ser336 for HKU-SZ-002a; Leu336 for HKU-SZ-005b) [1]. Approximately 66% of the amino acid residues of the N-terminal domain of Spike subunit 1 of SARS-CoV-2 are identical to that of the SARS-related CoV. The core region of the SARS-CoV-2 receptor-binding domain is approximately 68% similar to the virus at the amino acid level. It has the same acidity as that of the SARS-related CoV. The amino acid sequences in the outer subdomain of the Spike subunit 1 receptor binding domain show only 39% identity, which may influence the selection of human receptors and thus the biological behavior of the virus [1].
Phylogenetic relationship of SARS-CoV-2 with bat SARS-like CoV and SARS-CoV. The full genome sequences of SARS-CoV-2 and SARS-CoV were obtained from the National Center for Biotechnology Information search engine (http://www.ncbi.nim.nih.gov/). The phylogenetic tree was created using the MEGA 5.0 software package and neighbor-joining program (http://www.megasoftware.net) with 1000 boots trapped value support and a Poisson correction. The bootstrap values greater than 50% are given at the nodes. The scale bar shows the phylogenetic range determined from the number of variations. The NCBI GenBank accession numbers of the genome sequences are EU371564.1 (SARS coronavirus BJ182-12), EU371559.1 (SARS coronavirus ZJ02), AY559093.1 (SARS coronavirus Sin845), AY278554.2 (SARS coronavirus CUHK-W1), AY394985.1 (SARS coronavirus HSZ-Bb), AY394994.1 (SARS coronavirus HSZ-Bc), AY394986.1 (SARS coronavirus HSZ-Cb), AY390556.1 (SARS coronavirus GZ02), AY395003.1 (SARS coronavirus ZS-C), AY394996.1 (SARS coronavirus ZS-B), AY304488.1 (SARS coronavirus SZ16), AY304486.1 (SARS coronavirus SZ3), MK211376.1 (Coronavirus BtRs-BetaCoV/YN2018B), MG772933.1 (Coronavirus bat-SL-CoVZC45), MG772934.1 (Coronavirus bat-SL-CoVZXC21), MN975262.1 (2019-nCoV HKU-SZ-005b 2020), MN997409.1 (2019-nCoV/USA-AZ1/2020), MN938384.1 (2019-nCoV HKU-SZ-002a 2020), MN985325.1 (2019-nCoV/USA-WA1/2020), NC_045512.2 (Wuhan-Hu-1), MN988669.1 (2019-nCoV WHU02), MN988668.1 (2019-nCoV WHU01), MN988713.1 (2019-nCoV/USA-IL1/2020), MN994468.1 (2019-nCoV/USA-CA2/2020), and MN994467.1 (2019-nCoV/USA-CA1/2020).
Molecular signatures of SARS-CoV-2
According to its genetic characteristics, the CoV family consists of four genera, including Alphacoronavirus, Betacoronavirus, Gammacoronavirus, and Deltacoronavirus [22]. Of all RNA viruses, the CoV RNA genome (ranging from 26 to 32 kbp) is the largest one [23]. SARS-CoV and MERS-CoV are classified into the Betacoronavirus genus and are zoonotic pathogens that can cause critical respiratory infections in humans [24]. The SARS-CoV-2 genomic structure belongs to that of β-CoV (MN908947) [25]. In the previous study in early 2020, samples were collected from seven patients with severe pneumonia who had been exposed to SARS-CoV-2 at a seafood market in Wuhan and were placed in an intensive care unit at the onset of the outbreak. The samples were then sent to the WIV laboratory for pathogen diagnosis [26]. They observed five positive bands on PCR. A sample from bronchoalveolar lavage fluid (BALF) (WIV04) was analyzed using the next generation sequenced (NGS) to identify possible pathogens. Of the 1582 total readings obtained after filtering through 64 human genomes, 1,378 (87.1%) matched SARS-CoV sequences [26]. The SARS-CoV-2 was named because the novel coronavirus has a 79.5% sequence identity with SARS-CoV BJ01 (GenBank accession number AY278488.2) [26]. The full-length genome sequences of four SARS-CoV-2 (WIV02, WIV05, WIV06, and WIV07) (GISAID accession numbers EPI_ISL_402127-402130) were subsequently obtained from four other patients, with more than 99.9% homology being observed among them [26]. The viral genome consists of six main open reading frames (ORFs) as well as many other accessory genes [26]. Additional analysis showed that the SARS-CoV-2 genome had less than 80% nucleotide sequence identity with the SARS-CoV sequence. They obtained a short RdRp region from a bat CoV, known as RaTG13, that was previously found in Rhinolophus affinis in Yunnan Province, China, with a high degree of sequence identity with SARS-CoV-2 [26]. Physicochemical parameters of SARS-CoV-2 (Sequence ID: MN908947.3), including isoelectric point, instability index, estimated half-life, and grand average of hydropathicity (GRAVY), were investigated using the ProtParam tool of ExPASy [27]. Analysis of physicochemical parameters showed that the SARS-CoV-2 polypeptide is 9194 amino acids with a molecular weight of 1069577.59 Dalton, theoretical pI: 9.02, the estimated half-life is 20 hours (mammalian reticulocytes, in vitro), 30 min (yeast, in vivo), and >10 hours (Escherichia coli, in vivo), and a GRAVY score of -0.087 GRAVY, categorizing the protein as unstable (Table 2).
Physicochemical Parameters of SARS-CoV-2.
| Parameters | SARS-CoV-2 Sequence ID: (MN908947.3) |
|---|---|
| Molecular Weight | 1069577.59 Da |
| Number of amino acids | 9194 |
| Theoretical pI | 9.02 |
| Instability index (II) | 43.95 (unstable) |
| Number of Negatively Charged Residues (Asp + Glu) | 560 |
| Number of Positively Charged Residues (Arg + Lys) | 971 |
| Aliphatic Index | 77.44 |
| Estimated half-life | 20 hours (mammalian reticulocytes, in vitro) 30 min (yeast, in vivo) >10 hours (Escherichia coli, in vivo) |
| Grand average of Hydropathicity (GRAVY) | -0.087 |
Angiotensin-converting enzyme 2 (ACE2) is a membrane-bound aminopeptidase that has been shown to play a critical role in the cardiovascular and immune systems through two different important physiological mechanisms. ACE2 catalyzes the production of vasodilatory peptides, including angiotensin 1 to 7 [28]. Since its discovery in 2000, ACE2 has been extensively studied in the cardiovascular and metabolic fields, suggesting that it plays a critical role in the development of cardiac function, hypertension, and diabetes. In addition, it is noteworthy that ACE2 is a functional receptor for coronaviruses, such as SARS-CoV, and SARS infection is caused by the binding of the SARS-CoV protein trimer to the hydrophobic pockets of ACE2's extracellular catalytic domain [29]. This interaction enables endocytosis, membrane fusion, and SARS-CoV entry into host cells. When the virus entered, ACE2 protein was downregulated, leading to a localized increase in Angiotensin II levels, which provided a molecular explanation for the continual development of Acute Respiratory Distress Syndrome (ARDS) during SARS-CoV infection. ACE2 is highly expressed in both heart and lung tissues. Therefore, in addition to lowering blood pressure and improving heart function, ACE2 inhibitors are expected to prevent ACE2/SARS-CoV protein interaction and inhibit SARS-CoV infection [29]. The spike (S) protein of SARS-CoV-2 contains a SARS-CoV-like receptor binding domain, suggesting that its receptor should be ACE2, which is the same as one for SARS-CoV. Recently, it has been found that the expression level of ACE2 in Asian people is significantly higher than that in European and American people and that in men is higher than that in women [30-32]. This difference also explains the high incidence of new coronary pneumonia in Asia, where there are more men than women [30-32]. The SARA-CoV-2 primarily invades alveolar epithelial cells and interacts with ACE2, leading to respiratory symptoms in patients [33]. When patients complicated with underlying cardiovascular diseases, the symptoms tended to be severe, which may be related to increased renin secretion in these patients and increased ACE2 secretion through negative feedback regulation, providing more binding sites for SARA-CoV-2 [34].
Clinical characteristics of SARS-CoV-2
The common symptoms of COVID-19 are fever (91.7%), fatigue (75.0%), cough (75.0%), progressive dyspnea (36.7%), gastrointestinal symptoms (39.6%), and some patients (≤ 10%) start with mild symptoms or even no palpable fever (Fig. 3) [35]. From the current case situation, most patients have a good prognosis; a few patients are in critical conditions or even died [36]. In addition to the above symptoms, there may be "atypical" symptoms, such as (a) only the symptoms of the digestive system as the first manifestation: such as mild tolerance, weakness, general mental disorder, nausea and vomiting, diarrhea, etc.; (b) neurological symptoms as the first manifestation: such as headache; (c) cardiovascular symptoms as the first manifestation: such as palpitation, chest tightness, etc.; (d) ophthalmic symptoms as the first manifestation: such as conjunctivitis; (e) only mild muscle soreness in limbs or lower back [4].
The difference between COVID-19 (SARS-CoV-2) and flu, common cold symptoms.
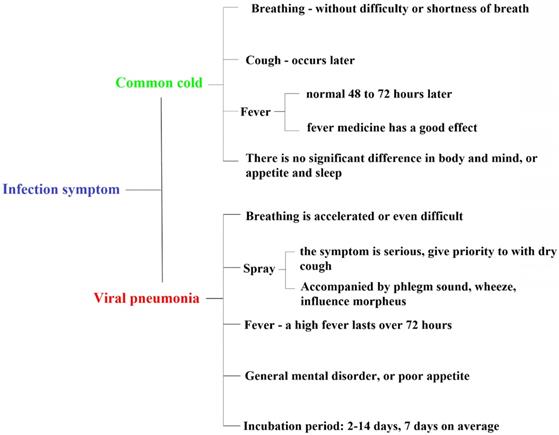
Many respiratory diseases are characterized by fever, fatigue, dry cough, and other symptoms. Diagnosis of COVID-19 requires a physician to make a comprehensive judgment based on whether he/she has been to an endemic area before the onset of the disease, whether he/she has been exposed to suspected or confirmed cases, and the results of laboratory tests [14]. A dry cough is one of the symptoms of COVID-19; the main difference between a dry cough and a cough is the presence of sputum [4, 14]. A dry cough is a cough in which there is no sputum or very little sputum. Common cold, acute bronchitis, inhalation of some irritant gas, and dust and so on can also cause a dry cough. Pneumonia may be accompanied, but early pneumonia may not be fever, only chills and respiratory infection symptoms, but computed tomography (CT) scan will show pneumonia. In severe cases, dyspnea and hypoxemia usually occur 1 week after infection [4, 37]. In severe cases, it rapidly progressed to ARDS, septic shock, metabolic acidosis that was difficult to treat, and bleeding and coagulation dysfunction [4, 36, 37]. It is worth noting that patients with severe or critical illness may have a moderate or low fever or even no palpable fever [4]. Patients with mild cases showed only low fever, mild fatigue, and no pneumonia usually recovered after 1 week [4]. A few infected patients exhibited no obvious clinical symptoms and only tested positive [4]. From the current case situation, most of the patients have a good prognosis, the child cases are relatively mild, and a few patients are in critical condition [14]. Deaths are more common in the elderly and those with chronic underlying diseases [14]. Of the 1023 deaths (fatality rate 2.3%), most were aged ≥ 60 years or had comorbidities such as hypertension, cardiovascular disease, and diabetes [38]. Laboratory examination showed that the total number of white blood cells in peripheral blood was normal or decreased, the lymphatic size was reduced, and the cell count decreased. Liver enzymes, muscle enzymes, and myoglobin increased in some patients. Most patient's C-reactive protein (CRP) and erythrocyte sedimentation rate were elevated, and procalcitonin (PCT) was normal. Normal or slightly higher levels of cytokines, inflammatory cytokines such as interleukins-2 (IL-2), tumor necrosis factor-α (TNF- α), IL-6, interferon-γ (IFN-γ), and so on, can be markedly increased in patients with organ failure. In severe cases, D-D dimer increased, and lymphocytes progressively decreased [4, 36, 37].
Diagnosis
On January 28, 2020, the fourth edition of the “Pneumonia Diagnostic Guidelines” for COVID-19 was officially released, and the pneumonia caused by SARS-CoV-2 was characterized by human-to-human transmission [39]. According to the "Pneumonia Diagnosis and Treatment” program for COVID-19 (trial version 4)", suspected cases can be defined as (1) epidemiological history: travel history or residence history in Wuhan within two weeks before the onset of the disease; or had been in contact with patients with fever and respiratory symptoms from Wuhan within 14 days before the onset, or had aggregated onset. (2) clinical manifestations: fever; (a) with typical pneumonia imaging characteristics; (b) the total number of early-onset white blood cells normal or decreased, or lymphocyte count decreased. Real-time fluorescence quantitative PCR (RT-PCR) tested confirmed cases: based on meeting the criteria for suspected cases, sputum, pharynx swabs, lower respiratory tract secretions, and other specimens for positive SARS-CoV-2 nucleic acid, or viral gene sequencing, highly homologous to SARS-CoV-2 [39]. As one of the methods for early diagnosis of suspected cases, chest imaging is widely used in a clinic because of its convenience and versatility [14]. It is easy to diagnose multiple ground glass density shadows in typical bilateral lungs [14, 40]. However, COVID-19 caused by SARS-CoA-2 has not been thoroughly analyzed, clinical features have not been described collectively [14, 41].
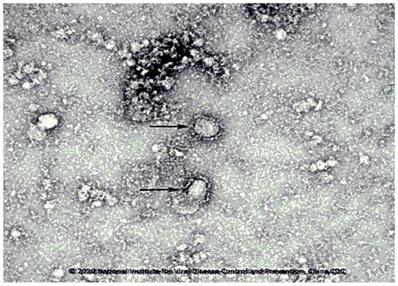
In the early stage of the disease, it was mainly manifested as scattered small flakes in the lung, and focal ground-glass density and consolidation shadows distributed under the pleura or along the bronchial tree [14]. In the progressive stage, most of the patients reached the scene of the most severe lung infiltration within 8-14 days after the onset, and the ground glass density shadows could be fused into scattered and multiple consolidation shadows, in which air bronchi signs could be seen [4]. When compared to SARS in 2003, the main manifestations of COVID-19 are: fever is evident at the initial stage (2-3 days after onset), and the imaging manifestations are usually small lamina in the lung, a ground-glass shadow is common [4, 14]. The distribution of the following lung and surrounding areas is universal [4, 14]. Progressive exacerbation occurs within 5-8 days after the onset of the disease, and the most severe symptoms appear 2-3 weeks after onset, which is called a progressive stage [42]. Ground glass shadow consolidation and mesh shadow appear, followed by interlobular septum thickening [42]. Small patchy lesions develop to large patchy ones, with single progression to multiple or diffuse lesions. The development is swift, and the change is rapid [14]. The number of affected pulmonary lobes is increased rapidly, and pathological changes can progress from a lung to several lobes, from one lung to two lungs, even "white lung" appears [14]. Two to three weeks after the onset of the disease, it enters the absorption phase and changes from multiple and diffuse lesions to the limitation [14]. The density begins to decrease, and the range gradually shrinks or even disappears. After inflammatory absorption, some patients leave behind some manifestations of pulmonary interstitial fibrosis [4, 14]. This new type of viral pneumonia is fierce, and the characteristics of human-to-human transmission have been identified (China's National Microbiology Data Center shared the image [43, 44], Fig. 4). "Early detection, early report, early isolation, and early treatment" is critical [45]. The recognition of atypical imaging manifestations plays a vital role in the early identification of COVID-19 caused by SARS-CoV-2 [45]. It is recommended to timely conduct multiple examinations and follow up with imaging to quickly and appropriately perform the diagnosis and observe the changes in the disease [45].
SARS-CoV-2 particles in ultrathin sections of human airway epithelial cells.
Treatment and Prevention
Treatment recommendations
Principles of treatment for suspected and confirmed cases should be isolated in designated hospitals with sufficient isolation conditions and protective conditions [14]. Suspected cases should be treated in isolation, while confirmed cases can be admitted to the same ward. Critical cases should be admitted to the ICU as soon as possible [14]. Bed rest, vital signs (heart rate, pulse oxygen saturation, respiratory rate, and blood pressure) should be observed. Supportive treatment should be strengthened to ensure adequate heat to maintain a stable internal environment such as water, electrolyte, and acid-base balance [14]. Complete blood count, C-reactive protein (CRP), procalcitonin (PCT) and dirt were monitored according to the conditions of organ functions (liver enzyme, bilirubin, myocardial enzyme, creatinine, urea nitrogen, urine volume, etc.), coagulation function, arterial blood gas analysis, and chest imaging [4]. Effective oxygen therapy, including a nasal catheter, mask oxygen, high flow oxygen therapy (HFNO), and non-invasive mechanical ventilation (NIV) or invasive mechanical ventilation is recommended [14]. Extracorporeal oxygenation (ECMO) should be considered for refractory hypoxemia that is difficult to manage after protective pulmonary ventilation [4].
Therapeutic use
There is no evidence from a randomized controlled trial (RCT) to support specific anti-SARS-CoV-2 drugs for suspected or confirmed cases up to now. In treating Severe Acute Respiratory Syndrome (SARS) and the Middle East respiratory syndrome (MERS) infections, some studies, such as retrospective cohort studies, historical control studies, case reports, or serial case reports, confirmed that lopinavir/ritonavir or, in combination with antivirals, has some therapeutic advantages, such as reducing the incidence of acute respiratory distress syndrome (ARDS) mortality [46-48]. Previous published systematic reviews show that lopinavir/ritonavir is primarily effective in early application, and can reduce mortality and glucocorticoid dosage [49, 50]. If the early treatment window is missed, however, the late application has no significant effect. We still need to carry out real-world clinical studies to further explore the clinical implications of their early use in the COVID-19 [14]. The effectiveness of antiviral combinations remains controversial [51-54]. Therefore, blind or inappropriate use of antibiotics, especially in combination with broad-spectrum antibiotics, should be avoided in the treatment of COVID-19. Strengthening bacteriological monitoring, when there are secondary bacterial infections appropriate, antimicrobial agents may timely be applied. Depending on the patient's clinical presentation, patients with mild cases can take oral antibiotics and antimicrobial agents, such as amoxicillin, azithromycin, or fluoroquinolones, against community-acquired pneumonia if the bacterial infection cannot be ruled out. Severe patients need to cover all possible pathogens for empirical treatment, after the pathogen identified descending stair therapy [51, 52]. Steroid therapy for severe ARDS is controversial, thus, systemic use of glucocorticoids should be cautious. Methylprednisolone can be used as appropriate in patients with rapid development or severe illness, and 40~80 mg daily can be considered depending on the severity of the disease, with the total daily dose not exceeding 2 mg/kg [55-57]. Previous studies on SARS showed that timely noninvasive continuous positive airway pressure ventilation and corticosteroids were used when lung shadows increase and dyspnea worsens effective solutions. Appropriate use of glucocorticoids can significantly improve the clinical symptoms of SARS patients, reduce the extent of disease progression, accelerate pulmonary disease, and absorption, but failed to shorten the length of hospitalization [55, 56]. Steroid therapy has a specific incidence of adverse reactions [57].
Possible new therapies for COVID-19
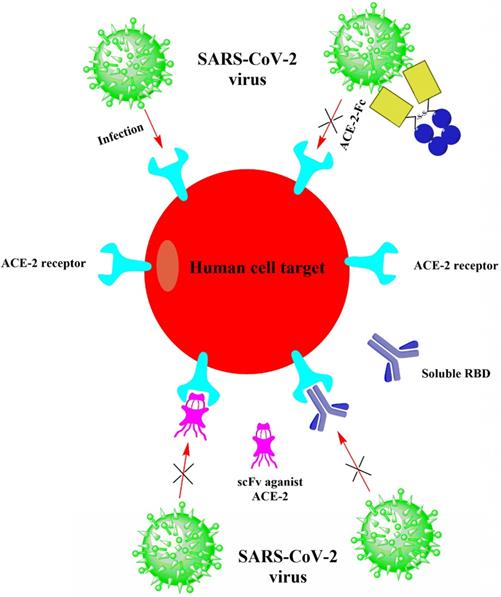
Neutralizing antibodies to the spike (S) protein on the surface of SARS-CoV-2 may be the first therapeutic strategy considered by biomedical researchers in academia and industry to provide passive immunity to illness [58]. The newly released SARS-CoV-2 genome sequence (GenBank: MN908947.3) enables researchers to conduct gene synthesis in the laboratory and estimate the expression of S proteins as immunogens. Conventional methods of screening mice or rabbits for neutralizing antibodies might be too slow for this outbreak. Still, rapid methods (such as phage or yeast display libraries that express antibody fragments) can be used to identify virally neutralizing antibody candidates quickly [59, 60]. The simplest, most direct way to combat SARS-CoV-2 during the outbreak is to interfere with the viral entry into cells, where antibodies usually work [61]. It won't be effortless to confirm the functions of neutralizing antibodies quickly. In addition, ensuring that mutated RNA viruses fail to enter the cells will be a challenge. A cocktail of antibodies applied to the Ebola pandemic could be used, but it would add complexity to the production manner [62]. However, another strategy that does not rely on directly targeting to viral glycoproteins can apply to the treatment of COVID-19. In this procedure, viral receptor proteins on the host cell surface are neutralized, thereby preventing the virus from binding and entering the cells. Fortunately, investigators have already identified the viral receptors expressed on the cell surface. Up to date, preprint publications have reported that SARS-CoV-2 employs angiotensin- converting enzyme 2 (ACE2) as the receptor for cell entry (Fig. 5) [26], which is the identical receptor that SARS-CoV uses for entry [63]. Both CoVs bind to ACE2 through their S proteins on the virions, after which the viral membrane and cell membrane fuse. The RNA virus SARS-CoV-2 then replicates its genome inside the host cell and new virions are eventually produced that are secreted to infect other cells. The fact that both SARS-CoV and SARS-CoV-2 utilize the same ACE2 receptor gives impetus to investigators to develop biologics for SARS-CoV promptly. Based on the SARS reports, some neutralizing antibodies are effective in preventing infections in the SARS prevention models [64].
Suggested therapeutic agents that can be utilized to prevent SARS-CoV-2 from infecting cells. Target cells expressing ACE2 involve human lung and gastrointestinal tissues. The huge spike protein on the surface of SARS-CoV-2 binds to ACE2 on the infected cell, making the cell to enter. The three suggested procedures would prevent this interaction and thus eliminate the infection. First, the receptor-binding domain (RBD) of the spike protein from SARS-CoV or SARS-CoV-2 will be applied to bind ACE2 and saturate the unoccupied sites. Second, an antibody or a single-chain antibody fragment (scFv) can be applied to ACE2 for the same purpose. The third procedure is to use ACE2 cell outfield directly as bait to bind spike proteins and immediately target coronavirus virions. The Fc domain combined with ACE2 will promote the prolonged cycle of the organism (ACE2-Fc).
SARS-CoV-2 vaccine under development
One important step to control COVID-19 is to develop a vaccine. In the first laboratory stage of a vaccine development, it is necessary to screen the virus strains, attenuate the virus strains as necessary [65]. Next steps are to study the adaptability of the virus strains to the cultured cell-matrix and the stability of the virus strains in the passage process, to explore the stability of process quality, and to establish animal models [66]. Dr. Paul Stoffels, vice chairman, and chief scientific officer of Johnson & Johnson, is one of the researchers who are engaged in the development of vaccines for SARS-CoV-2. According to his estimate, it takes eight months to a year for the SARS-CoV-2 vaccine to be rigorously tested before it is ready for public use [67]. It is also imperative to analyze the rate of SARS-CoV-2 genome evolution and to determine peptide vaccine targets [68]. There are eight CD4 T-cell epitopes with high binding affinity within S, E, M, and N proteins that are generally restricted by the human leukocyte antigen (HLA)-DR alleles found in Asian and Asian-pacific populations [69]. These immunodominant epitopes can be incorporated into the generic subunit coronavirus vaccine [69]. They used highly conserved and annotated structural protein sequences of the representative SARS-CoV-2 strain Wuhan-Hu-1 (MN908947.3) to predict possible CD4 T-cell epitopes [69]. Their prediction method recommended by the IEDB is based on the "immune epitope database and analysis resources (IEDB) consensus tool" [69]. However, standard mapping epitopes are essential to the design of globally effective subunit vaccines. They, therefore, examined common epitopes that are recognized by all dominant HLA-DR alleles that are prevalent in five ethnic groups [69]. Their analyses demonstrated that a subunit vaccine, including eight immunologically dominant HLA-DR epitopes, produced effective antiviral T-cell and antibody responses in populations of different ethnicities [69].
Previous studies have observed that only 23% and 16% of the known SARS-CoV T cell and B cell epitopes respectively are the same as that of SARS-CoV-2 and no mutant sequences have been recognized in the available SARS-CoV-2 epitopes [70]. This is a strong evidence of their potential to obtain a potent T cell or antibody response in SARS-CoV-2. In terms of T cells, the identification of the SARS-CoV-derived epitopes, located in the same location as SARS-CoV-2, and the large population assumed to be included, is especially encouraging [70]. It proposed further investigation into a vaccine designed to produce a protective T-cell response, which has been proved to afford long-term protection in SARS-CoV [70]. Linear SARS-CoV-derived B cell epitopes in the S2 subunit may be more likely applicants for producing protective antibody responses [70]. Many of these epitopes, despite their low exposure, map to the same locations as those of SARS-CoV and SARS-CoV-2, and preliminary outcomes have appeared indicating their potential for cross-reactive and antibody neutralization [70]. Therefore, vaccine solutions that endeavor to produce antibodies targeting S2 linear epitopes might be useful and should be more investigated.
Prevention and control
The current research data show that, like SARS-CoV, the SARS-CoV-2 receptor on human host cells is ACE2 [26, 71]. It is speculated that antibodies or drugs designed for SARS may also be suitable for SARS-CoV-2. Thus they can be used as priority product candidates for disease prevention and treatment. Vaccine research needs to be deployed for a long time. From the experience of SARS-CoV research, the inactivated virus vaccine may have an immunopathological enhancement effect [72]. Therefore, adverse reactions should be carefully examined and controlled. One of the candidates is an S-protein receptor-binding region-based vaccine [73]. It has a protective effect against SARS-CoV in an experimental animal model. When it comes to small molecular drugs for SARS-CoV-2, several teams from China have carried out the screening of compounds currently on the market as antivirals and found promising results in vitro, which are expected to be verified in an experimental animal model and clinical trials [74]. Military Academy of Medical Sciences Institute of Chinese Academy of Sciences and Toxic Drug Research Institute jointly found that three drugs, including Remdesivir, Chloroquine, and Ritonavir, inhibited the replication of SARS-CoV-2 in vitro [75,76]. In addition, in 2017, Zhang's group pointed to a host cell protease TMPRSS2 being a potentially important target for the treatment of influenza virus and coronavirus infections in the research on searching and screening the antiviral drugs [77]. More recently, results by Hoffmann's group from Germany demonstrate that SARS-CoV-2 spike proteins use the SARS-coronavirus receptor, ACE2, for entry and the cellular protease TMPRSS2 for SARS-CoV-2 spike protein priming [78]. A TMPRSS2 inhibitor blocked entry and might constitute a treatment option. Other studies have also shown that yogurt and probiotics can help prevent lung infections in the elderly [79]. Since the outbreak of SARS in 2003, China's scientific research capacity in the field of preventive medicine has been dramatically improved. In the outbreak of COVID-19 caused by SARS-CoV-2, rapid progress has been made in etiology identification, diagnostic reagent development, virus characteristic analysis, and clinical treatment strategy. At the same time, popular science education and media publicity have also played a decisive role, and public awareness and social responsibility for the prevention and control of the epidemic have become an essential factor in curbing the rise of the epidemic. However, still much has to be done in this outbreak, which is highlighted by the fact that essential questions, such as the virus's mutation characteristics, transmission pattern, and traceability, have yet to be answered. The scientific and public health community still needs to reflect deeply and act quickly to meet the challenge of SARS-CoV-2 and more new pathogens in the future.
Since ACE2 is a functional receptor of SARS-CoV-2, its safety and potential impact on disease should be fully considered when applying Angiotensin-converting enzyme inhibitors (ACEI) [74]. Therefore, for patients with the novel coronavirus pneumonia (COVID-19) combined with hypertension, Calcium channel blockers (CCBs) are recommended if they are taking ACEI or Angiotensin receptor blockers (ARBs) [80, 81]. In severe cases, Bradykinin (BK2) receptor blockers or direct renin inhibitor aliskiren can be used [82]. If hypotension occurs, antihypertensive drugs should be discontinued [14]. The application of ACEI drugs is still controversial and needs further discussion [80]. The SARS-CoV-2 causes COVID-19 by infecting host cells via ACE2 receptors, and also causes damage to the myocardium, although the mechanism is unclear. Patients with underlying cardiovascular disease had a poor prognosis after infection with SARS-CoV-2 [80, 81]. For acute myocardial injury, the primary disease should actively be treated and the myocardial protective treatment carried out [80, 81]. For patients with underlying heart disease, especially ST-Elevation Myocardial Infarction (STEMI), early reperfusion therapy should be performed under well-protected conditions. For patients with SARS-CoV-2 infection with cardiovascular disease, they should try to avoid using ACEI or ARB [14, 80, 81].
Conclusion
The novel coronavirus was first detected in December 2019 during the outbreak of severe pneumonia in Wuhan, Hubei Province, China. The disease caused by the virus outbreak was finally named COVID-19, which stands for the coronavirus (CoV) disease 2019, and the etiologic virus SARS- CoV-2, which stands for severe acute respiratory syndrome CoV 2 by WHO. Scientific investigation is underway to learn more about the virus, its origin, and how it affects humans, and the situation is changing rapidly. Most of the first patients in COVID-19 claimed to be related to the Huanan Seafood market in Wuhan, indicating that the virus had a zoonotic origin. However, the source of the virus remains to be determined. Bats, known as a reservoir for most viruses, might be the original host of the new strain, but the intermediate animal host is unknown. The virus is similar to SARS-CoV-like CoV in bats but is different from SARS-CoV and MERS-CoV. One study suggests that the virus may be a recombination of a bat CoV with a CoV of unknown origin, possibly from snakes [83], but there is no evidence to support this hypothesis. Human-to- human transmission has been confirmed in communities and medical institutions in China, as well as in Vietnam, Japan, Thailand, Germany, and the United States [84, 85]. Preliminary assessment of the transmission dynamics of the first 425 confirmed cases found that before January 1, 2020, 55% of the cases were related to the Huanan seafood market in Wuhan; after this date, only 8.6% of the cases were associated with this market [86]. This confirms that human-to-human transmission has occurred between close contacts since the middle of December 2019 [86]. Infections that occurred among health workers were also identified [86]. It is unclear how easily the virus can be transmitted from person to person. Asymptomatic carriers can become contagious to others, as there have been confirmed reports of transmission from asymptomatic contacts in Germany [87].
The full genome of the virus has been deposited in GenBank. Molecular assays using Reverse-Transcriptase Polymerase Chain Reaction (RT-PCR) are needed to confirm the diagnosis. There are no specific treatments available for COVID-19 up to now; therefore, the main body of treatment is optimized to support treatment to reduce symptoms and maintain organ functions in more severe diseases. Also, there are no specific anti-SARS-CoV-2 drugs. However, some existing antiviral medications have been used as part of clinical trials [88].
At present, China's National Health Commission has classified the new coronavirus as a class A infectious disease, requiring the most stringent prevention and control measures, including mandatory quarantine of patients and medical observation of those in close contact with them [4]. Many Chinese cities are currently on lockdown because of travel bans and the suspension of public transport services. It was reported that most of those who died were the elderly and individuals who had underlying illnesses [89]. According to the first case report, the overall case fatality rate is approximately 2.6% [2]; however, this estimate should be treated with caution as the actual number of cases of infection and its course are not yet known. We, thus, should carefully observe the current situation and take measures to control COVID-19.
Acknowledgements
We gratefully acknowledge the support by the National Natural Science Foundation of China (No. 21877101), the Zhejiang Leading Innovation and Entrepreneurship Team (2018R01015) and the Emergency Project of Key Research and Development Plan of Zhejiang Province (2020C03124).
Author contributions
AA designed research, wrote the manuscript and revised the manuscript. YLW conceived of the study. YT revised the manuscript. WZ designed the study, revised the manuscript and provided funding support. All authors have read and approved the final manuscript.
Competing Interests
The authors have declared that no competing interest exists.
References
1. Chan JF-W, Yuan S, Kok K-H, To KK-W, Chu H, Yang J. et al. A familial cluster of pneumonia associated with the 2019 novel coronavirus indicating person-to-person transmission: a study of a family cluster. The Lancet. 2020
2. Yang Y, Peng F, Wang R, Guan K, Jiang T, Xu G. et al. The deadly coronaviruses: The 2003 SARS pandemic and the 2020 novel coronavirus epidemic in China. Journal of Autoimmunity. 2020: 102434.
3. European Centre for Disease Prevention and Control 2020; https://. www.ecdc.europa.eu/en/geographical-distribution-2019-ncov-cases
4. Diagnosis and Treatment Protocol for Novel Coronavirus Pneumonia (Trial Version 6, Revised). The General Office of National Health Commission Office of State TCM Administration. 2020; Printed and distributed on February 18, 2020. http://www.kankyokansen.org/uploads/uploads/files/jsipc/protocol_V6.pdf
5. Khan N, Naushad M. Effects of Corona Virus on the World Community. Available at SSRN: https://ssrn.com/abstract=3532001 or. http://dx.doi.org/10.2139/ssrn.3532001
6. Corman VM, Landt O, Kaiser M, Molenkamp R, Meijer A, Chu DK. et al. Detection of 2019 novel coronavirus (2019-nCoV) by real-time RT-PCR. Eurosurveillance. 2020;25:2000045
7. Hui DS, Madani T, Ntoumi F, Kock R, Dar O, Ippolito G. et al. The continuing 2019-nCoV epidemic threat of novel coronaviruses to global health-The latest 2019 novel coronavirus outbreak in Wuhan, China. International journal of infectious diseases: IJID: official publication of the International Society for Infectious Diseases. 2020;91:264
8. Nishiura H, Jung S-m, Linton NM, Kinoshita R, Yang Y, Hayashi K. et al. The Extent of Transmission of Novel Coronavirus in Wuhan, China, 2020. Multidisciplinary Digital Publishing Institute. 2020
9. Carlos WG, Dela Cruz CS, Cao B, Pasnick S, Jamil S. Novel Wuhan (2019-nCoV) Coronavirus. American Journal of Respiratory and Critical Care Medicine. 2020
10. Pan Y, Guan H, Zhou S, Wang Y, Li Q, Zhu T. et al. Initial CT findings and temporal changes in patients with the novel coronavirus pneumonia (2019-nCoV): a study of 63 patients in Wuhan, China. European Radiology. 2020:1-4
11. Tian H. 2019-nCoV: new challenges from coronavirus. Zhonghua yu fang yi xue za zhi [Chinese journal of preventive medicine]. 2020;54:E001
12. Munster VJ, Koopmans M, van Doremalen N, van Riel D, de Wit E. A Novel Coronavirus Emerging in China-Key Questions for Impact Assessment. New England Journal of Medicine. 2020
13. Randhawa GS, Soltysiak MP, El Roz H, de Souza CP, Hill KA, Kari L. Machine learning-based analysis of genomes suggests associations between Wuhan 2019-nCoV and bat Betacoronaviruses. bioRxiv. 2020
14. Jin Y-H, Cai L, Cheng Z-S, Cheng H, Deng T, Fan Y-P. et al. A rapid advice guideline for the diagnosis and treatment of 2019 novel coronavirus (2019-nCoV) infected pneumonia (standard version). Military Medical Research. 2020;7:4
15. Sahin AR, Erdogan A, Agaoglu PM, Dineri Y, Cakirci AY, Senel ME. et al. 2019 Novel Coronavirus (COVID-19) Outbreak: A Review of the Current Literature. EJMO. 2020;4:1-7
16. Zhu N, Zhang D, Wang W, Li X, Yang B, Song J. et al. A Novel Coronavirus from Patients with Pneumonia in China, 2019. New England Journal of Medicine. 2020
17. Chen Z-M, Fu J-F, Shu Q, Chen Y-H, Hua C-Z, Li F-B. et al. Diagnosis and treatment recommendations for pediatric respiratory infection caused by the 2019 novel coronavirus. World Journal of Pediatrics. 2020:1-7
18. Chan JF-W, Kok K-H, Zhu Z, Chu H, To KK-W, Yuan S. et al. Genomic characterization of the 2019 novel human-pathogenic coronavirus isolated from a patient with atypical pneumonia after visiting Wuhan. Emerging Microbes & Infections. 2020;9:221-36
19. Zumla A, Hui DS, Perlman S. Middle East respiratory syndrome. The Lancet. 2015;386:995-1007
20. Ohmit SE, Thompson MG, Petrie JG, Thaker SN, Jackson ML, Belongia EA. et al. Influenza vaccine effectiveness in the 2011-2012 season: protection against each circulating virus and the effect of prior vaccination on estimates. Clinical Infectious Diseases. 2014;58:319-27
21. Liu J, Zheng X, Tong Q, Li W, Wang B, Sutter K. et al. Overlapping and discrete aspects of the pathology and pathogenesis of the emerging human pathogenic coronaviruses SARS-CoV, MERS-CoV, and 2019-nCoV. Journal of Medical Virology. 2020
22. Cui J, Li F, Shi Z-L. Origin and evolution of pathogenic coronaviruses. Nature reviews Microbiology. 2019;17:181-92
23. Schoeman D, Fielding BC. Coronavirus envelope protein: current knowledge. Virology journal. 2019;16:69
24. Gorbalenya AE. Severe acute respiratory syndrome-related coronavirus-The species and its viruses, a statement of the Coronavirus Study Group. bioRxiv. 2020
25. Mirza MU, Froeyen M. Structural Elucidation of SARS-CoV-2 Vital Proteins: Computational Methods Reveal Potential Drug Candidates Against Main Protease, Nsp12 RNA-dependent RNA Polymerase and Nsp13 Helicase. 2020.
26. Zhou P, Yang X-L, Wang X-G, Hu B, Zhang L, Zhang W. et al. Discovery of a novel coronavirus associated with the recent pneumonia outbreak in humans and its potential bat origin. bioRxiv. 2020
27. Gasteiger E, Gattiker A, Hoogland C, Ivanyi I, Appel RD, Bairoch A. ExPASy: the proteomics server for in-depth protein knowledge and analysis. Nucleic acids research. 2003;31:3784-8
28. Chappell MC, Marshall AC, Alzayadneh EM, Shaltout HA, Diz DI. Update on the angiotensin converting enzyme 2-angiotensin (1-7)-Mas receptor axis: fetal programing, sex differences, and intracellular pathways. Frontiers in endocrinology. 2014;4:201
29. Kuba K, Imai Y, Ohto-Nakanishi T, Penninger JM. Trilogy of ACE2: A peptidase in the renin-angiotensin system, a SARS receptor, and a partner for amino acid transporters. Pharmacology & therapeutics. 2010;128:119-28
30. Cai G. Bulk and single-cell transcriptomics identify tobacco-use disparity in lung gene expression of ACE2, the receptor of 2019-nCov. medRxiv. 2020
31. Cao Y, Li L, Feng Z, Wan S, Huang P, Sun X. et al. Comparative genetic analysis of the novel coronavirus (2019-nCoV/SARS-CoV-2) receptor ACE2 in different populations. Cell Discovery. 2020;6:1-4
32. Zhao Y, Zhao Z, Wang Y, Zhou Y, Ma Y, Zuo W. Single-cell RNA expression profiling of ACE2, the putative receptor of Wuhan 2019-nCov. bioRxiv. 2020
33. Yu Han HY. The transmission and diagnosis of 2019 novel coronavirus infection disease (COVID-19): A Chinese perspective. 2020.
34. Zhang H, Penninger JM, Li Y, Zhong N, Slutsky AS. Angiotensin-converting enzyme 2 (ACE2) as a SARS-CoV-2 receptor: molecular mechanisms and potential therapeutic target. Intensive Care Medicine. 2020:1-5
35. Zhang JJ, Dong X, Cao YY, Yuan YD, Yang YB, Yan YQ. et al. Clinical characteristics of 140 patients infected by SARS-CoV-2 in Wuhan, China. Allergy. 2020
36. Chen N, Zhou M, Dong X, Qu J, Gong F, Han Y. et al. Epidemiological and clinical characteristics of 99 cases of 2019 novel coronavirus pneumonia in Wuhan, China: a descriptive study. The Lancet. 2020
37. Shen K, Yang Y, Wang T, Zhao D, Jiang Y, Jin R. et al. Diagnosis, treatment, and prevention of 2019 novel coronavirus infection in children: experts' consensus statement. World Journal of Pediatrics. 2020:1-9
38. The Novel Coronavirus Pneumonia Emergency Response Epidemiology Team. The Epidemiological Characteristics of an Outbreak of 2019 Novel Coronavirus Diseases (COVID-19)-China, 2020. China CDC Weekly. 2020;2:113-22
39. General Office of National Health Committee. Office of State Administration oftraditional Chinese Medicine. Notice on the issuance of a programme for the diagnosis and treatment of novel coronavirus (2019-nCoV) infected pneumonia (trial fourth edition) (In Chinese) Released on 28 January 2020 (EB/OL) Available at:. http://bgs.satcm.gov.cn/zhengcewenjian/2020-01-28/12576.html
40. Xie X, Zhong Z, Zhao W, Zheng C, Wang F, Liu J. Chest CT for typical 2019-nCoV pneumonia: relationship to negative RT-PCR testing. Radiology. 2020: 200343.
41. Huang C, Wang Y, Li X, Ren L, Zhao J, Hu Y. et al. Clinical features of patients infected with 2019 novel coronavirus in Wuhan, China. The Lancet. 2020
42. Pan F, Ye T, Sun P, Gui S, Liang B, Li L. et al. Time course of lung changes on chest CT during recovery from 2019 novel coronavirus (COVID-19) pneumonia. Radiology. 2020: 200370.
43. China's National Microbiology Data Center https://ww2sinaimgcn/thumb150/bd29cf19gy1gb7ntt1h0oj20jg0grn00jpg or. https://iimgurcom/xOBHPNJjpg
44. https://www.reddit.com/r/China_Flu/comments/etkdcv/chinas_national_microbiology_data_
45. Wang F-S, Zhang C. What to do next to control the 2019-nCoV epidemic? The Lancet. 2020;395:391-3
46. Que T, Wong V, Yuen K. Treatment of severe acute respiratory syndrome with lopinavir/ritonavir: a multicentre retrospective matched cohort study. Hong Kong Med J. 2003;9:399-406
47. Chu C, Cheng V, Hung I, Wong M, Chan K, Chan K. et al. Role of lopinavir/ritonavir in the treatment of SARS: initial virological and clinical findings. Thorax. 2004;59:252-6
48. Lai S. Treatment of severe acute respiratory syndrome. European Journal of Clinical Microbiology and Infectious Diseases. 2005;24:583-91
49. Kwan AC, Chau TN, Tong WL, Tsang OT, Tso EY, Chiu MC. et al. Severe acute respiratory syndrome related diarrhea. Journal of gastroenterology and hepatology. 2005;20:606-10
50. Meyer B, Basra A, Aberle S, Aberle J, Robibaro B, Wenisch C. et al. 1225: MERS-COV Disease Associated Ards-A Case Report. Critical care medicine. 2015;43:308
51. Falzarano D, De Wit E, Rasmussen AL, Feldmann F, Okumura A, Scott DP. et al. Treatment with interferon-α2b and ribavirin improves outcome in MERS-CoV-infected rhesus macaques. Nature medicine. 2013;19:1313-7
52. Chan JFW, Yao Y-F, Yeung ML, Deng W, Bao L-L, Jia L-L. et al. Treatment with lopinavir/ritonavir or interferon-β1b improves outcome of MERS-CoV infection in a nonhuman primate model of common marmoset. The Journal of infectious diseases. 2015;212:1904-13
53. Omrani AS, Saad MM, Baig K, Bahloul A, Abdul-Matin M, Alaidaroos AY. et al. Ribavirin and interferon alfa-2a for severe Middle East respiratory syndrome coronavirus infection: a retrospective cohort study. The Lancet Infectious Diseases. 2014;14:1090-5
54. Khan PA, Nousheen B, Maryam N, Sultana K. Middle East Respiratory Syndrome (Mers): A Systematic Review. International Journal Of Pharmaceutical Sciences And Research. 2018;9:2616-25
55. Zhao Z, Zhang F, Xu M. Clinical analysis of 190 cases of outbreak with atypical pneumonia in Guangzhou in spring, 2003. Guangdong Medical Journal. 2003;24:73-6
56. Meng Q, Dong P, Guo Y, Zhang K, Liang L, Hou W. et al. Use of glucocorticoid in treatment of severe acute respiratory syndrome cases. Zhonghua yu fang yi xue za zhi [Chinese journal of preventive medicine]. 2003;37:233-5
57. Xiao J, Ma L, Gao J, Yang Z, Xing X, Zhao H. et al. Glucocorticoid-induced diabetes in severe acute respiratory syndrome: the impact of high dosage and duration of methylprednisolone therapy. Zhonghua Nei Ke Za Zhi. 2004;43:179-82
58. Casadevall A, Pirofski L-a. The Ebola epidemic crystallizes the potential of passive antibody therapy for infectious diseases. PLoS pathogens. 2015 11
59. Keck Z-y, Wang Y, Lau P, Foung SK. Isolation of HCV Neutralizing Antibodies by Yeast Display. Hepatitis C Virus Protocols: Springer. 2019 p. 395-419
60. Shin YW, Chang K-H, Hong G-W, Yeo S-G, Jee Y, Kim J-H. et al. Selection of vaccinia virus-neutralizing antibody from a phage-display human-antibody library. Journal of Microbiology and Biotechnology. 2019;29:651-7
61. Walker LM, Burton DR. Passive immunotherapy of viral infections: 'super-antibodies' enter the fray. Nature Reviews Immunology. 2018;18:297
62. Group TPIW, Team M-NPIS. A randomized, controlled trial of ZMapp for Ebola virus infection. The New England journal of medicine. 2016;375:1448
63. Li W, Moore MJ, Vasilieva N, Sui J, Wong SK, Berne MA. et al. Angiotensin-converting enzyme 2 is a functional receptor for the SARS coronavirus. Nature. 2003;426:450-4
64. Kruse RL. Therapeutic strategies in an outbreak scenario to treat the novel coronavirus originating in Wuhan, China. F1000Research. 2020;9:72
65. Han S. Clinical vaccine development. Clinical and experimental vaccine research. 2015;4:46-53
66. Buckland BC. The process development challenge for a new vaccine. Nature medicine. 2005;11:S16-S9
68. Kumar S. Drug and vaccine design against Novel Coronavirus (2019-nCoV) spike protein through Computational approach. 2020.
69. Ramaiah A, Arumugaswami V. Insights into cross-species evolution of novel human coronavirus 2019-nCoV and defining immune determinants for vaccine development. bioRxiv. 2020
70. Ahmed SF, Quadeer AA, McKay MR. Preliminary identification of potential vaccine targets for the COVID-19 coronavirus (SARS-CoV-2) based on SARS-CoV immunological studies. Viruses. 2020;12:254
71. Shen Q, Xiao X, Aierken A, Liao M, Hua J. The ACE2 Expression in Sertoli cells and Germ cells may cause male reproductive disorder after SARS-CoV-2 Infection. 2020.
72. Honda-Okubo Y, Barnard D, Ong CH, Peng B-H, Tseng C-TK, Petrovsky N. Severe acute respiratory syndrome-associated coronavirus vaccines formulated with delta inulin adjuvants provide enhanced protection while ameliorating lung eosinophilic immunopathology. Journal of virology. 2015;89:2995-3007
73. Zhu X, Liu Q, Du L, Lu L, Jiang S. Receptor-binding domain as a target for developing SARS vaccines. Journal of thoracic disease. 2013;5:S142
74. Pang J, Wang MX, Ang IYH, Tan SHX, Lewis RF, Chen JI-P. et al. Potential Rapid Diagnostics, Vaccine and Therapeutics for 2019 Novel Coronavirus (2019-nCoV): A Systematic Review. Journal of Clinical Medicine. 2020;9:623
75. Johnson M. Wuhan 2019 Novel Coronavirus-SARS-CoV-2. MATER METHODS. 2020
76. Wang M, Cao R, Zhang L, Yang X, Liu J, Xu M. et al. Remdesivir and chloroquine effectively inhibit the recently emerged novel coronavirus (2019-nCoV) in vitro. Cell research. 2020:1-3
77. Shen LW, Mao HJ, Wu YL, Tanaka Y, Zhang W. TMPRSS2: A potential target for treatment of influenza virus and coronavirus infections. Biochimie. 2017;142:1-10
78. Hoffmann M, Kleine-Weber H, Krueger N, Mueller MA, Drosten C, Poehlmann S. The novel coronavirus 2019 (2019-nCoV) uses the SARS-coronavirus receptor ACE2 and the cellular protease TMPRSS2 for entry into target cells. bioRxiv. 2020
79. Pu F, Guo Y, Li M, Zhu H, Wang S, Shen X. et al. Yogurt supplemented with probiotics can protect the healthy elderly from respiratory infections: a randomized controlled open-label trial. Clinical interventions in aging. 2017;12:1223
80. Liu Y, Yang Y, Zhang C, Huang F, Wang F, Yuan J. et al. Clinical and biochemical indexes from 2019-nCoV infected patients linked to viral loads and lung injury. Science China Life Sciences. 2020:1-11
81. Zhou Y, Hou Y, Shen J, Huang Y, Martin W, Cheng F. Network-based Drug Repurposing for Human Coronavirus. medRxiv. 2020
82. Riccioni G. The role of direct renin inhibitors in the treatment of the hypertensive diabetic patient. Therapeutic Advances in Endocrinology and Metabolism. 2013;4:139-45
83. Bassetti M, Vena A, Roberto Giacobbe D. The Novel Chinese Coronavirus (2019-nCoV) Infections: challenges for fighting the storm. European Journal of Clinical Investigation. 2020: e13209.
84. Phan LT, Nguyen TV, Luong QC, Nguyen TV, Nguyen HT, Le HQ. et al. Importation and human-to-human transmission of a novel coronavirus in Vietnam. New England Journal of Medicine. 2020;382:872-4
85. Manandhar S, Nakarmi P, Baniya N. A Novel Coronavirus Emerging in World—Key Questions for Developing Countries and Under Developed Countries
86. Li Q, Guan X, Wu P, Wang X, Zhou L, Tong Y. et al. Early transmission dynamics in Wuhan, China, of novel coronavirus-infected pneumonia. New England Journal of Medicine. 2020
87. Rothe C, Schunk M, Sothmann P, Bretzel G, Froeschl G, Wallrauch C. et al. Transmission of 2019-nCoV infection from an asymptomatic contact in Germany. The New England Journal of Medicine. 2020
88. Lu H. Drug treatment options for the 2019-new coronavirus (2019-nCoV). Bioscience Trends. 2020
89. Lai C-C, Shih T-P, Ko W-C, Tang H-J, Hsueh P-R. Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) and corona virus disease-2019 (COVID-19): the epidemic and the challenges. International journal of antimicrobial agents. 2020: 105924.
Author Biographies
 Dr. Annoor Awadasseid received his bachelor's degree in Biochemistry and Food Sciences, Faculty of Natural Resources and Environmental Studies, University of Kordofan, Sudan. He entered Prof. Yifa Zhou's group and received a Master's degree in Biochemistry and Molecular biology, School of Life Science, Northeast Normal University, China. Dr. Annoor entered Prof. Yi Xin's group and received his first Doctoral degree in Biochemistry and Molecular biology, College of Basic Medical Sciences, Dalian Medical University, China. After that, he moved to Center for Natural Products Research, Chengdu Institute of Biology, University of Chinese Academy of Sciences, China, to join Prof. Guolin Zhang's group as a second Ph. D research fellow working on the biosynthesis of natural products. Since 2020, he entered Prof. Zhang's group of College of Pharmaceutical Science, Zhejiang University of Technology, China, as a postdoctoral fellow working in the evaluation of the druggability of novel heterocyclic compounds as potential target drugs under the direction of Profs W. Zhang and Y.-L. Wu.
Dr. Annoor Awadasseid received his bachelor's degree in Biochemistry and Food Sciences, Faculty of Natural Resources and Environmental Studies, University of Kordofan, Sudan. He entered Prof. Yifa Zhou's group and received a Master's degree in Biochemistry and Molecular biology, School of Life Science, Northeast Normal University, China. Dr. Annoor entered Prof. Yi Xin's group and received his first Doctoral degree in Biochemistry and Molecular biology, College of Basic Medical Sciences, Dalian Medical University, China. After that, he moved to Center for Natural Products Research, Chengdu Institute of Biology, University of Chinese Academy of Sciences, China, to join Prof. Guolin Zhang's group as a second Ph. D research fellow working on the biosynthesis of natural products. Since 2020, he entered Prof. Zhang's group of College of Pharmaceutical Science, Zhejiang University of Technology, China, as a postdoctoral fellow working in the evaluation of the druggability of novel heterocyclic compounds as potential target drugs under the direction of Profs W. Zhang and Y.-L. Wu.
 Dr. Yanling Wu is a professor in Molecular Immunology and now heads the Cellular and Molecular Immunology Research Group. She received Master and Doctoral degrees in Applied Life Science in 2003 and in Medicine Science in 2006, respectively, from Tohoku University, Japan. After that, she entered to Professor Minato's group of School of Medicine, Kyoto University, Japan, as a senior researcher working in the field of molecular immunology. Her current research mainly focuses on understanding the molecular mechanisms of gene regulation related to diseases by immune inhibitory receptors. Dr. Wu have given oral presentations in international conferences and published related papers.
Dr. Yanling Wu is a professor in Molecular Immunology and now heads the Cellular and Molecular Immunology Research Group. She received Master and Doctoral degrees in Applied Life Science in 2003 and in Medicine Science in 2006, respectively, from Tohoku University, Japan. After that, she entered to Professor Minato's group of School of Medicine, Kyoto University, Japan, as a senior researcher working in the field of molecular immunology. Her current research mainly focuses on understanding the molecular mechanisms of gene regulation related to diseases by immune inhibitory receptors. Dr. Wu have given oral presentations in international conferences and published related papers.
 Yoshimasa Tanaka received his Ph. D in Hokkaido University Graduate School of Agriculture with a specialization in Enzymology and Biochemistry. After graduation, he continued his research in the field of Immunobiology. Since 2008, he is an associate professor and works in the Center for Innovation in Immunoregulative Immunology and Therapeutics that belongs to the Kyoto University Graduate School of Medicine. Now He is a professor in Molecular Immunology and works in the Center for Medical Innovation, Nagasaki University, Japan.
Yoshimasa Tanaka received his Ph. D in Hokkaido University Graduate School of Agriculture with a specialization in Enzymology and Biochemistry. After graduation, he continued his research in the field of Immunobiology. Since 2008, he is an associate professor and works in the Center for Innovation in Immunoregulative Immunology and Therapeutics that belongs to the Kyoto University Graduate School of Medicine. Now He is a professor in Molecular Immunology and works in the Center for Medical Innovation, Nagasaki University, Japan.
 Dr. Wen Zhang is a full professor with more than 30 years of research and teaching experience in Medicinal Chemistry and Chemical Biology. Dr. Zhang earned his doctorate degree in Bioorganic Chemistry from East China University of Science and Technology, China. Then, he entered Professor Ohrui's Lab of Tohoku University, Japan, working in the field of molecular recognition as a JSPS postdoctoral fellow. After that, he moved to Kyoto University, Japan, to join Professor Sugiyama's Chemical Biology group as a COE and JST research fellow working on biology and chemistry of polyamide-nucleic acids interaction. Now, Dr. Zhang has a special interest in elucidating the gene regulation mechanisms with small organic molecules and the development of gene-targeted drug. His group formed in 2008 and established an extremely fruitful collaboration with Prof. Sugiyama's Group of Kyoto University in order to better pursue aspects of gene-targeted drug research. To date, Dr. Zhang has published better papers as the first/corresponding author in excellent Journals including J Am Chem Soc, Eur J Med Chem, Cancer Lett, Int J Biol Sci and Sci Rep etc.
Dr. Wen Zhang is a full professor with more than 30 years of research and teaching experience in Medicinal Chemistry and Chemical Biology. Dr. Zhang earned his doctorate degree in Bioorganic Chemistry from East China University of Science and Technology, China. Then, he entered Professor Ohrui's Lab of Tohoku University, Japan, working in the field of molecular recognition as a JSPS postdoctoral fellow. After that, he moved to Kyoto University, Japan, to join Professor Sugiyama's Chemical Biology group as a COE and JST research fellow working on biology and chemistry of polyamide-nucleic acids interaction. Now, Dr. Zhang has a special interest in elucidating the gene regulation mechanisms with small organic molecules and the development of gene-targeted drug. His group formed in 2008 and established an extremely fruitful collaboration with Prof. Sugiyama's Group of Kyoto University in order to better pursue aspects of gene-targeted drug research. To date, Dr. Zhang has published better papers as the first/corresponding author in excellent Journals including J Am Chem Soc, Eur J Med Chem, Cancer Lett, Int J Biol Sci and Sci Rep etc.
Corresponding authors: Yanling Wu, Lab of Molecular Immunology, Virus Inspection Department of Zhejiang Provincial Center for Disease Control and Prevention, 630 Xincheng Road, Hangzhou, 310051, PR China; Tel: +86-571-87115282; Fax: +86-571-87115282; e-mail: ylwuzj.cn. Wen Zhang, Lab of Chemical Biology and Molecular Drug Design, College of Pharmaceutical Science, Zhejiang University of Technology, 18 Chaowang Road, Hangzhou, 310014, PR China; Tel: +86-571-88871507; Fax: +86-571-88871507; e-mail: wzhang63edu.cn.

 Global reach, higher impact
Global reach, higher impact