Impact Factor ISSN: 1449-2288
Int J Biol Sci 2020; 16(13):2283-2295. doi:10.7150/ijbs.33481 This issue Cite
Review
CRISPR/Cas9-related technologies in liver diseases: from feasibility to future diversity
1. Inflammation and Immune Mediated Diseases Laboratory of Anhui Province, Anhui Institute of Innovative Drugs, School of Pharmacy, Anhui Medical University, 81 Meishan Road, Hefei 230032, Anhui Province, China.
2. Institute for Liver Diseases of Anhui Medical University, Anhui Medical University, Hefei 230032, China.
3. Department of Pathology and Pathophysiology, and Bone Marrow Transplantation Center of the First Affiliated Hospital, Zhejiang University School of Medicine, Hangzhou, China.
4. Institute of Hematology, Zhejiang University & Zhejiang Engineering Laboratory for Stem Cell and Immunotherapy, Hangzhou, China.
5. Key Laboratory of Medical Reprogramming Technology, Shenzhen Second People's Hospital, First Affiliated Hospital of Shenzhen University, Shenzhen 518039, China.
6. Shenzhen Qianhai Icecold IT Co., Ltd. China.
7. Hefei Institutes of Physical Science, Chinese Academy of Sciences; Anhui Province Key Laboratory of Environmental Toxicology and Pollution Control Technology, Hefei, Anhui, PR China.
8. National Drug Clinical Trial Institution, the First Affiliated Hospital of Bengbu Medical College.
9. Anhui Provincial Cancer Hospital, West Branch of The First Affiliated Hospital of USTC, Division of Life Sciences and Medicine, University of Science and Technology of China, Hefei, Anhui, 230031, P.R. China.
*These authors contributed equally to this work.
Received 2019-2-8; Accepted 2019-12-17; Published 2020-6-1
Abstract
Liver diseases are one of the leading causes of mortality in the world, mainly caused by different etiological agents, alcohol consumption, viruses, drug intoxication, and malnutrition. The maturation of gene therapy has heralded new avenues for developing effective interventions for these diseases. Derived from a remarkable microbial defense system, clustered regularly interspaced short palindromic repeats/CRISPR-associated proteins 9 system (CRISPR/Cas9 system) is driving innovative applications from basic biology to biotechnology and medicine. Recently, the mutagenic function of CRISPR/Cas9 system has been widely adopted for genome and disease research. In this review, we describe the development and applications of CRISPR/Cas9 system on liver diseases for research or translational applications, while highlighting challenges as well as future avenues for innovation.
Keywords: CRISPR/Cas9, viral hepatitis, hepatocellular carcinoma (HCC), clinical research
Introduction
Genome-editing technologies such as zinc-finger nucleases (ZFNs) and transcription activator-like effector nucleases (TALENs) have begun to enable targeted genome modifications, but they still have some restrictions [1]. As far as ZFNs, Two ZFNs are needed at either side of the double-strand breaks (DSBs), increasing specificity while escalating the size of the complex hindering delivery at the same time. TALENs can almost target any DNA sequence. However, TALENs are not easy to deliver in vivo due to a large molecular size [2]. But the situation changed with the development of clustered regularly interspaced short palindromic repeats/CRISPR-associated proteins 9 (CRISPR/Cas9) genome editing. CRISPR/Cas9 genome editing is a natural RNA-mediated adaptive defense mechanism found in some prokaryotic organisms [3]. It has rapidly become the most promising genome editing tool with great potential to revolutionize biomedical treatment [4]. Studies have shown that the ease of retargeting the system to modify genomic sequences greatly exceeds the capability of ZFNs and TALEs, while offering similar or greater efficiencies. Many physicians and scientists are now searching for the best clinical applications for this promising technology [5].
On the other hand, liver disease incidence is increasing year by year [6]. Frequently, liver diseases are initiated by oxidative stress and inflammatory reaction which leads to the excessive production of extracellular matrix (ECM), followed by a progression to hepatitis, fibrosis, cirrhosis and hepatocellular carcinoma (HCC) [7]. Additionally, steatosis can cause nonalcoholic steatohepatitis (NASH) and this is also accompanied with advanced fibrosis, cirrhosis, and ultimately HCC [8]. A major milestone in the development of CRISPR/Cas9 was its adaptation for use in mammalian cells, which provided researchers with a powerful tool to study genetic perturbations in tissue homeostasis and disease [1]. In this review, we sum up recent progress using this genome editing technology and explore its potential clinical applications, strategies, and challenges in the liver.
Overview of CRISPR/Cas9 technique
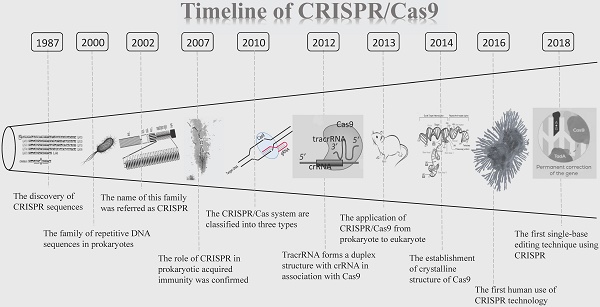
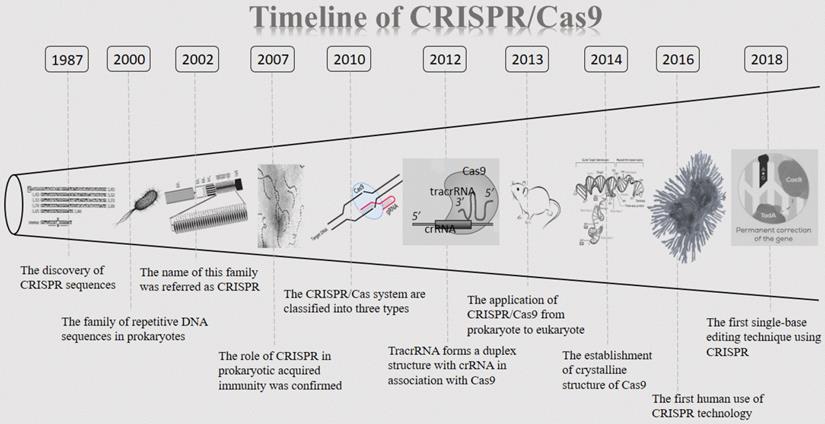
The CRISPR clustered repeats were first reported by Nakata and colleagues in 1987 [9]. They discovered a curious set of 29 nt repeats were interspaced by five intervening 32 nt nonrepetitive sequences when studying the iap enzyme involved in isozyme conversion of alkaline phosphatase in E. coli [9]. However, the significance of this discovery was not immediately recognized at the time by the scientific community. 13 years later, in 2000, this sequence was described again and was proposed to exist in prokaryotes. At that time, no such sequence has been named [10]. It was not until 2002 that this novel family of repetitive DNA sequences, present among both domains of the prokaryotes (Archaea and Bacteria), but absent from eukaryotes or viruses, once again caught the attention of researchers. This family is referred as CRISPR, which is characterized by direct repeats, varying in size from 21 to 37 bp, and interspaced by similarly sized nonrepetitive sequences [11]. CRISPR has since opened its legendary path. Then five years later, the role of CRISPR in prokaryotic acquired immunity was confirmed for the first time [12]. Between 2007 and 2011, CRISPR/Cas9 has shown great appeal to researchers and discoveries about CRISPR are springing up. In 2010, CRISPR/Cas systems are classified into three types (I- III) based on the structure and sequence of Cas proteins (Cas3 for type I, Cas9 for type II and Cas10 for type III), with a further division into several subtypes [3,13,14]. Afterward, one study proposed trans-activating crRNA (tracrRNA) forms a duplex structure with CRISPR-derived RNA (crRNA) in association with Cas9 [15]. Since then, CRISPR/Cas9 has entered an era of rapid development. Based on previous researches published, biochemical studies revealed that Cas9 is a programmable RNA-guided DNA endonuclease, and the Cas9-crRNA-tracrRNA complex cleaves double-stranded DNA (dsDNA) targets complementary to the 20-nucleotide guide sequence in the crRNA [16,17]. In 2013, the research on CRISPR/Cas9 finally moved from prokaryote to eukaryote [5]. It was reported that the CRISPR/Cas9 system can be used for genome editing in mammalian cells. The following studies found that the CRISPR/Cas9 system can be utilized to generate KO mice efficiently [18]. At this point, CRISPR/Cas9 ultimately came to the world as genetic engineering technology. With the deepening of exploration, the first high throughput screening of CRISPR long-chain non-coding RNA (lncRNA) gene was completed and a pregenome RNA (pgRNA) library was constructed using lentivirus as a vector. The genome-wide screening of nearly 700 genes in the human liver cancer cell line Huh 7.5 OC associated with cancer or other diseases with lncRNA was performed [19]. In October 2016, Lu you et al. launched the world's first human application of CRISPR, in which T cells were isolated from patients recruited, and CRISPR technology was used to KO the PD-1 gene in the cell [20]. Another breakthrough was achieved by the Zhang F team in 2017, which fused RNA-editing enzymes into targeted RNA-targeted Cas proteins, which can artificially edit specific nucleotides in human cells. This method called RNA Editing for Programmable A-to-I replacement (REPAIR) [21]. However, this technology has also caused controversy among some researchers, who believe it may be a double-edged sword. Treatment by CRISPR/Cas9 may increase the risk of cancer in patients [22,23]. Based on this, in 2018, single-base editing is a further development of gene-editing technology CRISPR, which may break the cancer panic of CRISPR [24]. In summary, as an important, emerging and cutting-edge gene-editing technology, CRISPR/Cas9 plays an important role in genetic engineering (Figure 1).
The timeline of CRISPR/Cas9. CRISPR/Cas9: Clustered regularly interspaced short palindromic repeats/CRISPR-associated proteins 9 system; crRNA: CRISPR-derived RNA.
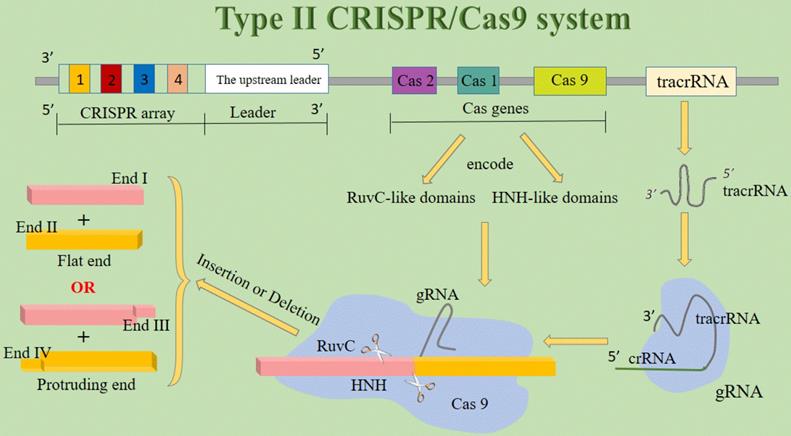
The working mechanism of the CRISPR/Cas system differs from other genome editing platforms, for it uses an RNA molecule more than a protein to recognize DNA [25]. CRISPR/Cas systems are classified into three types (I-III). All these types have three essential components in common: the CRISPR array, the upstream leader sequence, and the Cas genes. The CRISPR array consists of identical repeats with a length of 23-47 bp. The CRISPR/Cas leader region acts as a promoter for the transcription of the CRISPR array. The Cas genes encode the Cas proteins, containing RuvC- and HNH-like catalytic domains, which cleave the targeted DNA. Recognition of the target sequence by the Cas proteins is facilitated by the presence of the protospacer-associated motif (PAM). This sequence is usually 2-4 nucleotides long and flanks the target site. It is absent from the endogenous loci so it can prevent CRISPR/Cas auto cleavage and add specificity to targeting [3]. Types I and III CRISPR loci contain multiple Cas proteins, now known to form complexes with crRNA to facilitate the recognition and destruction of target nucleic acids. Type II CRISPR/Cas system is the simplest in terms of the number of genes for types I and III have more various cas genes [14]. The type II CRISPR/Cas system is the most commonly used CRISPR/Cas system for gene editing applications, which use the Cas9 protein, the only enzyme that mediates target DNA cleavage within the Cas gene cluster, through recognizing the relevant PAM sequence [26]. Therefore, the majority of CRISPR-based technology development has focused on the signature Cas9 nuclease from type II CRISPR systems [27]. In type II systems, a small non-coding RNA called the tracrRNA, is partially complementary to the CRISPR repeats, forming an RNA duplex with crRNA. This RNA hybrid is recognized and processed to form mature gRNA and then is combined with Cas9. The complex, including the tracrRNA, recognizes invading DNA and inactivates it by cleavage [3]. In addition to Cas9, all identified type II CRISPR-Cas loci contain Cas1 and Cas2 and most type II loci also encode a tracrRNA, which is partially complementary to the repeats within the respective CRISPR array [14]. Since the establishment of a new generation of CRISPR/Cas9 gene-editing system, it has been widely believed that Cas9 nuclease cleans DNA double strands to produce flat ends. But amazingly, DNA double-strand cleavage induced by Cas9 nuclease can produce protruding terminal. This latest study has fundamentally challenged the existing understanding of DNA cleavage by Cas9 nucleases and may become a turning point discovery and original innovation in the new generation of CRISPR/Cas9 gene-editing systems [28]. It lays a solid foundation for optimizing and modifying gene-editing technology (Figure 2).
The structure of type II CRISPR/Cas9 system. Abbreviations; crRNA: CRISPR-derived RNA; gRNA: guide RNA; tracrRNA: trans-activating crRNA.
In addition to generating frameshift-derived KO mice, the CRISPR/Cas9 system offers many other applications, such as point mutations, small insertions, large deletions, large insertions, and multiplex modification. In 2018, a significant breakthrough in gene therapy was achieve by Professor Zhang F and Professor David Liu, showing that single-base editing technology can accurately edit individual bases on DNA or RNA with a very high degree of flexibility and efficiency [24]. By using this technology, we can precisely repair a series of point mutations which lead to diseases, write genetic mutations that help prevent disease or regulate the expression of disease-causing genes.
The CRISPR/Cas9 has been particularly hot in the field of liver research. For example, the CRISPR/Cas9 system was capable of disrupting the intrahepatic HBV genome, with a significant increase of hepatitis B surface antigen (HBsAg) in an HBV hydrodynamics-mouse model [29]. At present, gene editing is being developed to treat a variety of chronic disorders and has exciting potential for curing liver diseases [3].
Current applications in liver diseases
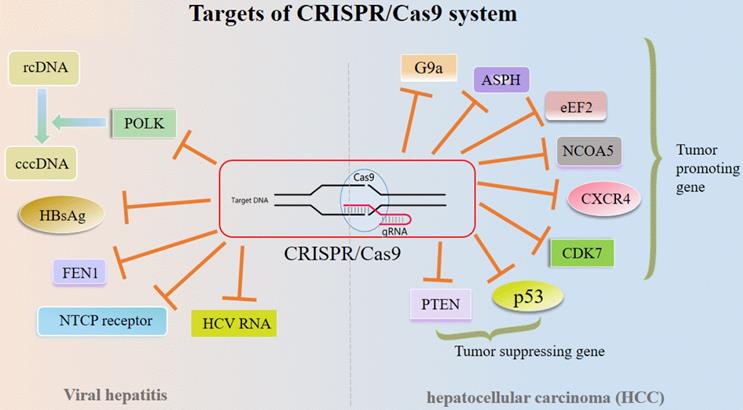
Recently, the CRISPR/Cas9 system is utilized as a precise and time- and cost-saving technique for gene KO, while liver diseases vary and suitable options for the therapy adopting CRISPR/Cas9 technique are often limited [3, 30]. Treatment outcomes depend not only on patient history and disease type but also on the progression of the disease and prior treatment. Overall, gene therapy using DNA modification is now possible and available, which may be exploited to achieve long term therapeutic benefit [3] (Figure 3).
Targets of CRISPR/Cas9 system in liver diseases. Abbreviations; ASPH: Aspartate β-hydroxylase; cccDNA: covalently closed circular DNA; CXCR4: CXC chemokine receptor 4; eEF2: eukaryotic elongation factor 2; FEN1: Flap structure-specific endonuclease; HbsAg: Hepatitis B surface antigen; NCOA5: Nuclear receptor coactivator 5; NTCP: Sodium-taurocholate co-transporting polypeptide; POLK: Polymerase κ; PTEN: Tensin homolog deleted on chromosome ten; rcDNA: relaxed circular DNA.
Viral hepatitis
Hepatitis B virus (HBV) infection is a major global health problem, despite the availability of effective vaccines [31]. Approximately 240 million people are chronically infected with HBV worldwide [32]. Viral infections are major risk factors for chronic liver diseases. With carriers exhibiting increased susceptibility to cirrhosis and HCC, more than 700, 000 people die from HBV-associated diseases each year [3,31]. Persistence of HBV covalently closed circular DNA (cccDNA) under current antiviral therapy is the major barrier to the eradication of chronic hepatitis B (CHB) [29]. For instance, although nucleos(t)ide analogs can inhibit HBV replication efficiently, they cannot eliminate cccDNA and it persists in hepatocyte nuclei [31]. Thus, HBV has emerged as an attractive target for CRISPR/Cas9 in the laboratory and curing CHB will require novel strategies for specific disruption of cccDNA [29].
Lin et al. were the first group to use CRISPR/Cas9 to target HBV [29]. Additionally, multiple studies have showed antiviral gRNAs can be promoted to expression by co-injecting HBV-encoding plasmid DNA and CRISPR/Cas9 vectors through the animal's tail vein in hydrodynamics-HBV in vivo models, which will eliminate the virus production efficiently [33]. Subsequently, Dong et al. used CRISPR/Cas9 target to establish cccDNA expression model in vivo [34]. Moreover, DNA polymerase κ (POLK), a Y-family DNA polymerase with maximum activity in non-dividing cells, substantially contributes to cccDNA formation during de novo HBV infection. Depleting gene expression of POLK by CRISPR/Cas9 KO inhibited the conversion of relaxed circular DNA (rcDNA) into cccDNA, thereby diminishing cccDNA formation and the viral infection [35]. A recent study indicated that HBV replication markers were effectively inhibited following the delivery of SaCas9 and S gene targeting gRNA into HepG2.2.15 cells [36].
HCV (hepatitis C virus) is one of the leading causes of chronic hepatitis, liver cirrhosis and HCC and infects approximately 170 million people worldwide [37]. The Cas9 variant from Francisella novicida (FnCas9) is reported to be capable of binding the mRNA from HCV, and thus inhibiting HCV protein expression. Interestingly, FnCas9 is also able to target DNA for double-strand cleavage, showing the possibility for a dual-targeting strategy [38]. Besides, CRISPR/Cas9 could be also used to target host factors that are essential for viral replication. These two targeting strategies may reduce the possibility of a severe liver injury for therapy trials [39] (Table 1).
Applications of CRISPR/Cas9 to viral hepatitis
| Species | Target | Cell type | Effect | Function | References (PMID) |
|---|---|---|---|---|---|
| hCas9 | HBV1.2 S1(3028-3050), P1(1292-1314), PS(261-283; 621-643;648-670), XCp(1742-1764), eE(1876-1898), PCE(2421-2443) | Huh7 cells | Diminish cccDNA and rcDNA production | Inhibit HBV | 25137139 |
| pX330-U6-Chimeric_BB-CBh-hSpCas9 | HBV1.3 X(1523-1542; 1661-1700; 2338-2357; 2416-2435), ORF X/L | Huh7 and HepG2.2.15 cells | Decrease cccDNA concentration, decrease serum levels of the HBsAg and HBeAg | Inhibit HBV | 25843425 |
| SaCas9 | HBV ORF S,P | Huh7, hNTCP-HepG2 and HepG2.2.15 cells | Inactivation and preferentially degradation of cccDNA | Inhibit HBV | 28785016 |
| SaCas9 | HBV Sa1(278-252), Sa3(1862-1889), Sa4(2405-2378) | Huh7, HepG2.2.15 and HepAD38 cells | Decrease HBsAg, HBV DNA and pgRNA levels | Inhibit HBV | 29458131 |
| pSpCas9 BB-2A-Puro (PX459) | HBV1.2 ORF S4(368-390), S5 (688-710), XP(1257-1278), CP-BCP(1868-1890), CP-URR(1682-1703) | HepG2.A64 cells | Reduce serum surface-antigen levels | Inhibit HBV | 27570484 |
| Cas9n | HBV ORF S1, S2, X1, X2 | HeLa, HEK293, HepG2-H1.3, HepG2-H2.2.15, hNTCP-HepG2 cells | Inactivate HBV in chronically and de novo infected cells | Inhibit HBV | 26334116 |
| Cas9 | POLK | Huh7 and HepG2-NTCP cells | Inhibit the conversion of rcDNA into cccDNA, diminish cccDNA formation and the viral infection | Inhibit HBV | 27783675 |
| pX330-U6-Chimeric_BB-CBh-hSpCas9 | FEN1 | Hep38.7-Tet cells, HepG2-hNTCP-C4 cells, Hep38.7-Tet cells, 293FT cells, PXB primary human hepatocytes | Inhibit conversion of rcDNA to cccDNA, reduce cccDNA levels | Inhibit HBV | 29928064 |
| FnCas9 | HCV RNA | Huh-7.5 cells | Inhibit HCV protein expression | Inhibit HCV | 25918406 |
Additional studies are needed to assess whether CRISPR/Cas9 holds the therapeutic potential to limit productive infections in vivo and are capable of eradicating multiple cccDNA copies present in infected hepatocytes [33]. In the future, the CRISPR/Cas9 system may be the most feasible approach for targeting HBV cccDNA. It is necessary to further improve the strategy to maximize the effects and minimize toxicity [31].
HCC
HCC is the fifth most common tumor and the third leading cause of cancer mortality worldwide [40]. CRISPR/Cas9-mediated gene KO is a powerful technique for precise gene modification in numerous cell types, which provides a theoretical basis for the development of novel therapies for many cancers, including HCC.
G9a is a lysine methyltransferase and its primary function is to di-methylate lysine 9 of histone H3 (H3K9me2). G9a-dependent H3K9me2 can recruit H3K9me2-binding proteins preventing transcriptional activation, which leads to gene silencing [41]. The up-regulation of G9a indicates poor prognosis in HCC [42]. Experimentally, G9a-KO by using CRISPR/Cas9 can suppress the proliferation and migration of HCC cells in vitro and inhibited HCC tumorigenicity in vivo, suggesting that targeted applications of CRISPR/Cas9 system could disrupt the HCC development both in vitro and in vivo [43]. Moreover, much of evidence proved that total eukaryotic elongation factor 2 (eEF2) and phosphorylated eEF2 at threonine 56 are prognostic markers for overall survival of HCC-patients and the regulating eEF2 kinase is a potential drug target for tumor therapy [44]. eEF2 plays an essential role in the GTP-dependent translocation of the ribosome along mRNA [45]. CRISPR/Cas9-mediated eEF2 kinase KO was performed in the HCC cell line, indicating that both the cell proliferation and the growth rate decrease with the elimination of eEF2 kinase by CRISPR/Cas9 in HCC cells [44]. Nuclear receptor coactivator 5 (NCOA5) plays an important role in the development of a variety of malignancies. NCOA5-KO HCC cells (LM3) by CRISPR/Cas9-mediated genome editing has been successfully generated, finding that NCOA5-KO suppresses epithelial-mesenchymal transition (EMT) in LM3, which leads to impaired cell proliferation and migration. EMT, a complex process, plays an important role in the advance of cancer [46]. Additionally, CXC chemokine receptor 4 (CXCR4) is a specific receptor of chemokine stromal cell-derived factor-1 (CXCL12). CXCL12 has a strong chemotaxis effect on lymphocytes. Numerous studies have demonstrated a marked association between high CXCR4 expression and the invasiveness, progression, and metastasis of HCC [47]. Targeting CXCR4 by CRISPR/Cas9 could inhibit HepG2 cell proliferation, migration and invasion, reversed EMT, increased chemosensitivity and decrease the malignancy of HCC in vitro and in vivo [48]. Another example comes to aspartate β-hydroxylase (ASPH), a ~86-kDa type II transmembrane protein, belonging to the α-ketoglutarate-dependent dioxygenase family. In HCC tumors, ASPH overexpresses and participates in the malignant transformation process. CRISPR/Cas9-mediated ASPH-knockout (KO) successfully trapped human HCC cells into senescence, thereby retarding HCC progression. It suggests that ASPH can be a potential therapeutic target, which shows a new mechanism that promotes HCC growth by regulating senescence of tumor cells [40] (Table 2).
Applications of CRISPR/Cas9 to HCC
| Species | Target | Cell type | Effect | Function | References (PMID) |
|---|---|---|---|---|---|
| Cas9 | HBV ORFpreS1/preS2/S | HCC cell lines (PLC/PRF/5, HepG2-2.15, Hep3B, SK-hep1, HLF, and Huh-7) and HEK293F cells | Decrease IL-6 production and inhibit STAT3 signaling | Inhibit HBV-associated HCC | 29904948 |
| Cas9 | MiR-3188 | HepG2 and SMMC7721 cells | Upregulate the expression of ZHX2 and block HBx-mediated activation of Notch signaling | Inhibit HBV-associated HCC | 28574502 |
| Cas9 | ASPH | Huh7 and HepG2 cells | Guide HCC cells to senescence | Inhibit HCC | 26683595 |
| Cas9 | eEF2 | JHH5 cell lines | Decrease cell proliferation and the growth rate | Inhibit HCC | 28060762 |
| Cas9 | NCOA5 | HCC cell lines (Huh-7, HepG2, Bel-7402, Bel-7404, LM3, SK-Hep-1) and LO2 | Suppress EMT | Inhibit HCC | 29626478 |
| Cas9 | CXCR4 | HepG2 cells | Inhibit proliferation, migration and invasion, reverse EMT, increase chemosensitivity, decrease the malignancy | Inhibit HCC | 28498420 |
| pSpCas9 BB-2A-Puro (PX459) | G9a | BEL7402, SMMC-7721, THLE-3 cells | Suppress the proliferation and migration, inhibit HCC tumorigenicity | Inhibit HCC | 28532996 |
| Dead SpCas9 | BAX and BCL2 | HepG2 cells | Induce cell apoptosis both in vitro and in vivo | Inhibit HCC | 27595406 |
| Cas9 | CDK7 | Hep3B and Huh7 cells | Impair proliferation of Hep3B and Huh7 cells | Inhibit HCC | 29507396 |
| pSpCas9 BB-2A-GFP (PX458) | p53 and PTEN | Mouse H2.35 cells | Accelerate HCC development | Promote HCC | 28584302 |
Most recently, by using CRISPR/Cas9-based functional genomics screening of the human kinome (including about 6,000 gRNAs for about 500 different kinases), researchers have found that a set of cyclin-dependent kinases (CDKs), which regulates gene transcription, may be a potential therapeutic target for HCC. Further studies have also indicated that there may be a subgroup of HCC cells with a profound dependence on CDK7 for survival and CDK7 may represent a novel therapeutic target in this subgroup. The high-throughput CRISPR/Cas9 screening technique was first applied to explore the treatment strategy of HCC, providing new approach towards the prevention and treatment of HCC [49]. In the future, we will strengthen our exploration in the field of liver and may have the capability to apply the CRISPR/Cas9 technique to HCC.
HTI
Metabolic liver diseases are better candidates for genome editing correction because many of them are typically severe, refractory to drug therapy and require orthotopic liver transplantation. For some, a low level of gene correction could significantly improve the disease phenotype [50,51]. Hereditary tyrosinemia type I (HTI) is a rare autosomal recessive disorder caused by a deficiency of fumarylacetoacetate hydrolase (Fah), the last enzyme that catalyzes the tyrosine catabolic pathway. The mutation of Fah will cause an accumulation of tyrosine and toxic catabolites in the body. The acute form of the disease is characterized by a hepatic failure while the chronic form is characterized by renal dysfunction and neurological crisis, and may lead to death [52]. HTI makes for better CRISPR/Cas9 targets because liver cells in which Fah has been repaired have a selective advantage and can expand and repopulate the liver [53].
The potential to correct a Fah mutation mediated by CRISPR/Cas9 has been demonstrated in hepatocytes in a mouse model of HTI. They injected the delivery of Cas9, sgRNA and a coinjected single-stranded DNA(ssDNA) containing the wild-type G nucleotide and homology arms flanking the sgRNA target region by non-viral hydrodynamic injection in the mouse model, which resulted in initial genetic correction rate of ~1/250 cells. The expansion of Fah-positive hepatocytes is sufficient to restore the weight loss of a mouse model of HTI [54]. Nevertheless, hydrodynamic injection yielded a low correction rate of 0.4% of hepatocytes. Then a safer and more efficient method of CRISPR delivery was reported. Systemic delivery of Cas9 mRNA by lipid nanoparticles and sgRNA/HDR template by adenovirus or adeno-associated virus (AAV) can correct a Fah mutation in the tyrosinemia mice liver and rescue their body weight loss. In this way, Cas9 nuclease can just express for a short time, making targeted gene editing more effective. The efficiency of correction was >6% of hepatocytes, which is significantly higher than the previous delivery system [55].
A new method called metabolic pathway reprogramming was used to cure HTI mice model successfully. Hydroxyphenylpyruvate dioxygenase (Hpd) is the enzyme that catalyzes the second step of tyrosine catabolism. By deleting Hpd with CRISPR/Cas9 in vivo, hepatacytes has been converted from tyrosinaemia type I into the benign tyrosinaemia type III. Then edited hepatocytes will replace the entire liver in only a few weeks. Hpd excision can change tyrosine catabolism, preventing the accumulation of tyrosine and toxic catabolites. Compared with gene therapy, metabolic pathway reprogramming doesn't need to express the wild-type protein of the disease-causing gene continuously and the protein may cause an immune response, limiting long-term expression [51].
It has brought up new ideas about treating HTI and the approach may be suitable for the treatment of a range of metabolic liver diseases. Because of its flexibility and ease in adjusting the target site, CRISPR/Cas9 system has many possibilities in the therapy of metabolic liver diseases.
Regulation of cellular signaling with CRISPR/Cas9 in liver diseases
Recently, novel designer enzymes, such as the CRISPR/Cas9 RNA-guided nuclease system, have provided technologies for developing advanced therapeutic strategies [56].
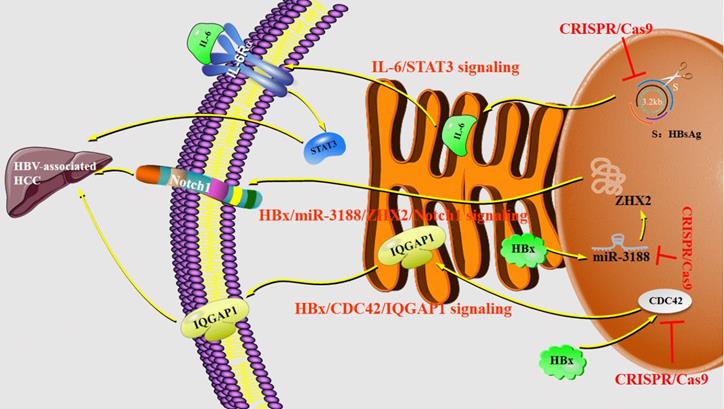
Researchers attempted to explore the mechanisms of the tumorigenicity suppression conferred by CRISPR/Cas9-mediated disruption of HBsAg in HCC cells. The results have demonstrated that HBsAg-KO in HCC cells can decrease interleukin (IL)-6 production and inhibite signal transducer and activator of transcription 3 (STAT3) signaling, which highlighted the tumorigenic role of HBsAg. It suggests that the IL-6/STAT3 pathway may be implicated in the HBsAg-mediated malignant potential of HBV-associated HCC [56]. The previous study has shown that microRNA-3188 (miR-3188) acts as the markedly overexpressed miRNA in HBV-associated HCC but the mechanism of miR-3188 regulation and cancer-related signaling pathways have not been elucidated. Then scientists manipulated miR-3188 expression in human HCC cell lines by the CRISPR/Cas9 systems [57]. The results have proved that miR-3188-KO by CRISPR/Cas9 can significantly increase the expression of zinc fingers and homeoboxes protein 2 (ZHX2), which is a direct target of miR-3188 in HCC cells, thus blocking HBV X protein (HBx)-mediated activation of miR-3188 and Notch signaling pathway [57]. HBx has been implicated in HBV-related hepatocarcinogenesis and is considered to be oncogenic [58,59]. The Notch signaling pathway is expected to become a new target for the biological treatment of HCC. Compared with non-HCC tissues, Notch1 associates with a higher expression level as well as Notch3, Notch4 [60], confirming that the HBx-miR-3188-ZHX2-Notch1 signaling pathway plays an important role in the pathogenesis and progression of HBV-related HCC with a family history of HCC. Furthermore, both the expression level and activity of cell division cycle 42 (CDC42) are up-regulated in HBx-expressing HuH-7 cells [61]. The analysis of CDC42 expression in 20 human liver samples has shown that HBV-related HCC tissues have a higher CDC42 expression [62]. Deficiency of CDC42 by the CRISPR/Cas9 system can significantly reduce the proliferation of HuH-7 cells promoted by HBx and truly down-regulate IQ Motif Containing GTPase Activating Protein 1 (IQGAP1), which by the way is a downstream effector of CDC42. Together, HBx/CDC42/IQGAP1 signaling pathways are downregulated in HuH-7-HBx CDC42 KO cells [61] (Figure 4).
Regulations of CRISPR/Cas9 system in liver diseases. Abbreviations; CDC42: cell division cycle 42; HBx: HBV X protein; IL-6: interleukin 6; IQGAP1: IQ Motif Containing GTPase Activating Protein 1; MiR-3188: microRNA-3188; STAT3: signal transducer and activator of transcription 3; ZHX2: zinc fingers and homeoboxes protein 2.
Current advantages and limitations
The treatment of liver diseases has been sophisticated, which requires and calls for new and better technologies. The ability to edit specific DNA sequences is of paramount importance to advance gene therapy for application to liver diseases [3].
The greatest advantage of the CRISPR/Cas9 system is its simplicity and wide applicability in genome manipulations of almost all biological systems tested to date, including cell lines, stem cells, yeasts, worms, insects, rodents, and mammals. Researchers can also mutate amino acids of Cas9 that are critical to DNA catalysis to produce dead Cas9 (dCas9), which still binds DNA but lacks endonuclease activity. When targeting the transcription start site (TSS) of genes, dCas9 can physically block the passage of RNA polymerase, leading to gene silencing [63]. Most importantly, this effect is both controllable and reversible.
To the best of our knowledge, the most exciting application of the CRISPR/Cas9 technology is its potential in the treatment of human diseases. There have been a growing number of preclinical trials targeting various human diseases even though it is currently still at its early stage for clinical applications [4]. Some corresponding limitations need to be addressed before clinical application of CRISPR/Cas9 for liver diseases can be achieved.
One of the foremost challenges is the off-target activity [4]. For example, the double-strand breaks (DSBs) may widely occur when the gRNA and DNA heteroduplex is formed without strict restrictions [4]. Due to the formation of DNA DSBs, Cas9 cutting may lead to cytotoxicity while the modification of Cas9 is irreversible. Bradley's team has shown that DNA breaks introduced by single-guide RNA/Cas9 can frequently resolve into deletions extending over many kilobases. Furthermore, lesions distal to the cut site and crossover events have been identified. The observed genomic damage in mitotically active cells caused by CRISPR/Cas9 editing may have pathogenic consequences [64]. For instance, a CRISPR/Cas9-system -direct-targeting p53 and Phosphatase and Tensin homolog deleted on chromosome ten (PTEN) in combination have been constructed and delivered by hydrodynamic tail vein injection to the liver of HBV-transgenic (HBV-Tg) mice [65]. The p53 is a tumor suppressor gene and more than 50% of all malignant tumors have mutations in the gene. PTEN, also a tumor suppressor gene, is a negative regulator of the phosphatidylinositol-3-kinase (PI3K)/Akt signaling pathway [66]. This study has demonstrated that the CRISPR/Cas9 mediated mutations of p53 and PTEN loci in adult mice are sufficient to accelerate HCC development in HBV-Tg mice without treatment of any chemical carcinogen [65]. In another experiment, the liver cancer models have been generated in wild-type mice by using the CRISPR/Cas9 system to target P53 and PTEN [66]. These researches have raised a serious concern for clinical applications that any off-target activity may cause undesirable consequences, which is not acceptable towards treatments [4]. To alleviate these concerns, several methods have been proposed including selecting gRNAs through various bioinformatics tools to use shorter gRNA, developing sensitive detection methods, improving Cas9 enzymes, constructing effective anti-Cas9 inhibitors [67,68], and amending delivery solutions [4,64]. In order to solve the off-target problem of Cas9 system and avoid side effects, many research teams try to add a safe and controllable switch to regulate the system, making the work of Cas9 system have spatial-temporal specificity and conditional restriction. Doxycycline-regulated Cas9 expression induction system has been constructed and gene knockout has been achieved in multiple tissues, which enables safe and controllable gene editing in vivo [69]. Nihongaki Y. et al. have split Cas9 into two inactive fragments and added a magnet protein to each fragment. When being irradiated by blue light, the magnet proteins come together and the separated Cas9 fragments recombine to activate RNA-guided nuclease. Importantly, the process is reversible: when the light is extinguished, the Cas9 nuclease splits again and its activity can be silenced [70]. Our team has also developed a similar CRISPR-Cas9-based light-controlled gene expression device that consists of genomic anchor (dCas9-CIB1 fusion protein), transcriptional activator (CRY2-AD fusion protein), sgRNA and reporter/effector gene. In the absence of blue light, the genomic anchor binds guided by the sgRNA to the targeted sequence (gene promoter sequence), while the activator is freely diffused within nucleus. Upon blue light illumination, CRY2 and CIBI combine with each other, recruiting transcriptional ADs to the targeted sequence (promoter) to activate gene transcription [71].
The second obstacle is the PAM restriction. The precision of the CRISPR/Cas9 DSB activity requires the presence of PAM. CRISPR/Cas9s from different bacterial species have been found to have different PAMs with various lengths and nucleotide compositions. Each type of PAM determines the cutting frequencies of the CRISPR/Cas9 for a given genome. None of the PAMs identified so far or even a combination of all of the known PAMs could cover any whole genome sequences, which may restrict the use of CRISPR/Cas9 technology in some cases. The ideal strategy would be to customize a PAM sequence tailored only to the desired target DNA site through alteration of the responsible PAM interacting amino acids, which has been proved possible by the identification of the amino acids responsible for PAM recognition in both SpCas9 and Staphylococcus aureus Cas9 (SaCas9) [4]. Most recently, phage-assisted continuous evolution has been used to evolve an expanded PAM SpCas9 variant (xCas9) that can recognize a broad range of PAM sequences including NG, GAA, and GAT. xCas9 not only broadened PAM compatibility but also has much greater DNA specificity than SpCas9 [72].
The third difficulty is the variations of efficiency. The efficiency of CRISPR/Cas9 directed DSB activities varies widely depending on the nucleotide compositions and genomic context of the target protospacer DNA sites as well as the sgRNA secondary structure. Additionally, the genome editing efficiency is also affected by non-homologous end joining (NHEJ) and homology-directed repair (HDR) DNA repair mechanisms [4]. Studies have demonstrated that the HDR rate is lower than the NHEJ rate in the liver, limiting the therapy of diseases requiring gene editing by HDR [25]. Thankfully, the latest research has revealed the Fanconi anemia pathway to make CRISPR/Cas9 work better in almost all cells [73]. For liver diseases requiring a high percentage of gene correction such as liver cancer and HBV, it is still unknown whether CRISPR can restore protein levels to a therapeutic threshold so increasing HDR efficiency is an essential task [25]. Great efforts have been made to address these impending issues, and the ongoing progress is encouraging, but much more is needed to fully realize the medical potential of CRISPR/Cas9.
Additionally, there are also some limitations such as the specific delivery to intended target cells and the limitation of immunogenicity of gene modifiers and its delivery agent [3]. Viral vectors, which have been widely used in other gene therapy applications, may be employed as an efficient delivery mechanism for gene modifiers. Besides, immune stimulation may result from either the gene modifier itself or from the mechanism of delivery. It is important to assess immune activation as it may diminish efficacy following repeat administrations and cause toxicity in either cases. Immune stimulation has significantly hampered gene therapies in the past [3]. But surprisingly, the latest research found that bacteriophages are equipped with “anti- CRISPR” molecules. When the number of anti CRISPR bacteriophages exceeds the “critical point”, it can cooperate to overcome CRISPR/Cas immunity [74]. This discovery is a key breakthrough and of great significance to the clinical application of gene editing. Although the application of CRISPR/Cas9 is far-reaching, not all liver diseases are currently amenable to this type of treatment. Reversal of certain hepatic disorders, such as alcohol-induced cirrhosis, is not feasible at present [3]. But we believe that the rapid pace of technical improvements and the development of new applications will undoubtedly make the CRISPR/Cas9 system an integral part of liver research in the future.
Future Expectations and Conclusions
Initially, CRISPR/Cas9, the bacterial defense system, had been used by microbiologists to understand bacterial immunity. Over the past five years, however, researchers have turned CRISPR/Cas9 into a powerful tool for biological research. The CRISPR/Cas9 genome editing system has accelerated scientific research. With further attempting to combine CRISPR/Cas9 system and the latest technology such as single-cell sequencing and single base editing, the treatment of liver diseases will be upgraded to a new level.
As a powerful genome-editing tool, the CRISPR/Cas9 system has been quickly developed into a large-scale function-based screening strategy in mammalian cells [75]. Researchers established Cas9/sgRNA screens as a powerful tool for systematic genetic analysis in mammalian cells [76]. For further research, the CRISPR screening technique can be used to identify the target gene efficiently, while single-cell sequencing can allow us to dig into the single-cell level. As we all know, even in the same kind of tumor, such as HCC, the heterogeneity of the cells in-focus areas of the same patient or among different patients can be very large. In the future, we may be able to combine the single-cell sequencing for cell detection and CRISPR for high-throughput screening of genes to treat liver diseases, which is in line with the idea of “accurate medicine”.
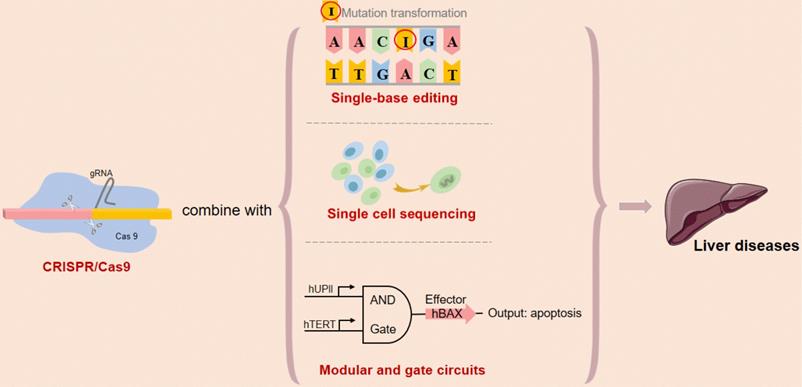
Recent works in mammalian synthetic biology have increased the applicability of synthetic logic circuits for human cancer therapy. In our previous study, we have constructed the modular AND gate circuits based on CRISPR/Cas9 system and validated novel genetic circuits that selectively and robustly mediate gene expression in human cancer cells, providing a safe, controllable and specific intelligent gene-editing system [77]. Although there is no precedent for the genetic circuit as a means of improving or treating liver disease currently, our researchers will strive to explore later. It would be of great interest to extend these circuits to clinical research on liver diseases when the challenges for in vivo DNA delivery are overcome (Figure 5).
Future expectations of type II CRISPR/Cas9 system. Abbreviations; CAR-T: chimeric antigen receptor T-cell.
Currently, the safest and most effective biological vector for DNA delivery is AAV. In recent years, AAV-based gene therapy drugs have been approved by the US FDA for pre-clinical trials. The All-in-one AAV-Cas9 system recently developed by our group can effectively regulate lipid metabolism in the liver of mice by targeting the Apao1 gene [78]. This provides a potentially useful vector to Cas9-based gene therapy for liver diseases.
To sum up, this review has discussed CRISPR/Cas9 and highlighted its advantages and disadvantages for applications to the treatment of liver diseases. Several studies have already demonstrated that these technologies may be used in a tissue- and patient-specific manner to disable, augment and correct gene function. Ultimately gene modifiers and cell-based therapies will increase the number of disorders that may be permanently corrected and alter how inborn liver diseases and viral hepatic infections are treated. These technologies present a novel, versatile tool for the treatment and cure of many hepatic illnesses. In the future, we even hope to achieve "rehabilitation" of cells, which is the transformation of tumor cells into normal cells through CRISPR/Cas9, and it will be a technology sweeping the world. When humans begin to modify their own source code of life, we know that a new era has come.
Highlights
- Liver diseases are one of the leading causes of mortality in the world and there is an urgent need for effective and safe treatment.
- CRISPR/Cas9 system has driven progress, improved our understanding of liver diseases and has the potential to revolutionize clinical treatment.
- The applications of CRISPR/Cas9 systems are rapidly expanding and limitations are being overcome.
- CRISPR/Cas9 can be readily applied to make basic science discoveries, and also offers exciting prospects for the treatment of liver diseases.
Abbreviations
AAV: adeno-associated virus; AFP: α-fetoprotein; ASPH: aspartate β-hydroxylase; Bax: Bcl2-associated X; Bcl2: B-cell lymphoma 2; cccDNA: covalently closed circular DNA; CDC42: cell division cycle 42; CDKs: cyclin-dependent kinases; CHB: chronic hepatitis B; CRISPR/Cas9: clustered regularly interspaced short palindromic repeats/CRISPR-associated proteins 9; crRNA: CRISPR-derived RNA; CXCR4: CXC chemokine receptor 4; CXCL12: chemokine stromal cell derived factor-1; dCas9: dead Cas9; DSBs: double strand breaks; dsDNA: double stranded DNA; ECM: extracellular matrix; eEF2: eukaryotic elongation factor 2; EMT: epithelial-mesenchymal transition; Fah: fumarylacetoacetate hydrolase; FnCas9: francisella novicida Cas9; H3K9me2: histone H3 lysine 9 di-methylation; HBsAg: hepatitis B surface antigen; HBV: hepatitis B virus; HBV-Tg: HBV-transgenic; HBx: HBV X protein; HCC: hepatocellular carcinoma; HCV: hepatitis C virus; HDR: homology-directed repair; Hpd: hydroxyphenylpyruvate dioxigenase; HTI: hereditary tyrosinemia type I; IL-6: interleukin-6; IQGAP1: IQ motif containing GTPase activating protein 1; KO: knockout; LncRNA: long-chain non-coding RNA; MiR-3188: microRNA-3188; NASH: nonalcoholic steatohepatitis; NCOA5: nuclear receptor coactivator 5; NHEJ: non-homologous end joining; NTCP: sodium taurocholate co-transporting polypeptide; PAM: protospacer-associated motif; pgRNA: pregenome RNA; PI3K: phosphatidylinositol-3-kinase; POLK: polymerase κ; PTEN: phosphatase and tensin homolog deleted on chromosome ten; rcDNA: relaxed circular DNA; REPAIR: RNA editing for programmable A-to-I replacement; RNP: ribonucleoprotein; SaCas9: staphylococcus aureus Cas9; sgRNA: small guide RNA; SpCas9: streptococcus pyogenes Cas9; ssDNA: single strand DNA; STAT3: signal transducer and activator of transcription 3; TALENs: transcription activator-like effector nucleases; tracrRNA: trans-activating crRNA; TSS: transcription start site; ZFNs: zinc finger nucleases; ZHX2: zinc fingers and homeoboxes protein 2.
Acknowledgements
Authors' Contributions
Tao Xu, Li Li, Yu-chen Liu and Wei Cao collected the related literature and drafted the manuscript. Jia-si Chen, Ying Liu and Huan Zhou participated in the design of the review and drafted the manuscript. Liang-yun Li and Hong Zhou revised and edited the manuscript. Xiao-ming Meng, Cheng Huang, Lei Zhang, Jun Li and Huan Zhou supervised the review process. All authors have read and approved the final manuscript.
Grant support
This project was supported by the National Natural Science Foundation of China (Nos. 81700522, 81602344), the fund of Anhui medical university doctoral start research (No.0601067101), Anhui Provincial Natural Science Foundation (1704a0802161), Soft science project of Anhui provincial science and technology department (1607a0202062), Natural Science Foundation of Anhui Province (1808085MH235), National Natural Science Foundation of China (81773257), Natural Science Foundation of Guangdong (2018B030306023), Key Special Project of Translational Medicine of Bengbu Medical University(BYTM2019021), the General Medicine Clinical Scientific Research Project of Anhui Provincial Health Committee (2016QK090), and the training objects of excellent young teachers in the 512 talent plan of Bengbu Medical College.
Competing Interests
The authors have declared that no competing interest exists.
References
1. Cong L, Ran FA, Cox D, Lin S, Barretto R, Habib N. et al. Multiplex genome engineering using CRISPR/Cas systems. Science. 2013;339:819-23
2. Ruiz de Galarreta M, Lujambio A. Therapeutic editing of hepatocyte genome in vivo. J Hepatol. 2017;67:818-28
3. Nicholson SA, Moyo B, Arbuthnot PB. Progress and prospects of engineered sequence-specific DNA modulating technologies for the management of liver diseases. World J Hepatol. 2015;7:859-73
4. Zhang JH, Adikaram P, Pandey M, Genis A, Simonds WF. Optimization of genome editing through CRISPR-Cas9 engineering. Bioengineered. 2016;7:166-74
5. Mali P, Yang L, Esvelt KM, Aach J, Guell M, DiCarlo JE. et al. RNA-guided human genome engineering via Cas9. Science. 2013;339:823-6
6. Zhang Z, Zhang YH, Zhang T. [Research progress of liver disease treatment by Zhuang medicines]. Zhongguo Zhong Yao Za Zhi. 2018;43:2224-9
7. Hernandez-Aquino E, Muriel P. Beneficial effects of naringenin in liver diseases: Molecular mechanisms. World J Gastroenterol. 2018;24:1679-707
8. Yu Y, Cai J, She Z, Li H. Insights into the Epidemiology, Pathogenesis, and Therapeutics of Nonalcoholic Fatty Liver Diseases. Adv Sci (Weinh). 2019;6:1801585
9. Ishino Y, Shinagawa H, Makino K, Amemura M, Nakata A. Nucleotide sequence of the iap gene, responsible for alkaline phosphatase isozyme conversion in Escherichia coli, and identification of the gene product. J Bacteriol. 1987;169:5429-33
10. Mojica FJ, Diez-Villasenor C, Soria E, Juez G. Biological significance of a family of regularly spaced repeats in the genomes of Archaea, Bacteria and mitochondria. Mol Microbiol. 2000;36:244-6
11. Jansen R, Embden JD, Gaastra W, Schouls LM. Identification of genes that are associated with DNA repeats in prokaryotes. Mol Microbiol. 2002;43:1565-75
12. Barrangou R, Fremaux C, Deveau H, Richards M, Boyaval P, Moineau S. et al. CRISPR provides acquired resistance against viruses in prokaryotes. Science. 2007;315:1709-12
13. Makarova KS, Haft DH, Barrangou R, Brouns SJ, Charpentier E, Horvath P. et al. Evolution and classification of the CRISPR-Cas systems. Nat Rev Microbiol. 2011;9:467-77
14. Makarova KS, Wolf YI, Alkhnbashi OS, Costa F, Shah SA, Saunders SJ. et al. An updated evolutionary classification of CRISPR-Cas systems. Nat Rev Microbiol. 2015;13:722-36
15. Deltcheva E, Chylinski K, Sharma CM, Gonzales K, Chao Y, Pirzada ZA. et al. CRISPR RNA maturation by trans-encoded small RNA and host factor RNase III. Nature. 2011;471:602-7
16. Czajkowski J, Tozzi V, Strannegard O. Suppression of HIV replication in vitro by fusidic acid parallels cell toxicity. J Antimicrob Chemother. 1989;23:155-6
17. Gasiunas G, Barrangou R, Horvath P, Siksnys V. Cas9-crRNA ribonucleoprotein complex mediates specific DNA cleavage for adaptive immunity in bacteria. Proc Natl Acad Sci U S A. 2012;109:E2579-86
18. Higashijima Y, Hirano S, Nangaku M, Nureki O. Applications of the CRISPR-Cas9 system in kidney research. Kidney Int. 2017;92:324-35
19. Zhu S, Li W, Liu J, Chen CH, Liao Q, Xu P. et al. Genome-scale deletion screening of human long non-coding RNAs using a paired-guide RNA CRISPR-Cas9 library. Nat Biotechnol. 2016;34:1279-86
20. Cyranoski D. CRISPR gene-editing tested in a person for the first time. Nature. 2016;539:479
21. Cox DBT, Gootenberg JS, Abudayyeh OO, Franklin B, Kellner MJ, Joung J. et al. RNA editing with CRISPR-Cas13. Science. 2017;358:1019-27
22. Haapaniemi E, Botla S, Persson J, Schmierer B, Taipale J. CRISPR-Cas9 genome editing induces a p53-mediated DNA damage response. Nat Med. 2018;24:927-30
23. Ihry RJ, Worringer KA, Salick MR, Frias E, Ho D, Theriault K. et al. p53 inhibits CRISPR-Cas9 engineering in human pluripotent stem cells. Nat Med. 2018;24:939-46
24. Koblan LW, Doman JL, Wilson C, Levy JM, Tay T, Newby GA. et al. Improving cytidine and adenine base editors by expression optimization and ancestral reconstruction. Nat Biotechnol. 2018;36:843-6
25. Song CQ, Xue W. CRISPR-Cas-related technologies in basic and translational liver research. Nat Rev Gastroenterol Hepatol. 2018;15:251-2
26. Garneau JE, Dupuis ME, Villion M, Romero DA, Barrangou R, Boyaval P. et al. The CRISPR/Cas bacterial immune system cleaves bacteriophage and plasmid DNA. Nature. 2010;468:67-71
27. Hsu PD, Lander ES, Zhang F. Development and applications of CRISPR-Cas9 for genome engineering. Cell. 2014;157:1262-78
28. Shou J, Li J, Liu Y, Wu Q. Precise and Predictable CRISPR Chromosomal Rearrangements Reveal Principles of Cas9-Mediated Nucleotide Insertion. Mol Cell. 2018;71:498-509 e4
29. Lin SR, Yang HC, Kuo YT, Liu CJ, Yang TY, Sung KC. et al. The CRISPR/Cas9 System Facilitates Clearance of the Intrahepatic HBV Templates In vivo. Mol Ther Nucleic Acids. 2014;3:e186
30. Huang P, Tong D, Sun J, Li Q, Zhang F. Generation and characterization of a human oral squamous carcinoma cell line SCC-9 with CRISPR/Cas9-mediated deletion of the p75 neurotrophin receptor. Arch Oral Biol. 2017;82:223-32
31. Ohno M, Otsuka M, Kishikawa T, Yoshikawa T, Takata A, Koike K. Novel therapeutic approaches for hepatitis B virus covalently closed circular DNA. World J Gastroenterol. 2015;21:7084-8
32. Sarin SK, Kumar M, Lau GK, Abbas Z, Chan HL, Chen CJ. et al. Asian-Pacific clinical practice guidelines on the management of hepatitis B: a 2015 update. Hepatol Int. 2016;10:1-98
33. Soppe JA, Lebbink RJ. Antiviral Goes Viral: Harnessing CRISPR/Cas9 to Combat Viruses in Humans. Trends Microbiol. 2017;25:833-50
34. Dong C, Qu L, Wang H, Wei L, Dong Y, Xiong S. Targeting hepatitis B virus cccDNA by CRISPR/Cas9 nuclease efficiently inhibits viral replication. Antiviral Res. 2015;118:110-7
35. Qi Y, Gao Z, Xu G, Peng B, Liu C, Yan H. et al. DNA Polymerase kappa Is a Key Cellular Factor for the Formation of Covalently Closed Circular DNA of Hepatitis B Virus. PLoS Pathog. 2016;12:e1005893
36. Scott T, Moyo B, Nicholson S, Maepa MB, Watashi K, Ely A. et al. ssAAVs containing cassettes encoding SaCas9 and guides targeting hepatitis B virus inactivate replication of the virus in cultured cells. Sci Rep. 2017;7:7401
37. Ren Q, Li C, Yuan P, Cai C, Zhang L, Luo GG. et al. A Dual-reporter system for real-time monitoring and high-throughput CRISPR/Cas9 library screening of the hepatitis C virus. Sci Rep. 2015;5:8865
38. Price AA, Sampson TR, Ratner HK, Grakoui A, Weiss DS. Cas9-mediated targeting of viral RNA in eukaryotic cells. Proc Natl Acad Sci U S A. 2015;112:6164-9
39. Yan H, Zhong G, Xu G, He W, Jing Z, Gao Z. et al. Sodium taurocholate cotransporting polypeptide is a functional receptor for human hepatitis B and D virus. Elife. 2012 3
40. Iwagami Y, Huang CK, Olsen MJ, Thomas JM, Jang G, Kim M. et al. Aspartate beta-hydroxylase modulates cellular senescence through glycogen synthase kinase 3beta in hepatocellular carcinoma. Hepatology. 2016;63:1213-26
41. Scheer S, Zaph C. The Lysine Methyltransferase G9a in Immune Cell Differentiation and Function. Front Immunol. 2017;8:429
42. Qin J, Li Q, Zeng Z, Wu P, Jiang Y, Luo T. et al. Increased expression of G9A contributes to carcinogenesis and indicates poor prognosis in hepatocellular carcinoma. Oncol Lett. 2018;15:9757-65
43. Wei L, Chiu DK, Tsang FH, Law CT, Cheng CL, Au SL. et al. Histone methyltransferase G9a promotes liver cancer development by epigenetic silencing of tumor suppressor gene RARRES3. J Hepatol. 2017;67:758-69
44. Pott LL, Hagemann S, Reis H, Lorenz K, Bracht T, Herold T. et al. Eukaryotic elongation factor 2 is a prognostic marker and its kinase a potential therapeutic target in HCC. Oncotarget. 2017;8:11950-62
45. Shi N, Chen X, Liu R, Wang D, Su M, Wang Q. et al. Eukaryotic elongation factors 2 promotes tumor cell proliferation and correlates with poor prognosis in ovarian cancer. Tissue Cell. 2018;53:53-60
46. He J, Zhang W, Li A, Chen F, Luo R. Knockout of NCOA5 impairs proliferation and migration of hepatocellular carcinoma cells by suppressing epithelial-to-mesenchymal transition. Biochem Biophys Res Commun. 2018;500:177-83
47. Jeng KS, Jeng CJ, Jeng WJ, Chang CF, Sheen IS. Role of C-X-C chemokine ligand 12/C-X-C chemokine receptor 4 in the progression of hepatocellular carcinoma. Oncol Lett. 2017;14:1905-10
48. Wang X, Zhang W, Ding Y, Guo X, Yuan Y, Li D. CRISPR/Cas9-mediated genome engineering of CXCR4 decreases the malignancy of hepatocellular carcinoma cells in vitro and in vivo. Oncol Rep. 2017;37:3565-71
49. Wang C, Jin H, Gao D, Wang L, Evers B, Xue Z. et al. A CRISPR screen identifies CDK7 as a therapeutic target in hepatocellular carcinoma. Cell Res. 2018;28:690-2
50. Aponte JL, Sega GA, Hauser LJ, Dhar MS, Withrow CM, Carpenter DA. et al. Point mutations in the murine fumarylacetoacetate hydrolase gene: Animal models for the human genetic disorder hereditary tyrosinemia type 1. Proc Natl Acad Sci U S A. 2001;98:641-5
51. Pankowicz FP, Barzi M, Legras X, Hubert L, Mi T, Tomolonis JA. et al. Reprogramming metabolic pathways in vivo with CRISPR/Cas9 genome editing to treat hereditary tyrosinaemia. Nat Commun. 2016;7:12642
52. Giguere Y, Berthier MT. Newborn Screening for Hereditary Tyrosinemia Type I in Quebec: Update. Adv Exp Med Biol. 2017;959:139-46
53. Paulk NK, Wursthorn K, Wang Z, Finegold MJ, Kay MA, Grompe M. Adeno-associated virus gene repair corrects a mouse model of hereditary tyrosinemia in vivo. Hepatology. 2010;51:1200-8
54. Yin H, Xue W, Chen S, Bogorad RL, Benedetti E, Grompe M. et al. Genome editing with Cas9 in adult mice corrects a disease mutation and phenotype. Nat Biotechnol. 2014;32:551-3
55. Yin H, Song CQ, Dorkin JR, Zhu LJ, Li Y, Wu Q. et al. Therapeutic genome editing by combined viral and non-viral delivery of CRISPR system components in vivo. Nat Biotechnol. 2016;34:328-33
56. Song J, Zhang X, Ge Q, Yuan C, Chu L, Liang HF. et al. CRISPR/Cas9-mediated knockout of HBsAg inhibits proliferation and tumorigenicity of HBV-positive hepatocellular carcinoma cells. J Cell Biochem. 2018;119:8419-31
57. Zhou SJ, Deng YL, Liang HF, Jaoude JC, Liu FY. Hepatitis B virus X protein promotes CREB-mediated activation of miR-3188 and Notch signaling in hepatocellular carcinoma. Cell Death Differ. 2017;24:1577-87
58. Tang H, Oishi N, Kaneko S, Murakami S. Molecular functions and biological roles of hepatitis B virus x protein. Cancer Sci. 2006;97:977-83
59. Martin-Vilchez S, Lara-Pezzi E, Trapero-Marugan M, Moreno-Otero R, Sanz-Cameno P. The molecular and pathophysiological implications of hepatitis B X antigen in chronic hepatitis B virus infection. Rev Med Virol. 2011;21:315-29
60. Zhang Y, Li D, Feng F, An L, Hui F, Dang D. et al. Progressive and Prognosis Value of Notch Receptors and Ligands in Hepatocellular Carcinoma: A Systematic Review and Meta-analysis. Sci Rep. 2017;7:14809
61. Xu Y, Qi Y, Luo J, Yang J, Xie Q, Deng C. et al. Hepatitis B Virus X Protein Stimulates Proliferation, Wound Closure and Inhibits Apoptosis of HuH-7 Cells via CDC42. Int J Mol Sci. 2017 18
62. Stengel K, Zheng Y. Cdc42 in oncogenic transformation, invasion, and tumorigenesis. Cell Signal. 2011;23:1415-23
63. Yeo NC, Chavez A, Lance-Byrne A, Chan Y, Menn D, Milanova D. et al. An enhanced CRISPR repressor for targeted mammalian gene regulation. Nat Methods. 2018;15:611-6
64. Kosicki M, Tomberg K, Bradley A. Repair of double-strand breaks induced by CRISPR-Cas9 leads to large deletions and complex rearrangements. Nat Biotechnol. 2018;36:765-71
65. Liu Y, Qi X, Zeng Z, Wang L, Wang J, Zhang T. et al. CRISPR/Cas9-mediated p53 and Pten dual mutation accelerates hepatocarcinogenesis in adult hepatitis B virus transgenic mice. Sci Rep. 2017;7:2796
66. Xue W, Chen S, Yin H, Tammela T, Papagiannakopoulos T, Joshi NS. et al. CRISPR-mediated direct mutation of cancer genes in the mouse liver. Nature. 2014;514:380-4
67. Rauch BJ, Silvis MR, Hultquist JF, Waters CS, McGregor MJ, Krogan NJ. et al. Inhibition of CRISPR-Cas9 with Bacteriophage Proteins. Cell. 2017;168:150-8 e10
68. Pawluk A, Amrani N, Zhang Y, Garcia B, Hidalgo-Reyes Y, Lee J. et al. Naturally Occurring Off-Switches for CRISPR-Cas9. Cell. 2016;167:1829-38 e9
69. Dow LE, Fisher J, O'Rourke KP, Muley A, Kastenhuber ER, Livshits G. et al. Inducible in vivo genome editing with CRISPR-Cas9. Nat Biotechnol. 2015;33:390-4
70. Nihongaki Y, Kawano F, Nakajima T, Sato M. Photoactivatable CRISPR-Cas9 for optogenetic genome editing. Nat Biotechnol. 2015;33:755-60
71. Lin F, Dong L, Wang W, Liu Y, Huang W, Cai Z. An Efficient Light-Inducible P53 Expression System for Inhibiting Proliferation of Bladder Cancer Cell. Int J Biol Sci. 2016;12:1273-8
72. Hu JH, Miller SM, Geurts MH, Tang W, Chen L, Sun N. et al. Evolved Cas9 variants with broad PAM compatibility and high DNA specificity. Nature. 2018;556:57-63
73. Richardson CD, Kazane KR, Feng SJ, Zelin E, Bray NL, Schafer AJ. et al. CRISPR-Cas9 genome editing in human cells occurs via the Fanconi anemia pathway. Nat Genet. 2018;50:1132-9
74. Landsberger M, Gandon S, Meaden S, Rollie C, Chevallereau A, Chabas H. et al. Anti-CRISPR Phages Cooperate to Overcome CRISPR-Cas Immunity. Cell. 2018;174:908-16 e12
75. Peng J, Zhou Y, Zhu S, Wei W. High-throughput screens in mammalian cells using the CRISPR-Cas9 system. FEBS J. 2015;282:2089-96
76. Wang T, Wei JJ, Sabatini DM, Lander ES. Genetic screens in human cells using the CRISPR-Cas9 system. Science. 2014;343:80-4
77. Liu Y, Zeng Y, Liu L, Zhuang C, Fu X, Huang W. et al. Synthesizing AND gate genetic circuits based on CRISPR-Cas9 for identification of bladder cancer cells. Nat Commun. 2014;5:5393
78. Zhan H, Zhou Q, Gao Q, Li J, Huang W, Liu Y. Multiplexed promoterless gene expression with CRISPReader. Genome Biol. 2019;20:113
Author contact
Corresponding authors: Jun Li, PhD. School of Pharmacy, Anhui Medical University, 81 Meishan Road, Hefei, Anhui Province, 230032, China. Tel./Fax: +86 551 65161001. E-mail: ljedu.cn; Huan Zhou, PhD, MD. National Drug Clinical Trial Institution, the First Affiliated Hospital of Bengbu Medical College, Anhui Province, China. E-mail: zhouhuanbestcom.

 Global reach, higher impact
Global reach, higher impact