Impact Factor ISSN: 1449-2288
Int J Biol Sci 2021; 17(1):89-96. doi:10.7150/ijbs.52619 This issue Cite
Review
The Regulatory Effect of SIRT1 on Extracellular Microenvironment Remodeling
College of Basic Medical Science, Institute of Translational Medicine, Key Laboratory of Medical Cell Biology, Ministry of Education, Key Laboratory of Liaoning Province, China Medical University, Shenyang, Liaoning Province, P.R. China, 110122.
#These authors contributed equally to this work.
Received 2020-8-30; Accepted 2020-10-22; Published 2021-1-1
Abstract
The sirtuins family is well known by its unique nicotinamide adenine dinucleotide (NAD+)-dependent deacetylase function. The most-investigated member of the family, Sirtuin 1 (SIRT1), accounts for deacetylating a broad range of transcription factors and coregulators, such as p53, the Forkhead box O (FOXO), and so on. It serves as a pivotal regulator in various intracellular biological processes, including energy metabolism, DNA damage response, genome stability maintenance and tumorigenesis. Although the most attention has been focused on its intracellular functions, the regulatory effect on extracellular microenvironment remodeling of SIRT1 has been recognized by researchers recently. SIRT1 can regulate cell secretion process and participate in glucose metabolism, neuroendocrine function, inflammation and tumorigenesis. Here, we review the advances in the understanding of SIRT1 on remodeling the extracellular microenvironment, which may provide new ideas for pathogenesis investigation and guidance for clinical treatment.
Keywords: SIRT1, microenvironment remodeling, cell secretion, endocrine, inflammation, tumorigenesis
Introduction
Extracellular microenvironment has been considered as a crucial element that influences cellular proliferation and metabolism [1, 2]. It mediates the communication and interaction between neighboring cells, different tissue cells, and even distinct organs for the insurance of normal functions [3, 4]. Meanwhile, the extracellular microenvironment can be reformed by intracellular proteins through regulating the processes and cargos of cell secretion [5]. As the significant nicotinamide adenine dinucleotide-dependent protein deacetylase, sirtuin 1 (SIRT1) is extensively involved in various cellular processes and metabolism [6, 7]. Our previous work revealed that SIRT1 played important roles in genome stability maintenance [8], DNA damage response [9] and autophagy [10]. We further realize that SIRT1 not only affects intracellular homeostasis, but also participates in the extracellular microenvironment remodeling. In this study, we review the related literatures focusing on SIRT1 function as a cell secretion regulator, hoping to find new ways and targets for future research and clinical treatment.
SIRT1 modulates glucose metabolism through cell secretion
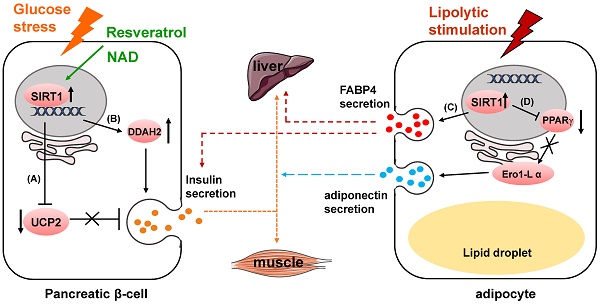
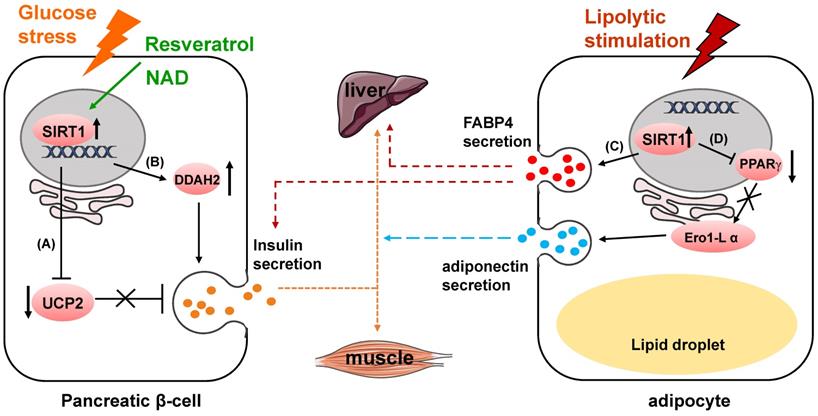
SIRT1 has got long-term attention and well known for playing a pivotal role in glucose homeostasis and Type 2 Diabetes [11, 12]. One of its functions is to regulate insulin secretion. In pancreatic beta cells, SIRT1 promotes insulin secretion in response to glucose stress by suppressing the expression of uncoupling protein 2 (UCP2) [13-15] (Fig. 1A). As a negative regulator of insulin, increased expression of UCP2 in the pancreatic β-cells results in the decrease of glucose-stimulated insulin secretion (GSIS) leading to pancreatic β-cell dysfunction and development of type-II diabetes [16, 17]. UCP2 is shown to regulate glucose metabolism and insulin secretion through many biological activities, including reducing NADH levels, decreasing ATP production, weakening mitochondrial membrane potential and suppressing generation of superoxide [18-20]. SIRT1 can also enhance insulin secretion of pancreatic β-cells through the hydrolase dimethylarginine dimethylaminohydrolase 2 (DDAH2)/secretagogin pathway [21] (Fig. 1B). SIRT1 boosts DDAH2 expression on the transcriptional level by activating promoter of 5' deletion constructs. The overexpression of DDAH2 induces the upregulation of secretagogin, an EF-hand Ca2+-binding protein which is involved in vesicle secretion [22]. Besides, DDAH2 was reported to have directly interaction with secretagogin [23].
SIRT1 modulates glucose metabolism through regulating cell secretion. (A) SIRT1 suppresses UCP2 expression to upregulate insulin secretion. (B) SIRT1 enhances insulin secretion through DDAH2/secretagogin pathway. (C) SIRT1 increases FABP4 secretion to regulate hepatic glucose production and glucose-stimulated insulin secretion. (D) SIRT1 inhibits PPARγ activity to downregulate Ero1-L α expression, and suppresses adiponectin secretion.
Furthermore, SIRT1 itself can be regulated by in the process of insulin secretion [24]. MicroRNA mir-9 targets and reduces SIRT1 protein level during glucose-dependent insulin secretion. Some protein and compounds can also influence insulin secretion through regulating SIRT1 activity. For instance, Wallerian degeneration slow (WldS), a fusion protein with NAD biosynthesis activity, can increase NAD level, which leads to the enhanced SIRT1 activity to downregulate UCP2 [25, 26]. Resveratrol, currently a most potent natural compound SIRT1 activator, enhances insulin secretion in human islets in response to both glucose and high fat diet [27, 28]. Additionally, resveratrol improves insulin induced NO secretion partly through activating SIRT1 in endothelial cells [29]. This effect might be related to improvement of endothelial cell function in animal models and in humans [30-32].
In mouse models, β-cell specific SIRT1-overexpression transgenic mice have presented enhanced glucose-stimulated insulin secretion and improved glucose tolerance at age of 3 and 8 months [13]. However, the phenotype completely vanishes when the transgenic mice reach 18 to 24 months [33]. Meanwhile, in these mouse models, decreased SIRT1 expression impairs glucose sensing as well as insulin secretion [34, 35].
In mature adipocytes, SIRT1 negatively regulates adiponectin secretion through inhibiting peroxisome proliferator activated receptor γ (PPARγ) activity [36] (Fig. 1D). Suppression of SIRT1 or activation of PPARγ upregulates the protein level of endoplasmic reticulum oxidoreductase 1 α (Ero1-L α) and stimulates secretion of high-molecular-weight adiponectin. The secreted complexes of adiponectin was reported to sensitize liver and muscle cells to insulin in response to various metabolic states [37, 38]. Secretion of fatty acid binding protein 4 (FABP4), a lipid carrier protein, from white adipose tissue is dependent on SIRT1 in response to lipolytic stimulation [39]. The secretion process also requires early components of autophagy such as beclin-1. The secreted FABP4 in circulating system is a signal molecule transmitted from adipose tissue to liver to augment the production of hepatic glucose [40] (Fig. 1C). Circulating FABP4 level is positively correlated with glucose-stimulated insulin secretion [41].
SIRT1 regulates lipid metabolism through cell secretion
Various researches have proved that SIRT1 plays a role in regulating lipid metabolism [42]. Among those reports, SIRT1 was found to be a protective factor in the development of atherosclerosis [43-45]. SIRT1 inhibits secretion of thrombosis promoting factors, von Willebrand factor (vWF) and P-selectin, from vascular endothelial cells, thus preventing thrombosis formation [46]. This effect is probably relevant to the regulation of autophagy through SIRT1/FOXO1 pathway. According to Miranda's work, SIRT1 activation reduces hepatic secretion of a serine protease, proprotein convertase subtilisin/kexin type 9 (PCSK9). The secreted mediates lysosomal degradation of hepatic low-density lipoprotein receptor (LDLR) and prevents its internalizing recycle to cell surface [47]. PCSK9 accumulation increases LDLR protein degradation and then enhances LDL-cholesterol plasma clearance, leading to decreased plaque formation [48, 49]. These results are consistent with reduced levels of blood cholesterol and adipokines in SIRT1 transgenic mice [50-52].
SIRT1 contributes to neuroendocrine secretion
In clinical work, Diabetes mellitus is often found to be coexisted with hypothyroidism [53]. This might be partially related to the upregulation of advanced glycation end products (AGEs), high glucose induce advanced glycation end products receptor (RAGE) and the inactivation of SIRT1/nuclear factor erythroid 2-related factor (NRF2) pathway [54-56] (Fig. 2B). High glucose and AGEs induce upregulation of RAGE, downregulation of SIRT1 and NRF2, and decrease of proteins related to thyroid hormone (TH) secretion, thus finally resulting in decreased TH secretion and circulating TH deficiency (Fig. 2C). Sayaka et al reveals that in pituitary gland, SIRT1 multiples exocytosis of thyroid stimulating hormone (TSH) containing granules by deacetylating phosphatidylinositol-4-phosphate 5-kinase γ (PIP5Kγ) [57], which mainly mediates the large dense-core vesicle fusion [58] (Fig. 2C). Consistently, high acetylated PIP5Kγ and decreased thyroid hormone in the plasma were observed in SIRT1 knock out mice [57, 59]. Hormone disorder displayed in neuron-specific SIRT1 knock out mice showed that more work still needs to be done to address how SIRT1 modulates neural cell secretion and somatotropic signaling [60].
SIRT1 regulates neuroendocrine secretion. (A) SIRT1 upregulates S100β release. (B) AGEs bind to RAGE and inactivates SIRT1/NRF2 pathway, and decreases TH secretion. (C) SIRT1 deacetylates PIP5Kγ to increase TSH release, and stimulates TH secretion.
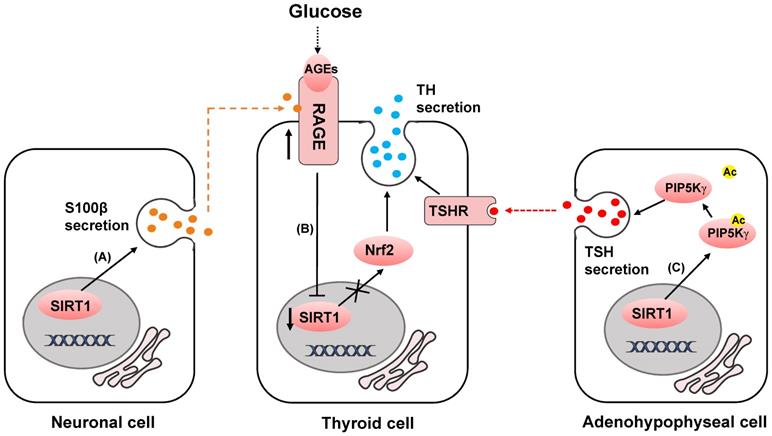
In previous studies, SIRT1 was found to be predominantly expressed in neurons and highly involved in neurodegenerative diseases [61, 62]. SIRT1 is revealed to be neuroprotective in amyloid-β-induced ROS production, DNA damage and oxidative modifications [63, 64]. Qin et al identify in C6 rat glioma cells that enhanced expression of SIRT1 can upregulate the release of a calcium-binding protein S100β [65]. The extracellular S100β functions as cytokines with both neurotrophic and neurotoxic effects and affects the activity of several cell types, such as neurons [66-68], astrocytes [69, 70], and microglia [71, 72], through the surface receptor RAGE [73-76] (Fig. 2A).
Moreover, SIRT1 positively affects the NAD biosynthesis in the hypothalamus by distinctly regulating the release of Nicotinamide phosphoribosyl transferase (NAMPT) in adipose tissue. In this process, SIRT1 predisposes NAMPT to be secreted to the extracellular enzyme pool by deacetylating it at lysine 53 [77].
SIRT1 and inflammatory microenvironment
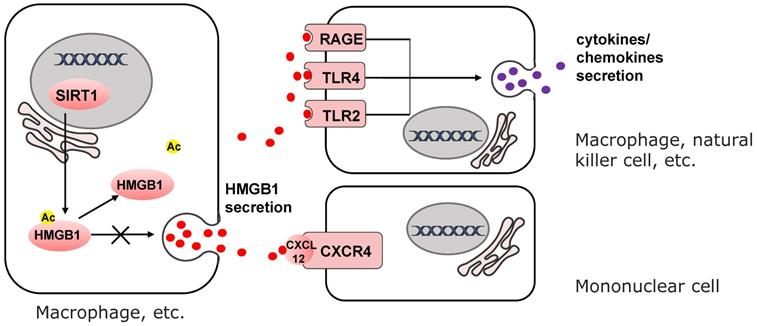
High mobility group box 1 (HMGB1) was identified as a structural protein of chromatin which functioning in transcription [78, 79]. But when secreted into the extracellular microenvironment, it induces acute and chronic inflammation as a proinflammatory cytokine [80-83]. HMGB1 mediates the recruitment of mononuclear cells [84], the release of cytokines or chemokines [85-87], and the activation of effector T cells and suppression of regulatory T cells[88, 89] (Fig. 3). The exocytosis of HMGB1 is highly dependent on its level of acetylation [90]. SIRT1 attenuates the exocytosis of HMGB1 through deacetylation [91-93]. The secretion of HMGB1 into the extracellular microenvironment can be downregulated by upregulating the activity or expression of SIRT1 [91, 94]. This SIRT1-mediated mechanism has been described in Zeng's research and can prevent non-alcoholic fatty liver disease induced by high fat diet [95]. In response to the stimulation of H2O2, suppression of SIRT1 leads to the upregulation of HMGB1 released from hepatocytes [96]. Intriguingly, secretion of HMGB1 from kidney cells increases in early stage of hemorrhagic shock by downregulation of SIRT1 expression level [97]. In turn, downregulation of circulating HMGB1 by SIRT1 protects liver from ischemic injury [98]. This phenomenon portends that the secretion of HMGB1 changes in different organs and extracellular microenvironment dependent on SIRT1 activity.
SIRT1 mediates inflammatory microenvironment remodeling by deacetylating HMGB1 to reduce its secretion. Released HMGB1 forms complex with CXCL12 and CXCR4 to induce the recruitment of mononuclear cells. HMGB1 also binds to TLR2, TLR4 and RAGE to mediate cytokines or chemokines secretion of immune cells.
SIRT1 and tumor microenvironment
SIRT1 is proved to be highly involved in cancer because of its underlying functions in tumorigenesis [99-101], senescence [102, 103], immunity [104, 105] and inflammation [106]. Chronic lymphocytic leukemia creates its suitable microenvironment for survival through the release of aforementioned protein, HMGB1 [107]. Besides, the extracellular S100β, which can be regulated by SIRT1, participates in the recruitment and activity modulation of monocytes in tumor microenvironment [65, 108, 109]. A recent study exhibits that inhibition of SIRT1 by caveolin-1 in senescent fibroblasts promotes the secretion of interleukin 6 (IL-6) and stimulates tumor growth [110].
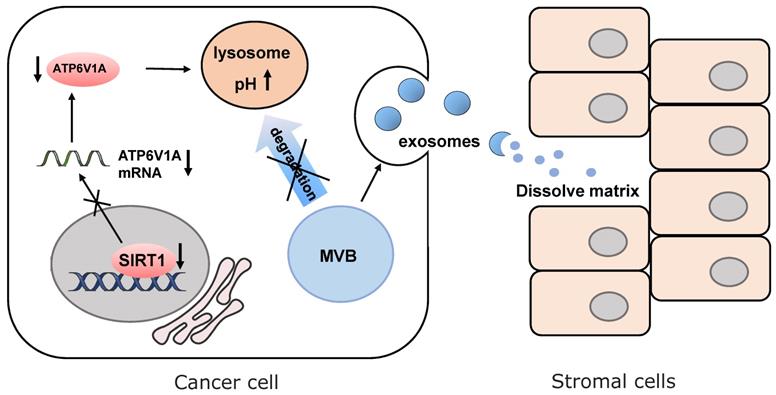
The latest progress discovers that SIRT1 is responsible for the change of microenvironment by regulating the secretion of exosomes [111] (Fig. 4). In breast cancer, loss of SIRT1 enhances the secretion of pro-tumorigenic exosomes and promotes cancer invasion [112]. In triple-negative breast cancer, down-regulating SIRT1 levels decreases the expression of ATPase H+ transporting V1 subunit A (ATP6V1A), a particular subunit of the vacuolar-type H+ ATPase (V-ATPase). It is responsible for acidification of lysosomes and degradation of protein. Disruption of the degradation process leads to the reduction of multi-vesicular bodies (MVBs) and the formation of larger MVBs. Finally, the imbalanced MVB formation promotes enhancing protein cargos released by exosomes [113, 114]. The released cargos dissolve the extracellular matrix of normal cells adjacent to tumor cells and destroy para cancer tissue structures. This SIRT1-dependent mechanism enables cancer cells to create the suitable extracellular microenvironment for the expansion of themselves. In senescent stromal cells, SIRT1-loss also causes impairment of lysosomes acidification and protein degradation. That is why senescent cells presumably prefer to release small extracellular vesicles into the tumor microenvironment, which enhances the aggressiveness and drug resistance of recipient cancer cells mediated by ATP binding cassette subfamily B member 4 (ABCB4) [115].
SIRT1 increases the expression of ATP6V1A to maintain pH level of lysosomes. Down-regulation of SIRT1 promotes MVBs formation and enhancing protein cargos released by exosomes to dissolve the cell matrix of tumor microenvironment.
Conclusions
Accumulating evidences suggest that SIRT1 has wide effects on regulating extracellular microenvironment through cell secretion. Changes of expression or activity of SIRT1 can result in functional variations of the neuroendocrine system, inflammatory and tumor microenvironment through driving proteins, cargos in exocytosis vesicles and exosomes secreting into extracellular microenvironment. The released proteins functions as enzymes, cytokines, neuroendocrine factors and ligands of cell surface receptors. These molecules regulate downstream target cells bringing about different effects. These variations are involved in glucose metabolism, TH/TSH secretion, lipid metabolism, inflammation in thrombosis, tumorigenesis and metastasis.
We should give more concern to the extracellular microenvironment remodeling function of SIRT1 in scientific researches and clinical treatment. Further research on new drug design may focus on its regulatory effect of extracellular mechanisms, targeting SIRT1 activity or the secreted downstream proteins [116, 117].
It is worth noting that the SIRT1 dependent HMGB1 secretion in kidney and liver shows different alterations in ischemic injury [97, 98]. This prompts that SIRT1-dependent extracellular microenvironment regulation may be variant in different organs and cause opposite results. Thus, in using drugs targeted for SIRT1 in clinical treatment or assessing the medical effects, the specific extracellular effects should be taken into consideration.
Abbreviations
ABCB4: ATP binding cassette subfamily B member 4; AGEs: advanced glycation end products; ATP6V1A: ATPase H+ transporting V1 subunit A; CXCL12: C-X-C motif chemokine ligand 12; CXCR4: C-X-C motif chemokine receptor 4; DDAH2: dimethylarginine dimethylaminohydrolase 2; Ero1-L α: endoplasmic reticulum oxidoreductase 1 alpha; FABP4: fatty acid binding protein 4; FOXO: Forkhead box O; GSIS: glucose-stimulated insulin secretion; HMGB1: High mobility group box 1; IL-6: interleukin 6; LDLR: low-density lipoprotein receptor; LPS: lipopolysaccharide; MVBs: multi-vesicular bodies; NAD: nicotinamide adenine dinucleotide; NAMPT: nicotinamide phosphoribosyl transferase; NRF2: nuclear factor erythroid 2-related factor; PCSK9: proprotein convertase subtilisin/kexin type 9; PIP5Kγ: phosphatidylinositol-4-phosphate 5-kinase γ; PPARγ: peroxisome proliferator activated receptor γ; RAGE: advanced glycation end products receptor; SIRT1: sirtuin 1; TH: thyroid hormone; TLR2: toll like receptor 2; TLR4: toll like receptor 4; TSH: thyroid stimulating hormone; UCP2: uncoupling protein 2; V-ATPase: vacuolar-type H+ ATPase; vWF: von Willebrand factor; WldS: Wallerian degeneration slow.
Acknowledgements
Author Contributions
XS and LC designed and conceived the general idea and context of this review. ZW, TZ and XS wrote and integrated all sections. QG, HX and WG contributed to the “SIRT1 and tumor microenvironment” section. FY, YG, XL and YF relevant references of this manuscript. All the authors read and approved the final manuscript.
Funding
This study was supported by grants from the National Key R&D Program of China (2016YFC1302400) and the Ministry of Education Innovation Team Development plan (IRT13101&17R107) to Liu Cao; the National Science Foundation of China to Liu Cao (81770001), Xiaoyu Song (31300963, 2018225083) and Tingting Zhou (81502438); the Science and Technology Research Project of Liaoning Provincial Education Department to Zhuo Wang (LQNK201747) and Hongde Xu (LK201627).
Competing Interests
The authors have declared that no competing interest exists.
References
1. Hynes RO. The extracellular matrix: not just pretty fibrils. Science. 2009;326:1216-9
2. Fane M, Weeraratna AT. How the ageing microenvironment influences tumour progression. Nat Rev Cancer. 2020;20:89-106
3. Ten Hacken E, Burger JA. Microenvironment interactions and B-cell receptor signaling in Chronic Lymphocytic Leukemia: Implications for disease pathogenesis and treatment. Biochim Biophys Acta. 2016;1863:401-13
4. Mavrogonatou E, Pratsinis H, Papadopoulou A, Karamanos NK, Kletsas D. Extracellular matrix alterations in senescent cells and their significance in tissue homeostasis. Matrix Biol. 2019;75-76:27-42
5. Baker BM, Trappmann B, Wang WY, Sakar MS, Kim IL, Shenoy VB. et al. Cell-mediated fibre recruitment drives extracellular matrix mechanosensing in engineered fibrillar microenvironments. Nat Mater. 2015;14:1262-8
6. Finkel T, Deng CX, Mostoslavsky R. Recent progress in the biology and physiology of sirtuins. Nature. 2009;460:587-91
7. Chang HC, Guarente L. SIRT1 and other sirtuins in metabolism. Trends Endocrinol Metab. 2014;25:138-45
8. Wang RH, Sengupta K, Li C, Kim HS, Cao L, Xiao C. et al. Impaired DNA damage response, genome instability, and tumorigenesis in SIRT1 mutant mice. Cancer Cell. 2008;14:312-23
9. Zhang W, Feng Y, Guo Q, Guo W, Xu H, Li X. et al. SIRT1 modulates cell cycle progression by regulating CHK2 acetylation-phosphorylation. Cell Death Differ. 2020;27:482-96
10. Lee IH, Cao L, Mostoslavsky R, Lombard DB, Liu J, Bruns NE. et al. A role for the NAD-dependent deacetylase Sirt1 in the regulation of autophagy. Proc Natl Acad Sci U S A. 2008;105:3374-9
11. Ali S, Nafis S, Kalaiarasan P, Rai E, Sharma S, Bamezai RN. Understanding Genetic Heterogeneity in Type 2 Diabetes by Delineating Physiological Phenotypes: SIRT1 and its Gene Network in Impaired Insulin Secretion. Rev Diabet Stud. 2016;13:17-34
12. Liang F, Kume S, Koya D. SIRT1 and insulin resistance. Nat Rev Endocrinol. 2009;5:367-73
13. Moynihan KA, Grimm AA, Plueger MM, Bernal-Mizrachi E, Ford E, Cras-Meneur C. et al. Increased dosage of mammalian Sir2 in pancreatic beta cells enhances glucose-stimulated insulin secretion in mice. Cell Metab. 2005;2:105-17
14. Rai E, Sharma S, Koul A, Bhat AK, Bhanwer AJ, Bamezai RN. Interaction between the UCP2-866G/A, mtDNA 10398G/A and PGC1alpha p.Thr394Thr and p.Gly482Ser polymorphisms in type 2 diabetes susceptibility in North Indian population. Hum Genet. 2007;122:535-40
15. Bordone L, Motta MC, Picard F, Robinson A, Jhala US, Apfeld J. et al. Sirt1 regulates insulin secretion by repressing UCP2 in pancreatic beta cells. PLoS Biol. 2006;4:e31
16. Robson-Doucette CA, Sultan S, Allister EM, Wikstrom JD, Koshkin V, Bhattacharjee A. et al. Beta-cell uncoupling protein 2 regulates reactive oxygen species production, which influences both insulin and glucagon secretion. Diabetes. 2011;60:2710-9
17. Krauss S, Zhang CY, Scorrano L, Dalgaard LT, St-Pierre J, Grey ST. et al. Superoxide-mediated activation of uncoupling protein 2 causes pancreatic beta cell dysfunction. J Clin Invest. 2003;112:1831-42
18. Mills EM, Xu D, Fergusson MM, Combs CA, Xu Y, Finkel T. Regulation of cellular oncosis by uncoupling protein 2. J Biol Chem. 2002;277:27385-92
19. Chan CB, De Leo D, Joseph JW, McQuaid TS, Ha XF, Xu F. et al. Increased uncoupling protein-2 levels in beta-cells are associated with impaired glucose-stimulated insulin secretion: mechanism of action. Diabetes. 2001;50:1302-10
20. Patane G, Anello M, Piro S, Vigneri R, Purrello F, Rabuazzo AM. Role of ATP production and uncoupling protein-2 in the insulin secretory defect induced by chronic exposure to high glucose or free fatty acids and effects of peroxisome proliferator-activated receptor-gamma inhibition. Diabetes. 2002;51:2749-56
21. Hasegawa K, Wakino S, Kimoto M, Minakuchi H, Fujimura K, Hosoya K. et al. The hydrolase DDAH2 enhances pancreatic insulin secretion by transcriptional regulation of secretagogin through a Sirt1-dependent mechanism in mice. FASEB J. 2013;27:2301-15
22. Wagner L, Oliyarnyk O, Gartner W, Nowotny P, Groeger M, Kaserer K. et al. Cloning and expression of secretagogin, a novel neuroendocrine- and pancreatic islet of Langerhans-specific Ca2+-binding protein. J Biol Chem. 2000;275:24740-51
23. Bauer MC, O'Connell DJ, Maj M, Wagner L, Cahill DJ, Linse S. Identification of a high-affinity network of secretagogin-binding proteins involved in vesicle secretion. Mol Biosyst. 2011;7:2196-204
24. Ramachandran D, Roy U, Garg S, Ghosh S, Pathak S, Kolthur-Seetharam U. Sirt1 and mir-9 expression is regulated during glucose-stimulated insulin secretion in pancreatic beta-islets. FEBS J. 2011;278:1167-74
25. Wu J, Zhang F, Yan M, Wu D, Yu Q, Zhang Y. et al. WldS enhances insulin transcription and secretion via a SIRT1-dependent pathway and improves glucose homeostasis. Diabetes. 2011;60:3197-207
26. Joseph JW, Koshkin V, Zhang CY, Wang J, Lowell BB, Chan CB. et al. Uncoupling protein 2 knockout mice have enhanced insulin secretory capacity after a high-fat diet. Diabetes. 2002;51:3211-9
27. Vetterli L, Brun T, Giovannoni L, Bosco D, Maechler P. Resveratrol potentiates glucose-stimulated insulin secretion in INS-1E beta-cells and human islets through a SIRT1-dependent mechanism. J Biol Chem. 2011;286:6049-60
28. Kong W, Chen LL, Zheng J, Zhang HH, Hu X, Zeng TS. et al. Resveratrol supplementation restores high-fat diet-induced insulin secretion dysfunction by increasing mitochondrial function in islet. Exp Biol Med (Maywood). 2015;240:220-9
29. Yang J, Wang N, Li J, Zhang J, Feng P. Effects of resveratrol on NO secretion stimulated by insulin and its dependence on SIRT1 in high glucose cultured endothelial cells. Endocrine. 2010;37:365-72
30. Fitzpatrick DF, Hirschfield SL, Coffey RG. Endothelium-dependent vasorelaxing activity of wine and other grape products. Am J Physiol. 1993;265:H774-8
31. Lekakis J, Rallidis LS, Andreadou I, Vamvakou G, Kazantzoglou G, Magiatis P. et al. Polyphenolic compounds from red grapes acutely improve endothelial function in patients with coronary heart disease. Eur J Cardiovasc Prev Rehabil. 2005;12:596-600
32. Ndiaye M, Chataigneau M, Lobysheva I, Chataigneau T, Schini-Kerth VB. Red wine polyphenol-induced, endothelium-dependent NO-mediated relaxation is due to the redox-sensitive PI3-kinase/Akt-dependent phosphorylation of endothelial NO-synthase in the isolated porcine coronary artery. FASEB J. 2005;19:455-7
33. Ramsey KM, Mills KF, Satoh A, Imai S. Age-associated loss of Sirt1-mediated enhancement of glucose-stimulated insulin secretion in beta cell-specific Sirt1-overexpressing (BESTO) mice. Aging Cell. 2008;7:78-88
34. Luu L, Dai FF, Prentice KJ, Huang X, Hardy AB, Hansen JB. et al. The loss of Sirt1 in mouse pancreatic beta cells impairs insulin secretion by disrupting glucose sensing. Diabetologia. 2013;56:2010-20
35. Herranz D, Serrano M. SIRT1: recent lessons from mouse models. Nat Rev Cancer. 2010;10:819-23
36. Qiang L, Wang H, Farmer SR. Adiponectin secretion is regulated by SIRT1 and the endoplasmic reticulum oxidoreductase Ero1-L alpha. Mol Cell Biol. 2007;27:4698-707
37. Kadowaki T, Yamauchi T. Adiponectin and adiponectin receptors. Endocr Rev. 2005;26:439-51
38. Trujillo ME, Scherer PE. Adiponectin-journey from an adipocyte secretory protein to biomarker of the metabolic syndrome. J Intern Med. 2005;257:167-75
39. Josephrajan A, Hertzel AV, Bohm EK, McBurney MW, Imai SI, Mashek DG. et al. Unconventional Secretion of Adipocyte Fatty Acid Binding Protein 4 Is Mediated By Autophagic Proteins in a Sirtuin-1-Dependent Manner. Diabetes. 2019;68:1767-77
40. Cao H, Sekiya M, Ertunc ME, Burak MF, Mayers JR, White A. et al. Adipocyte lipid chaperone AP2 is a secreted adipokine regulating hepatic glucose production. Cell Metab. 2013;17:768-78
41. Wu LE, Samocha-Bonet D, Whitworth PT, Fazakerley DJ, Turner N, Biden TJ. et al. Identification of fatty acid binding protein 4 as an adipokine that regulates insulin secretion during obesity. Mol Metab. 2014;3:465-73
42. Kemper JK, Choi SE, Kim DH. Sirtuin 1 deacetylase: a key regulator of hepatic lipid metabolism. Vitam Horm. 2013;91:385-404
43. Zhang QJ, Wang Z, Chen HZ, Zhou S, Zheng W, Liu G. et al. Endothelium-specific overexpression of class III deacetylase SIRT1 decreases atherosclerosis in apolipoprotein E-deficient mice. Cardiovasc Res. 2008;80:191-9
44. Stein S, Lohmann C, Schafer N, Hofmann J, Rohrer L, Besler C. et al. SIRT1 decreases Lox-1-mediated foam cell formation in atherogenesis. Eur Heart J. 2010;31:2301-9
45. Gorenne I, Kumar S, Gray K, Figg N, Yu H, Mercer J. et al. Vascular smooth muscle cell sirtuin 1 protects against DNA damage and inhibits atherosclerosis. Circulation. 2013;127:386-96
46. Wu Q, Hu Y, Jiang M, Wang F, Gong G. Effect of Autophagy Regulated by Sirt1/FoxO1 Pathway on the Release of Factors Promoting Thrombosis from Vascular Endothelial Cells. Int J Mol Sci. 2019 20
47. Zaid A, Roubtsova A, Essalmani R, Marcinkiewicz J, Chamberland A, Hamelin J. et al. Proprotein convertase subtilisin/kexin type 9 (PCSK9): hepatocyte-specific low-density lipoprotein receptor degradation and critical role in mouse liver regeneration. Hepatology. 2008;48:646-54
48. Miranda MX, van Tits LJ, Lohmann C, Arsiwala T, Winnik S, Tailleux A. et al. The Sirt1 activator SRT3025 provides atheroprotection in Apoe-/- mice by reducing hepatic Pcsk9 secretion and enhancing Ldlr expression. Eur Heart J. 2015;36:51-9
49. Osono Y, Woollett LA, Herz J, Dietschy JM. Role of the low density lipoprotein receptor in the flux of cholesterol through the plasma and across the tissues of the mouse. J Clin Invest. 1995;95:1124-32
50. Bordone L, Cohen D, Robinson A, Motta MC, van Veen E, Czopik A. et al. SIRT1 transgenic mice show phenotypes resembling calorie restriction. Aging Cell. 2007;6:759-67
51. Purushotham A, Schug TT, Xu Q, Surapureddi S, Guo X, Li X. Hepatocyte-specific deletion of SIRT1 alters fatty acid metabolism and results in hepatic steatosis and inflammation. Cell Metab. 2009;9:327-38
52. Chen D, Bruno J, Easlon E, Lin SJ, Cheng HL, Alt FW. et al. Tissue-specific regulation of SIRT1 by calorie restriction. Genes Dev. 2008;22:1753-7
53. Tirosh D, Benshalom-Tirosh N, Novack L, Press F, Beer-Weisel R, Wiznitzer A. et al. Hypothyroidism and diabetes mellitus - a risky dual gestational endocrinopathy. PeerJ. 2013;1:e52
54. Yamagishi S. Role of advanced glycation end products (AGEs) and receptor for AGEs (RAGE) in vascular damage in diabetes. Exp Gerontol. 2011;46:217-24
55. Ramasamy R, Yan SF, Schmidt AM. Receptor for AGE (RAGE): signaling mechanisms in the pathogenesis of diabetes and its complications. Ann N Y Acad Sci. 2011;1243:88-102
56. Chen XJ, Gong XH, Jie JP, Yu WH, Chen X, Du X. et al. Receptor for advanced glycation end products reveals a mechanism regulating thyroid hormone secretion through the SIRT1/Nrf2 pathway. J Cell Biochem. 2019;120:4582-98
57. Akieda-Asai S, Zaima N, Ikegami K, Kahyo T, Yao I, Hatanaka T. et al. SIRT1 Regulates Thyroid-Stimulating Hormone Release by Enhancing PIP5Kgamma Activity through Deacetylation of Specific Lysine Residues in Mammals. PLoS One. 2010;5:e11755
58. Gong LW, Di Paolo G, Diaz E, Cestra G, Diaz ME, Lindau M. et al. Phosphatidylinositol phosphate kinase type I gamma regulates dynamics of large dense-core vesicle fusion. Proc Natl Acad Sci U S A. 2005;102:5204-9
59. Boily G, Seifert EL, Bevilacqua L, He XH, Sabourin G, Estey C. et al. SirT1 regulates energy metabolism and response to caloric restriction in mice. PLoS One. 2008;3:e1759
60. Cohen DE, Supinski AM, Bonkowski MS, Donmez G, Guarente LP. Neuronal SIRT1 regulates endocrine and behavioral responses to calorie restriction. Genes Dev. 2009;23:2812-7
61. Ramadori G, Lee CE, Bookout AL, Lee S, Williams KW, Anderson J. et al. Brain SIRT1: anatomical distribution and regulation by energy availability. J Neurosci. 2008;28:9989-96
62. Donmez G, Outeiro TF. SIRT1 and SIRT2: emerging targets in neurodegeneration. EMBO Mol Med. 2013;5:344-52
63. Furukawa A, Tada-Oikawa S, Kawanishi S, Oikawa S. H2O2 accelerates cellular senescence by accumulation of acetylated p53 via decrease in the function of SIRT1 by NAD+ depletion. Cell Physiol Biochem. 2007;20:45-54
64. Pallas M, Verdaguer E, Tajes M, Gutierrez-Cuesta J, Camins A. Modulation of sirtuins: new targets for antiageing. Recent Pat CNS Drug Discov. 2008;3:61-9
65. Qin B, Panickar KS, Anderson RA. Cinnamon polyphenols attenuate the hydrogen peroxide-induced down regulation of S100beta secretion by regulating sirtuin 1 in C6 rat glioma cells. Life Sci. 2014;102:72-9
66. Huttunen HJ, Kuja-Panula J, Sorci G, Agneletti AL, Donato R, Rauvala H. Coregulation of neurite outgrowth and cell survival by amphoterin and S100 proteins through receptor for advanced glycation end products (RAGE) activation. J Biol Chem. 2000;275:40096-105
67. Businaro R, Leone S, Fabrizi C, Sorci G, Donato R, Lauro GM. et al. S100B protects LAN-5 neuroblastoma cells against Abeta amyloid-induced neurotoxicity via RAGE engagement at low doses but increases Abeta amyloid neurotoxicity at high doses. J Neurosci Res. 2006;83:897-906
68. Kogel D, Peters M, Konig HG, Hashemi SM, Bui NT, Arolt V. et al. S100B potently activates p65/c-Rel transcriptional complexes in hippocampal neurons: Clinical implications for the role of S100B in excitotoxic brain injury. Neuroscience. 2004;127:913-20
69. Hu J, Castets F, Guevara JL, Van Eldik LJ. S100 beta stimulates inducible nitric oxide synthase activity and mRNA levels in rat cortical astrocytes. J Biol Chem. 1996;271:2543-7
70. Lam AG, Koppal T, Akama KT, Guo L, Craft JM, Samy B. et al. Mechanism of glial activation by S100B: involvement of the transcription factor NFkappaB. Neurobiol Aging. 2001;22:765-72
71. Adami C, Sorci G, Blasi E, Agneletti AL, Bistoni F, Donato R. S100B expression in and effects on microglia. Glia. 2001;33:131-42
72. Petrova TV, Hu J, Van Eldik LJ. Modulation of glial activation by astrocyte-derived protein S100B: differential responses of astrocyte and microglial cultures. Brain Res. 2000;853:74-80
73. Donato R, Sorci G, Riuzzi F, Arcuri C, Bianchi R, Brozzi F. et al. S100B's double life: intracellular regulator and extracellular signal. Biochim Biophys Acta. 2009;1793:1008-22
74. Hofmann MA, Drury S, Fu C, Qu W, Taguchi A, Lu Y. et al. RAGE mediates a novel proinflammatory axis: a central cell surface receptor for S100/calgranulin polypeptides. Cell. 1999;97:889-901
75. Donato R. RAGE: a single receptor for several ligands and different cellular responses: the case of certain S100 proteins. Curr Mol Med. 2007;7:711-24
76. Ostendorp T, Leclerc E, Galichet A, Koch M, Demling N, Weigle B. et al. Structural and functional insights into RAGE activation by multimeric S100B. EMBO J. 2007;26:3868-78
77. Yoon MJ, Yoshida M, Johnson S, Takikawa A, Usui I, Tobe K. et al. SIRT1-Mediated eNAMPT Secretion from Adipose Tissue Regulates Hypothalamic NAD+ and Function in Mice. Cell Metab. 2015;21:706-17
78. Pallier C, Scaffidi P, Chopineau-Proust S, Agresti A, Nordmann P, Bianchi ME. et al. Association of chromatin proteins high mobility group box (HMGB) 1 and HMGB2 with mitotic chromosomes. Mol Biol Cell. 2003;14:3414-26
79. Stros M, Ozaki T, Bacikova A, Kageyama H, Nakagawara A. HMGB1 and HMGB2 cell-specifically down-regulate the p53- and p73-dependent sequence-specific transactivation from the human Bax gene promoter. J Biol Chem. 2002;277:7157-64
80. Lotze MT, Tracey KJ. High-mobility group box 1 protein (HMGB1): nuclear weapon in the immune arsenal. Nat Rev Immunol. 2005;5:331-42
81. Muller S, Scaffidi P, Degryse B, Bonaldi T, Ronfani L, Agresti A. et al. New EMBO members' review: the double life of HMGB1 chromatin protein: architectural factor and extracellular signal. EMBO J. 2001;20:4337-40
82. Andersson U, Tracey KJ. HMGB1 is a therapeutic target for sterile inflammation and infection. Annu Rev Immunol. 2011;29:139-62
83. Wang H, Bloom O, Zhang M, Vishnubhakat JM, Ombrellino M, Che J. et al. HMG-1 as a late mediator of endotoxin lethality in mice. Science. 1999;285:248-51
84. Schiraldi M, Raucci A, Munoz LM, Livoti E, Celona B, Venereau E. et al. HMGB1 promotes recruitment of inflammatory cells to damaged tissues by forming a complex with CXCL12 and signaling via CXCR4. J Exp Med. 2012;209:551-63
85. Li J, Kokkola R, Tabibzadeh S, Yang R, Ochani M, Qiang X. et al. Structural basis for the proinflammatory cytokine activity of high mobility group box 1. Mol Med. 2003;9:37-45
86. Chen GY, Tang J, Zheng P, Liu Y. CD24 and Siglec-10 selectively repress tissue damage-induced immune responses. Science. 2009;323:1722-5
87. Yang H, Hreggvidsdottir HS, Palmblad K, Wang H, Ochani M, Li J. et al. A critical cysteine is required for HMGB1 binding to Toll-like receptor 4 and activation of macrophage cytokine release. Proc Natl Acad Sci U S A. 2010;107:11942-7
88. Dumitriu IE, Baruah P, Valentinis B, Voll RE, Herrmann M, Nawroth PP. et al. Release of high mobility group box 1 by dendritic cells controls T cell activation via the receptor for advanced glycation end products. J Immunol. 2005;174:7506-15
89. Wild CA, Bergmann C, Fritz G, Schuler P, Hoffmann TK, Lotfi R. et al. HMGB1 conveys immunosuppressive characteristics on regulatory and conventional T cells. Int Immunol. 2012;24:485-94
90. Bonaldi T, Talamo F, Scaffidi P, Ferrera D, Porto A, Bachi A. et al. Monocytic cells hyperacetylate chromatin protein HMGB1 to redirect it towards secretion. EMBO J. 2003;22:5551-60
91. Hwang JS, Lee WJ, Kang ES, Ham SA, Yoo T, Paek KS. et al. Ligand-activated peroxisome proliferator-activated receptor-delta and -gamma inhibit lipopolysaccharide-primed release of high mobility group box 1 through upregulation of SIRT1. Cell Death Dis. 2014;5:e1432
92. Walko TD 3rd, Di Caro V, Piganelli J, Billiar TR, Clark RS, Aneja RK. Poly(ADP-ribose) polymerase 1-sirtuin 1 functional interplay regulates LPS-mediated high mobility group box 1 secretion. Mol Med. 2015;20:612-24
93. Kim YM, Park EJ, Kim JH, Park SW, Kim HJ, Chang KC. Ethyl pyruvate inhibits the acetylation and release of HMGB1 via effects on SIRT1/STAT signaling in LPS-activated RAW264.7 cells and peritoneal macrophages. Int Immunopharmacol. 2016;41:98-105
94. Qi Z, Zhang Y, Qi S, Ling L, Gui L, Yan L. et al. Salidroside Inhibits HMGB1 Acetylation and Release through Upregulation of SirT1 during Inflammation. Oxid Med Cell Longev. 2017;2017:9821543
95. Zeng W, Shan W, Gao L, Gao D, Hu Y, Wang G. et al. Inhibition of HMGB1 release via salvianolic acid B-mediated SIRT1 up-regulation protects rats against non-alcoholic fatty liver disease. Sci Rep. 2015;5:16013
96. Ye TJ, Lu YL, Yan XF, Hu XD, Wang XL. High mobility group box-1 release from H2O2-injured hepatocytes due to sirt1 functional inhibition. World J Gastroenterol. 2019;25:5434-50
97. Xu S, Zeng Z, Zhao M, Huang Q, Gao Y, Dai X. et al. Evidence for SIRT1 Mediated HMGB1 Release From Kidney Cells in the Early Stages of Hemorrhagic Shock. Front Physiol. 2019;10:854
98. Rickenbacher A, Jang JH, Limani P, Ungethum U, Lehmann K, Oberkofler CE. et al. Fasting protects liver from ischemic injury through Sirt1-mediated downregulation of circulating HMGB1 in mice. J Hepatol. 2014;61:301-8
99. Ford J, Jiang M, Milner J. Cancer-specific functions of SIRT1 enable human epithelial cancer cell growth and survival. Cancer Res. 2005;65:10457-63
100. Ota H, Tokunaga E, Chang K, Hikasa M, Iijima K, Eto M. et al. Sirt1 inhibitor, Sirtinol, induces senescence-like growth arrest with attenuated Ras-MAPK signaling in human cancer cells. Oncogene. 2006;25:176-85
101. Abdelmohsen K, Pullmann R Jr, Lal A, Kim HH, Galban S, Yang X. et al. Phosphorylation of HuR by Chk2 regulates SIRT1 expression. Mol Cell. 2007;25:543-57
102. Fu M, Liu M, Sauve AA, Jiao X, Zhang X, Wu X. et al. Hormonal control of androgen receptor function through SIRT1. Mol Cell Biol. 2006;26:8122-35
103. Saunders LR, Verdin E. Sirtuins: critical regulators at the crossroads between cancer and aging. Oncogene. 2007;26:5489-504
104. Jeng MY, Hull PA, Fei M, Kwon HS, Tsou CL, Kasler H. et al. Metabolic reprogramming of human CD8(+) memory T cells through loss of SIRT1. J Exp Med. 2018;215:51-62
105. Kwon HS, Lim HW, Wu J, Schnolzer M, Verdin E, Ott M. Three novel acetylation sites in the Foxp3 transcription factor regulate the suppressive activity of regulatory T cells. J Immunol. 2012;188:2712-21
106. Yeung F, Hoberg JE, Ramsey CS, Keller MD, Jones DR, Frye RA. et al. Modulation of NF-kappaB-dependent transcription and cell survival by the SIRT1 deacetylase. EMBO J. 2004;23:2369-80
107. Jia L, Clear A, Liu FT, Matthews J, Uddin N, McCarthy A. et al. Extracellular HMGB1 promotes differentiation of nurse-like cells in chronic lymphocytic leukemia. Blood. 2014;123:1709-19
108. Shanmugam N, Kim YS, Lanting L, Natarajan R. Regulation of cyclooxygenase-2 expression in monocytes by ligation of the receptor for advanced glycation end products. J Biol Chem. 2003;278:34834-44
109. Shanmugam N, Ransohoff RM, Natarajan R. Interferon-gamma-inducible protein (IP)-10 mRNA stabilized by RNA-binding proteins in monocytes treated with S100b. J Biol Chem. 2006;281:31212-21
110. Volonte D, Zou H, Bartholomew JN, Liu Z, Morel PA, Galbiati F. Oxidative stress-induced inhibition of Sirt1 by caveolin-1 promotes p53-dependent premature senescence and stimulates the secretion of interleukin 6 (IL-6). J Biol Chem. 2015;290:4202-14
111. Lee BR, Sanstrum BJ, Liu Y, Kwon SH. Distinct role of Sirtuin 1 (SIRT1) and Sirtuin 2 (SIRT2) in inhibiting cargo-loading and release of extracellular vesicles. Sci Rep. 2019;9:20049
112. McAndrews KM, LeBleu VS, Kalluri R. SIRT1 Regulates Lysosome Function and Exosome Secretion. Dev Cell. 2019;49:302-3
113. Latifkar A, Ling L, Hingorani A, Johansen E, Clement A, Zhang X. et al. Loss of Sirtuin 1 Alters the Secretome of Breast Cancer Cells by Impairing Lysosomal Integrity. Dev Cell. 2019;49:393-408 e7
114. Kalluri R. The biology and function of exosomes in cancer. J Clin Invest. 2016;126:1208-15
115. Han L, Long Q, Li S, Xu Q, Zhang B, Dou X. et al. Senescent stromal cells promote cancer resistance through SIRT1 loss-potentiated overproduction of small extracellular vesicles. Cancer Res. 2020
116. Guo X, Williams JG, Schug TT, Li X. DYRK1A and DYRK3 promote cell survival through phosphorylation and activation of SIRT1. J Biol Chem. 2010;285:13223-32
117. Guo X, Kesimer M, Tolun G, Zheng X, Xu Q, Lu J. et al. The NAD(+)-dependent protein deacetylase activity of SIRT1 is regulated by its oligomeric status. Sci Rep. 2012;2:640
Author contact
Corresponding authors: Liu Cao: College of Basic Medical Science, Institute of Translational Medicine, Key Laboratory of Medical Cell Biology, Ministry of Education, Key Laboratory of Liaoning Province, China Medical University, Shenyang, Liaoning Province, China, 110122. Email: lcaoedu.cn. Tel: +86-18900911888, Fax: +86-24-31939636, Xiaoyu Song: College of Basic Medical Science, Institute of Translational Medicine, Key Laboratory of Medical Cell Biology, Ministry of Education, Key Laboratory of Liaoning Province, China Medical University, Shenyang, Liaoning Province, China, 110122; Email: xysongedu.cn; Tel: +86-18900910183, Fax: +86-24-31939636

 Global reach, higher impact
Global reach, higher impact