Impact Factor ISSN: 1449-2288
Int J Biol Sci 2021; 17(9):2157-2166. doi:10.7150/ijbs.59648 This issue Cite
Research Paper
BMP and Notch Signaling Pathways differentially regulate Cardiomyocyte Proliferation during Ventricle Regeneration
1. School of Life Sciences, Fudan University, Shanghai, China.
2. Department of Medicine, Li Ka Shing Faculty of Medicine, The University of Hong Kong, Pokfulam, Hong Kong, China.
3. School of Biomedical Sciences, Li Ka Shing Faculty of Medicine, The University of Hong Kong, Pokfulam, Hong Kong, China.
4. Institute of Molecular Medicine, Beijing Key Laboratory of Cardiometabolic Molecular Medicine, Peking University, Beijing, China.
5. School of Basic Medical Sciences, Wuhan University, Wuhan, China.
#These authors contributed equally to this study.
Received 2021-2-22; Accepted 2021-5-14; Published 2021-5-27
Abstract
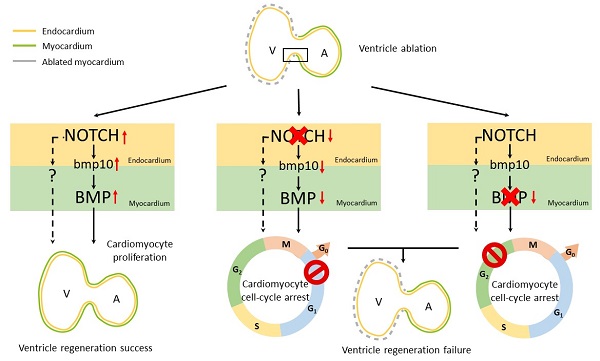
Adult mammalian hearts show limited capacity to proliferate after injury, while zebrafish are capable to completely regenerate injured hearts through the proliferation of spared cardiomyocytes. BMP and Notch signaling pathways have been implicated in cardiomyocyte proliferation during zebrafish heart regeneration. However, the molecular mechanism underneath this process as well as the interaction between these two pathways remains to be further explored. In this study we showed BMP signaling was activated after ventricle ablation and acted epistatic downstream of Notch signaling. Inhibition of both signaling pathways differentially influenced ventricle regeneration and cardiomyocyte proliferation, as revealed by time-lapse analysis using a cardiomyocyte-specific FUCCI (fluorescent ubiquitylation-based cell cycle indicator) system. Further experiments revealed that inhibition of BMP and Notch signaling led to cell-cycle arrest at different phases. Overall, our results shed light on the interaction between BMP and Notch signaling pathways and their functions in cardiomyocyte proliferation during cardiac regeneration.
Keywords: heart regeneration, BMP signaling, Notch signaling, cardiomyocyte proliferation, cell-cycle arrest
Introduction
Myocardial infarction (MI) is a leading cause of death worldwide [1, 2]. In adult mammals, the heart, as one of the least regenerative organs, replaces the infarcted myocardium with noncontractile scar tissue [3]. On the contrast, zebrafish are capable to completely regenerate the injured hearts [4-6]. New cardiomyocytes have been shown to arise from proliferation of pre-existing cardiomyocytes through genetic fate-mapping experiments [7, 8]. Though natural cardiomyocyte turnover rate is low in adult humans [9, 10], accelerating such process after acute insult is therapeutically valuable. Thus, utilizing cellular and genetic factors that stimulate cardiomyocyte proliferation may provide viable solutions for promoting human cardiac regeneration.
Zebrafish cardiac regeneration involves multiple signaling pathways [11-13]. BMP signaling is vital for vertebrate cardiovascular development [14, 15], and its function in cardiac injury and repair has become more appreciated. Since inhibitory effects of BMP2 on fibroblast function have been reported [16], exogenous BMP2 reduced the size of infarcted tissue by diminishing apoptotic cardiomyocytes in the border zone in a mouse permanent coronary occlusion model [17]. While BMP4 enhances apoptosis and hypertrophy of cultured mammalian cardiomyocytes [18], BMP7 signaling attenuates myocardial fibrosis by inhibiting TGF-β responses in a rat MI model [19]. BMP10 has been demonstrated to enhance inflammation activity after MI and exogenous BMP10 results in cardiomyocyte cell-cycle re-entry [20]. Thus, BMP signaling has been suggested as a good candidate pathway for modulating cardiac regeneration [21], but its role in cardiomyocyte proliferation requires further investigation.
Notch signaling pathway also plays important roles in cardiac regeneration [22, 23]. Notch signaling is activated in the endocardium of injured zebrafish hearts and its inhibition impedes cardiomyocyte proliferation and heart regeneration [6, 24-26]. Endocardial Notch signaling has been shown to promote myocardial proliferative events through inducing BMP10 expression in adjacent cardiomyocytes [27-29]. However, the interaction between these two pathways and their differential regulation of cardiomyocyte proliferation remains elusive. In this study we utilized the zebrafish larval ventricle ablation model and a cardiomyocyte-specific FUCCI (fluorescent ubiquitylation-based cell cycle indicator) system to tackle these questions. We demonstrated the epistatic relationship between BMP and Notch signaling pathways during ventricle regeneration and further revealed their differential functions in cardiomyocyte proliferation, blocking of these pathways lead to cell-cycle arrest at different phases. Overall, our results shed light on molecular mechanism of BMP and Notch signaling pathways in cardiomyocyte proliferation during cardiac regeneration, which could lay a foundation for future development of therapeutic interventions.
Results
BMP signaling is activated during larval ventricle regeneration
Recent research suggested that BMP signaling activation was an injury response in adult zebrafish hearts [21]. To investigate the role of BMP signaling in the larval ventricle regeneration model, we treated the ventricle-specific genetic ablation line Tg(vmhc:mCherry-NTR) with metronidazole (MTZ) at 3 days post fertilization (dpf) [6]. Whole-mount in situ hybridization (WISH) staining revealed that gene expressions of several BMP signaling components were dramatically increased in ablated hearts compared to that in control hearts at 5 dpf/ 2 days post MTZ-treatment (dpt) (Fig. 1A-D'). BMP ligand bmp10 was weakly expressed in the ventricle of control hearts, probably due to the trabeculae formation at this stage. After ablation, bmp10 expression was strongly induced, especially in the atrium (Fig. 1A, A'). Expression of BMP receptor bmpr1aa and signal transducer smad1 was also up-regulated in ablated hearts, while expression of BMP signaling effector id2b showed a much dramatic increment after ablation (Fig. 1B-D').
Activation of BMP signaling pathway during ventricle regeneration. (A-D') Whole-mount in situ hybridizations showed upregulated expression of BMP signaling components in ablated hearts (A'-D') compared to control hearts (A-D) at 5 dpf/2 dpt. (E-G) Confocal stack projections of Tg(vmhc:mCherry-NTR; Bre:dGFP) hearts showed BMP signaling activation at 1-2 dpt. (H-J) Confocal stack projections of immunofluorescence showed BMP signaling activation at 1-2 dpt. Green, phospho-Smad1/5/9; red, myosin heavy chain (MF20). (K-N') Overlay of Bre:dGFP and pSmad1/5/9 signals in myocardium and epicardium at 2 dpt. (K-N) stack projections, (K'-N') optical sections. Scale bars, (A-D') 25 µm, (E-N') 50 µm. dpf, days post fertilization; dpt, days post MTZ-treatment; A, atrium; V, ventricle. Dashed lines outline the hearts.
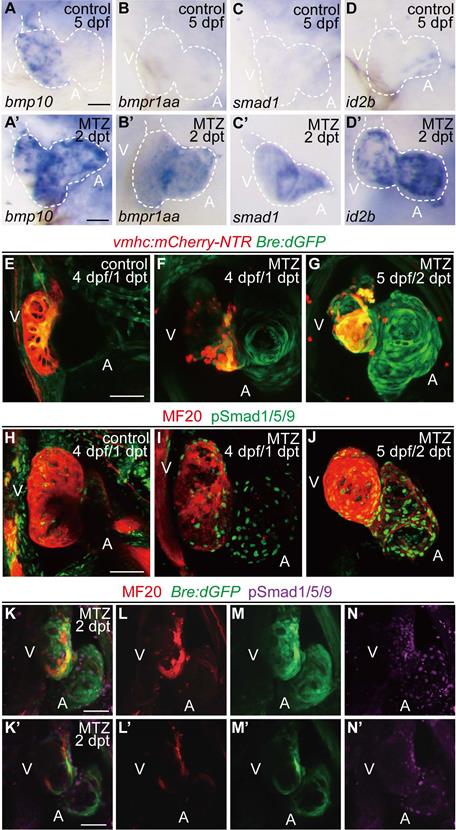
To visualize the transient activation of BMP signaling in regenerating hearts, we used a reporter line Tg(Bre:dGFP) which expressed destabilized GFP in BMP-activated cells [30, 31]. Control Tg(vmhc:mCherry-NTR; Bre:dGFP) larvae displayed weak GFP signal in the ventricle at 4 dpf, besides other extracardiac signals. After ventricle injury, BMP signaling was activated in the ablated heart, mainly in the atrium, at 1 dpt. The signal intensity was gradually enhanced in the atrium and extended to the ventricle at 2 dpt (Fig. 1E-G). The activation of BMP signaling during ventricle regeneration was also confirmed by immunofluorescence staining of another marker, phosphorylation of Smad1/5/9 (pSmad1/5/9), which showed a similar pattern as Bre:dGFP (Fig. 1H-J). To further validate these results, we also performed immunostaining of pSmad1/5/9 in Tg(vmhc:mCherry-NTR; Bre:dGFP) larvae. The results suggested that BMP signaling was strongly activated in myocardium during ventricle regeneration, while a weaker epicardial BMP signal could be detected occasionally (Fig. 1K-N'). Overall, our results confirmed that BMP signaling was activated during larval ventricle regeneration.
BMP signaling acts downstream of Notch signaling during ventricle regeneration
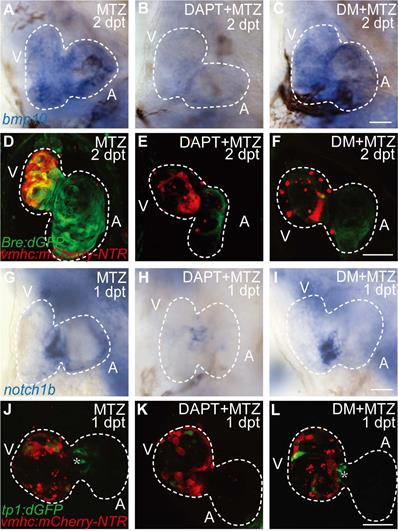
Previous study showed post-injury upregulation of bmp10 expression was blocked after Notch signaling inhibition [29], which suggested that BMP signaling was regulated by Notch signaling during regeneration. Whether BMP signaling regulated Notch signaling in a reciprocal manner remained unclear. We used pharmacological approach to study the epistatic relationship between these two signaling pathways. Inhibition of Notch signaling by DAPT treatment significantly reduced bmp10 expression in ablated hearts at 2 dpt as revealed by WISH staining (Fig. 2A, B). BMP signaling activation was also blocked as revealed by diminished Bre:dGFP signals in DAPT-treated ablated hearts (Fig. 2D, E). Treatment of BMP signaling inhibitor Dorsomorphin (DM) blocked downstream transduction of BMP pathway, but had no effect on ligand expression (Fig. 2C, F). On the other hand, WISH analysis of notch1b expression in ablated hearts at 1 dpt showed no difference between DM-treated group and control group, while notch1b upregulation was attenuated after DAPT treatment (Fig. 2G-I). We observed a similar pattern in the Notch signaling reporter line Tg(tp1:dGFP) that inhibition of BMP signaling by DM treatment did not affect the activation of Notch signaling in the endocardium around atrioventricular canal (Fig. 2J-L, asterisk). Thus, our results suggested that BMP signaling acted epistatic downstream of Notch signaling during ventricle regeneration.
BMP signaling acts downstream of Notch signaling during ventricle regeneration. (A-C) Whole-mount in situ hybridizations showed upregulated bmp10 expression in ablated hearts at 2 dpt could be blocked by Notch signaling inhibitor DAPT but not BMP signaling inhibitor DM. (D-F) Confocal stack projections of Tg(vmhc:mCherry-NTR; Bre:dGFP) hearts showed BMP signaling activation at 2 dpt could be blocked by both DAPT and DM. (G-I) Whole-mount in situ hybridizations showed upregulated notch1b expression in ablated hearts at 1 dpt could be blocked by DAPT but not DM. (J-L) Confocal stack projections of Tg(vmhc:mCherry-NTR; tp1:dGFP) hearts showed Notch signaling activation at 1 dpt could be attenuated by DAPT but not DM. Asterisk, Notch signal at the atrioventricular canal. Scale bars, (A-C, G-I) 25 µm, (D-F, J-L) 50 µm. dpt, days post MTZ-treatment; A, atrium; V, ventricle; DM, dorsomorphin. Dashed lines outline the hearts.
BMP and Notch signaling pathways differentially influence ventricle regeneration and cardiogenic factor re-activation
To compare the functions of Notch and BMP signaling in ventricle regeneration, we first quantified the heart recovery rate at 4 dpt. DAPT treatment during the whole course (0-4 dpt) or early stage (0-2 dpt) of ventricle regeneration significantly reduced the heart recovery rates (Fig. 3A, 39.8%, N=118, and 48.8%, N=41, respectively) compared to control group (81.2%, N=186), which was in accordance with previous report [6, 32]. A similar pattern of reduced heart recovery rate was observed when DM treatment was applied during 0-4 dpt (Fig. 3B, 48.6%, N=72 vs 87.6%, N=178 in control group) and 0-2 dpt (55.9%, N=36). We further assessed the effects of pathway inhibition during late stage of ventricle regeneration. DAPT treatment during 1-4 dpt and 2-4 dpt also decreased the heart recovery rates, though to a lesser degree of reduction (Fig. 3A, 55.8%, N=43, and 61.3%, N=31, respectively), while DAPT treatment during 3-4 dpt did not obviously change the recovery rate (78.4%, N=37). On the contrast, DM treatment showed reduction effect only when applied during 1-4 dpt (Fig. 3B, 58.1%, N=43) but not during 2-4 dpt (81.1%, N=37) or 3-4 dpt (86.0%, N=50). Such difference between DAPT and DM treatments indicated that BMP and Notch signaling pathways influence ventricle regeneration in different time windows.
BMP and Notch signaling pathways differentially influence ventricle regeneration and cardiogenic factor re-activation. (A, B) Quantification of heart recovery rate in control and DAPT-treated or DM-treated ablated hearts at 4 dpt. The numbers of larvae analyzed were indicated. Binomial test, n.s, non-significant, *, P < 0.05, **, P <0.01, ***, P < 0.001. (C-T) Whole-mount in situ hybridizations showed different effects on the re-activation of cardiogenic factor nkx2.5 (C-H), gata4 (I-N), tbx20 (O-T) after DAPT or DM treatment at 5 dpf/2 dpt. Scale bar, 50 µm. dpf, days post fertilization; dpt, days post MTZ-treatment; A, atrium; V, ventricle; Ctrl, control; DM, dorsomorphin. Dashed lines outline the hearts.
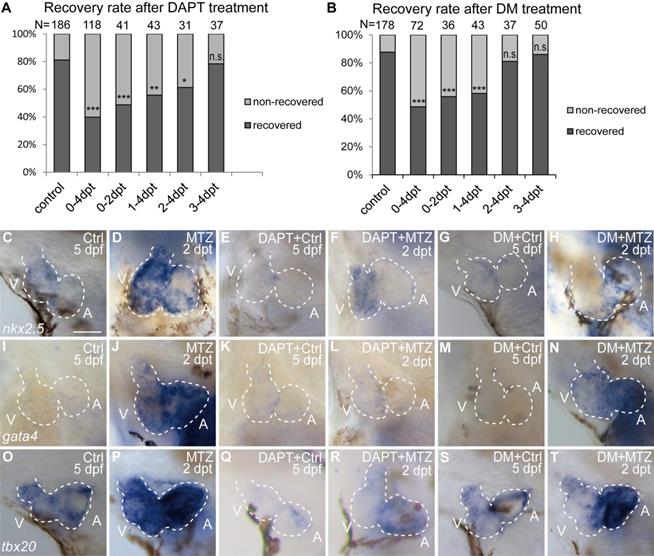
We then examined the effect of Notch or BMP signaling blockage on re-activation of early cardiogenic transcription factors. Expression of nkx2.5, gata4 and tbx20 was dramatically increased in ablated hearts compared to control hearts at 2 dpt as revealed by WISH experiments (Fig. 3C, D, I, J, O, P). After DAPT treatment, cardiogenic factor re-activation was blocked in both chambers (Fig. 3E, F, K, L, Q, R) similar as previously reported [6, 29]. However, DM treatment showed heterogeneous effect on cardiogenic factor re-activation in the two chambers, with an apparent stronger reduction in the ventricle (Fig. 3G, H, M, N, S, T). These results suggested BMP and Notch signaling pathways might have differential functions in cardiogenic factor re-activation.
FUCCI as a cardiomyocyte proliferation indicator during ventricle regeneration
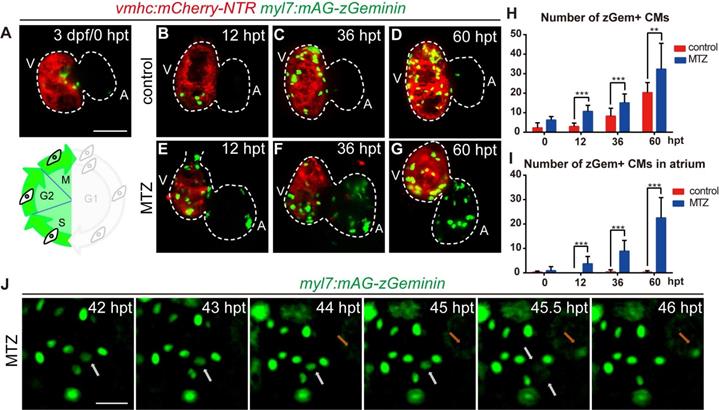
Next, we focused on the regulation of cardiomyocyte proliferation, which is an important aspect of heart regeneration [33, 34]. FUCCI was developed as a tool to monitor cell-cycle behavior in vivo [35-37]. To visualize the dynamic process of cardiomyocyte proliferation, we crossed cardiomyocyte-specific FUCCI line Tg(myl7:mAG-zGeminin) with Tg(vmhc:mCherry-NTR) so that cardiomyocytes in S/G2/M phases would be labelled with green fluorescence (Fig. 4A). Few zGem+ cardiomyocytes were observed in control hearts at 3 dpf, mainly in the ventricle. The number of zGem+ cardiomyocytes gradually increased (Fig. 4A-D), probably due to trabeculae formation in the ventricle at this stage. After MTZ treatment zGem+ cardiomyocytes first showed up in the atrium of ablated hearts at 12 hours post treatment (hpt) with increasing numbers during regeneration (Fig. 4E-G). Quantification of zGem+ cardiomyocyte numbers at 0, 12, 36, 60 hpt indicated that myocardial proliferation in ablated hearts was enhanced (6.2 ± 1.8, 10.7 ± 3.1, 15.1 ± 4.5, 32.3 ± 13.3, N=8, 15, 15, 10, respectively) compared to control hearts (2.2 ± 2.6, 2.9 ± 1.8, 8.2 ± 4.1, 20.3 ± 5.1, N=8, 15, 13, 12, respectively) (Fig. 4H). Consistent with this pattern, the atrium responded to ventricle injury and the quantity of zGem+ cardiomyocytes in the atrium of ablated hearts escalated markedly (0.9 ± 1.6, 3.7 ± 3.0, 8.9 ± 4.4, 22.5 ± 8.3, N=8, 15, 15, 10, respectively) compared to control atriums (0.2 ± 0.5, 0.0 ± 0.0, 0.4 ± 0.9, 0.2 ± 0.6, N=8, 15, 13, 12, respectively) (Fig. 4I). Short period time-lapse fluorescence imaging captured how FUCCI system reflected the cell cycle state with loss of zGem+ signals after cell division (Fig. 4J, white arrows), as well as newly appeared zGem+ signal suggesting the cardiomyocyte just entered the proliferating stage (orange arrows).
FUCCI as a cardiomyocyte proliferation indicator during ventricle regeneration. (A-G) Confocal stack projections of Tg(vmhc:mCherry-NTR; myl7:mAG-zGeminin) hearts showed pattern of zGem+ CMs at 3 dpf/0 hpt, which are in the S, G2 and M phases of the cell-cycle (A). Numbers of zGem+ CMs dramatically increased in the atrium of ablated hearts (E-G) compared to control hearts (B-D). (H, I) Quantification of zGem+ CM numbers in the whole heart (H) or in the atrium only (I) of control and ablated groups at 0, 12, 36 and 60 hpt. Mean + SD, N=8-15 for each time point, Student's t-test, **, P <0.01, ***, P < 0.001. (J) Time-lapse images of Tg(myl7:mAG-zGeminin) hearts showed the dynamic pattern of zGem+ CMs during 42-46 hpt. White arrows point to a CM which lost zGem+ signal after division, while orange arrows point to a CM with newly appeared zGem+ signal. Scale bars, (A-G) 50 µm, (J) 20 µm. dpf, days post fertilization; hpt, hours post MTZ treatment; A, atrium; V, ventricle; CM, cardiomyocyte. Dashed lines outline the hearts.
Inhibition of BMP and Notch signaling pathways differentially suppresses FUCCI expression
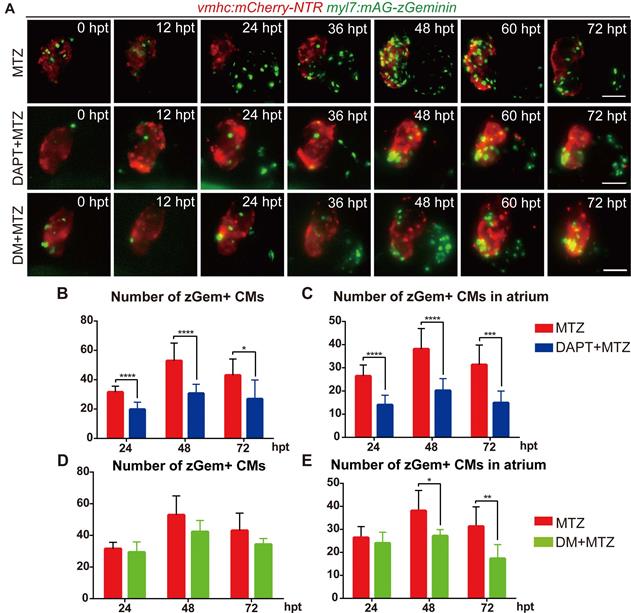
To further explore the functions of BMP and Notch signaling in cardiomyocyte proliferation, we performed pharmacological inhibition experiments in Tg(vmhc:mCherry-NTR; myl7:mAG-zGeminin) and recorded the change in zGem+ cardiomyocyte pattern by 12-hour-interval living imaging (Fig. 5A). Compared with control hearts, the zGem+ signal decreased in DAPT-treated ablated hearts from 24 hpt, which was more obvious in the atrium. DM treatment showed a lesser inhibitory effect on zGem+ signals. We then quantified the numbers of zGem+ cardiomyocytes in the ablated hearts at 24, 48, 72 hpt. DAPT treatment significantly reduced the numbers from 31.6 ± 4.1 (N=9) to 19.8 ± 4.9 (N=13) at 24 hpt, from 52.9 ± 12.0 (N=11) to 30.7 ± 6.2 (N=10) at 48 hpt, and from 43.0 ± 11.1 (N=9) to 26.9 ± 13.0 (N=8) at 72 hpt (Fig. 5B). Considering the numbers of zGem+ cardiomyocytes in the atrium only, we found a similar trend upon DAPT treatment that the numbers dropped from 26.4 ± 4.8 (N=9) to 14.0 ± 4.2 (N=13) at 24 hpt, from 38.1 ± 8.9 (N=11) to 20.2 ± 5.2 (N=10) at 48 hpt, and from 31.3 ± 8.5 (N=9) to 14.9 ± 5.1 (N=8) at 72 hpt (Fig. 5C). However, the numbers of zGem+ cardiomyocytes did not show statistically significant reduction upon DM treatment in the whole hearts at different timepoints with the only differences observed in the atrial zGem+ signals at 48 hpt (38.1 ± 8.9 vs. 27.2 ± 2.8, N=11, 6, respectively) and 72 hpt (31.3 ± 8.5 vs. 17.3 ± 6.0, N=9, 6, respectively) (Fig. 5D, E). In short, these results suggested BMP and Notch signaling might have differential functions in cardiomyocyte proliferation.
Inhibition of BMP and Notch signaling pathways differentially suppresses FUCCI expression. (A) Time-lapse images of Tg(vmhc:mCherry-NTR; myl7:mAG-zGeminin) hearts showed differential patterns of zGem+ CMs in ablated, DAPT-treated ablated and DM-treated ablated groups at 0-72 hpt. Scale bars, 50 µm. hpt, hours post MTZ-treatment; CM, cardiomyocyte; DM, dorsomorphin. (B, C) Quantification of zGem+ CM numbers in the whole heart (B) or in the atrium only (C) of ablated and DAPT-treated ablated groups at 24, 48 and 72 hpt. Mean + SD, N=8-15 for each group, Student's t-test, *, P <0.05, ***, P < 0.001, ****, P < 0.0001. (D, E) Quantification of zGem+ CM numbers in the whole heart (D) or in the atrium only (E) of ablated and DM-treated ablated groups at 24, 48 and 72 hpt. Mean + SD, N=6-13 for each group, Student's t-test, *, P <0.05, **, P < 0.01.
BMP and Notch signaling pathways differentially influence cardiomyocyte cell-cycle progression
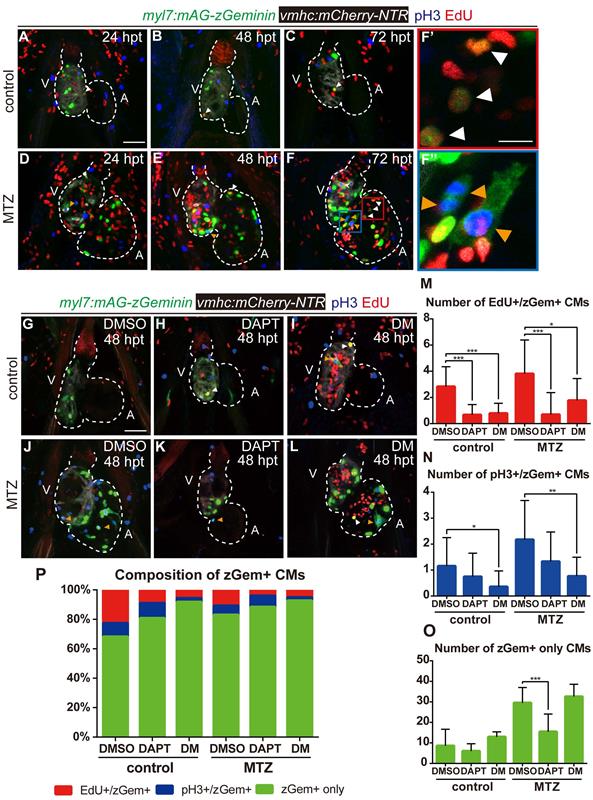
Previous studies reported that both BMP and Notch signaling regulated cardiomyocyte proliferation [29]. To resolve the discrepancy in the zGem+ signals observed above, we examined other proliferation markers, such as immunostaining of phospho-histone H3 (referred as pH3) and EdU pulsed labelling, in Tg(vmhc:mCherry-NTR; myl7:mAG-zGeminin) fish. The overlapped signals of pH3 and EdU with zGem+ could help to eliminate the interference caused by proliferative cells in adjacent tissues, such as endocardium, epicardium and blood cells. Proliferating cardiomyocytes were in different cell-cycle stages (Fig. 6A-F''), an EdU+/zGem+ cell represented a cardiomyocyte at S phase (white arrowheads), a pH3+/zGem+ cell represented a cardiomyocyte at M phase (orange arrowheads), while a zGem+ only cell without pH3 or EdU signals was considered in G2 phase. We then assessed the marker distribution in fish treated with DAPT or DM at 48 hpt (Fig. 6G-L) and quantified the numbers of proliferating cardiomyocytes in different cell-cycle phases (Fig. 6M-O). In terms of BMP signaling inhibition, DM treatment reduced the numbers of EdU+/zGem+ cells in control hearts (2.8 ± 1.5 vs. 0.8 ± 0.8, N=12, 14, respectively) and ablated hearts (3.8 ± 2.6 vs. 1.8 ± 1.7, N=16, 13, respectively). Similarly, the numbers of pH3+/zGem+ cells decreased in control hearts (1.2 ± 1.1 vs. 0.4 ± 0.6, N=13, 14, respectively) and ablated hearts (2.2 ± 1.5 vs. 0.8 ± 0.8, N=17, 17, respectively) upon DM treatment. However, the numbers of zGem+ only cells remained unchanged between groups with or without DM treatment. On the contrast, Notch signaling inhibition with DAPT treatment significantly reduced the number of zGem+ only cells in the ablated heart (29.5 ± 7.5 vs. 15.5 ± 8.6, N=18, 17, respectively), reflecting a significant reduction in the number of G2 phase cardiomyocytes. DAPT treatment also reduced the numbers of EdU+/zGem+ cells in control hearts (2.8 ± 1.5 vs. 0.7 ± 0.8, N=12, 15, respectively) and ablated hearts (3.8 ± 2.6 vs. 0.7 ± 1.7, N=16, 16, respectively). The reduction in the numbers of pH3+/zGem+ cells upon DAPT treatment was not statistically significant. We also plotted the data based on the proportions of marked cardiomyocytes in different cell-cycle phases (Fig. 6P). Reduced percentages of cardiomyocytes in S and M phases and increased percentage of G2 phase cardiomyocytes implied that DM treatment may cause cell-cycle arrest at G2 phase. Blocking Notch signaling pathway may possibly led to G0/G1 phase arrest because of the universally dwindled number of proliferative cardiomyocytes. Taken together, our results suggested that BMP and Notch signaling pathways regulated myocardium proliferation through differentially influence cardiomyocyte cell-cycle progression.
BMP and Notch signaling pathways differentially influence cardiomyocyte cell-cycle progression. (A-F'') Confocal stack projections of Tg(vmhc:mCherry-NTR; myl7:mAG-zGeminin) hearts with immunofluorescence of phospho-histone H3 and EdU showed proliferating CMs in different phases of cell cycle in control or ablated groups at 24, 48 and 72 hpt. (F', F”) magnified red and blue box areas in F. White arrowheads point to EdU+/ zGem+ CMs, orange arrowheads point to pH3+/ zGem+ CMs. (G-L) Confocal stack projections of Tg(vmhc:mCherry-NTR; myl7:mAG-zGeminin) hearts with immunofluorescence of phospho-histone H3 and EdU in control or ablated groups without or with DAPT or DM treatment at 48 hpt. White arrowheads point to EdU+/ zGem+ CMs, orange arrowheads point to pH3+/ zGem+ CMs. (M-P) Quantification of EdU+/zGem+ (M), pH3+/zGem+ (N), and zGem+ only (O) CM number in control or ablated groups without or with DAPT or DM treatment at 48 hpt. The respective proportions in total zGem+ CMs were shown in P. Mean + SD, N=12-18 for each group, Student's t-test, *, P < 0.05, **, P < 0.01, ***, P < 0.001. Scale bars, (A-L) 50 µm, (F', F”) 20 µm. hpt, hours post MTZ-treatment; A, atrium; V, ventricle; CM, cardiomyocyte; DM, dorsomorphin. Dashed lines outline the hearts.
Discussion
In this study we affirmed the upregulated expression of multiple BMP signaling components by WISH and the activation of BMP signaling in the myocardium and epicardium by pSmad1/5/9 immunostaining and Bre:dGFP reporter signals during larval ventricle regeneration. In zebrafish larvae, bmp10 has been reported to express in the endocardium, while bmp10l is expressed in the myocardium [38, 39]. The expression of bmp6 and id2b is endocardial specific during heart regeneration [40]. Wu et al. reported that bmp2b and bmp7 was expressed in the endocardium and epicardium of cryo-injured hearts, while pSmad1/5/9 signals were up-regulated in the myocardium [21]. Thus, we speculate that BMP signaling activation in myocardium may be promoted by enhanced ligand secretion from the endocardium after ventricle ablation, probably through interaction with other endocardial signaling pathways [26, 29, 41].
It is suggested that multiple signaling pathways eventually converge together and control cardiomyocyte cell cycle progression through certain function axes [42], implying the importance to understand the complex crosstalk among different signaling pathways. Cardiomyocyte proliferation in injured hearts with defective endocardial Notch signaling can be partially restored by WNT antagonist treatment, demonstrating the crosstalk between Notch and WNT/β-catenin signaling [43]. Moreover, endocardial Notch signaling promotes cardiomyocyte reprogramming and cardiac regeneration through activating myocardial Erbb2 and BMP signaling [29]. Our results proved that inhibition of Notch signaling by DAPT abolished the upregulated expression of ligand bmp10 and blocked BMP signaling activation. BMP pathway blockage by DM treatment did not reciprocally affect expression of notch1b nor signals of Notch signaling reporter, suggesting BMP signaling acts epistatic downstream of Notch signaling. Whether this regulation is cell autonomous or not requires further investigation. Our results showed a broader expression of bmp10 by WISH than the more confined Notch signaling revealed by tp1:dGFP fluorescence, suggesting at least certain cell non-autonomous mechanism is involved in this process. Interestingly, we also observed differential influence of Notch and BMP signaling on the re-activation of cardiogenic factors during regeneration. Bmp signaling exerts opposite effects on chamber differentiation during early heart development [44, 45], so one possible explanation is that BMP pathway regulates this process in a chamber-specific way by which the ventricle, instead of the atrium, is more susceptible. Another possibility is that the BMP pathway may not be directly involved in the regulation of cardiogenic factor re-activation, but rather that its inhibition on cardiomyocyte proliferation leads to a decrease in the number of ventricular cardiomyocytes as well as the expression level of cardiogenic factors. This is consistent with a previous report that BMP10 is responsible for cardiomyocyte proliferation while NRG1 regulates cardiomyocyte differentiation downstream of Notch signaling during mice ventricle trabeculation [27].
Many studies on the regulation of cell proliferation focus on the change in cell cycle progression [42, 46, 47]. Blocking Notch signaling pathway has been reported in several models to cause G0/G1 phase arrest by influencing p27Kip1 and other elements, thereby affecting proliferation [48-50]. On the contrary, few studies have been conducted on the mechanism of BMP pathway in regulating proliferative events. Overexpression of Id1 increases Cdk4 levels and reduces p21Clip1, thus promoting cell cycle progression in mouse cardiomyocytes [51]. BMP signaling responses to laminar shear stress by activation of p21 which leads to G2/M phase arrest of human bladder transitional carcinoma cells [52]. In our larval ventricle regeneration model, inhibition of BMP signaling reduced the numbers of EdU+/zGem+ and pH3+/zGem+ cardiomyocytes but had no effect on the number of zGem+ only cardiomyocytes, implying cell-cycle arrest at G2 phase. Blocking Notch signaling pathway may possibly led to G0/G1 phase arrest because of the universally dwindled number of proliferative cardiomyocytes. Our study provides novel insights into and potential direction for the understanding of cardiomyocyte proliferation in zebrafish ventricle regeneration process. The molecular mechanisms how these signaling pathways regulate proliferation, and whether it is conserved in different species and under different situations warrants further investigation.
Materials and methods
Zebrafish husbandry
Zebrafish were maintained under standard conditions [53]. Zebrafish embryos were raised at 28 °C and were staged according to Kimmel et al. [54]. All experiments were performed in accordance with institutional and national animal welfare guidelines. The transgenic lines used in this study were as follows: Tg(vmhc:mCherry-NTR), Tg(Bre:dGFP), Tg(tp1:dGFP), Tg(myl7:mAG-zGeminin).
In situ hybridization
Whole-mount in situ hybridization was performed as previously described [6], including notch1b, nkx2.5, gata4, tbx20. The probes for bmp10, bmpr1aa, id2b and smad1 were amplified by PCR with the following primers: bmp10 F-GCAGCCAGCAAGTAAGAGGA and R-GGTAGAGCAGGGAGATGGGA, bmpr1aa F-TAGCCAACCCCAATGCTTAC and R-GCCCATTTGTCTCGCAGGTAT, id2b F-TCGTGCCGAGTTTACCG and R-GCAATACCATACAGCTCCAGAT, smad1 F-CACGCTTCCGTAACCCACTCC and R-GCTCAAACATTCGGCATACACCT.
Chemical treatment
Tg(vmhc:mCherry-NTR) larvae were treated with 6 mM MTZ (metronidazole, Sigma) in E3 water at 3 dpf for four hours as previously described [6]. After washing with fresh E3 water for three times, the larvae were then incubated in 100 μM DAPT (Sigma-Aldrich), 7.5 μM Dorsomorphin (Sigma-Aldrich) or 1% DMSO (dimethyl sulfoxide, Thermo Fisher Scientific) as control for 0-2 dpt or other time frames as indicated.
EdU labeling
The larvae were incubated with 500 μM EdU for 1 h in E3 water with 2% DMSO to facilitate EdU solubilization. After pulse labeling, larvae were rinsed with E3 water, anaesthetized with 0.2% tricaine and fixed in 4% PFA. The CLICK-IT reaction for EdU labeling was performed according to the manufacturer's instruction (Thermo Fisher Scientific).
Immunostaining
Immunofluorescence staining on whole-mount larvae was performed as previously described [6], using the following primary antibodies: anti-phospho-histone H3(Ser10) (rabbit; Merck Millipore, 06570), anti-phospho-Smad1/5/9 (rabbit; CST, 13820), anti-MHC (mouse; DSHB, MF20). The secondary antibodies used were Alexa Fluor 405 goat anti-rabbit IgG, Alexa Fluor 488 goat anti-rabbit IgG and Alexa Fluor 555 goat anti-mouse IgG from Thermo Fisher Scientific.
Imaging
Live imaging and immunostaining images were obtained using a Zeiss LSM880 confocal microscope or an Olympus IX83 inverted microscope. The numbers of differentially marked cardiomyocytes were counted with ZEN software or cellSens software.
Statistical analysis
Values were presented as mean ± SD. Statistical significance was defined as a threshold of P < 0.05 determined by Student's t-test between two groups, ANOVA analysis between more than two groups or Binomial test in quantification of the percentage of recovered hearts.
Abbreviations
A: atrium; CM: cardiomyocyte; Ctrl: control; DM: Dorsomorphin; dpf: days post fertilization; dpt: days post MTZ-treatment; FUCCI: fluorescent ubiquitylation-based cell cycle indicator; hpt: hours post MTZ-treatment; MI: myocardial infarction; MTZ: metronidazole; pH3: phospho-histone H3; pSmad1/5/9: phospho-Smad1/Smad5/Smad9; V: ventricle; WISH: whole-mount in situ hybridization.
Acknowledgements
We thank Haitao Zhou and Lifeng Li for fish care and other lab members for in depth discussion.
Funding
This study was supported by National Key R&D Program of China grants 2020YFA0803900 and 2018YFA0801000 to R.Z.
Author Contributions
W.W, Y.H, Y.P and R.Z conceived and designed the project. W.W, Y.H, C.Y, Q.L and Y.P conducted the experiments. M.P, N.C and J.X provided critical reagents. W.W and R.Z wrote and revised the manuscript. All authors reviewed the manuscript.
Ethics statement
All procedures involving animals were approved by Fudan University Institutional Animal Care and Use Committee (IACUC) and in accordance with the NIH Guide for the care and use of Laboratory Animals.
Competing Interests
The authors have declared that no competing interest exists.
References
1. Thygesen K, Alpert JS, White HD. Universal definition of myocardial infarction. Journal of the American College of Cardiology. 2007;50:2173-95
2. Virani SS, Alonso A, Benjamin EJ, Bittencourt MS, Callaway CW, Carson AP. et al. Heart Disease and Stroke Statistics-2020 Update: A Report From the American Heart Association. Circulation. 2020;141:e139-e596
3. Porrello ER, Mahmoud AI, Simpson E, Hill JA, Richardson JA, Olson EN. et al. Transient regenerative potential of the neonatal mouse heart. Science. 2011;331:1078-80
4. Poss KD, Wilson LG, Keating MT. Heart regeneration in zebrafish. Science. 2002;298:2188-90
5. Gonzalez-Rosa JM, Martin V, Peralta M, Torres M, Mercader N. Extensive scar formation and regression during heart regeneration after cryoinjury in zebrafish. Development. 2011;138:1663-74
6. Zhang R, Han P, Yang H, Ouyang K, Lee D, Lin YF. et al. In vivo cardiac reprogramming contributes to zebrafish heart regeneration. Nature. 2013;498:497-501
7. Jopling C, Sleep E, Raya M, Marti M, Raya A, Izpisua Belmonte JC. Zebrafish heart regeneration occurs by cardiomyocyte dedifferentiation and proliferation. Nature. 2010;464:606-9
8. Kikuchi K, Holdway JE, Werdich AA, Anderson RM, Fang Y, Egnaczyk GF. et al. Primary contribution to zebrafish heart regeneration by gata4(+) cardiomyocytes. Nature. 2010;464:601-5
9. Bergmann O, Bhardwaj RD, Bernard S, Zdunek S, Barnabe-Heider F, Walsh S. et al. Evidence for cardiomyocyte renewal in humans. Science. 2009;324:98-102
10. Bergmann O, Zdunek S, Felker A, Salehpour M, Alkass K, Bernard S. et al. Dynamics of Cell Generation and Turnover in the Human Heart. Cell. 2015;161:1566-75
11. Zhen YS, Wu Q, Xiao CL, Chang NN, Wang X, Lei L. et al. Overlapping cardiac programs in heart development and regeneration. J Genet Genomics. 2012;39:443-9
12. Cai WF, Liu GS, Wang L, Paul C, Wen ZL, Wang Y. Repair Injured Heart by Regulating Cardiac Regenerative Signals. Stem Cells Int. 2016;2016:6193419
13. Fernandez CE, Bakovic M, Karra R. Endothelial Contributions to Zebrafish Heart Regeneration. J Cardiovasc Dev Dis. 2018 5
14. van Wijk B, Moorman AFM, van den Hoff MJB. Role of bone morphogenetic proteins in cardiac differentiation. Cardiovasc Res. 2007;74:244-55
15. Morrell NW, Bloch DB, ten Dijke P, Goumans MJTH, Hata A, Smith J. et al. Targeting BMP signalling in cardiovascular disease and anaemia. Nat Rev Cardiol. 2016;13:106-20
16. Yang YL, Liu YS, Chuang LY, Guh JY, Lee TC, Liao TN. et al. Bone Morphogenetic Protein-2 Antagonizes Renal Interstitial Fibrosis by Promoting Catabolism of Type I Transforming Growth Factor-beta Receptors. Endocrinology. 2009;150:727-40
17. Ebelt H, Hillebrand I, Arlt S, Zhang Y, Kostin S, Neuhaus H. et al. Treatment with Bone Morphogenetic Protein 2 Limits Infarct Size after Myocardial Infarction in Mice. Shock. 2013;39:353-60
18. Sun B, Huo R, Sheng Y, Li Y, Xie X, Chen C. et al. Bone morphogenetic protein-4 mediates cardiac hypertrophy, apoptosis, and fibrosis in experimentally pathological cardiac hypertrophy. Hypertension. 2013;61:352-60
19. Jin YL, Cheng XY, Lu JP, Li X. Exogenous BMP-7 Facilitates the Recovery of Cardiac Function after Acute Myocardial Infarction through Counteracting TGF-beta 1 Signaling Pathway. Tohoku J Exp Med. 2018;244:1-6
20. Sun LJ, Yu J, Qi S, Hao YW, Liu Y, Li ZW. Bone Morphogenetic Protein-10 Induces Cardiomyocyte Proliferation and Improves Cardiac Function After Myocardial Infarction. J Cell Biochem. 2014;115:1868-76
21. Wu CC, Kruse F, Vasudevarao MD, Junker JP, Zebrowski DC, Fischer K. et al. Spatially Resolved Genome-wide Transcriptional Profiling Identifies BMP Signaling as Essential Regulator of Zebrafish Cardiomyocyte Regeneration. Dev Cell. 2016;36:36-49
22. MacGrogan D, Munch J, de la Pompa JL. Notch and interacting signalling pathways in cardiac development, disease, and regeneration. Nat Rev Cardiol. 2018;15:685-704
23. Li H, Chang C, Li X, Zhang R. The roles and activation of endocardial Notch signaling in heart regeneration. Cell Regen. 2021;10:3
24. Raya A, Koth CM, Buscher D, Kawakami Y, Itoh T, Raya RM. et al. Activation of Notch signaling pathway precedes heart regeneration in zebrafish. P Natl Acad Sci USA. 2003;100:11889-95
25. Zhao L, Borikova AL, Ben-Yair R, Guner-Ataman B, MacRae CA, Lee RT. et al. Notch signaling regulates cardiomyocyte proliferation during zebrafish heart regeneration. P Natl Acad Sci USA. 2014;111:1403-8
26. Munch J, Grivas D, Gonzalez-Rajal A, Torregrosa-Carrion R, de la Pompa L. Notch signalling restricts inflammation and serpine1 expression in the dynamic endocardium of the regenerating zebrafish heart. Development. 2017;144:1425-40
27. Grego-Bessa J, Luna-Zurita L, del Monte G, Bolos V, Melgar P, Arandilla A. et al. Notch signaling is essential for ventricular chamber development. Dev Cell. 2007;12:415-29
28. Luxan G, D'Amato G, MacGrogan D, de la Pompa JL. Endocardial Notch Signaling in Cardiac Development and Disease. Circ Res. 2016;118:E1-E18
29. Gálvez-Santisteban M, Chen D, Zhang R, Serrano R, Nguyen C, Zhao L. et al. Hemodynamic-mediated endocardial signaling controls in vivo myocardial reprogramming. Elife. 2019;8:e44816
30. Collery RF, Link BA. Dynamic smad-mediated BMP signaling revealed through transgenic zebrafish. Dev Dyn. 2011;240:712-22
31. Han P, Bloomekatz J, Ren J, Zhang R, Grinstein JD, Zhao L. et al. Coordinating cardiomyocyte interactions to direct ventricular chamber morphogenesis. Nature. 2016;534:700-4
32. Li X, Lu Q, Peng Y, Geng F, Shao X, Zhou H. et al. Primary cilia mediate Klf2-dependant Notch activation in regenerating heart. Protein Cell. 2020;11:433-45
33. Matrone G, Tucker CS, Denvir MA. Cardiomyocyte proliferation in zebrafish and mammals: lessons for human disease. Cell Mol Life Sci. 2017;74:1367-78
34. Foglia MJ, Poss KD. Building and re-building the heart by cardiomyocyte proliferation. Development. 2016;143:729-40
35. Sakaue-Sawano A, Kurokawa H, Morimura T, Hanyu A, Hama H, Osawa H. et al. Visualizing spatiotemporal dynamics of multicellular cell-cycle progression. Cell. 2008;132:487-98
36. Sugiyama M, Sakaue-Sawano A, Iimura T, Fukami K, Kitaguchi T, Kawakami K. et al. Illuminating cell-cycle progression in the developing zebrafish embryo. P Natl Acad Sci USA. 2009;106:20812-7
37. Choi WY, Gemberling M, Wang J, Holdway JE, Shen MC, Karlstrom RO. et al. In vivo monitoring of cardiomyocyte proliferation to identify chemical modifiers of heart regeneration. Development. 2013;140:660-6
38. Capasso TL, Li BJ, Volek HJ, Khalid W, Rochon ER, Anbalagan A. et al. BMP10-mediated ALK1 signaling is continuously required for vascular development and maintenance. Angiogenesis. 2020;23:203-20
39. Laux DW, Young S, Donovan JP, Mansfield CJ, Upton PD, Roman BL. Circulating Bmp10 acts through endothelial Alk1 to mediate flow-dependent arterial quiescence. Development. 2013;140:3403-12
40. Fang Y, Lai KS, She P, Sun J, Tao W, Zhong TP. Tbx20 Induction Promotes Zebrafish Heart Regeneration by Inducing Cardiomyocyte Dedifferentiation and Endocardial Expansion. Front Cell Dev Biol. 2020;8:738
41. Kikuchi K, Holdway JE, Major RJ, Blum N, Dahn RD, Begemann G. et al. Retinoic Acid Production by Endocardium and Epicardium Is an Injury Response Essential for Zebrafish Heart Regeneration. Dev Cell. 2011;20:397-404
42. Hashmi S, Ahmad HR. Molecular switch model for cardiomyocyte proliferation. Cell Regen. 2019;8:12-20
43. Zhao L, Ben-Yair R, Burns CE, Burns CG. Endocardial Notch Signaling Promotes Cardiomyocyte Proliferation in the Regenerating Zebrafish Heart through Wnt Pathway Antagonism. Cell Rep. 2019;26:546-54
44. Marques SR, Yelon D. Differential requirement for BMP signaling in atrial and ventricular lineages establishes cardiac chamber proportionality. Dev Biol. 2009;328:472-82
45. de Pater E, Ciampricotti M, Priller F, Veerkamp J, Strate I, Smith K. et al. Bmp signaling exerts opposite effects on cardiac differentiation. Circ Res. 2012;110:578-87
46. Muralidhar SA, Sadek HA. Meis1 Regulates Postnatal Cardiomyocyte Cell Cycle Arrest. In: Nakanishi T, Markwald RR, Baldwin HS, Keller BB, Srivastava D, Yamagishi H, editors. Etiology and Morphogenesis of Congenital Heart Disease: From Gene Function and Cellular Interaction to Morphology. Tokyo. 2016 p. 93-101
47. Ponnusamy M, Li PF, Wang K. Understanding cardiomyocyte proliferation: an insight into cell cycle activity. Cell Mol Life Sci. 2017;74:1019-34
48. Wu JR, Yeh JL, Liou SF, Dai ZK, Wu BN, Hsu JH. Gamma-secretase Inhibitor Prevents Proliferation and Migration of Ductus Arteriosus Smooth Muscle Cells through the Notch3-HES1/2/5 Pathway. Int J Biol Sci. 2016;12:1063-73
49. Niapour A, Hamidabadi HG, Niapour N, Mohammadi P, Pasandi MS, Malekzadeh V. Pharmacological Notch pathway inhibition leads to cell cycle arrest and stimulates ascl1 and neurogenin2 genes expression in dental pulp stem cells-derived neurospheres. Biotechnol Lett. 2019;41:873-87
50. Velaithan V, Okuda KS, Ng MF, Samat N, Leong SW, Faudzi SMM. et al. Zebrafish phenotypic screen identifies novel Notch antagonists (vol 35, pg 166, 2017). Invest New Drug. 2017;35:250 -
51. Cheng YY, Yan YT, Lundy DJ, Lo AHA, Wang YP, Ruan SC. et al. Reprogramming-derived gene cocktail increases cardiomyocyte proliferation for heart regeneration. Embo Mol Med. 2017;9:251-64
52. Lee YH, Lai CW, Cheng YC. Fluid Shear Stress Induces Cell Cycle Arrest in Human Urinary Bladder Transitional Cell Carcinoma Through Bone Morphogenetic Protein Receptor-Smad1/5 Pathway. Cell Mol Bioeng. 2018;11:185-95
53. Westerfield M. The Zebrafish Book. A Guide for the Laboratory Use of Zebrafish (Danio rerio), 5th Edition. University of Oregon Press, Eugene. 2007
54. Kimmel CB, Ballard WW, Kimmel SR, Ullmann B, Schilling TF. Stages of Embryonic-Development of the Zebrafish. Dev Dynam. 1995;203:253-310
Author contact
Corresponding authors: Y. Peng, E-mail: yuanyuanpeng16edu.cn; R. Zhang, E-mail: zhangruilinedu.cn.

 Global reach, higher impact
Global reach, higher impact