Impact Factor ISSN: 1449-2288
Int J Biol Sci 2021; 17(9):2308-2322. doi:10.7150/ijbs.59876 This issue Cite
Review
miR-223: An Effective Regulator of Immune Cell Differentiation and Inflammation
1. School of Agriculture, Ningxia University, Yinchuan 750021, China.
2. Key Laboratory of Ruminant Molecular Cell Breeding, Ningxia Hui Autonomous Region, Yinchuan 750021, China.
#These authors contributed equally to this work.
Received 2021-2-27; Accepted 2021-5-21; Published 2021-6-4
Abstract
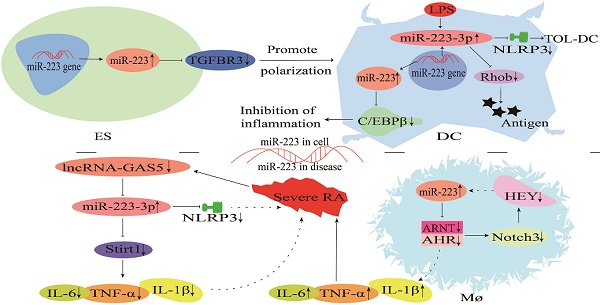
MicroRNAs (miRNAs) play a critical role in regulating various biological processes, such as cell differentiation and immune modulation by binding to their target genes. miR-223 is a miRNA with important functions and has been widely investigated in recent years. Under certain physiological conditions, miR-223 is regulated by different transcription factors, including sirtuin1 (Sirt1), PU.1 and Mef2c, and its biological functions are mediated through changes in its cellular or tissue expression. This review paper summarizes miR-223 biosynthesis and its regulatory role in the differentiation of granulocytes, dendritic cells (DCs) and lymphocytes, macrophage polarization, and endothelial and epithelial inflammation. In addition, it describes the molecular mechanisms of miR-223 in regulating lung inflammation, rheumatoid arthritis, enteritis, neuroinflammation and mastitis to provide insights into the existing molecular regulatory networks and therapies for inflammatory diseases in humans and animals.
Keywords: miR-223, Cellular inflammation, Inflammatory diseases, Immunity, Molecular regulatory network
Introduction
Inflammatory diseases are commonly caused by pathogen infection or external factors (e.g., trauma) [1]. Chronic inflammation could result in metabolic disorders, organ damages, and severe inflammatory diseases that could even cause human or animal death. Therefore, exploring different molecular mechanisms on the development and progression of inflammatory diseases could improve the breeding of new animal varieties with strong disease resistance. Besides, it could help develop early diagnostic tools and biological or chemical therapies for human inflammatory diseases. Notably, many factors could trigger inflammatory diseases. Recent studies have shown that the development and progression of inflammatory diseases are very complex and are regulated by a molecular network involving multiple genes or proteins [2, 3]. miRNAs are a newly identified class of non-coding RNAs that could influence immune responses, cancer development, and cell proliferation, differentiation, and apoptosis by regulating the expression of target mRNAs [2]. Monitoring their level is critical for early disease diagnosis and prognostic observation [3]. Many researchers have screened and identified inflammation-related miRNAs, and conducted their functional analyses in human and animal cells and tissue inflammation. Of these miRNAs, miR-223 has been identified to exhibit multiple regulatory functions during inflammation [4]. This review paper has discussed the modulatory effects of miR-233 in the differentiation of multiple cell types and inflammation. Also, it provides a systematic overview of the molecular mechanisms of miR-223 in regulating lung inflammation, rheumatoid arthritis, enteritis, neuroinflammation and mastitis to offer new insights into treatment development for inflammatory diseases in humans and animals.
Biosynthesis of miR-223
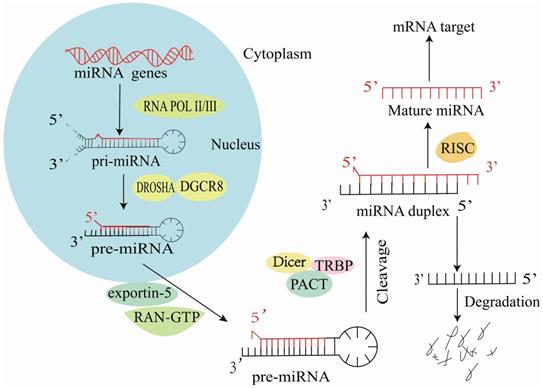
Mature miRNAs are short non-coding RNA molecules comprising 19-25 nucleotides [5]. In the nucleus, RNA polymerase II synthesizes most miRNAs into primary miRNAs (pri-miRNAs) [6] and most miRNAs are subsequently cleaved by Drosha RNase III into 70-80 nt long hairpin precursor miRNAs (pre-miRNAs) [7]. Next, exportin-5 transports pre-miRNAs into the cytosol, where they are cleaved by the endoribonuclease Dicer [8] to form double-stranded RNA molecules. These double-stranded RNA molecules are modified by the RNA-induced silencing complex (RISC), and one strand is retained to target and regulate mRNA expression (Figure 1) [9].
Synthesis of miR-223 in cells.
miR-223 is an important member of the miRNA family, first identified by quantitative polymerase chain reaction (qPCR) in 2003 [10]. Its genes are located in the X chromosome of humans, mice and cows. Under certain physiological conditions, miR-223 expression is promoted by different transcription factors, C/EBPα, PU.1 and C/EBPβ [11, 12] and inhibited by nuclear factor I A (NFIA), Mef2c and KLF6 [13-17], and the importin-α4 and importin-α5 transporters [18].
miR-223 expression during cellular and tissue inflammation
Several studies have been conducted to understand the miR-223 functions that determine its cellular and tissue expression [13, 15, 19-22]. Based on these studies, miR-223 expression is altered during the inflammatory response of various cell types, including granulocytes, macrophages, dendritic cells (DCs), T cells, endothelial cells and epithelial cells. This change in miR-223 expression regulates the various functions of these cells and attenuates or exacerbates the associated tissue inflammation (Table 1).
Change in miR-223 expression during the inflammation of various cell types
| Cell type | miR-223 expression | Affected tissue | Reference |
|---|---|---|---|
| Granulocytes | Significantly up-regulated | Lung tissue | [13, 23] |
| Macrophages | Significantly down-regulated | Fat and muscle tissues | [15, 24, 25] |
| Dendritic cells | Down-regulated | Small intestinal tissue | [19, 26] |
| T cells | Highly expressed | Nerve tissues | [20,27,28] |
| Endothelial cells | Mass delivery via microvesicles highly expressed | Tissues supplied by arteries | [21, 29] |
| Epithelial cells | Up-regulated | Kidney tissues | [22, 30] |
Molecular mechanism of action of miR-223 at the cellular level
miR-233 regulates the differentiation and proliferation of granulocytes, macrophages and DCs by binding to specific targets. In addition, miR-223 regulates pro-inflammatory or anti-inflammatory macrophage polarization. miR-223 can also bind specific target genes to inhibit pro-inflammatory cytokines or inflammatory signals in these cells (Table 2).
Intracellular targets and functions of miR-223
| Cell | Target | Function | Reference |
|---|---|---|---|
| Granulocytes | NFIA, C/EBPα, and Mef2c | Regulates granulocyte proliferation, and differentiation | [11, 13, 14] |
| Macrophages | STATS, Pknox1, and TRAF6 | Regulates macrophage differentiation, polarization, and pro-inflammatory cytokine release, and promotes NF-kB-induced inflammatory injury | [31-33] |
| Dendritic cells | NLRP3, C/EBPβ, TGFBR3, MR, Rhob, Rasa1, Cfla, and Kras | Regulates dendritic cell functions and influences immune-related protein networks | [19, 26, 34-36] |
| Endothelial cells | importin-α4, importin-α5, NLRP3, and IL-6 | Attenuates endothelial cell injury | [18, 37, 38] |
| Epithelial cells | NLRP3, Sirt1, and ST1M1 | Attenuates epithelial cell injury | [30, 39, 40] |
Role of miR-223 in granulocyte differentiation
miR-223 plays a critical role in the differentiation and activation of granulocytes. According to Johnnidis et al. [13] miR-223 could target the myocyte enhancer factor 2C (Mef2c) to regulate neutrophil progenitor proliferation and granulocyte differentiation and activation in mice. miR-223-deficient mice had hyperactive granulocytes that were highly sensitive to activating stimuli [13]. Additionally, the lipopolysaccharide (LPS) challenge resulted in rapid accumulation of endotoxin, and the inflammatory lung pathology induction was characterized by excessive lung tissue injury [13]. During Group B Streptococcus (GBS)-induced lung inflammation, miR-223 was rapidly up-regulated in lung-infiltrating granulocytes at 3-6 h post-GBS infection and this attenuated lung tissue injury [41].
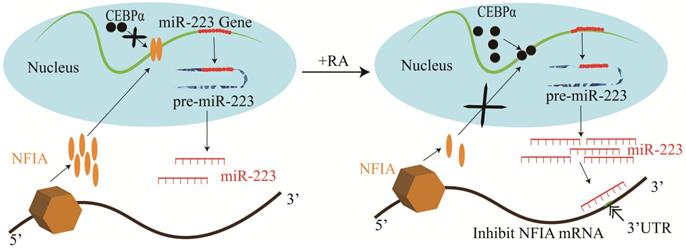
Studies of human hematopoietic progenitor cell (HPC) differentiation have revealed that miR-223 is significantly up-regulated by the myeloid transcription factors, PU.1 and C/EBPβ during HPC differentiation into granulocytes and monocytopoiesis [12, 23]. However, during erythropoiesis, they are expressed at a low level [23]. miR-223 overexpression has been shown to increase granulocytopoiesis, whereas it impairs erythropoiesis and monocyte-macrophage differentiation [23]. Other studies have discovered that NFIA could bind to the miR-223 gene promoter and repress its expression during granulocyte differentiation [11, 17]. However, retinoic acid triggers C/EBPα to bind to the miR-223 promoter competitively and up-regulates miR-223 expression, inhibiting NFIA expression in a targeted manner and promotes granulocyte differentiation (Figure 2) [11, 14].
Mechanisms of miR-223 in the regulation of granulocyte differentiation.
Role of miR-223 in macrophage polarization
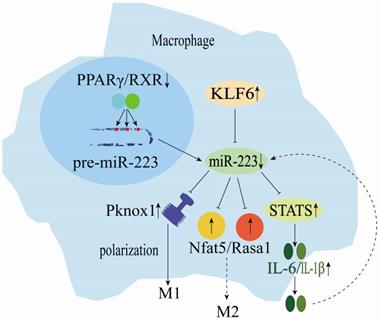
miRNAs are key regulators of various biological processes and have regulated macrophage (Mø) polarization and promoted inflammatory activities. miR-223 is significantly down-regulated during human monocyte-macrophage differentiation [25]. On the other hand, macrophage (M1)-mediated inflammation in adipose and muscle tissues could cause low-grade systemic inflammation development. Macrophages are vital coordinators of immune activity and homeostasis. They could change polarization direction based on temporal and environmental cues and play a central role in promoting host immune defense mechanisms [24]. Furthermore, PPARγ regulates the miR-223 expression by directly binding onto PPARγ regulatory elements (PPREs) in the pre-miR-223 promoter (Figure 3) [42]. Rasa1 and NFAT5 real targets of miR-223 play a crucial role in controlling selective macrophage activation (Figure 3) [42]. The miR-223 expression could induce the polarization of inflammatory macrophages (M1), as its down-regulation in macrophages reduces the inhibition of STAT genes, promoting the release of LPS-induced interleukin 6 (IL-6) and IL-1β. These cytokines can regulate miR-223 expression negatively and ultimately promote muscle tissue inflammation exacerbation and injury (Figure 3) [31, 43, 44]. Kruppel-like factor 6 (KLF6) has been identified as a new transcription factor involved in macrophage polarization (Figure 3) [16]. KLF6 inhibits miR-223 expression by occupying the miR-223 promoter, and KLF6 over-expression has been shown to down-regulate miR-223 expression in macrophages. Furthermore, KLF6-mediated the miR-223 down-regulation in macrophages and has been reported to promote adipose tissue inflammation [15]. Moreover, low lncRNA MEG3 expression inhibits M1 macrophage polarization, whereas its deletion could up-regulate miR-223 expression and promote M2 macrophage polarization. High miR-223 expression inhibits TNF receptor-associated factor 6 (TRAF6), suppressing the NF-κB signaling pathway and alleviating myocarditis-associated injury [33]. Zhuang et al. [32] showed that miR-223 overexpression could prevent diet-induced adipose tissue inflammation and systemic insulin resistance by inhibiting the Pknox1 gene expression in mice (Figure 3). However, macrophages could use microvesicles (MVs) to deliver miR-223, which exerts specific functions in the target cells [45]. In summary, miR-223 is a key regulator of the dynamic balance between M1/M2 macrophages and inflammatory diseases.
Mechanisms of miR-223 in the regulation of macrophage differentiation.
Role of miR-223 in dendritic cell differentiation
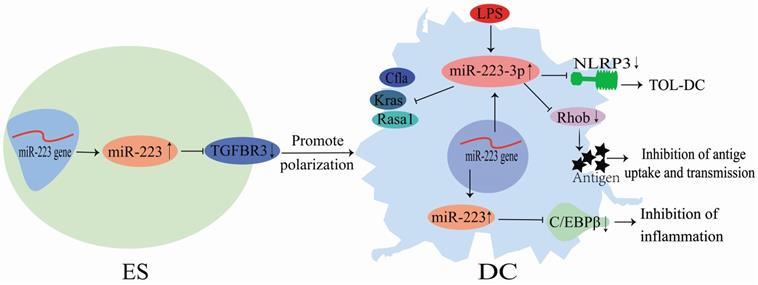
Although miR-223 does not directly act on DCs, it could regulate DC differentiation via several pathways (Figure 4). During the differentiation of mouse HSCs into DCs, the miR-223 expression is altered in HSCs, myeloid stem cells and DCs, indicating that miR-223 could play a role in DC differentiation [46]. LPS stimulation could up-regulate miR-223-3p expression in DCs, and its high expression could subsequently down-regulate Rasa1, Cfla and Kras mRNA expression and influence immune-related protein regulatory networks [19]. Also, miR-223-3p could regulate DC differentiation by binding to Rhob and inhibiting antigen uptake and presentation by DCs [36]. Chen et al. [35] showed that miR-223-3p expression was significantly lower in mice with autoimmune myocarditis than in normal mice. Consequently, miR-223-3p could inhibit the NLR family pyrin domain containing 3 (NLRP3) inflammasome to promote the polarization of tolerogenic DCs [35]. Furthermore, it could regulate the differentiation and function of mice and human intestinal DCs by targeting C/EBPβ and reducing inflammatory injury (Figure 4) [26]. Zhu et al. [34] demonstrated that miR-223 directly targeted TGFBR3 to promote the human embryonic stem cells (ESCs) differentiation into DCs. These findings demonstrate that miR-223 is an essential regulator of DC differentiation. It can regulate DC polarization and functions through binding to specific targets, improving tissue inflammation, and preventing the development of inflammatory diseases (Figure 4).
Mechanisms of miR-223 in the regulation of dendritic cell differentiation.
Role of miR-223 in T cell-mediated inflammation
Mature T cells travel through the blood, and reside and proliferate in the T cell zone of peripheral tissues. These cells can be circulated through the body via lymphatics, peripheral blood, and tissue fluids to exert their cellular immune functions. Hosseini et al. [28] showed that miR-223 expression was up-regulated in CD4+ T cells during multiple sclerosis. Besides, they could modulate chemokine signaling to promote T helper 17 (Th17) cell expression and suppress regulatory T cell (Treg) differentiation, highlighting a potential miR-223 role in maintaining the Th17/Treg balance. Another study demonstrated that miR-233 is highly expressed in immature CD4+ T cells and participates in the proliferation and differentiation of these cells during rheumatoid arthritis [47]. Moreover, miR-233 could regulate myeloid DCs (mDCs) to activate and promote the pathological Th17 cells differentiation during autoimmunity [20]. Furthermore, miR-223 promotes T helper 1 (Th1) and Th17 cell differentiation and the experimental autoimmune encephalomyelitis (EAE) progression, and its deficiency prevents the infiltration of Th1 and Th17 cells into the spinal cord [27]. Additionally, the interaction between miR-223 and SRY-box 11 (SOX11) has gained importance in research towards treating Mantle Cell Lymphoma (MCL) and inflammatory diseases [48]. In chicken T cells, both Marek's disease (MD) and overexpressed lnc-GALMD3 result in low miR-223 expression, leading to malignant T cell proliferation [49,50]. Taken together, miR-233 regulates T cell proliferation and differentiation and modulates inflammatory diseases by promoting helper T cell proliferation.
Role of miR-223 in endothelial inflammation
Endothelial MVs play a vital role in treating numerous cardiovascular diseases, including atherosclerosis (AS) [21, 51-54]. Moreover, in horse, miR-223 in MV participates in immune system regulation by modulating inter leukocyte signaling and inflammatory processes [55]. Thrombopoietin could stimulate platelets to release numerous miR-223-expressing MVs [29]. Similarly, miR-223 up-regulation has been detected in peripheral MVs (P-MV) in the plasma samples of enteritis, hepatitis, nephritis, and AS patients [29]. Platelet-derived miR-223 could be delivered to human umbilical vascular endothelial cells (HUVECs) through P-MVs. Also, it down-regulates the insulin-like growth factor 1 (IGF-1R) expression, promoting HUVEC apoptosis induced by advanced glycation end products (Figure 5) [29]. Li et al. [56] showed that P-MV-derived miR-223 could inhibit NF-κB and MAPK signaling pathways. Besides, it down-regulates ICAM-1 expression in HUVECs, demonstrating that miR-223 is a critical factor in platelet-derived exosomes that plays essential roles during inflammation and AS (Figure 5). Bao et al. [18] demonstrated that IL-6 expression in glomerular endothelial cells (GEnCs) of immunoglobulin A nephropathy (IgAN) patients could induce miR-223 down-regulation. Subsequently, they could promote its binding to importin-α4 and importin-α5, activating GEnCs, and inhibiting the nuclear translocation of P56 and STAT3 (Figure 5). STAT3 is critical during the induction of IL-6 in HUVECs, and its expression is positively correlated with IL-6 expression [57], indicating that IL-6/miR-223/importin-α4 (-α5)/STAT3 constitutes a feedback regulatory network in endothelial cells (Figure 5).
Mechanisms of miR-223 in the regulation of endothelial cells.
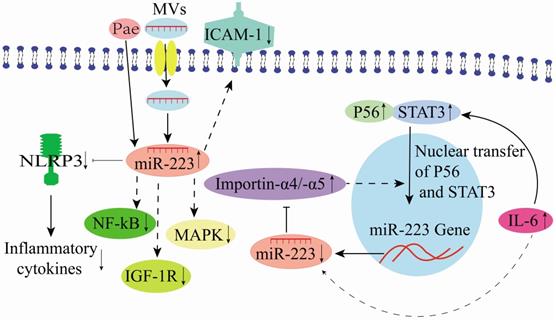
Intriguingly, the Chinese medicine tree peony bark (Pae) could be a potential therapeutic agent for AS. It could increase monocyte exosome-derived miR-223 [57]. Additionally, Pae could increase the miR-223 expression level in exosomes derived from the plasma of hyperlipidemic rats to inhibit the NLRP3 inflammasome pathway in endothelial cells, supporting its therapeutic potentials in AS [37, 57]. In an endothelial cell study in pigs, miR-223 targeting NLRP3 alleviated inflammation development in porcine endothelial cells and triggered the aorta inflammatory injury [58]. Moreover, miR-223 could target β1 integrin to prevent endothelial cell proliferation [59]. Differential miR-223 expression in vascular endothelial cell (VEC) MVs influences VEC generation and apoptosis that impacts the functioning of other cells when delivered into the surrounding through VEC-MVs. In contrast, the decrease in miR-223 expression in plasma-derived MVs could attenuate VEC apoptosis [60]. Previously, different studies have shown that miR-223-3p is an important regulator of vascular endothelial injury in Kawasaki Disease (KD) [38, 61, 62]. It could further reduce vascular endothelial injury by inhibiting IL-6 and TNF-α production [38], and its overexpression could reduce the apoptotic rate of VECs [63]. Therefore, there is evidence that some therapeutic agents could modulate miR-223 expression to attenuate endothelial cell injury (Figure 5).
Role of miR-223 in epithelial inflammation
miR-223 plays essential functions in epithelial cells. For instance, it could control the growth and morphology of mammary epithelial cells [64]. Bio-informatic analysis has revealed that NLRP3 is a target of miR-223 and could directly act on renal tubular epithelial cells (RTECs) to induce renal tissue injury in mice [30]. Sun et al. [22] reported that in LPS-treated RTECs, baicalin could up-regulate miR-223-3p to inhibit TXNIP and NLRP3 gene expression, resulting in attenuated LPS-induced injury in the proximal tubule epithelial cells (HK-2 cell line) (Figure 6). Sirt1 is a direct target of miR-223 in HK2 cells. Notably, low expression of lncRNA TUG1 could increase miR-223 expression, which negatively regulates Sirt1 and reduces PI3K and AKT phosphorylation [39]. Also, low expression of lncRNA TUG1 activates the NF-κB signaling pathway and protects HK-2 cells from LPS-induced injury (Figure 6) [39]. In an in vitro model of endometritis, Zhao et al. [65] discovered that LPS treatment could activate the NF-κB signaling that promotes the miR-223 up-regulation in bovine endometrial epithelial cells (BEND). Hence, inhibiting NLRP3 activation and IL-1β production to prevent inflammatory-induced damage (Figure 6). Therefore, miR-223 restricts NLRP3 activation and acts as a protective factor during inflammatory responses [65]. Liu et al. [40] illustrated that the stromal interaction molecule 1 (STIM1) regulated NLRP3 expression by binding the AACUGA motif in miR-223. On the other hand, silencing STIM1 alleviates influenza A virus (IAV)-induced inflammation in lung epithelial cells by inactivating the NLRP3 and inflammasome by promoting the miR-223 expression [40].
Mechanisms of miR-223 in the regulation of epithelial inflammation.
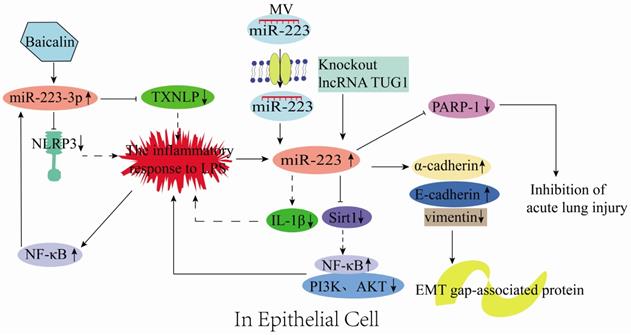
Vesicles are the main structures for information transfer between cells, and 80% of them are derived from epithelial cells. miRNAs are the main components in vesicles and, are transferred between cells along with the vesicles. However, miR-223 in mouse neutrophils could be transferred through MVs to lung epithelial cells and inhibits PARP-1 to prevent acute lung injury (Figure 6) [66]. In addition, inflammatory mediators could stimulate various signaling pathways to induce the expression of epithelial-mesenchymal transition-related transcription factors (EMT-TFs; e.g., Snail, Zeb, and Twist) and epigenetic regulators (e.g., miRNA, DNA, and histone-modifying enzymes). The expression of miRNA- or DNA-based epigenetic regulators could regulate the expression of genes related to these inflammation signaling pathways, ultimately forming a gene regulatory network [67, 68]. Tang et al. [69] reported that miR-223 inhibited cancer cell metastasis by regulating the EMT-related protein expression, which involves up-regulating the epithelial markers (E-cadherin and α-cadherin) and down-regulating the mesenchymal marker vimentin (Figure 6). Briefly, the miR-223 expression is up-regulated in epithelial cells, produced by neutrophils, and delivered to epithelial cells via MVs during inflammation. Subsequently, miR-223 regulates the epithelial inflammation process through various signaling pathways to reduce tissue damage (Figure 6).
Role of miR-223 in inflammatory diseases
Recent studies have established that miR-223 could bind to specific target genes to inhibit the production of inflammatory mediators or block inflammation signaling pathways, which protect the body from inflammation-induced injury [11, 70-81]. miR-223 plays a vital role in various inflammatory diseases, including acute lung injury, rheumatoid arthritis, enteritis, nervous system inflammation and mastitis (Table 3). However, the regulatory networks through which miR-223 modulates immune responses are unclear.
Target and role of miR-223 in various inflammatory diseases
| Inflammatory disease | Target | Function | Reference |
|---|---|---|---|
| Acute lung injury (ALI) | RHOB, NLRP3, and PARP-1 | Alleviates ALI | [11,70-73] |
| Rheumatoid arthritis (RA) | ARNT and Stirt1 | Alleviates RA | [74,82] |
| Enteritis | NLRP3, CLDN8, and IKK-α | Alleviates enteritis | [75, 76] |
| Nervous system inflammation | ATG16L1 and NLRP3 | Alleviates nervous system inflammation | [78, 79] |
| Mastitis | HMGB1 and CBLB | Alleviates mastitis | [80, 81] |
Role of miR-223 during lung injury
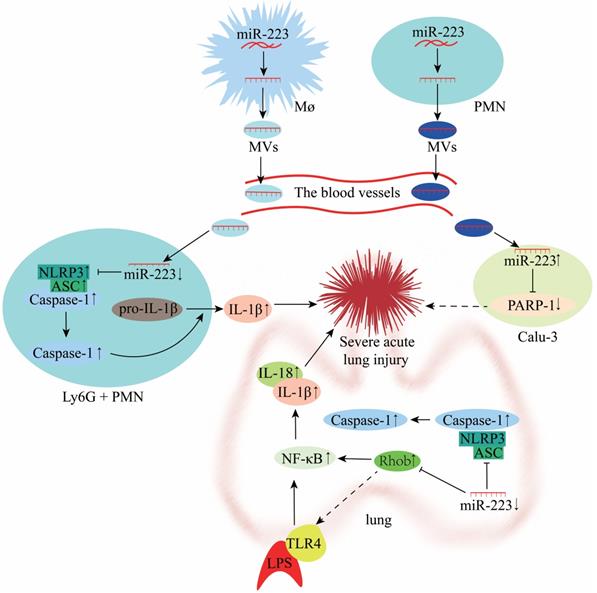
Lung injury involves two processes, namely inflammatory damage and lung fibrosis. Acute lung injury (ALI) is a type of lung injury that has been widely investigated in recent years. Accumulating evidence indicates that miR-223 regulates the NLRP3 inflammasome and plays an important role during ALI [73]. However, the TLR4 or NLRP3 inhibitors could impair the anti-inflammatory effect of miR-223 in ALI [11, 72]. Moreover, a study showed that the miR-223 response was the fastest in the porcine lung tissue H1N2 infection and highly predicted that NLRP3 could be a miR-223 target, affecting the injury of porcine lung tissue through inflammatory factors [83]. In a mouse model of macrophage-mediated lung inflammation, miR-223 regulates macrophage differentiation by targeting the NLRP3 inflammasome. It is transferred by MVs to inhibit lung inflammation (Figure 7) [70]. Overexpression of miR-223 can directly inhibit NF-κB in bronchial epithelial cells to alleviate lung inflammation [84]. Furthermore, miR-223 expression was reported to be down-regulated in LPS-treated A549 cells (lung adenocarcinoma cells) [72]. In vitro experiments also demonstrated that decreased miR-223 expression resulted in diminished inhibition of RHOB, NLRP3 inflammasome and TLR4/NF-κB pathway, exacerbating lung injury (Figure 7) [72, 85]. In a mouse study of hippocampal inflammation, aerobic exercise can induce miR-223 expression in the hippocampus, which negatively regulates the TLR4/MyD88-NF-κB pathway to improve inflammation-induced injury [86]. miR-223 can also alleviate neutrophilic airway inflammation by inhibiting NLRP3 inflammasome and IL-1β release [87]. However, miR-223 was reported to attenuate ALI induced by mitochondrial damage-associated molecular patterns (MTDs) by limiting the differentiation of bone marrow-derived Ly6G+ neutrophils and inhibiting NLRP3 inflammasome activity and IL-1β production [71]. On the other hand, miR-223 deficiency results in persistent NLRP3 and IL-1β activation and exacerbates lung injury [88]. MV-mediated transfer of miR-223 from neutrophils to lung epithelial cells (Calu-3) attenuates ALI via inhibition of poly (ADP-ribose) polymerase-1 (PARP-1) (Figure 7) [89, 90]. However, the miR-223 expression is significantly down-regulated during lung fibrosis, and this is attributed to the antioxidative properties of the amino acid hydroxyproline (HYP) [91]. miR-223 can alleviate ALI by regulating the process of inflammation, which serves as a new potential therapeutic target and prognostic marker for ALI (Figure 7).
Mechanisms of miR-223 in the modulation of acute lung injury (ALI).
Role of miR-223 in rheumatoid arthritis
Rheumatoid arthritis (RA) is a chronic inflammatory disease. Studies have demonstrated that miR-223 is significantly higher in the plasma, serum and peripheral mononuclear cells (PBMCs) of RA patients than those of controls, indicating that miR-223 could be associated with the development, progression and severity of RA [92, 93]. The reliability of the biological functions of miR-223 in RA has been confirmed by accurately measuring the AUC value, and sensitivity and specificity of miR-223 [94]. Also, other studies have shown that the expression of miR-223 in PBMCs and plasma is positively correlated with the level of rheumatoid factor (RF) in RA patients [95-98]. miR-223 is differentially expressed between the PBMCs and plasma in RA patients, suggesting that miR-223 could be a biological marker for RA diagnosis [95-98].
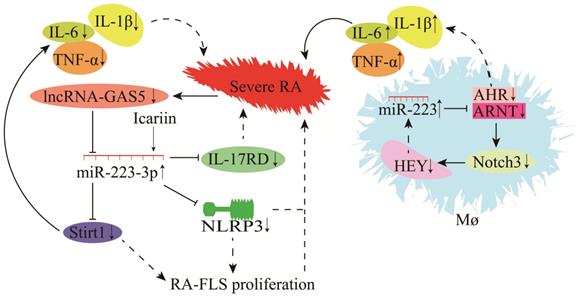
The role of miR-223 in RA is highly complex (Figure 8). miR-223 is significantly up-regulated during RA in mice. Some target genes regulated by miR-223 include actin alpha 1 (ACTA1), (ACVR2A), cholecystokinin B receptor (CCKBR), dual-specificity phosphatase 10 (DUSP10), forkhead box O1 (FOXO1), heat shock protein 90 beta family member 1 (HSP90B1), interleukin 6 cytokine family signal transducer (IL6ST), inositol polyphosphate-5-phosphatase B (INPP5B), MX dynamin-like GTPase 1 (MX1), protein tyrosine phosphatase non-receptor type 2 (PTPN2), and tyrosine 3-monooxygenase/tryptophan 5-monooxygenase activation protein gamma (YWHAG) [99]. According to a recent study in the arthritis fibroblast cell line MH7A, IL-17 receptor D (IL-17RD) was identified as a target gene of miR-223-3p [100]. Overexpression of miR-223-3p could down-regulate IL-17RD expression and alleviate RA-induced injury [100, 101]. Furthermore, the mouse model also showed that icariin could inhibit NLRP3 by up-regulating miR-223-3p expression, reducing the apoptosis of RA joint fibroblast-like synovial cells (RA-FLSCs), and attenuates RA-induced injury [102, 103]. Sirt1 is also a target gene of miR-223-3p, and lncRNA-GAS5 is a molecular sponge for miR-223-3p [82, 104]. Down-regulation of lncRNA-GAS5 during RA leads to the up-regulation of miR-223-3p and down-regulation of Sirt1. It decreases the secretion of TNF-α, IL-6, IL-8, and IL-1β and reduces RA-FLSC apoptosis, hence slowing RA progression [82, 104]. Macrophages are critical players in RA pathogenesis, and these are the most abundant inflammatory cells in RA. Ogando et al. [74] discovered that miR-223 was significantly up-regulated in the macrophages of RA patients. Increased miR-223 expression inhibited ARNT protein synthesis (a co-receptor for AHR) [105] and prevented AHR/ARNT-mediated pro-inflammatory cytokine expression. AHR agonists could suppress the gene expression of pro-inflammatory cytokines (TNF-α, IL-1β, and IL-6), Notch3, and HEY. A low level of HEY could up-regulate miR-223 expression in macrophages (Figure 8) [74]. These findings demonstrate that miR-223 regulates RA progression by increasing the macrophage sensitivity to pro-inflammatory cytokines and dampening their response to anti-inflammatory signals during RA [74].
Mechanisms of miR-223 in the modulation of rheumatoid arthritis (RA).
Role of miR-223 in enteritis
Enteritis could be classified into bacterial enteritis, parasitic enteritis and viral enteritis based on pathogens. Bacteria cause it, and microbes and viruses in the intestine could also induce enteritis [106, 107]. Several studies have demonstrated that miR-223 is highly expressed in the intestinal tissues, serum, and feces of ulcerative colitis (UC) and inflammatory bowel disease (IBD) patients, suggesting that miR-223 is involved in its regulation and could serve as a biomarker for UC and IBD [108-110]. Besides, NLRP3 inflammasome expression in mice and humans is gradually elevated as IBD exacerbates [111, 112], demonstrating that the expression of inflammasomes (NLRP1, NLRP2 and NLRP3) dictates the severity of IBD [113].
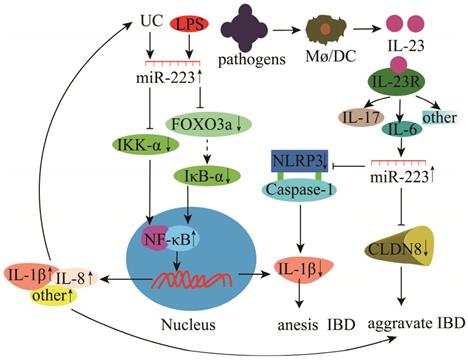
miR-223 plays a vital role in the development and progression of IBD and UC. Studies have found that miR-223 expression is significantly increased in the mucosal biopsy tissues of UC patients [114]. miR-223 regulates UC by inhibiting the expression of IKK-α, a negative regulator of NF-κB, and promotes the release of p56 and pro-inflammatory cytokines (IL-1β and IL-8) (Figure 9) [75]. In the case of IBD, LPS stimulation induces the inhibition of FOXO3a by miR-223, which subsequently down-regulates IκB-α and activates the NF-κB signaling pathway to release pro-inflammatory cytokines, thus promoting IBD progression (Figure 9) [115]. Neudecker et al. [76] showed that miR-223 expression was up-regulated during experimental IBD in mice. The increase in miR-223 expression caused the down-regulation of NLRP3 inflammasome and IL-1β secretion, reducing IBD severity (Figure 9) [76]. In the dextran sodium sulfate (DSS) colitis model, miR-223 can alleviate intestinal inflammation by reducing the release of inflammatory mediators via inhibition of IL-6 and STAT3 signaling [116]. Claudin-8 (CLDN8) is a member of the claudin multigene family and the primary tight junction protein. CLDN8 expression has been significantly down-regulated in the inflamed colonic mucosa of IBD patients and mice with trinitrobenzene sulfonic acid (TNBS)-induced colitis. Its level could be restored in colitic mice treated with IL-23 antibody. However, miR-223 could bind to CLDN8, and its up-regulation exacerbates IBD (Figure 9). Therefore, the interaction among IL-23, miR-223, and CLDN8 could serve as a new therapeutic strategy for IBD [77].
Mechanisms of miR-223 in the modulation of enteritis.
Role of miR-223 in central nervous system inflammation
Inflammation of the central nervous system (CNS) mainly includes EAE, multiple sclerosis (MS), and bacterial meningitis [117-119]. Microglial cells (bv-2) are resident macrophages of the CNS and have critical physiological functions in attenuating CNS inflammation and maintaining tissue homeostasis [120, 121]. miR-223 deficiency increases the autophagy of resting microglia and microglia in the brain, which significantly reduces demyelination and improves CNS inflammation and the clinical symptoms of EAE [78]. Notably, normal cell proliferation and autophagy are metabolic processes required to maintain homeostasis, and abnormal proliferation and autophagy could cause inflammatory diseases. Different, studies have shown that miR-223 could inhibit P21 by inhibiting NFIA expression, resulting in abnormal microglia proliferation [78, 122]. In mouse EAE and MS, the miR-223 expression was significantly increased as the disease duration increased [123, 124]. Deleting miR-223 (miR-223-/-) could significantly delay EAE onset, alleviate spinal injury and decrease neurological symptoms in mice, demonstrating that miR-223 is a potential marker and therapeutic target for EAE (Figure 10) [123, 124]. Also, miR-223 promotes MS progression by inhibiting bv-2 autophagy via the down-regulation of ATG16L1 (Figure 10) [78]. Galloway et al. [125] found that miR-223 effectively regulated M2 polarization and promoted bone marrow activation and CNS remyelination. These findings demonstrate a critical pathophysiological relationship between miR-223 and MS, and other neurodegenerative diseases, thus providing new insights into MS diagnosis and treatment [78]. During BM, resveratrol could modulate the miR-223-3p/NLRP3 pathway to inhibit downstream caspase-1 activation and IL-1β and IL-18 processing in neurons and bv-2 cells, protecting cortical neurons from inflammatory damage and death (Figure 10) [79]. However, findings from bv-2 cells have shown that lncRNA GAS5 and NLRP3 are ceRNAs that sponge miR-223-3p and a high expression of lncRNA GAS5 promotes NLRP3 activation and pro-inflammatory cytokine release [126].
Mechanisms of miR-223 in the modulation of experimental autoimmune encephalomyelitis (EAE).
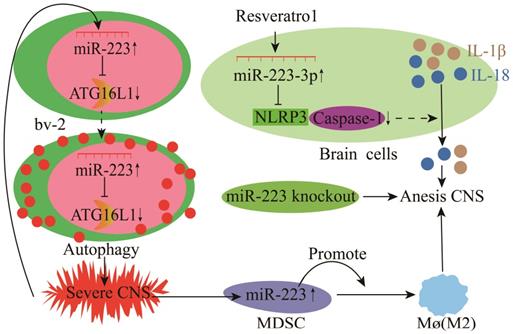
Myeloid-derived suppressor cells (MDSCs) play essential regulatory and effector functions in MS and EAE. miR-223 knockout (miR-223-/-) mice have increased MDSCs in the spleen and spinal cord and milder EAE, supporting the critical role of miR-223 in the regulation of MDSCs during EAE and MS and further highlights the possibility of miR-223 as a new therapeutic agent [127].
Role of miR-223 in mastitis
Mastitis could be caused by several factors, including pathogen infection and environmental changes and has complex pathogenesis. Staphylococcus aureus, Streptococcus. uberis, and Escherichia. coli are common causative agents of mastitis [128]. LPS is a surface component of E. coli that induces inflammation. Hao et al. [129] demonstrated that miR-223 was significantly up-regulated during LPS-induced mammary epithelial inflammation in mice. In addition, miR-223 expression was 2.5-3 times higher in the mammary tissue of cows with mastitis than that of healthy cows [130-134], suggesting that it could be a biomarker for mastitis [135, 136].
The molecular mechanism of miR-223 in regulating mastitis has been investigated in cows, humans, and mice. Moreover, miR-223 in MVs released by platelets could effectively inhibit the carcinogenesis of human breast cells [137]. miR-223 could participate in TLR signaling and down-regulate IL-6 protein expression (not gene expression) to inhibit inflammation (Figure 11) [138]. Studies of Staphylococcus. aureus or lipoteichoic acid (LTA)-induced mastitis have identified CBLB as a target gene for miR-223 [81, 139]. Bovine miR-223 could target CBLB and inhibit the PI3K/AKT/NF-κB signaling pathway to suppress IL-6 expression and attenuate inflammation (Figure 11) [81]. Furthermore, miR-223 could inhibit CXCL14 and regulate bovine mastitis through a series of complex gene regulatory networks (GRNs) (Figure 11) [140].
Mechanisms of miR-223 in the modulation of mastitis.
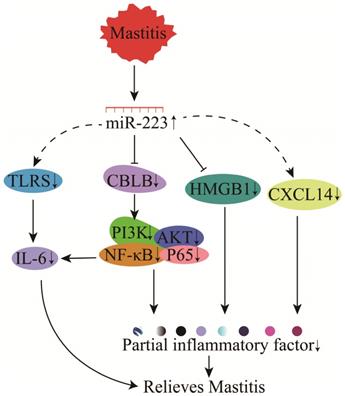
High mobility group box 1 (HMGB1) is involved in the pathogenesis of multiple inflammatory diseases, including bovine mastitis [141]. The functional SNP in the 3'-UTR of HMGB1 could affect its binding to miR-223, thereby influencing bovine mastitis regulation through miR-223 (Figure 11) [80]. Also, miR-223 has demonstrated protein-protein interactions with the predicted candidate genes (F-box protein 30 (FBXO30), SMAD specific E3 ubiquitin-protein ligase 2 (SMURF2), F-box and WD repeat domain containing 7 (FBXW7), and ubiquitin-like modifier activating enzyme 2 (UBA2), suggesting that miR-223 could regulate mastitis through bacterial invasion, endocytosis, antigen processing, immune response, and TGF-β and MAPK signaling in mammary epithelial cells [142-144].
Conclusions and future perspectives
miR-223 is a multifunctional miRNA regulated by several transcription factors, and its expression is significantly elevated during cellular or tissue inflammation. miR-223 regulates immune cell proliferation, differentiation and polarization, and has immunomodulatory effects in certain tissues. It also serves as a messenger and a regulator of inflammation in immune and resident cells. Furthermore, it plays a critical role in cell-cell, cell-tissue, and cell-tissue-inflammatory disease interactions. Elucidation of its molecular mechanisms in immune modulation could provide insights in developing early diagnosis or treatment for inflammatory diseases. Although several miR-223 target genes have been identified in humans and animals have effectively mediated the regulatory functions of miR-223, some of these functional studies have only been conducted using in vitro setups or at the cellular level. Thus, it is imperative to establish in vivo studies to validate these findings. In addition, the miR-223 target genes could be used to construct a detailed multi-species miR-223 regulatory network based on their sequence homology to identify precise miR-223-targeting drugs. These drugs could range from bioactive ingredients to DNA, mRNA, and translated proteins, and drugs identified using this method have high precision. Lastly, the drugs could exhibit better efficacy in treating inflammatory diseases, especially autoimmune disorders.
Abbreviations
Sirt1: sirtuin1; DCs: dendritic cells; RISC: RNA-induced silencing complex; qPCR: quantitative polymerase chain reaction; NFIA: nuclear factor I A; Mef2c: myocyte enhancer factor 2C; LPS: lipopolysaccharide; GBS: Group B Streptococcus; HPC: hematopoietic progenitor cell; Mø: macrophage; PPREs: PPARγ regulatory elements; MVs: microvesicles; IL-6: interleukin 6; KLF6: Kruppel-like factor 6; TRAF6: TNF receptor associated factor 6; NLRP3: NLR family pyrin domain containing 3; SOX11: SRY-box 11; MCL: Mantle Cell Lymphoma; MD: Marek's disease; ESCs: embryonic stem cells; Th17: T helper 17; Treg: regulatory T cell; EAE: experimental autoimmune encephalomyelitis; MCL: Mantle Cell Lymphoma; AS: atherosclerosis; P-MV: peripheral microvesicles; IGF-1R: insulin-like growth factor 1; HUVECs: human umbilical vascular endothelial cells; GEnCs: glomerular endothelial cells; IgAN: immunoglobulin A nephropathy; VEC: vascular endothelial cell; KD: Kawasaki Disease; RTECs: renal tubular epithelial cells; BEND: bovine endometrial epithelial cells; STIM1: stromal interaction molecule 1; IAV: influenza A virus; EMT-TFs: epithelial-mesenchymal transition-related transcription factors; ALI: Acute lung injury; MTDs: mitochondrial damage-associated molecular patterns; RA: Rheumatoid arthritis; PBMCs: peripheral mononuclear cells; RF: rheumatoid factor; ACTA1: actin alpha 1; ACVR2A: A receptor type 2A; CCKBR: cholecystokinin B receptor; DUSP10: dual specificity phosphatase 10; FOXO1: forkhead box O1; HSP90B1: heat shock protein 90 beta family member 1; IL6ST: interleukin 6 cytokine family signal transducer; INPP5B: inositol polyphosphate-5-phosphatase B; MX1: MX dynamin like GTPase 1; PTPN2: protein tyrosine phosphatase non-receptor type 2; YWHAG: tyrosine 3-monooxygenase/tryptophan 5-monooxygenase activation protein gamma; RA-FLSCs: RA joint fibroblast-like synovial cells; DSS: dextran sodium sulfate; CLDN8: Claudin-8; TNBS: trinitrobenzene sulfonic acid; IBD: inflammatory bowel disease; UC: ulcerative colitis; CNS: Inflammation of the central nervous system; GRNs: gene regulatory networks; FBXO30: F-box protein 30; SMURF2: SMAD specific E3 ubiquitin protein ligase 2; FBXW7: F-box and WD repeat domain containing 7; UBA2: ubiquitin like modifier activating enzyme 2; HMGB1: High mobility group box 1.
Acknowledgements
Funding
This research was supported by grants from the National Natural Science Foundation of China (32060749, 31960652), the Key Research and Development Project (Talent Introduction Project) of Ningxia Hui Autonomous Region (2019BEB04002), the Science and Technology research project of Ningxia Higher Education School (NGY2020007), the Introducing Talent Research Project of Ningxia University (030900001926).
Author contributions
P.J, X.-P.W. and Z.-M.L. designed and wrote the manuscript. Z.-M.L. and X.-P.W. helped to revise the content. J.Y. and L.J. helped to draw the images. Z.-M.L., X.-P.W., Y.M. and D.-W.W. revised the language.
Competing Interests
The authors have declared that no competing interest exists.
References
1. Mollaei M, Abbasi A, Hassan ZM & Pakravan N. The intrinsic and extrinsic elements regulating inflammation. Life Sci. 2020;260:118-258
2. Xu J, Xiao X. & Yang D. In vitro methods for analyzing miRNA roles in cancer cell proliferation, invasion, and metastasis. New York, NY: Springer New York. 2018;1733:159-171
3. Pogribny IP. MicroRNAs as biomarkers for clinical studies. Exp Biol Med. 2018;243:283-290
4. Jeffries J, Zhou W, Hsu AY. & Deng Q. MiRNA-223 at the crossroads of inflammation and cancer. Cancer Lett. 2019;451:136-141
5. Zhang N, Hu G, Myers TG & Williamson PR. Protocols for the analysis of microRNA expression, biogenesis, and function in immune cells. Curr Protoc Immunol. 2019;126:e78
6. Nepal C, Coolen M, Hadzhiev Y, Cussigh D & Mydel P. Transcriptional, post-transcriptional and chromatin-associated regulation of pri-miRNAs, pre-miRNAs and moRNAs. Nucleic Acids Res. 2016;44:3070-3081
7. Lee D & Shin C. Emerging roles of Drosha beyond primary microRNA processing. RNA Biol. 2018;15:186-193
8. Liu Z, Wang J, Cheng H, Ke X, Sun L & Zhang QC. Cryo-EM structure of human Dicer and its complexes with a pre-miRNA substrate. Cell. 2018;173:1191-1203
9. Winter J, Jung S, Keller S, Gregory RI. & Diederichs S. Many roads to maturity: microRNA biogenesis pathways and their regulation. Nat Cell Biol. 2009;11:228-234
10. Lim LP, Glasner ME, Yekta S, Burge CB & Bartel DP. Vertebrate microRNA genes. Science. 2003;299:15-40
11. Fazi F, Rosa A, Fatica A, Gelmetti V, Nervi C & Bozzoni I. A minicircuitry comprised of microRNA-223 and transcription factors NFI-A and C/EBPalpha regulates human granulopoiesis. Cell. 2005;123:819-831
12. Fukao T, Fukuda Y, Kiga K. An evolutionarily conserved mechanism for microRNA-223 expression revealed by microRNA gene profiling. Cell. 2007;129:617-631
13. Johnnidis JB, Harris MH, Wheeler RT, Stehling-Sun S, Lam MH, Kirak O, Brummelkamp TR. et al. Regulation of progenitor cell proliferation and granulocyte function by microRNA-223. Nature. 2008;451:1125-1129
14. Zardo G, Ciolfi A, Vian L, Starnes LM, Billi M, Racanicchi S, Maresca C. et al. Polycombs and microRNA-223 regulate human granulopoiesis by transcriptional control of target gene expression. Blood. 2012;119:4034-4046
15. Kim GD, N G, H P, Patel N & Mahabeleshwar GH. Kruppel-like factor 6 and miR-223 signaling axis regulates macrophage-mediated inflammation. FASEB J. 2019;33:10902-10915
16. Date D, Das R, Narla G, Simon DI, Jain MK & Mahabeleshwar GH. Kruppel-like transcription factor 6 regulates inflammatory macrophage polarization. J Biol Chem. 2014;289:10318-10329
17. O'Connell R M. MicroRNAs function on a new level. Blood. 2012;119:3875-3876
18. Bao H, Chen H, Zhu X, Zhang M, Yao G, Yu Y. et al. MiR-223 downregulation promotes glomerular endothelial cell activation by upregulating importin α4 and α5 in IgA nephropathy. Kidney Int. 2014;85:624-635
19. Bros M, Youns M, Kollek V, Buchmüller D, Bollmann F, Seo E J. et al. Differentially tolerized mouse antigen presenting cells share a common mirna signature including enhanced mmu-miR-223-3p expression which is sufficient to imprint a protolerogenic state. Front Pharmacol. 2018;9:915
20. Ifergan I, Chen S, Zhang B & Miller SD. Cutting Edge: MicroRNA-223 regulates myeloid dendritic cell-driven Th17 responses in experimental autoimmune encephalomyelitis. J Immunol. 2016;196:1455-1459
21. Mohajeri M, Banach M, Atkin SL, Butler AE, Ruscica M, Watts GF. et al. MicroRNAs: novel molecular targets and response modulators of statin therapy. Trends Pharmacol Sci. 2018;39:967-981
22. Sun Y, Liu MW, Zhao YH, Lu YX, Wang YA & Tong CW. Baicalin attenuates lipopolysaccharide-induced renal tubular epithelial cell injury by inhibiting the TXNIP/NLRP3 signalling pathway via increasing miR-223-3p expression. J Biol Regul Homeost Agents. 2020;34:69-82
23. Vian L, Di-Carlo M, Pelosi E, Fazi F, Santoro S, Cerio AM. et al. Transcriptional fine-tuning of microRNA-223 levels directs lineage choice of human hematopoietic progenitors. Cell Death Differ. 2014;21:290-301
24. Li C, Xu MM, Wang K, Adler AJ, Vella AT & Zhou B. Macrophage polarization and meta-inflammation. Transl Res. 2018;191:29-44
25. Li T, Morgan MJ, Choksi S, Zhang Y, Kim YS & Liu ZG. MicroRNAs modulate the noncanonical transcription factor NF-kappaB pathway by regulating expression of the kinase IKKalpha during macrophage differentiation. Nat Immunol. 2010;11:799-805
26. Zhou H, Xiao J, Wu N, Liu C, Xu J, Liu F. et al. MicroRNA-223 regulates the differentiation and function of intestinal dendritic cells and macrophages by targeting C/EBPβ. Cell Rep. 2015;13:1149-1160
27. Satoorian T, Li B, Tang X, Xiao J, Xing W, Shi W. et al. MicroRNA223 promotes pathogenic T-cell development and autoimmune inflammation in central nervous system in mice. Immunology. 2016;148:326-338
28. Hosseini A, Ghaedi K, Tanhaei S, Ganjalikhani-Hakemi M, Teimuri S, Etemadifar M. et al. Upregulation of CD4+ T-Cell derived miR-223 in the relapsing phase of multiple sclerosis patients. Cell J. 2016;18:371-380
29. Pan Y, Liang H, Liu H, Li D, Chen X, Li L. et al. Platelet-secreted microRNA-223 promotes endothelial cell apoptosis induced by advanced glycation end products via targeting the insulin-like growth factor 1 receptor. J Immunol. 2014;192:437-446
30. Yuan X, Wang X, Chen C, Zhou J & Han M. Bone mesenchymal stem cells ameliorate ischemia/reperfusion-induced damage in renal epithelial cells via microRNA-223. Stem Cell Res. 2017;8:146
31. Chen Q, Wang H, Liu Y, Song Y, Lai L, Han Q. et al. Inducible microRNA-223 down-regulation promotes TLR-triggered IL-6 and IL-1β production in macrophages by targeting STAT3. PLoS One. 2012;7:e42971
32. Zhuang G, Meng C, Guo X, Cheruku PS, Shi L, Xu H, Li H. et al. A novel regulator of macrophage activation: miR-223 in obesity-associated adipose tissue inflammation. Circulation. 2012;125:2892-2903
33. Xue YL, Zhang SX, Zheng CF, Li YF, Zhang LH, Su QY, Hao YF. et al. Long non-coding RNA MEG3 inhibits M2 macrophage polarization by activating TRAF6 via microRNA-223 down-regulation in viral myocarditis. J Cell Mol Med. 2020;24:12341-12354
34. Zhu MX, Wan WL, Hu K, Wang YF, Wang J, Zhu XW, Yan XX. et al. MicroRNA-223 regulates the differentiation of human embryonic stem cells to dendritic cells. Journal of Experimental Hematology. 2017;25:1275-1282 (in Chinese)
35. Chen L, Hou X, Zhang M, Zheng Y, Zheng X, Yang Q, Li J. et al. MicroRNA-223-3p modulates dendritic cell function and ameliorates experimental autoimmune myocarditis by targeting the NLRP3 inflammasome. Mol Immunol. 2020;117:73-83
36. Tang HC, Lai YY, Zheng J, Jiang HY & Xu G. MiR-223-3p inhibits antigen endocytosis and presentation and promotes the tolerogenic potential of dendritic cells through targeting mannose receptor signaling and Rhob. J Immunol Res. 2020;2020:1379458
37. Shi XY, Xie X, Sun Y, He H, Huang H, Liu Y. et al. Paeonol inhibits NLRP3 mediated inflammation in rat endothelial cells by elevating hyperlipidemic rats plasma exosomal miRNA-223. Eur J Pharmacol. 2020;885:173473
38. Wang X, Ding YY, Chen Y, Xu QQ, Qian GH, Qian WG. et al. MiR-223-3p alleviates vascular endothelial injury by targeting IL6ST in Kawasaki disease. Front Pediatr. 2019;7:288
39. Xu Y, Deng W & Zhang W. Long non-coding RNA TUG1 protects renal tubular epithelial cells against injury induced by lipopolysaccharide via regulating microRNA-223. Biomed Pharmacother. 2018;104:509-519
40. Liu CC, Miao Y, Chen RL, Zhang YQ, Wu H, Yang SM. et al. STIM1 mediates IAV-induced inflammation of lung epithelial cells by regulating NLRP3 and inflammasome activation via targeting miR-223. Life Sci. 2020;266:118845
41. Deny M, Romano M, Denis O, Casimir G & Chamekh M. Progressive control of Streptococcus agalactiae-induced innate inflammatory response is associated with time course expression of microRNA-223 by neutrophils. Infect Immun. 2020;88:e00563-20
42. Ying W, Tseng A, Chang RC, Morin A, Brehm T, Triff K, Nair V. et al. MicroRNA-223 is a crucial mediator of PPARγ-regulated alternative macrophage activation. J Clin Invest. 2015;125:4149-4159
43. Liu G & Abraham E. MicroRNAs in immune response and macrophage polarization. Arterioscler Thromb Vasc Biol. 2013;33:170-177
44. Cheng N, Liu C, Li Y, Gao S, Han YC, Wang X. et al. MicroRNA-223-3p promotes skeletal muscle regeneration by regulating inflammation in mice. J Biol Chem. 2020;295:10212-10223
45. Ismail N, Wang Y, Dakhlallah D, Moldovan L, Agarwal K, Batte K. et al. Macrophage microvesicles induce macrophage differentiation and miR-223 transfer. Blood. 2013;121:984-995
46. Johanson TM, Cmero M, Wettenhall J, Lew AM, Zhan Y & Chong MM. A microRNA expression atlas of mouse dendritic cell development. Immunol Cell Biol. 2015;93:480-485
47. Fulci V, Scappucci G, Sebastiani GD, Giannitti C, Franceschini D, Meloni F. et al. MiR-223 is overexpressed in T-lymphocytes of patients affected by rheumatoid arthritis. Hum Immunol. 2010;71:206-211
48. Zhou K, Feng X, Wang Y, Liu Y, Tian L, Zuo W. et al. MiR-223 is repressed and correlates with inferior clinical features in mantle cell lymphoma through targeting SOX11. Exp Hematol. 2018;58:27-34
49. Yao Y, Zhao Y, Smith L P, Lawrie C H, Saunders N J, Watson M. et al. Differential expression of microRNAs in Marek's disease virus-transformed T-lymphoma cell lines. J Gen Virol. 2009;90:1551-1559
50. Han B, He Y, Zhang L, Ding Y, Lian L, Zhao C. et al. Long intergenic non-coding RNA GALMD3 in chicken Marek's disease. Sci Rep. 2017;7:10294
51. Khalyfa A, Kheirandish-Gozal L, Khalyfa AA, Philby MF, Alonso-Álvarez ML, Mohammadi M. et al. Circulating plasma extracellular microvesicle microRNA cargo and endothelial dysfunction in children with obstructive sleep apnea. Am J Respir Crit Care Med. 2016;194:1116-1126
52. McGinn CM, MacDonnell BF, Shan CX, Wallace R, Cummins PM & Murphy RP. Microparticles: A pivotal nexus in vascular homeostasis and disease. Curr Clin Pharmacol. 2016;11:28-42
53. Shu Z, Tan J, Miao Y & Zhang Q. The role of microvesicles containing microRNAs in vascular endothelial dysfunction. J Cell Mol Med. 2019;23:7933-7945
54. Alexandru N, Andrei E, Safciuc F, Dragan E, Balahura A M, Badila E. et al. Intravenous administration of allogenic cell-derived microvesicles of healthy origins defend against atherosclerotic cardiovascular disease development by a direct action on endothelial progenitor cells. Cells. 2020;9:423
55. Lange CA, Lazzari B, Perrini C, Pizzi F, Stella A, Cremonesi F. et al. MicroRNAs of equine amniotic mesenchymal cell-derived microvesicles and their involvement in anti-inflammatory processes. Cell Transplant. 2018;27:45-54
56. Li J, Tan M, Xiang Q, Zhou Z & Yan H. Thrombin-activated platelet-derived exosomes regulate endothelial cell expression of ICAM-1 via microRNA-223 during the thrombosis-inflammation response. Thromb Res. 2017;154:96-105
57. Zegeye MM, Lindkvist M, Falker K, Kumawat AK, Paramel G, Grenegard M. et al. Activation of the JAK/STAT3 and PI3K/AKT pathways are crucial for IL-6 trans-signaling-mediated pro-inflammatory response in human vascular endothelial cells. Cell Commun Signal. 2018;16:55
58. Qing Z, Kaixin Z, Yanfei H, Yiming Z, Hua X, Ling Z. et al. MicroRNA-223 triggers inflammation in porcine aorta by activating NLRP3 inflammasome under selenium deficiency. J Cell Physiol. 2021;236:4555-4564
59. Shi L, Fisslthaler B, Zippel N, Frömel T, Hu J, Elgheznawy A. et al. MicroRNA-223 antagonizes angiogenesis by targeting β1 integrin and preventing growth factor signaling in endothelial cells. Circ Res. 2013;113:1320-1330
60. Cavallari C, Dellepiane S, Fonsato V, Medica D, Marengo M, Migliori MQ. et al. Online hemodiafiltration inhibits inflammation-related endothelial dysfunction and vascular calcification of uremic patients modulating miR-223 expression in plasma extracellular vesicles. J Immunol. 2019;202:2372-2383
61. Chu M, Wu R, Qin S, Hua W, Shan Z, Rong X, Zeng J, Hong L. et al. Bone marrow-derived microRNA-223 works as an endocrine genetic signal in vascular endothelial cells and participates in vascular injury from kawasaki disease. J Am Heart Assoc. 2017;6:e004878
62. Zhang Y, Wang Y, Zhang L, Xia L, Zheng M, Zeng Z. et al. Reduced platelet miR-223 induction in Kawasaki disease leads to severe coronary artery pathology through a miR-223/PDGFRβ vascular smooth muscle cell axis. Circ Res. 2020;127:855-873
63. Deng B, Hu Y, Sheng X, Zeng H & Huo Y. MiR-223-3p reduces high glucose and high fat-induced endothelial cell injury in diabetic mice by regulating NLRP3 expression. Exp Ther Med. 2020;20:1514-1520
64. Citron F, Segatto I, Vinciguerra GLR, Musco L, Russo F, Mungo G. et al. Downregulation of miR-223 expression is an early event during mammary transformation and confers resistance to CDK4/6 inhibitors in luminal breast cancer. Cancer Res. 2020;80:1064-1077
65. Zhao G, Jiang K, Yang Y, Zhang T, Wu H, Shaukat A. et al. The Potential therapeutic role of miR-223 in bovine endometritis by targeting the NLRP3 inflammasome. Front Immunol. 2018;9:1916
66. Neudecker V, Fletcher AA, Barthel L, Masterson JC, Furuta GT, Cai C. et al. Neutrophil transfer of miR-223 to lung epithelial cells dampens acute lung injury in mice. Sci Transl Med. 2017;9:eaah5360
67. Dominguez C, David JM & Palena C. Epithelial-mesenchymal transition and inflammation at the site of the primary tumor. Semin Cancer Biol. 2017;47:177-184
68. Markopoulos GS, Roupakia E, Marcu KB & Kolettas E. Epigenetic regulation of inflammatory cytokine-induced epithelial-to-mesenchymal cell transition and cancer stem cell generation. Cells. 2019;8:1143
69. Tang Y, Wang Y, Chen Q, Qiu N, Zhao Y & You X. MiR-223 inhibited cell metastasis of human cervical cancer by modulating epithelial mesenchymal transition. Int J Clin Exp Pathol. 2015;8:11224-11229
70. Zhang D, Lee H, Wang X, Groot M, Sharma L, Jin Y. et al. A potential role of microvesicle-containing miR-223/142 in lung inflammation. Thorax. 2019;74:865-874
71. Feng Z, Qi S, Zhang Y, Qi Z, Yan L, Zhou J. et al. Ly6G+ neutrophil-derived miR-223 inhibits the NLRP3 inflammasome in mitochondrial DAMP-induced acute lung injury. Cell Death Disease. 2017;8:e3170
72. Yan Y, Lu K, Ye T & Zhang Z. MicroRNA-223 attenuates LPS-induced inflammation in an acute lung injury model via the NLRP3 inflammasome and TLR4/NF-κB signaling pathway via RHOB. Int J Mol Med. 2019;43:1467-1477
73. Haneklaus M, ONeill LA & Coll RC. Modulatory mechanisms controlling the NLRP3 inflammasome in inflammation: recent developments. Curr Opin Immunol. 2013;25:40-45
74. Ogando J, Tardaguila M, Diaz-Alderete A, Usategui A, Miranda-Ramos V, Martinez-Herrera DJ. et al. Notch-regulated miR-223 targets the aryl hydrocarbon receptor pathway and increases cytokine production in macrophages from rheumatoid arthritis patients. Sci Rep. 2016;6:20223
75. Valmiki S, Ahuja V, Puri N & Paul J. MiR-125b and miR-223 contribute to inflammation by targeting the key molecules of NFκB pathway. Front Med Lausanne. 2019;6:313
76. Neudecker V, Haneklaus M, Jensen O, Khailova L, Masterson JC, Tye H. et al. Myeloid-derived miR-223 regulates intestinal inflammation via repression of the NLRP3 inflammasome. J Exp Med. 2017;214:1737-1752
77. Wang H, Chao K, Ng SC, Bai AH, Yu Q, Yu J. et al. Pro-inflammatory miR-223 mediates the cross-talk between the IL23 pathway and the intestinal barrier in inflammatory bowel disease. Genome Biol. 2016;17:58
78. Li Y, Zhou D, Ren Y, Zhang Z, Guo X, Ma M. et al. MiR-223 restrains autophagy and promotes CNS inflammation by targeting ATG16L1. Autophagy. 2019;15:478-492
79. Gao M & Huang J. Resveratrol exerts a protective effect on cortical neurons of juvenile rat meningitis model by regulating miR-223-3p/NLRP3 pathway. Chinese Journal of Pathophysiology. 2020;36:1161-1169 (in Chinese)
80. Li L, Huang J, Zhang X, Ju Z, Qi C, Zhang Y. et al. One SNP in the 3'-UTR of HMGB1 gene affects the binding of target bta-miR-223 and is involved in mastitis in dairy cattle. Immunogenetics. 2012;64:817-824
81. Han S, Li X, Liu J, Zou Z, Luo L, Wu R. et al. Bta-miR-223 targeting CBLB contributes to resistance to Staphylococcus aureus mastitis through the PI3K/AKT/NF-kappaB pathway. Front Vet Sci. 2020;7:529
82. Yang Z, Lin SD, Zhan F, Liu Y & Zhan YW. LncRNA GAS5 alleviates rheumatoid arthritis through regulating miR-222-3p/Sirt1 signalling axis. Autoimmunity. 2021;54:13-22
83. Skovgaard K, Cirera S, Vasby D, Podolska A, Breum S, Dürrwald R. et al. Expression of innate immune genes, proteins and microRNAs in lung tissue of pigs infected experimentally with influenza virus (H1N2). Innate Immun. 2013;19:531-544
84. Zhou W, Pal AS, Hsu AY, Gurol T, Zhu X, Freeman JL. et al. MicroRNA-223 suppresses the canonical NF-κB pathway in basal keratinocytes to dampen neutrophilic inflammation. Cell Rep. 2018;22:1810-1823
85. Zeng Y, Zhang X, Kang K, Chen J, Wu Z, Huang J. et al. MicroRNA-223 attenuates hypoxia-induced vascular remodeling by targeting RhoB/MLC2 in pulmonary arterial smooth muscle cells. Sci Rep. 2016;6:24900
86. Qu H, Liu R, Chen J, Zheng L & Chen R. Aerobic exercise inhibits cums-depressed mice hippocampal inflammatory response via activating hippocampal miR-223/TLR4/MyD88-NF-κB pathway. Int J Environ Res Public Health. 2020;17:2676
87. Xu W, Wang Y, Ma Y & Yang J. MiR-223 plays a protecting role in neutrophilic asthmatic mice through the inhibition of NLRP3 inflammasome. Resp Res. 2020;21:116
88. Haneklaus M, Gerlic M, Kurowska-Stolarska M, Rainey AA, Pich D, McInnes IB. et al. Cutting edge: miR-223 and EBV miR-BART15 regulate the NLRP3 inflammasome and IL-1β production. J Immunol. 2012;189:3795-3799
89. Nie Y, Nirujogi TS, Ranjan R, Reader BF, Chung S, Ballinger MN. et al. PolyADP-Ribosylation of NFATc3 and NF-κB transcription factors modulate macrophage inflammatory gene expression in LPS-induced acute lung injury. J Innate Immun. 2021;13:83-93
90. Ferguson KT, McQuattie PA, Malsin ES & Sporn P. Dynamics of influenza-induced lung-resident memory t cells, anatomically and functionally distinct lung mesenchymal populations, and dampening of acute lung injury by neutrophil transfer of microRNA-223 to lung epithelial cells. Am J Respir Cell Mol Biol. 2018;59:397-399
91. Qu SJ, Zhao L, Song ZZ, Shen WP, Ju P & Li YM. Expression and significance of miR-223 in rats with pulmonary fibrosis. Eur Rev Med Pharmacol Sci. 2020;24:3951-3958
92. Dunaeva M, Blom J, Thurlings R & Pruijn G. Circulating serum miR-223-3p and miR-16-5p as possible biomarkers of early rheumatoid arthritis. Clin Exp Immunol. 2018;193:376-385
93. Taha M, Shaker OG, Abdelsalam E & Taha N. Serum a proliferation-inducing ligand and microRNA-223 are associated with rheumatoid arthritis: diagnostic and prognostic implications. Mol Med. 2020;26:92
94. Kriegsmann M, Randau TM, Gravius S, Lisenko K, Altmann C, Arens N. et al. Expression of miR-146a, miR-155, and miR-223 in formalin-fixed paraffin embedded synovial tissues of patients with rheumatoid arthritis and osteoarthritis. Virchows Arch. 2016;469:93-100
95. Lu MC, Yu CL, Chen HC, Yu HC, Huang HB & Lai NS. Increased miR-223 expression in T cells from patients with rheumatoid arthritis leads to decreased insulin-like growth factor-1-mediated interleukin-10 production. Clin Exp Immunol. 2014;177:641-651
96. Evangelatos G, Fragoulis GE, Koulouri V & Lambrou GI. MicroRNAs in rheumatoid arthritis: From pathogenesis to clinical impact. Autoimmun Rev. 2019;18:102391
97. Rezaeepoor M, Pourjafar M, Tahamoli-Roudsari A, Basiri Z, Hajilooi M & Solgi G. Altered expression of microRNAs may predict therapeutic response in rheumatoid arthritis patients. Int Immunopharmacol. 2020;83:106404
98. Filková M, Aradi B, Senolt L, Ospelt C, Vettori S, Mann H, Filer A. et al. Association of circulating miR-223 and miR-16 with disease activity in patients with early rheumatoid arthritis. Ann Rheum Dis. 2014;73:1898-1904
99. Dudics S, Venkatesha SH & Moudgil KD. The microRNA expression profiles of autoimmune arthritis reveal novel biomarkers of the disease and therapeutic response. Int J Mol Sci. 2018;19:2293
100. Moriya N, Shibasaki S, Karasaki M & Iwasaki T. The impact of microRNA-223-3p on IL-17 receptor D expression in synovial cells. PLoS One. 2017;12:e169702
101. Liu S, Fu Y, Mei K, Jiang Y, Sun X, Wang Y. et al. A shedding soluble form of interleukin-17 receptor D exacerbates collagen-induced arthritis through facilitating TNF-α-dependent receptor clustering. Cell Mol Immunol. 2020 Online ahead of print
102. Wu ZM, Luo J, Shi XD, Zhang SX, Zhu XB & Guo J. Icariin alleviates rheumatoid arthritis via regulating miR-223-3p/NLRP3 signalling axis. Autoimmunity. 2020;53:450-458
103. Cai P, Jiang T, Li B, Qin X, Lu Z, Shen C. et al. Comparison of rheumatoid arthritis (RA) and osteoarthritis (OA) based on microarray profiles of human joint fibroblast-like synoviocytes. Cell Biochem Funct. 2019;37:31-41
104. Li G, Xia Z, Liu Y, Meng F, Wu X, Fang Y. et al. SIRT1 inhibits rheumatoid arthritis fibroblast-like synoviocyte aggressiveness and inflammatory response via suppressing NF-κB pathway. Biosci Rep. 2018;38:BSR20180541
105. Furue M. Regulation of filaggrin, loricrin, and involucrin by IL-4, IL-13, IL-17A, IL-22, AHR, and NRF2: pathogenic implications in atopic dermatitis. Int J Mol Sci. 2020;21:5382
106. Gibbons JF, Boland F, Buckley JF, Butler F, Egan J, Fanning S. et al. Patterns of antimicrobial resistance in pathogenic Escherichia coli isolates from cases of calf enteritis during the spring-calving season. Vet Microbiol. 2014;170:73-80
107. Gomez DE & Weese JS. Viral enteritis in calves. Vet J. 2017;58:1267-1274
108. Wang H, Zhang S, Yu Q, Yang G, Guo J, Li M. et al. Circulating microRNA223 is a new biomarker for inflammatory bowel disease. Medicine. 2016;95:e2703
109. Hrabar J, Trumbic Z, Bocina I, Buselic I, Vrbatovic A & Mladineo I. Interplay between proinflammatory cytokines, miRNA, and tissue lesions in anisakis infected sprague-dawley rats. PLoS Negl Trop Dis. 2019;13:e7397
110. Schönauen K, Le N, Arnim U, Schulz C & Malfertheiner P. Circulating and fecal microRNAs as biomarkers for inflammatory bowel diseases. Inflamm Bowel Dis. 2018;24:1547-1557
111. Mao L, Kitani A, Strober W & Fuss IJ. The role of NLRP3 and IL-1β in the pathogenesis of inflammatory bowel disease. Front Immunol. 2018;9:2566
112. Liu L, Dong Y, Ye M, Jin S, Yang J, Joosse ME. et al. The pathogenic role of NLRP3 inflammasome activation in inflammatory bowel diseases of both mice and humans. J Crohns Colitis. 2017;11:737-750
113. Poli G, Fabi C, Bellet MM, Costantini C, Nunziangeli L, Romani L. et al. Epigenetic mechanisms of inflammasome regulation. Int J Mol Sci. 2020;21:5758
114. Valmiki S, Ahuja V & Paul J. MicroRNA exhibit altered expression in the inflamed colonic mucosa of ulcerative colitis patients. World J Gastroenterol. 2017;23:5324-5332
115. Kim HY, Kwon HY, Ha HT, Lee HJ, Kim GI, Hahm KB. et al. MicroRNA-132 and microRNA-223 control positive feedback circuit by regulating FOXO3a in inflammatory bowel disease. J Gastroenterol Hepatol. 2016;31:1727-1735
116. Zhang J, Wang C & Guo Z. MiR-223 improves intestinal inflammation through inhibiting the IL-6/STAT3 signaling pathway in dextran sodium sulfate-induced experimental colitis. Immun Inflamm Dis. 2021;9:319-327
117. Bolton C & Smith PA. The influence and impact of ageing and immunosenescence (ISC) on adaptive immunity during multiple sclerosis (MS) and the animal counterpart experimental autoimmune encephalomyelitis (EAE). Ageing Res Rev. 2018;41:64-81
118. Chitnis T & Weiner HL. CNS inflammation and neurodegeneration. J Clin Invest. 2017;127:3577-3587
119. Shemer A, Grozovski J, Tay TL, Tao J, Volaski A, Kim JS. et al. Engrafted parenchymal brain macrophages differ from microglia in transcriptome, chromatin landscape and response to challenge. Nat Commun. 2018;9:5206
120. Voet S, Prinz M & Van LG. Microglia in central nervous system inflammation and multiple sclerosis pathology. Trends Mol Med. 2019;25:112-123
121. Wu Y, Dissing-Olesen L, MacVicar BA & Stevens B. Microglia: dynamic mediators of synapse development and plasticity. Trends Immunol. 2015;36:605-613
122. Glasgow SM, Laug D, Brawley VS, Zhang Z, Corder A, Yin Z. et al. The miR-223/nuclear factor I-A axis regulates glial precursor proliferation and tumorigenesis in the CNS. J Neurosci. 2013;33:13560-13568
123. Ebrahimkhani S, Vafaee F, Young PE, Hawke S, Devenney E, Beadnall H. et al. Exosomal microRNA signatures in multiple sclerosis reflect disease status. Sci Rep. 2017;7:14293
124. Juźwik CA, Drake S, Johnson RM, Morquette B, Zhang Y, Charabati M. et al. Neuronal microRNA regulation in experimental autoimmune encephalomyelitis. Sci Rep. 2018;8:13437
125. Galloway DA, Blandford SN, Berry T, Williams JB, Stefanelli M, Ploughman M. et al. MiR-223 promotes regenerative myeloid cell phenotype and function in the demyelinated central nervous system. Glia. 2019;67:857-869
126. Xu W, Zhang L, Geng Y, Liu Y & Zhang N. Long noncoding RNA GAS5 promotes microglial inflammatory response in Parkinson's disease by regulating NLRP3 pathway through sponging miR-223-3p. Int Immunopharmacol. 2020;85:106614
127. Cantoni C, Cignarella F, Ghezzi L, Mikesell B, Bollman B, Ireland AR. et al. MiR-223 regulates the number and function of myeloid-derived suppressor cells in multiple sclerosis and experimental autoimmune encephalomyelitis. Acta Neuropathol. 2017;133:61-77
128. Klaas IC & Zadoks RN. An update on environmental mastitis: challenging perceptions. Transbound Emerg Disease. 2018;65:166-185
129. Hao JF, Huang K, Wang YY, Du LL & Zhong K. Expression of inflammation-associated miRNAs in LPS-induced inflammatory response of mouse mammary epithelial cells and construction of eukaryotic expression vector miR-223. Acta Veterinaria et Zootechinca Sinica. 2016;47:183-189 (in Chinese)
130. Pu J, Li R, Zhang C, Chen D, Liao X, Zhu Y. et al. Expression profiles of miRNAs from bovine mammary glands in response to streptococcus agalactiae-induced mastitis. J Dairy Res. 2017;84:300-308
131. Cai M, He H, Jia X, Chen S, Wang J, Shi Y. et al. Genome-wide microRNA profiling of bovine milk-derived exosomes infected with Staphylococcus aureus. Cell Stress Chaperon. 2018;23:663-672
132. Naeem A, Zhong K, Moisa SJ, Drackley JK, Moyes KM & Loor JJ. Bioinformatics analysis of microRNA and putative target genes in bovine mammary tissue infected with Streptococcus uberis. J Dairy Sci. 2012;95:6397-6408
133. Sun J, Aswath K, Schroeder SG, Lippolis JD, Reinhardt TA & Sonstegard TS. MicroRNA expression profiles of bovine milk exosomes in response to Staphylococcus aureus infection. BMC Genomics. 2015;16:806
134. Li R, Zhang CL, Liao XX, Chen D, Wang WQ, Zhu YH. et al. Transcriptome microRNA profiling of bovine mammary glands infected with Staphylococcus aureus. Int J Mol Sci. 2015;16:4997-5013
135. Dilda F, Gioia G, Pisani L, Restelli L, Lecchi C, Albonico F. et al. Escherichia coli lipopolysaccharides and Staphylococcus aureus enterotoxin B differentially modulate inflammatory microRNAs in bovine monocytes. Vet J. 2012;192:514-516
136. Chen L, Liu X, Li Z, Wang H, Liu Y, He H. et al. Expression differences of miRNAs and genes on NF-κB pathway between the healthy and the mastitis Chinese Holstein cows. Gene. 2014;545:117-125
137. Gasperi V, Vangapandu C, Savini I, Ventimiglia G, Adorno G & Catani MV. Polyunsaturated fatty acids modulate the delivery of platelet microvesicle-derived microRNAs into human breast cancer cell lines. J Nutr Biochem. 2019;74:108242
138. Hao JF. Preliminary study on the expression of miRNAs in LPS-induced inflammatory response of breast epithelial cells and the involvement of miR-223 in the regulation of Toll-like receptor signaling pathway. Henan Agricultural University. 2016 (in Chinese)
139. Han S. Screening of miRNAs related to mastitis resistance of Staphylococcus aureus in dairy cows and the regulatory mechanism of their target genes. HLAU. 2020 (in Chinese)
140. Fang L, Hou Y, An J, Li B, Song M, Wang X. et al. Genome-wide transcriptional and post-transcriptional regulation of innate immune and defense responses of bovine mammary gland to Staphylococcus aureus. Front Cell Infect Microbiol. 2016;6:193
141. Yang EJ, Lee W, Ku SK, Song KS & Bae JS. Anti-inflammatory activities of oleanolic acid on HMGB1 activated HUVECs. Food Chem Toxicol. 2012;50:1288-1294
142. Han S. Bioinformatics analysis of target genes and regulatory networks of bta-miR-223. Acta Veterinaria et Zootechinca Sinica. 2019;50:2171-2174 (in Chinese)
143. Yang Y, Sun H, Mo X, Liu Y, Jia H, Li X. et al. Prediction of novel genes associated with negative regulators of toll-like receptors-induced inflammation based on endotoxin tolerance. Inflammation. 2012;35:1889-1899
144. Balamurugan K, Sharan S, Klarmann KD, Zhang Y, Coppola V, Summers GH. et al. FBXW7α attenuates inflammatory signalling by downregulating C/EBPδ and its target gene TLR4. Nat Commun. 2013;4:1662
Author contact
Corresponding author: Zhuo-Ma Luoreng (E-mail: luorenzhuomaedu.cn).

 Global reach, higher impact
Global reach, higher impact