Impact Factor
ISSN: 1449-2288
Int J Biol Sci 2021; 17(12):3158-3172. doi:10.7150/ijbs.62553 This issue Cite
Research Paper
DDX39 as a predictor of clinical prognosis and immune checkpoint therapy efficacy in patients with clear cell renal cell carcinoma
1. Department of Urology, Changhai Hospital, Naval Medical University (Second Military Medical University), Shanghai, China
2. Department of Urology, Changzheng Hospital, Naval Medical University (Second Military Medical University), Shanghai, China.
*These authors have contributed equally to this work and share first authorship
Abstract
DEAD-box protein 39 (DDX39) has been demonstrated to be a tumorigenic gene in multiple tumor types, but its role in the progression and immune microenvironment of clear cell renal cell cancer (ccRCC) remains unclear. The aim of the present study was to investigate the role of DDX39 in the ccRCC tumor progression, immune microenvironment and efficacy of immune checkpoint therapy.
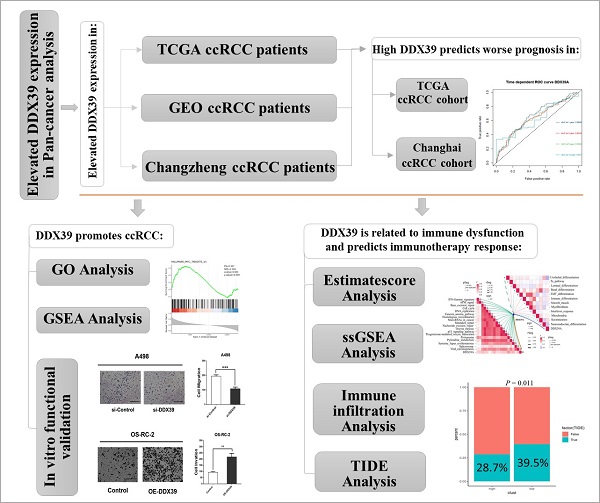
The DDX39 expression level was first detected in tumors in the public data and then verified in ccRCC samples from Changzheng Hospital. The prognostic value of DDX39 expression was assessed in the Cancer Genome Atlas (TCGA) and ccRCC patients from Changhai Hospital. The role of DDX39 in promoting ccRCC was analyzed by bioinformatic analysis and in vitro experiments. The association between DDX39 expression and immune cell infiltration and immune inhibitory markers was analyzed, and its value in predicting the immune checkpoint therapy efficacy in ccRCC were evaluated in the public database. DDX39 expression was elevated in Oncomine, GEO and TCGA ccRCC databases, as well as in Changzheng ccRCC samples. In TCGA ccRCC patients, increased DDX39 expression predicted worse overall survival (OS) (p<0.0001) and progression-free interval (PFI) (p<0.0001), and was shown as an independent predictive factor for OS (p=0.002). These findings were consistent with those from Changhai ccRCC patients. In addition, GO and GSEA analysis identified DDX39 as a pro-ccRCC gene. In vitro experiments confirmed the role of DDX39 in promoting ccRCC cell. Finally, DDX39 was found to be positively correlated with a variety of immune inhibitory markers, and could predict the adverse efficacy of immune checkpoint therapy in TIDE analysis. In conclusion, Increased DDX39 in ccRCC patients predicted worse clinical prognosis, promoted ccRCC cell proliferation, migration and invasion, and also predicted adverse efficacy of immune checkpoint therapy.
Keywords: ccRCC, DEAD Box protein 39, prognosis, cancer progression, immune microenvironment, immune checkpoint therapy

 Global reach, higher impact
Global reach, higher impact