Impact Factor ISSN: 1449-2288
Int J Biol Sci 2021; 17(13):3441-3455. doi:10.7150/ijbs.58872 This issue Cite
Review
The Function and related Diseases of Protein Crotonylation
1. Department of Biochemistry and Molecular Biology, Beijing Key Laboratory of Protein Posttranslational Modifications and Cell Function, School of Basic Medical Sciences, School of Basic Medical Sciences, Peking University Health Science Center, Beijing 100191, China.
2. China Institute of Veterinary Drugs Control, Beijing 100181, China.
3. Department of Radiation Medicine, Institute of Systems Biomedicine, School of Basic Medical Sciences, School of Basic Medical Sciences, Peking University Health Science Center, Beijing 100191, China.
Received 2021-1-31; Accepted 2021-7-14; Published 2021-8-9
Abstract
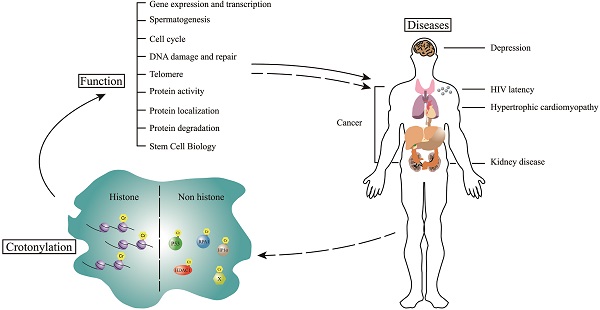
Crotonylation is a kind of newly discovered acylation modification. Thousands of crotonylation sites have been identified in histone and non-histone proteins over the past decade. As a modification closely related to acetylation, crotonylation was reported to share many universal enzymes with acetylation. Crotonylated proteins have important roles in the regulation of various biological processes, such as gene expression, process of spermatogenesis, cell cycle, and also in the pathogenesis of different diseases, which range from depression to cancer. In this review, we summarize the research processes of crotonylation and discuss the advances of regulation mechanism of both histone and non-histone proteins crotonylation in difference physiological processes. Also, we focus on the alteration of the crotonylation under certain pathological conditions and its role in the pathogenesis of each disease.
Keywords: crotonylation, histone crotonylation, non-histone crotonylation, disease
Introduction
Scientists have focused on epigenetic mechanism study since the concept of Epigenetics was firstly proposed by Austrian developmental biologist Conrad Waddington in 1942 [1]. In addition to DNA methylation, histone modifications have been found as a main component of the epigenetic code. Over the last ten years, a variety of histone acylation modifications have been discovered, including histone propionylation (Kpr) [2], butyrylation (Kbu) [2], 2-hydroxyisobutylacylation (Khib) [3], succinylation (Ksucc) [4], malonylation (Kma) [4], glutarylation (Kglu) [5], crotonylation (Kcr) [6], β-hydroxybutyrylation (Kbhb) [7], and benzoylation (Kbz) [8]. These modifications arise from their corresponding acyl-CoAs. Familiar with lysine acetylation, acylation mainly occurs on lysine residues through the addition of an acyl group from an acyl-CoA donor to the ε-amino group of the lysine side chain. However, there still have distinctions in chemical properties between different acylation. Kpr, Kbu, Kcr and Kbz belong to hydrophobic groups which can neutralise the positive charge of lysine residues (like acetylation) [9]. KSucc, Kmal and Kglu belong to acidic groups which change the positive charge to a negative charge. Khib and Kbhb belong to polar groups which allow hydrogen bond formation with interacting molecules [9]. Kbz stands out as the only known histone PTM with an aromatic acyl group, while Kcr is planar and Kbhb and Khib are branched [9]. The effect of structural biochemistry of acylation needs to further explore. Some data have shown that the known HAT and HDAC families have wide-ranging acylation and deacylation capabilities. Also, some acylation modifications were also found in non-histone proteins, indicating that protein acylation modifications are widely distributed in cells and play important roles in physiological and pathological processes.
Lysine crotonylation (Kcr) is one of histone lysine acylation modifications [6]. Crotonylation mainly occurs on the ε-amino group of lysine in histones and was recently reported to occurs on serine residue [10]. As mass spectrometry and proteomics developed, the existence of crotonylation has been confirmed in various microorganism (bacteria and fungus), plants and animals [11-13]. For example, Zhu J et al. found that Brucella secreted protein F (BspF) owned both acetylase and decrotonylase activity in Brucella. The data of LC-MS shown that a total of 5,559 crotonylation sites were identified on 1,525 different proteins, of which 331 sites on 265 proteins were significantly changed in BspF overexpression cells. So the authors speculated that BspF might influence the function of host proteins through its crotonylation to promote the intracellular propagation of Brucella [14]. This revealed a change of protein crotonylation in host cell mediated by microorganisms and provided a new perspective for further exploring the mechanism of microbial infection. In addition to the universality of crotonylation in various organisms, researches also indicated that crotonylation is a common protein modification which is not only limited to histone proteins, but also has potential effects on non-histone proteins. Histone Kcr chiefly enriches in the enhancer or transcription start site (TSS) of genes in human somatic genome and mouse male germ cell genome, which may activate specific genes [15]. What's more, crotonylation has been disclosed to involve in the physiological processes, such as RNA processing, nucleotide metabolism, chromatin recombination, regulation of protein activity and protein localization. Furthermore, crotonylation has countless ties with numerous diseases, like acute kidney injury (AKI) [16], IgA nephropathy [17], depression [18], HIV latency [19] and cancer [20]. This review summarizes the recent studies of crotonylation in detail and focuses on the mechanism and function of crotonylation both in histone and non-histone proteins in various physiological processes and different diseases.
The Research Progress of Crotonylation
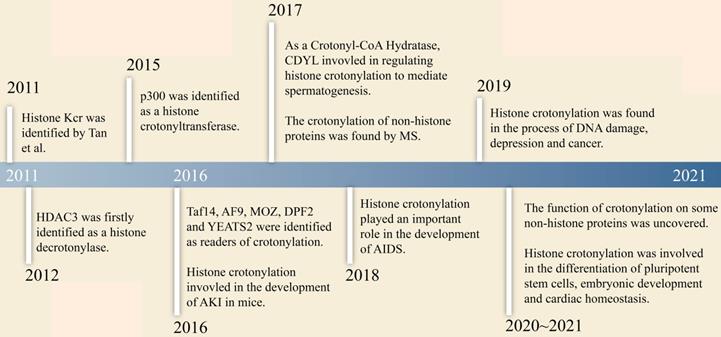
The research history of crotonylation is shown in Figure 1. In 2011, Tan et al. firstly reported that histone Kcr labelled either active promoters or potential enhancers in both human somatic cell and male mouse germ cell genomes [6]. Further research found that histone Kcr marked certain specific X-linked genes that escaped inactivation of sex chromosomes in haploid cells right after meiosis [6]. This study revealed a possible relationship between crotonylation and gene activation. Furthermore, Montellier et al. reported that histone Kcr was a dominant element in maintaining some genes activation in the globally repressive environment of haploid cell sex chromosomes by conferring resistance to transcriptional repressors [15]. This indicated that histone Kcr might be an indicator of the male haploid cell gene expression program. Subsequently, Sin et al. disclosed that RNF8 (an E3 ubiquitin-protein ligase) induced multiple modifications including Kcr, which led to gene activation from inactive sex chromosomes in post-meiotic spermatids. Kcr accumulated at transcriptional start sites of sex-linked genes in an RNF8-dependent manner, and increased the expression of RNF8 related genes [21]. Since then, the mechanism of histone Kcr during meiosis and post-meiosis in male germ cells was set forth. In 2017, histone crotonylation was found to play a key role in regulating spermatogenesis. Liu et al. showed that chromodomain Y-like transcription corepressor (CDYL) acted as a crotonyl-CoA hydratase and downregulated histone crotonylation. The level of histone crotonylation in round spermatids cells was significantly higher than those in spermatocytes cells, indicating that CDYL negatively regulated histone lysine crotonylation in spermatogenesis [22].
The research history of crotonylation.
The Writers/ Crotonyltransferases of crotonylation
In 2015, Sabari et al. showed that the coactivator p300 had both acetyltransferase and crotonyltransferase activities and found that p300-catalyzed histone crotonylation directly stimulated transcription to regulate gene expression [23]. Thus, a dynamic relationship between crotonylation and acetylation was speculated. Another study revealed that histone acetyltransferase (HAT) KAT8 (MOF) owns histone crotonyltransferase (HCT) activity, catalyzing the crotonylation in histone H3K4, H3K9, H3K18, H3K23, and histone H4K8 and H4K12 [24]. Also, CBP and p300 were found to be the major HCT in mammalian cells. CBP/p300 mutants with deficient HAT but competent HCT activity were able to substitute the endogenous CBP/p300 to enhance transcriptional activation, which correlated with enhanced promoter crotonylation and the recruitment of histone crotonylation reader protein Zinc finger protein ubi-d4 (DPF2) [24]. Another HAT, KAT2B (PCAF), was also confirmed as a HCT [25]. Also, Kollenstart et al. identified that HAT complex Gcn5-Ada2-Ada3 (ADA) and Esa1-Yng2-Epl1 (Piccolo NuA4) crotonylated histones in the N-terminal tails of histone H3 and H4 in budding yeast to promote crotonylation-dependent transcription [26].
The Erasers/Decrotonylases of crotonylation
The decrotonylases remove the covalent modification of lysine crotonylation. In 2012, histone deacetylase 3 (HDAC3) was firstly reported to own the activity of histone decrotonylase (HDCR) [27]. By systematic screening of the activities of the eleven human zinc-dependent lysine deacylases, HDAC3 in complex with nuclear receptor corepressor 1 (HDAC3-NCoR1) was shown to harbor decrotonylase activity in vitro [27]. Afterwards, SIRT1, SIRT2 and SIRT3 were indicated to act as histone decrotonylases in vitro [28, 29]. Wei et al. claimed that HDACs, rather than SIRTs, exerted the main activity of HDCR [30]. Histone crotonylation is a dynamic process as histone acetylation in mammalian cells and both catalytic centers of HDAC Class Ι and SIRTs have activities of HDAC and HDCR [30]. Kelly et al. found HDAC1/CoREST1/LSD1 complex was involved in both decrotonylation and deacetylation. In embryonic stem cells, knockdown HDAC1/2 increased global crotonylation levels of histone and caused a great reduction in total decrotonylase activity [31]. This study deepened the investigation of decrotonylase and further explored the mechanisms of decrotonylase.
The Readers of crotonylation
Certain specific domains are identified to participate in the process of transcriptional regulation induced by crotonylation. YEATS, Bromodomain and Double PHD finger (DPF) are three classes of domains that recognized acylation. In 2016, Andrews et al. found that transcription initiation factor TFIID subunit 14 (Taf14) was a reader of histone crotonylation [32]. Through π-π-π stacking mechanism, the YEATS domain of Taf14 bound to histone H3K9cr [32]. Gowans et al. discovered that histone Kcr was dynamically regulated during the yeast cell metabolic cycle (YMC) [33]. Under the condition of deficient nutrition, acetylation and crotonylation on the histone H3K9 site was dynamically changed in subsequence of Taf14 recognition, thus suppressing the expression of growth-related genes [33]. Several proteins with YEATS domain are identified, including protein AF-9 homolog (Yaf9), YEATS domain-containing protein 1 (protein ENL), protein AF-9 (AF-9), Taf14, and something about silencing protein 5 (Sas5). Li et al. disclosed that AF9 had stronger affinity with Kcr than Kac and regulated gene expression and positively activated transcription in YEATS domain-dependent manner [34]. Also, they found that YEATS domain-containing protein 2 (YEATS2) bound to a repertoire of acylated histone peptides with the best preference for histone H3 with crotonylation on lysine 27 (H3K27cr) [35]. Except for the YEATS domain, Bromodomain and DPF domain were also involved in the recognition of histone crotonylation. Researchers found that histone acetyltransferase KAT6A (MOZ) and DPF2 with DPF domain performed broad acylation of histone H3K14, including crotonylation (Kcr), butyrylation (Kbu), and propionylation (Kpr) with a preference for H3K14 crotonylation [36]. Immunofluorescence and chromatin immunoprecipitation quantitative PCR (ChIP-qPCR) showed that MOZ colocalized with histone H3K14cr in a DPF domain-dependent manner [36].
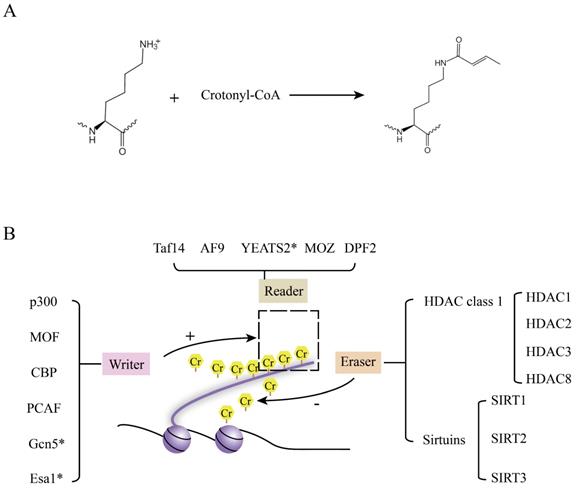
The chemical formula and the regulators of crotonylation are shown in Figure 2A and 2B. Writers (crotonyltransferases) and erasers (decrotonylases) regulated the dynamic balance of lysine crotonylation, playing different roles in a broad spectrum of cellular processes.
The chemical formula and regulators of crotonyaltion. A. The chemical formula and enzymatic reaction of crotonylation [66]. B. Readers, Writers (crotonyltransferases), and erasers (decrotonylases) are shown in this review. The readers are recruited by protein crotonylation. Writers and Erasers balance the protein crotonylation in vitro and in vivo. Esa1 and Gcn5 were found in budding yeast. YEATS2 was confirmed in vitro. Except for these, the remaining proteins were detected in human.
Other regulation factors of crotonylation
In addition to writers, erasers and readers, there are also other regulatory factors involved in the regulation of crotonylation. As a non-coding RNA, nuclear paraspeckle assembly transcript 1 (NEAT1) was associated with p300/CBP complex and its inhibition affected the location of H3K27 acetylation (H3K27Ac) and H3K27 crotonylation (H3K27Cr) to the transcription start site of many genes, including endocytosis related genes, which promotes the progression of disease [37]. Also, acyl-CoA synthetase short-chain family member 2(ASCC2) was a kind of crotonyl-CoA-producing enzyme and it was induced during the infection of HIV. The crotonylation regulated by ASCC2 was involved in the process of HIV reactivation in vivo [19]. In addition, CDYL was shown to act as a crotonyl-CoA hydratase and downregulated histone crotonylation to play an important role in regulation of crotonylation [22].
The Function of Crotonylation
The Function of Histone Crotonylation
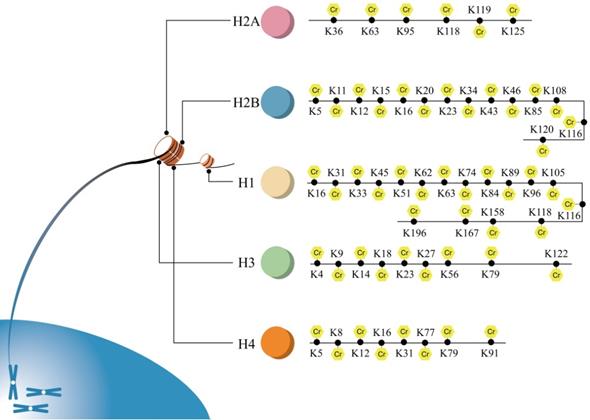
The identified sites of histone crotonylation are exhibited in Figure 3. Histone crotonylation was firstly reported to regulate gene transcription [6]. Histone crotonylation was shown to play roles in gene expression [6, 23], cell cycle [38], spermatogenesis [15, 21], aging [39] and DNA damage [40].
All sites of histone crotonylation. Sites of crotonylation were firstly discovered in histone. Until now, all of the sites of histone Kcr of H1, H2A, H2B, H3 and H4 are exhibited.
Gene expression and transcription
Transcription is a crucial process in the expression of coding genes [41]. Acetylation has been revealed to play a key role in gene expression in past decades. Through locating in TSS and enhancer region of target genes, acetylation relaxed highly condensed chromosomal structure to activate gene expression. Besides histone lysine acetylation, a repertoire of acylation types has been identified to regulate gene expression including histone crotonylation. The role of histone Kcr on gene transcription was firstly revealed by Tan et al. in haploid cells [6]. Sabari et al. showed that p300 catalyzed histone crotonylation in TSS and other regulatory elements of target genes such as Interleukin-6 (Il6), Guanylate-binding protein 2 (Gbp2), Interferon-induced protein with tetratricopeptide repeats 1 (Ifit1) and Radical S-adenosyl methionine domain-containing protein 2 (Rsad2) in response to LPS-induced macrophage inflammatory, thereby changing chromosome structure and promoting their expression [23] (Figure 4). Kollenstart et al. disclosed that Gcn5 and Esa1, as histone crotonylases, participated in cell metabolism through regulating gene expression, which was dependent on crotonate dosage [26]. During the self-renewal process of ESCs, histone crotonylation mediated by HDAC1 was reported to maintain the expression of transcription factor SOX-2, octamer-binding transcription factor 4 protein (Oct4) and Nanog [30]. However, researchers found that certain histone decrotonylation also promoted ESC differentiation by upregulating related gene expression [30]. These data suggested that crotonylation may have dual effects on gene expression.
The function of histone crotonylation in gene expression and transcription. In the LPS-induced inflammatory response, p300 induces histone crotonylation in TSS and other regulatory elements of target genes such as Il6, Gbp2, Ifit1 and Rsad2, thereby changing chromosome structure and promoting specific gene expression. AF9, as a reader, recognizes p300-catalyzed crotonylation to participate in the process of gene expression in macrophage.
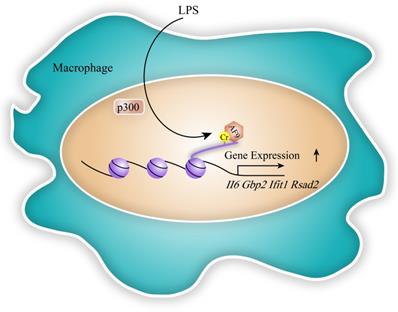
Meanwhile, some readers of histone Kcr were proved to involve in gene expression and transcription. Li et al. discovered that AF9, a reader of histone Kcr, was involved in the recognition of p300-catalyzed crotonylation and gene expression process in the LPS-induced inflammatory response (Figure 4). They showed that AF9 co-localizes with crotonylated histone H3K18 and positively regulates gene expression of Rsad2, II6, Ifit1 in a YEATS domain-dependent manner [34]. In addition, histone acetylation-binding DPF domains of human MOZ and DPF2 (also known as BAF45d) was shown to be the strongest preference for Kcr. MOZ and H3K14cr colocalized in a DPF-dependent manner, while H3K14cr was enriched in the genes which MOZ targeted in HEK 293T cells [36]. Moreover, YEATS2, a subunit of histone acetyltransferase ATAC complex, specifically recognized H3K27cr in vitro [35]. The significance of histone crotonylation sites involved in gene expression are shown in Table 1.
The significance of histone crotonylation sites reported in gene expression
| Histone crotonylation Sites | Activation/Inactivation | Description | References |
|---|---|---|---|
| H2B | |||
| K12 | activation | CDYL down-regulated gene expression through decreasing H2BK12cr on the promoter of genes such as BDNF, NEUROD1, SCG10, and MYT1. Downregulation of ECHS1 increased the level of histone crotonylation of H2BK12 and enhanced the expression of genes related to myocardial hypertrophy. | [22,59] |
| H3 | |||
| K9 | inactivation | Increased H3K9cr by crotonic acid suppressed the expression of growth-related genes. | [33] |
| K18 | activation | Downregulation of ECHS1 increased the level of histone crotonylation of H3K18 and enhanced the expression of genes related to myocardial hypertrophy. | [59] |
| K27 | activation | H3K27cr located in the regulatory elements of genes, and participated in the expression of genes, which are important for the proper differentiation of round spermatids into final sperm. | [67] |
Spermatogenesis
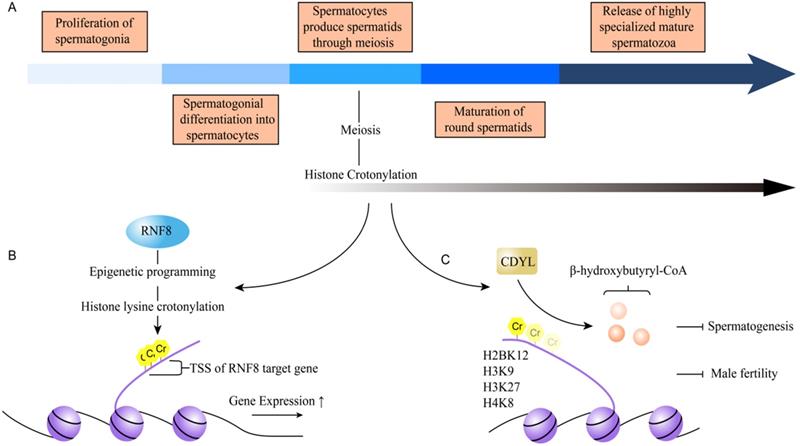
Spermatogenesis is a highly conserved physiological process. The complicated process contains five steps: the proliferation of spermatogonia, spermatogonial differentiation into spermatocytes, spermatids produced by spermatocytes through meiosis, maturation of round spermatids, and the release of highly specialized mature spermatozoa [42] (Figure 5A). Numerous factors are identified to affect spermatogenesis, such as obesity, diabetes, environmental chemicals, varicocele and epigenetic factors, especially. DNA content in the male germ cells (GCs) was packed in a very small volume to fit into the sperm head. Specific histones were replaced by protamine via hyperacetylation of histone H4 during spermatid stages, which plays an important role in spermatogenesis [43, 44]. Recently, histone Kcr was reported to play an important role in male haploid gene expression and sperm formation after meiosis [15]. Histone Kcr does not only mark the sex chromosome-related genes which are active in male germ cells after meiosis, but also mark the activated genes of autosomes after meiosis [15]. These studies indicated that histone Kcr may be a dominant factor in maintaining active gene activity in the overall suppressive environment of haploid sex chromosomes due to its resistance to transcription repressor [15]. Also, Sin et al. found that RNF8 brought about variation of chromosome conformation in epigenetic programming manner and regulated gene activation from inactive sex chromosomes in post-meiotic spermatids [21]. The epigenetic modifications included trimethylation of H3K4, histone Kcr, and incorporation of the histone variant H2AFZ [21]. It's worth noting that histone Kcr exceedingly accumulated around TSSs of sex-linked genes in a RNF8-dependent manner and activated gene expression [21] (Figure 5B). The removal of chromatin histones across the genome and their replacement by transition proteins (Tnps) and protamines (Prms) are another unique epigenetic event during spermatogenesis. Liu et al. reported that CDYL acts as a crotonyl-CoA hydratase which converts crotonyl-CoA to β-hydroxybutyryl-CoA and negatively regulates histone crotonylation [22]. The main histone crotonylation sites regulated by CDYL were H2BK12, H3K9, H3K27, and H4K8 (Figure 5C). They found that the gene expressions of peptidyl-prolyl cis-trans isomerase NIMA-interacting 4 (Pin4), coiled-coil domain-containing protein 160 (Ccdc160) and transcription elongation factor A protein-like 1 (Tceal) were greatly reduced in RS of CDYL transgenic mice, compared to that in wild-type mice. In addition, ChIP-qPCR assays detected the reduced levels of total histone Kcr and H2BK12cr on the promoter of related genes, showing that CDYL regulated the expression of sex chromosome-linked escaped genes through mainly influencing histone Kcr on the gene promoters to play an important role in spermatogenesis and thus male fertility [22]. Together, these results provide us a new insight to understand the role of histone crotonylation in spermatogenesis.
The mechanism of crotonylation involves in spermatogenesis. A. The process of spermatogenesis contains the proliferation of spermatogonia, spermatogonial differentiation into spermatocytes, spermatids produced by spermatocytes through meiosis, maturation of round spermatids, and the release of highly specialized mature spermatozoa. B. Histone Kcr is one of epigenetic programming caused by RNF-8. It is exceedingly accumulated around TSSs of sex-linked genes in a RNF8-dependent manner and activates gene expression. C. CDYL negatively regulates histone crotonylation, to affect spermatogenesis, and the normal function of testicular and male fertility. The main histone crotonylation sites regulated by CDYL are H2BK12, H3K9, H3K27 and H4K8.
Cell cycle
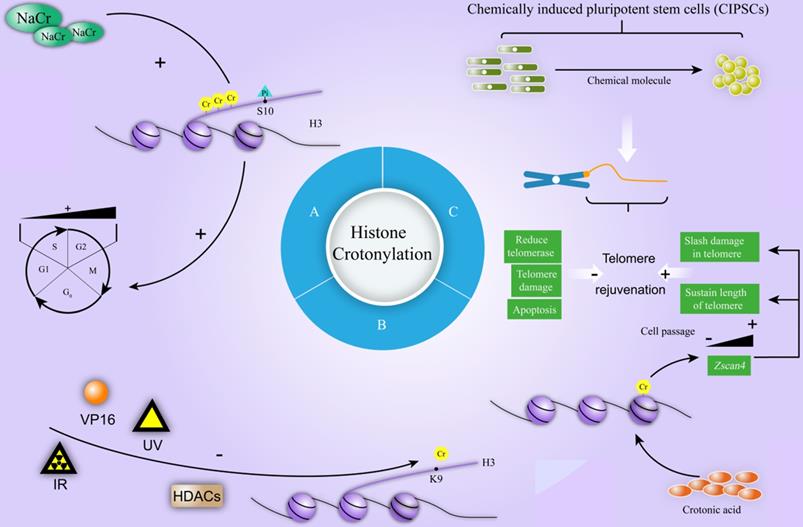
Cell cycle contains four phases (G1, S, G2, and M). Many intracellular and extracellular factors influence this process. During the cell cycle, certain protein has been dynamically regulated by modification, including ubiquitination, methylation, and phosphorylation. Serine phosphorylation, which most appears in the late of G2 stage and surges in H3 in the M stage, has been mostly studied [45]. Recently, researchers have initially explored the effect of crotonylation modification during cell cycle. Sodium Crotonate (NaCr) was found to enhance the crotonylation level of histone H3, and influence the phosphorylation of histone H3 at the Ser10 site in a dose-dependent manner, which is the mark of G2 / M phase cells [38]. Flow cytometry results further disclosed that the amount of S phase cells declined and G2 phase cells raised after NaCr treatment [38] (Figure 6A). Considering cell cycle comprises four phases and involves complex biological processes, the mechanism of histone crotonylation during cell cycle needs to further investigation.
The mechanism of crotonylation involves in cell cycle, DNA damage repair and CIPSCs. A. NaCr increases the crotonylation of histone H3 and influences the phosphorylation of histone H3 at the Ser10 site in a dose-dependent manner, which is the mark of G2 / M phase cells. NaCr treatment decreases the amount of S phase cells and raises cells in G2 phase. B. Ionizing Radiation (IR), Ultraviolet Radiation (UV), or Etopside (VP16) decreases the levels of H3K9cr. During this process, HDACs are the major HDCR in U2OS cells. C. During chemically induced pluripotent stem cells (CIPSCs), telomere rejuvenation goes through a dynamic process. Long-term induction by small molecules reduces telomerase, increases telomere damage and apoptosis, and finally contributes to negatively regulate telomere rejuvenation (shortened telomeres, limited the formation of CIPSCs). Crotonic acid positively regulates telomere rejuvenation through promoting the expression of Zscan4 with cell passage, which slashes damage in telomere, and sustains the length of telomere during chemically induced reprogramming.
DNA damage response
Post-translational modification plays an important role in DNA damage Response (DDR) [46]. Recently, a study shown the role of HDACs in regulating histone crotonylation in DDR [40]. After exposure to ionizing radiation (IR), ultraviolet radiation (UV), or treatment with Etopside (VP16), the rapid decrease of H3K9cr levels was detected and the reduction of H3K9cr levels in U2OS cells were transient and could be restored to basal level after few hours [40]. The treatment with HDACs inhibitor trichostatin A (TSA) induced severe increase in the levels of H3K9cr, indicating that HDACs are the major lysine decrotonylases (Figure 6B) [40]. These data only revealed that H3K9 crotonylation is reduced at damage sites in a HDAC-dependent manner under various DNA damage stimulation and the exact mechanism of histone crotonylation in the process of DNA damage is need to further explore.
Telomere
Telomere is a small DNA-protein complex at the ends of the linear chromosomes of eukaryotic cell. Cap-like structure covers the ends of chromosomes to maintain the stability of the genome [47]. Telomere length is mainly determined by genes, environmental factors and molecular pathways [48]. It is associated with infinite self-renewal and pluripotency, while shortened telomere causes cell senescence or tumorigenesis. Fu et al. disclosed that telomere length strongly correlated with the degree of reprogramming, pluripotency, and differentiation capacity of chemically induced pluripotent stem cells (CIPSCs). Further exploration found that long-term induction by small molecules reduced telomerase, increased telomere damage and apoptosis, and finally contributed to shorten telomeres and limited the formation of CIPSCs [39]. Mechanically, crotonic acid induced histone crotonylation activated the expression of Zinc finger and SCAN domain-containing protein 4 (Zscan4) gene by decreasing the abundance of heterochromatic H3K9me3 and HP1α at subtelomere. The induced expression of Zscan4 in turn slashed damage in telomere, and sustained length of telomere during chemically induced reprogramming [39] (Figure 6C).
Stem Cell Biology
In mouse ES cells, histone decrotonylation was found to promote ES cell differentiation. Histone Kcr was elevated compared to differentiated cells [30]. Also, Fu et al. identified the role for histone crotonylation on dynamics of telomere rejuvenation during chemical reprogramming in pluripotent stem cells [39]. In 2021, Y. Fang et al reported that histone crotonylation had an important role in the endoderm commitment of human embryonic stem cells [49]. They found that H4K77cr and H4K91cr were increased during endoderm differentiation of hESCs [49]. In the meanwhile, the expression of key enzymes involved in the metabolism of crotonyl CoA, such as peroxisomal acyl-coenzyme A oxidase 3 (ACOX3), short-chain specific acyl-CoA dehydrogenase (ACADS) and ACSS2, were significantly increased in endoderm differentiation [49]. These data indicated that crotonyl-CoA-producing enzymes modulated histone crotonylation and regulated endoderm differentiation. Also, the data of global profiling of the lysine crotonylome in different pluripotent states revealed that the important role of protein crotonylation in the maintenance and transformation of pluripotent stem cells [50]. They identified 3628 high-confidence crotonylated sites in 1426 proteins, which are involved in functions/processes related to pluripotency such as RNA biogenesis, central carbon metabolism, and proteasome function [50]. These reports indicated the crucial role of histone crotonylation on stem cell biology.
The Function of non-histone Crotonylation
Protein activity
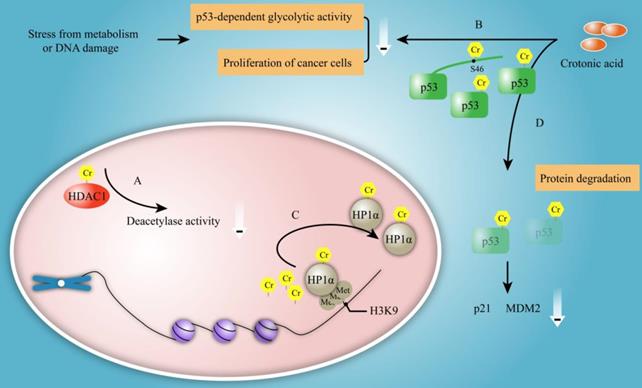
The function of crotonylation on non-histone protein is paid more and more attention except for its roles on histone. Hundreds of crotonylated proteins and lysine residues have been identified using specific antibody enrichment followed by high-resolution mass spectrometry analysis. Bioinformatics analysis revealed that crotonylated proteins were particularly enriched for nuclear proteins involved in several physiological processes [38]. The crotonylation of HDAC1 was enhanced by NaCr and the crotonylated HDAC1 exhibited reduction in its deacetylase activity compared with unmodified HDAC1 (Figure 7A). Also, p53 was found to be crotonylated at serine 46 after crotonic acid treatment that led to the inhibition of p53 activity in a dose-dependent fashion [10]. The crotonylation of p53 lowered the activity of p53 protein to enhance the p53-dependent glycolytic activity and the proliferation of cancer cells in response to stress from metabolism or DNA damage [10] (Figure 7B).
The functional of non-histone crotonylation. A. The crotonylation of HDAC1 can be enhanced by NaCr and crotonylated HDAC1 reduces its deacetylase activity. B. p53 is crotonylated at its serine 46 after CA treatment that leads to the inhibition of p53 activity in a dose dependent fashion. The crotonylation lowered activity of p53 enhances p53-dependent glycolytic activity and proliferation of cancer cells in response to stress from metabolism or DNA damage. C. Crotonylation of HP1α is shown to alter its localization from heterochromatin to nuclear plasm and dramatically reduces its binding of trimethylated H3 at K9 position, which results in variation of localization in vitro. D. CA treatment induces p53 crotonylation with the reduction of the protein level of p53 and its downstream protein p21 and MDM2.
Protein localization
Crotonylation has been reported to affect the localization of proteins. HP1α (also named chromobox protein homolog 5, CBX5) belongs to the heterochromatin family and enriches in heterochromatin by binding to methylated histones [38]. HP1α is primarily localized at nucleus and accumulates at bright dot-like heterochromatin regions in HeLa cells. The localization of HP1α was changed from the heterochromatin to the nuclear plasm after treatment with either TSA or NaCr for 72 hours [38]. For mechanism study, the authors found that NaCr treatment resulted in crotonylation of ectopically expressed Flag-HP1α by western blotting analysis. Also, the results of pulldown experiment showed that crotonylated HP1α dramatically reduced its binding of trimethylated H3K9 residue in vitro [38]. These data provided evidence that crotonylation might affect the localization of proteins. Researcher speculated that crotonylated HP1α might be resulted in the variation of localization by reducing its binding of trimethylated H3 at K9 residue in vitro [38] (Figure 7C).
Protein degradation
There are multiple pathways to break down protein. Lysosomal pathway is an approach for protein degradation entered cell in vitro; while for intracellular proteins, the way of degradation is ubiquitin proteasome pathway. Many researches have shown that protein acylation is closely related to proteasome-dependent protein degradation [51, 52]. Similarity to acetylation, whether crotonylation is associated with protein degradation or not? Liao et al. reported that crotonic acid induced the crotonylation of p53 and thus affected its degradation [10] (Figure 7D). Yet, the degradation pathways that were involved in the regulation of p53 degradation have not been identified. Furthermore, whether there is any association between crotonylation, acetylation and ubiquitination in protein degradation is poor understand. In the proteolytic pathway that relies on ubiquitin proteasome, acetylated protein was protected from ubiquitination and proteasome degradation [53]. One possible mechanism is that acetylation and ubiquitination may compete for same lysine sites. In proteasome-independent degradation pathways, acetylation can also influence the stability of protein. Due to the similarity of crotonylation and acetylation, it needs further exploration concerning the effect of crotonylation on protein degradation.
DNA repair
The repair of DNA damage is a complex process that relies on particular pathways to remedy specific types of damage to DNA. Crotonylation also plays a key role in DNA repair. In 2020, Yu et al. had identified replication protein A 70 kDa DNA-binding subunit (RPA1) as one of the downstream substrates of CDYL [54]. They found that CDYL negatively regulated Kcr of RPA1 at K88, K379 and K595 sites, which were involved in DNA damage [54]. Under the treatment of CPT, CDYL participated in up-regulation of Kcr in RPA1, which enhanced the interaction of RPA1 with ssDNA and/or HR factors. Combined with the physiological processes in the body, the researchers found that Kcr of RPA1 was important to cell survival and anti-apoptotic response under DNA-damaging conditions [54]. The data above suggested that non-histone as well as histone crotonylation exhibited a key role in the process of DNA repair.
Crotonylation is an important epigenetic modification form identified as a novel evolutionarily conserved histone PTM. Both histone and non-histone proteins were found to be crotonylated. Crotonylation is involved in various biological pathways that regulate diverse cellular functions ranging from gene expression to DNA damage. It is interesting to determine how their functions are mechanistically regulated by crotonylation in future.
Protein Crotonylation Associated Diseases
Kcr is a newly discovered post-translational modification, hence the associations between crotonylation and diseases are not fully clarified. More studies are needed to investigate the role of crotonylation in different diseases, which may provide new target for clinical therapy. The protein crotonylation associated diseases were summarized in Table 2.
The diseases associated with crotonylation
| Diseases | Protein crotonylation | Description | References |
|---|---|---|---|
| Depression | Histone | CDYL may inhibit structural synaptic plasticity mainly by transcriptional repression of neuropeptide VGF. CDYL-mediated histone crotonylation played a critical role in regulating stress-induced depression | [18] |
| HIV latency | H3K4 | Histone H3K4 crotonylation by ACSS2 inducted reprogramed histone tails at the HIV LTR and suppressed ACSS2 dampened reactivation of latent HIV. | [19] |
| Kidney disease | |||
| Acute kidney injury (AKI) | Histone | Crotonate protected experimental mice from AKI by preventing the decline of renal function, increased expression of renal PGC-1α and SIRT3, and decreased of CCL2 though histone crotonylation. | [16] |
| IgA nephropathy | 353 crotonylated proteins | Bioinformatics analysis identified 353 crotonylated proteins. Genomes and functional enrichment analyses suggested significant enrichment of crotonylated proteins displaying important relationships with IgA nephropathy. | [17] |
| Hemodialysis | 347 crotonylated proteins | Proteome of crotonylation was confirmed to evaluate the role of lysine crotonylation in maintenance hemodialysis patients (MHP) and found that lysine crotonylation played important regulatory roles in pathophysiological processes in MHP. | [58] |
| Hypertrophic cardiomyopathy | H3K18; H2BK12 | Downregulation of ECHS1 increased the level of histone crotonylation of H3K18 and H2BK12, and enhanced the expression of genes related to myocardial hypertrophy, such as natriuretic peptides B (Nppb), ultimately leading to the development of HCM. | [59] |
| Cancer | Numerous non-histone proteins | Quantitative proteomics study showed that p300-targeted Kcr substrates owned the potentially linked to cancer. Also, numerous non-histone proteins were shown to involve in the process of tumorigenesis by LC-MS/MS. | [20,62] |
Depression
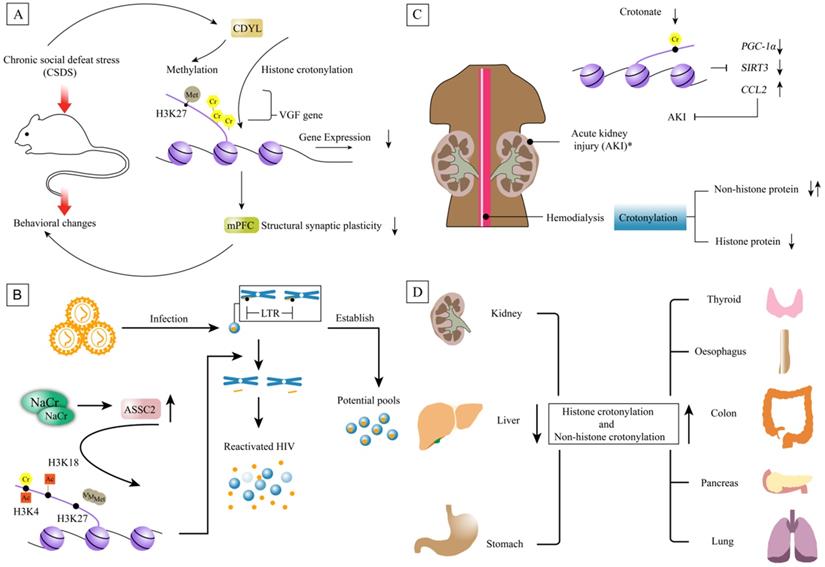
Depression is a psychological disease and the pathogenesis of depression is perplexing, but it is currently believed that depression is mainly determined by related genes and environmental factors. Also, PTM was shown to involve in this process, including acetylation, methylation, phosphorylation, and so on. For example, H3K14ac levels in hippocampus of male C57/Bl6J mice showed a significant increase after ten continuous days of social defeat stress [55]. Another study revealed that HDAC was altered in the mood disorder patients. The expression of HDAC2, 3, 4 and 5 mRNA were increased in a depressive state in major depressive disorder patients. The expression of HDAC4 mRNA was increased only in a depressive state, and the expression of HDAC6, 7 and 8 were decreased in both depressive and remissive states in bipolar disorder patients, indicating that the altered expression of HDACs was associated with the pathophysiology of mood disorders [56]. Interestingly, histone Kcr was reduced in the stress-susceptible rodents exposed in the medial prefrontal cortex concurrent with selective upregulation of CDYL [18]. Mechanistically, they found that CDYL inhibited structural synaptic plasticity mainly by transcriptional repression of neuropeptide VGF nerve growth factor inducible through its dual effect on histone crotonylation and H3K27 trimethylation on the VGF promoter [18]. CDYL-VGF axis inhibited the structural synaptic plasticity of medial prefrontal cortex (mPFC), eventually leading to behavioral changes in susceptible individuals (Figure 8A).
Non-histone and histone crotonylation associated diseases. A. The behavioral changes of rodents are caused by chronic social defeat stress. Because CDYL can make dual effect on histone crotonylation and H3K27 trimethylation on the VGF promoter. One of mechanism is that CDYL inhibits structural synaptic plasticity mainly by transcriptional repression of neuropeptide VGF. CDYL-VGF axis inhibits the structural synaptic plasticity of medial prefrontal cortex (mPFC), eventually leading to behavioral changes in susceptible individuals. B. NaCr increases the expression of ASSC2, thereby enhancing the expression of histone H3K4 crotonylation, H3K4 and H3K18 acetylation, and reducing H3K27 trimethylation. ACSS2-driven crotonylation of HIV LTR histones reshapes histone and reactivates HIV from the incubation period. ACSS2-driven crotonylation of histone where HIV LTR combine with DNA reshapes histone and reactivates HIV from the incubation period. C. Crotonate can protect experimental mice from AKI by preventing the decline of renal function, increase expression of renal PGC-1α and SIRT3, and decrease of CCL2 though histone crotonylation. Note: this experiment was confirmed in mice. Proteome of crotonylation in maintenance hemodialysis patients (MHP) find that histone crotonylation is decreased in MHP. D. The whole level of crotonylation is down-regulated in liver, stomach and kidney cancers. However, it is upregulated in thyroid, oesophagus, colon, pancreas and lung carcinomas.
HIV latency
Human immunodeficiency virus (HIV) is a retrovirus which causes a multisystemic disease called acquired immunodeficiency syndrome (AIDS). HIV are divided into two types: HIV-1 and HIV-2, of which the former is more pathogenic. People who get infection has a long incubation period and its long latency is a main reason for its incurability. Researchers have found that the epigenetic regulation of histone played a key role in the process of HIV latency. NaCr is a strong latent reversal agent (LRAs) to increase the expression of acyl-CoA synthetase short-chain family member 2 (ASSC2) in vitro, thereby enhancing histone H3K4 crotonylation, H3K4 and H3K18 acetylation, and reducing H3K27 trimethylation [19]. ACSS2-driven crotonylation of histone where HIV long terminal repeat (LTR) combined with DNA reshaped histone and reactivated HIV from the incubation period [19]. A similar phenomenon occurred in CD4+ T cells, and histone crotonylation combined with other LRAs interfered with HIV latency. In the rhesus monkey-AIDS animal model infected with simian immunodeficiency virus (SIV), the expression of ACSS2 was highly induced in the intestine during acute primary SIV infection in vivo. Entering the chronic stage, the expression of ACSS2 decreased [19] (Figure 8B). These results suggested a potential role of histone decrotonylation in the establishment of HIV latency.
Kidney disease
Various epigenetic mechanisms are involved in different kidney diseases, such as acute kidney injury (AKI), chronic kidney disease (CKD) and AKI-CKD conversion [57]. Histone crotylation is regulated by the concentration of crotonic acid and crotyl coenzyme A, and the crotonate has been applied to the folate-induced AKI model. By increasing histone crotonylation, crotonate reduced the inflammatory response and increased kidney function [16]. Nevertheless, the mechanism regulated by crotonylation in kidney disease is still not clear and need to further explore.
Acute kidney injury (AKI)
Acute kidney injury (AKI) refers to sudden (within 1-7d) and sustained (>24 h) declines in renal function, manifested by azotemia, water electrolyte and acid-base imbalance, and systemic symptoms. Post-translational histone modifications modulate gene expression under the circumstance of AKI. Recently, Olga et al. observed that histone H3k9cr was increased in folic-acid-induced AKI tissue and disclosed that inflammatory factors may participate in regulation of histone crotonylation in kidney tubular cells [16]. The results of ChIP-seq revealed enrichment of histone crotonylation at the genes encoding the mitochondrial biogenesis regulator PGC-1α and the decrotonylase SIRT3 in both TWEAK-stimulated tubular cells and in AKI kidney tissue [16]. Following these results, researchers evaluated the effect of variation in histone crotonylation on AKI. Crotonate heightened expression of PGC-1α and SIRT3 and limited the expression of C-C motif ligand 2 (CCL2) in healthy kidneys and renal tubular cell. Consistent with these results, crotonate protected experimental mice from AKI by preventing the decrease of renal PGC-1α and SIRT3 and increase of CCL2 [16] (Figure 8C). Cell stress and crotonate availability increased histone crotonylation in vivo and increasing histone crotonylation might have a protective effect from AKI.
IgA nephropathy
Immunoglobulin A nephropathy (IgAN) is one of the most common glomerular disease, characterized by IgA deposition, with or without the deposition of other immunoglobulins in the mesangial region. A proteomics analysis of crotonylation between healthy controls and IgAN patients was performed. By analyzing results in a bioinformatics manner, researchers focused on the characteristics of the crotonylayed proteins. Also, the functions of the differential crotonylayed proteins and the characteristics of the crotonylation sites in the amino acid sequence of proteins were analyzed. Integrated analysis revealed that crotonylated proteins were mainly involved in the humoral immune response in patients, especially in antigen processing and presentation [17].
Hemodialysis
Hemodialysis (HD) is one of the renal replacement treatments for patients with acute and chronic renal failure. Liquid chromatography tandem mass spectrometry (LC-MS/MS) coupled with highly sensitive immune-affinity purification was used to comparatively evaluate the crotonylation proteome of normal controls and maintenance hemodialysis patients (Figure 8C). There were 96 increased and 253 decreased crotonylated proteins have been confirmed, the decrease level of crotonylation in histones was disclosed in patients with kidney failure undergoing maintenance hemodialysis [58]. With KEGG analysis, upregulated crotonylated proteins were found to involve in different physiological process of HD, such as complement and coagulation cascades, cardiac muscle contraction, and hematopoietic cell lineage [58]. More researches are needed to disclose the relationships between crotonylation and the physiological and pathological processes in HD patients, which might provide a possible therapeutic target against HD complications and hopefully improve the quality of life of HD patients.
Hypertrophic cardiomyopathy
Hypertrophic cardiomyopathy (HCM) is the most common genetic cardiovascular disease characterized by unexplained non dilated left ventricular hypertrophy (LVH). Recently, downregulation of short-chain enoyl-CoA hydratase (ECHS1) was showed in human hearts with hypertrophic cardiomyopathy. ECHS1 was reported to mediate histone crotonylation and contributed to cardiac homeostasis [59]. They found that downregulation of ECHS1 increased the level of histone crotonylation of H3K18 and H2BK12, and enhanced the expression of genes related to myocardial hypertrophy, such as natriuretic peptides B (Nppb), ultimately leading to the development of HCM [59]. Hopefully, it provides a potential target for treatment of HCM.
Cancer
Cancer is a major public health problem and owns high leading cause of death worldwide[60]. Epigenetics is one of the major focus and extensive research on histone methylation, acetylation, and phosphorylation has been performed. Recently, a quantitative proteomics study characterized the p300-regulated lysine crotonylome and showed that p300-targeted Kcr substrates the potentially linked to cancer [61]. These suggest that crotonylation may act as a carcinogenic factor to promote tumor progress. In the EDRN database, 4.5% (20 out of 443) of tumor biomarkers have been crotonylated and 32 crotonylated proteins are associated with tumor genes [61]. The crotonylated proteins associated tumor genes are important for tumorigenesis and tumor development. In adrenal aldosterone producing adenoma, the crotonylated protein associated gene is ATPase Na+/K+ transporting subunit alpha 1 (ATP1A1). In Spitzoid tumor, the genes are lamin A/C (LMNA) and lamin B2 (LMNB2). In anaplastic large cell lymphoma, the genes related protein crotonylation are moesin (MSN), myosin heavy chain 9 (MYH9) and myosin heavy chain 10 (MYH10) [61]. The role of crotonylation on these proteins associated with tumor genes needs to be further explored[61]. Also, p300 was shown to mediate the expression of HnRNPA1 by lysine crotonylation to further promote the proliferation, invasion and migration of HeLa cells [62]. Furthermore, the level of crotonylation in several tumors were detected and found that the level of crotonylation was down-regulated in liver cancer, gastric cancer and renal cancer, while it was upregulated in thyroid, oesophagus, colon, pancreas and lung carcinoma [20] (Figure 8D). In hepatocellular carcinoma, the expression of Kcr was correlated with cancer staging [20]. In addition to histone crotonylation, numerous non-histone proteins are involved in the process of tumorigenesis. 2 696 crotonylation sites were identified in total on 1024 proteins of the human lung adenocarcinoma H1299 cell line [25]. Enrichment analysis of biological processes indicated that these crotonylated proteins are enriched in a variety of biological processes including transport, nucleobase-containing compound metabolic process and heterocycle metabolic process of lung adenocarcinoma cell line [25]. These data suggested that non-histone crotonylation might be play a critical role in tumorigenesis.
Conclusion and Perspectives
Kcr is one of the non-acetyl lysine acylation modifications, which caused more and more concern. This article generalizes the related researches on crotonylation over decades by discussing the research processes of crotonylation and its involved physiological and pathological function. Particularly, we summarized histones and non-histones protein crotonylation in combination with specific modification sites, which shows that protein crotonylation is widely distributed in cells and it has significant roles in countless physiological processes. Lysine histone and non-histone proteins crotonylation is involved in many physiological processes from gene expression to protein stability. Also, the histone and non-histone proteins crotonylation associated diseases were discussed deeply here. It will be interesting to determine how their functions are mechanistically regulated by crotonylation in the near future.
Studies have shown that acetylation and crotonylation coexist in TSS or other regulatory elements of certain genes. This phenomenon suggests that there are intricate correlations between histone crotonylation and other types of histone modifications in the regulation of gene expression in vivo. The mechanism of crotonylation and its dynamic relationship with other acylation needs further exploration.
Until now, drugs that target histone crotonylation are still not available. Countless reasons contribute to its perplexity. Due to complex biochemical reactions of ester drugs metabolism in body, the lack of specific crotonylated enzyme is a main challenge, which limits further investigation of drugs targeting histone crotonylation. Despite the obstacles, studies on non-histone crotonylation made breakthroughs with the developments in high‐resolution LC‐MS/MS approaches. In 2017, Z Ju et al. designed the new coding schemes to predict protein crotonylation sites through databases [63, 64]. In 2018, J Bos et al. designed a chemical probe for crotonylation on endogenous protein as a supplement to specific antibodies to detect crotonylation modifications [65]. The combination of traditional biochemical technology with bioinformatics method will be benefit to investigate the function of protein crotonylation on many cellular processes mechanistically. While understanding the roles of crotonylation in physiological processes and in the development of diseases, it will guide the development of drugs that target of protein crotonylation and provide us a new theoretical basis for clinical diseases therapy.
Acknowledgements
This work was supported by National Natural Science Foundation of China Grants 81672778 and 81372165. The authors would like to thank Dr. Xin Ye for language editing the manuscript prior to submission.
Competing Interests
The authors have declared that no competing interest exists.
References
1. Waddington CH. The epigenotype. 1942. Int J Epidemiol. 2012;41:10-3
2. Chen Y, Sprung R, Tang Y, Ball H, Sangras B, Kim SC. et al. Lysine propionylation and butyrylation are novel post-translational modifications in histones. Mol Cell Proteomics. 2007;6:812-9
3. Dai L, Peng C, Montellier E, Lu Z, Chen Y, Ishii H. et al. Lysine 2-hydroxyisobutyrylation is a widely distributed active histone mark. Nat Chem Biol. 2014;10:365-70
4. Xie Z, Dai J, Dai L, Tan M, Cheng Z, Wu Y. et al. Lysine succinylation and lysine malonylation in histones. Mol Cell Proteomics. 2012;11:100-7
5. Tan M, Peng C, Anderson KA, Chhoy P, Xie Z, Dai L. et al. Lysine glutarylation is a protein posttranslational modification regulated by SIRT5. Cell Metab. 2014;19:605-17
6. Tan M, Luo H, Lee S, Jin F, Yang JS, Montellier E. et al. Identification of 67 histone marks and histone lysine crotonylation as a new type of histone modification. Cell. 2011;146:1016-28
7. Xie Z, Zhang D, Chung D, Tang Z, Huang H, Dai L. et al. Metabolic Regulation of Gene Expression by Histone Lysine beta-Hydroxybutyrylation. Mol Cell. 2016;62:194-206
8. Ren X, Zhou Y, Xue Z, Hao N, Li Y, Guo X. et al. Histone benzoylation serves as an epigenetic mark for DPF and YEATS family proteins. Nucleic Acids Res. 2020;49:114-126
9. Barnes CE, English DM, Cowley SM. Acetylation & Co: an expanding repertoire of histone acylations regulates chromatin and transcription. Essays Biochem. 2019;63:97-107
10. Liao P, Bhattarai N, Cao B, Zhou X, Jung JH, Damera K. et al. Crotonylation at serine 46 impairs p53 activity. Biochem Biophys Res Commun. 2020;524:730-735
11. Sun H, Liu X, Li F, Li W, Zhang J, Xiao Z. et al. First comprehensive proteome analysis of lysine crotonylation in seedling leaves of Nicotiana tabacum. Sci Rep. 2017;7:3013
12. Kwon OK, Kim SJ, Lee S. First profiling of lysine crotonylation of myofilament proteins and ribosomal proteins in zebrafish embryos. Sci Rep. 2018;8:3652
13. Yang Q, Li Y, Apaliya MT, Zheng X, Serwah BNA, Zhang X. et al. The Response of Rhodotorula mucilaginosa to Patulin Based on Lysine Crotonylation. Front Microbiol. 2018;9:2025
14. Zhu J, Dong Q, Dong C, Zhang X, Zhang H, Chen Z. Global Lysine Crotonylation Alterations of Host Cell Proteins Caused by Brucella Effector BspF. Front Cell Infect Microbiol. 2020;10:603457
15. Montellier E, Rousseaux S, Zhao Y, Khochbin S. Histone crotonylation specifically marks the haploid male germ cell gene expression program: post-meiotic male-specific gene expression. Bioessays. 2012;34:187-93
16. Ruiz-Andres O, Sanchez-Nino MD, Cannata-Ortiz P, Ruiz-Ortega M, Egido J, Ortiz A. et al. Histone lysine crotonylation during acute kidney injury in mice. Dis Model Mech. 2016;9:633-45
17. Lin H, Tang D, Xu Y, Zhang R, Ou M, Zheng F. et al. Quantitative analysis of protein crotonylation identifies its association with immunoglobulin A nephropathy. Mol Med Rep. 2020;21:1242-1250
18. Liu Y, Li M, Fan M, Song Y, Yu H, Zhi X. et al. Chromodomain Y-like Protein-Mediated Histone Crotonylation Regulates Stress-Induced Depressive Behaviors. Biol Psychiatry. 2019;85:635-49
19. Jiang G, Nguyen D, Archin NM, Yukl SA, Mendez-Lagares G, Tang Y. et al. HIV latency is reversed by ACSS2-driven histone crotonylation. J Clin Invest. 2018;128:1190-8
20. Wan J, Liu H, Ming L. Lysine crotonylation is involved in hepatocellular carcinoma progression. Biomed Pharmacother. 2019;111:976-82
21. Sin HS, Barski A, Zhang F, Kartashov AV, Nussenzweig A, Chen J. et al. RNF8 regulates active epigenetic modifications and escape gene activation from inactive sex chromosomes in post-meiotic spermatids. Genes Dev. 2012;26:2737-48
22. Liu S, Yu H, Liu Y, Liu X, Zhang Y, Bu C. et al. Chromodomain Protein CDYL Acts as a Crotonyl-CoA Hydratase to Regulate Histone Crotonylation and Spermatogenesis. Mol Cell. 2017;67:853-66 e5
23. Sabari BR, Tang Z, Huang H, Yong-Gonzalez V, Molina H, Kong HE. et al. Intracellular crotonyl-CoA stimulates transcription through p300-catalyzed histone crotonylation. Mol Cell. 2015;58:203-15
24. Liu X, Wei W, Liu Y, Yang X, Wu J, Zhang Y. et al. MOF as an evolutionarily conserved histone crotonyltransferase and transcriptional activation by histone acetyltransferase-deficient and crotonyltransferase-competent CBP/p300. Cell Discov. 2017;3:17016
25. Xu W, Wan J, Zhan J, Li X, He H, Shi Z. et al. Global profiling of crotonylation on non-histone proteins. Cell Res. 2017;27:946-9
26. Kollenstart L, de Groot AJL, Janssen GMC, Cheng X, Vreeken K, Martino F. et al. Gcn5 and Esa1 function as histone crotonyltransferases to regulate crotonylation-dependent transcription. J Biol Chem. 2019;294:20122-34
27. Madsen AS, Olsen CA. Profiling of substrates for zinc-dependent lysine deacylase enzymes: HDAC3 exhibits decrotonylase activity in vitro. Angew Chem Int Ed Engl. 2012;51:9083-7
28. Feldman JL, Baeza J, Denu JM. Activation of the protein deacetylase SIRT6 by long-chain fatty acids and widespread deacylation by mammalian sirtuins. J Biol Chem. 2013;288:31350-6
29. Bao X, Wang Y, Li X, Li XM, Liu Z, Yang T. et al. Identification of 'erasers' for lysine crotonylated histone marks using a chemical proteomics approach. Elife. 2014 3
30. Wei W, Liu X, Chen J, Gao S, Lu L, Zhang H. et al. Class I histone deacetylases are major histone decrotonylases: evidence for critical and broad function of histone crotonylation in transcription. Cell Res. 2017;27:898-915
31. Kelly RDW, Chandru A, Watson PJ, Song Y, Blades M, Robertson NS. et al. Histone deacetylase (HDAC) 1 and 2 complexes regulate both histone acetylation and crotonylation in vivo. Sci Rep. 2018;8:14690
32. Andrews FH, Shinsky SA, Shanle EK, Bridgers JB, Gest A, Tsun IK. et al. The Taf14 YEATS domain is a reader of histone crotonylation. Nat Chem Biol. 2016;12:396-8
33. Gowans GJ, Bridgers JB, Zhang J, Dronamraju R, Burnetti A, King DA. et al. Recognition of Histone Crotonylation by Taf14 Links Metabolic State to Gene Expression. Mol Cell. 2019;76:909-21 e3
34. Li Y, Sabari BR, Panchenko T, Wen H, Zhao D, Guan H. et al. Molecular Coupling of Histone Crotonylation and Active Transcription by AF9 YEATS Domain. Mol Cell. 2016;62:181-93
35. Zhao D, Guan H, Zhao S, Mi W, Wen H, Li Y. et al. YEATS2 is a selective histone crotonylation reader. Cell Res. 2016;26:629-32
36. Xiong X, Panchenko T, Yang S, Zhao S, Yan P, Zhang W. et al. Selective recognition of histone crotonylation by double PHD fingers of MOZ and DPF2. Nat Chem Biol. 2016;12:1111-8
37. Wang Z, Zhao Y, Xu N, Zhang S, Wang S, Mao Y. et al. NEAT1 regulates neuroglial cell mediating Abeta clearance via the epigenetic regulation of endocytosis-related genes expression. Cell Mol Life Sci. 2019;76:3005-18
38. Wei W, Mao A, Tang B, Zeng Q, Gao S, Liu X. et al. Large-Scale Identification of Protein Crotonylation Reveals Its Role in Multiple Cellular Functions. J Proteome Res. 2017;16:1743-52
39. Fu H, Tian CL, Ye X, Sheng X, Wang H, Liu Y. et al. Dynamics of Telomere Rejuvenation during Chemical Induction to Pluripotent Stem Cells. Stem Cell Reports. 2018;11:70-87
40. Abu-Zhayia ER, Machour FE, Ayoub N. HDAC-dependent decrease in histone crotonylation during DNA damage. J Mol Cell Biol. 2019;11:804-6
41. de Klerk E, t Hoen PA. Alternative mRNA transcription, processing, and translation: insights from RNA sequencing. Trends Genet. 2015;31:128-39
42. Swerdloff RS, Bhasin S. Male Reproductive Physiology. In: Aiman J, editor. Infertility: Diagnosis and Management. New York, NY: Springer New York. 1984 p. 177-84
43. Neto FT, Bach PV, Najari BB, Li PS, Goldstein M. Spermatogenesis in humans and its affecting factors. Semin Cell Dev Biol. 2016;59:10-26
44. Goudarzi A, Shiota H, Rousseaux S, Khochbin S. Genome-scale acetylation-dependent histone eviction during spermatogenesis. J Mol Biol. 2014;426:3342-9
45. Leal-Esteban LC, Fajas L. Cell cycle regulators in cancer cell metabolism. Biochim Biophys Acta Mol Basis Dis. 2020;1866:165715
46. Machour FE, Ayoub N. Transcriptional Regulation at DSBs: Mechanisms and Consequences. Trends Genet. 2020;36:981-97
47. Morgan RG, Donato AJ, Walker AE. Telomere uncapping and vascular aging. Am J Physiol Heart Circ Physiol. 2018;315:H1-H5
48. Fasching CL. Telomere length measurement as a clinical biomarker of aging and disease. Crit Rev Clin Lab Sci. 2018;55:443-65
49. Fang Y, Xu X, Ding J, Yang L, Doan MT, Karmaus PWF. et al. Histone crotonylation promotes mesoendodermal commitment of human embryonic stem cells. Cell Stem Cell. 2021;28:748-63 e7
50. Lv Y, Bu C, Meng J, Ward C, Volpe G, Hu J. et al. Global Profiling of the Lysine Crotonylome in Different Pluripotent States. Genomics Proteomics Bioinformatics. 2021 S1672-0229(21)00073-5
51. You Z, Jiang WX, Qin LY, Gong Z, Wan W, Li J. et al. Requirement for p62 acetylation in the aggregation of ubiquitylated proteins under nutrient stress. Nat Commun. 2019;10:5792
52. Qian MX, Pang Y, Liu CH, Haratake K, Du BY, Ji DY. et al. Acetylation-mediated proteasomal degradation of core histones during DNA repair and spermatogenesis. Cell. 2013;153:1012-24
53. Li M, Luo J, Brooks CL, Gu W. Acetylation of p53 inhibits its ubiquitination by Mdm2. J Biol Chem. 2002;277:50607-11
54. Yu H, Bu C, Liu Y, Gong T, Liu X, Liu S. et al. Global crotonylome reveals CDYL-regulated RPA1 crotonylation in homologous recombination-mediated DNA repair. Science advances. 2020;6:eaay4697
55. Covington HE 3rd, Vialou VF, LaPlant Q, Ohnishi YN, Nestler EJ. Hippocampal-dependent antidepressant-like activity of histone deacetylase inhibition. Neurosci Lett. 2011;493:122-6
56. Hobara T, Uchida S, Otsuki K, Matsubara T, Funato H, Matsuo K. et al. Altered gene expression of histone deacetylases in mood disorder patients. J Psychiatr Res. 2010;44:263-70
57. Fontecha-Barriuso M, Martin-Sanchez D, Ruiz-Andres O, Poveda J, Sanchez-Nino MD, Valino-Rivas L. et al. Targeting epigenetic DNA and histone modifications to treat kidney disease. Nephrol Dial Transplant. 2018;33:1875-86
58. Chen W, Tang D, Xu Y, Zou Y, Sui W, Dai Y. et al. Comprehensive analysis of lysine crotonylation in proteome of maintenance hemodialysis patients. Medicine (Baltimore). 2018;97:e12035
59. Tang X, Chen XF, Sun X, Xu P, Zhao X, Tong Y. et al. Short-Chain Enoyl-CoA Hydratase Mediates Histone Crotonylation and Contributes to Cardiac Homeostasis. Circulation. 2021;143:1066-9
60. Siegel RL, Miller KD, Jemal A. Cancer statistics, 2019. CA: a cancer journal for clinicians. 2019;69:7-34
61. Huang H, Wang DL, Zhao Y. Quantitative Crotonylome Analysis Expands the Roles of p300 in the Regulation of Lysine Crotonylation Pathway. Proteomics. 2018;18:e1700230
62. Han X, Xiang X, Yang H, Zhang H, Liang S, Wei J. et al. p300-Catalyzed Lysine Crotonylation Promotes the Proliferation, Invasion, and Migration of HeLa Cells via Heterogeneous Nuclear Ribonucleoprotein A1. Anal Cell Pathol (Amst). 2020;2020:5632342
63. Ju Z, He JJ. Prediction of lysine crotonylation sites by incorporating the composition of k-spaced amino acid pairs into Chou's general PseAAC. J Mol Graph Model. 2017;77:200-4
64. Qiu WR, Sun BQ, Tang H, Huang J, Lin H. Identify and analysis crotonylation sites in histone by using support vector machines. Artif Intell Med. 2017;83:75-81
65. Bos J, Muir TW. A Chemical Probe for Protein Crotonylation. J Am Chem Soc. 2018;140:4757-60
66. Wan J, Liu H, Chu J, Zhang H. Functions and mechanisms of lysine crotonylation. J Cell Mol Med. 2019;23:7163-9
67. Crespo M, Damont A, Blanco M, Lastrucci E, Kennani SE, Ialy-Radio C. et al. Multi-omic analysis of gametogenesis reveals a novel signature at the promoters and distal enhancers of active genes. Nucleic Acids Res. 2020;48:4115-38
Author contact
Corresponding author: Yang Yang, Department of Biochemistry and Molecular Biology, School of Basic Medical Sciences, Peking University Health Science Center, #38 Xueyuan Road, Beijing 100191, China. Phone: 86-10-82801602; E-mail: yangshedu.cn.

 Global reach, higher impact
Global reach, higher impact