Impact Factor ISSN: 1449-2288
Int J Biol Sci 2021; 17(13):3646-3658. doi:10.7150/ijbs.51376 This issue Cite
Review
The Role of Respiratory Microbiota in Lung Cancer
1. Key Laboratory of Carcinogenesis and Cancer Invasion, Ministry of Education, Department of Pathology, Xiangya Hospital, Central South University, Hunan, 410078 China.
2. NHC Key Laboratory of Carcinogenesis (Central South University), Cancer Research Institute and School of Basic Medicine, Central South University, Changsha, Hunan, 410078 China.
3. Hunan Key Laboratory of Tumor Models and Individualized Medicine, Department of Thoracic Surgery, Second Xiangya Hospital, Central South University, Changsha, 410011 China.
*These authors contributed equally to this work.
Received 2020-8-1; Accepted 2021-2-9; Published 2021-8-25
Abstract
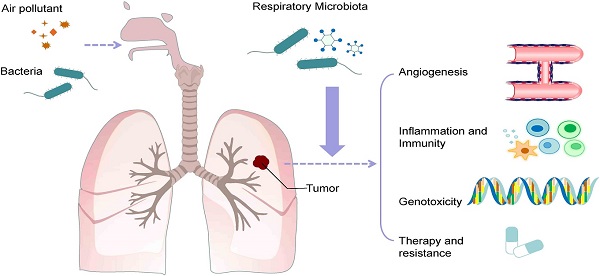
Recently, the impact of microorganisms on tumor growth and metastasis has attracted great attention. The pathogenesis and progression of lung cancer are related to an increase in respiratory bacterial load as well as changes in the bacterial community because the microbiota affects tumors in many ways, including canceration, metastasis, angiogenesis, and treatment. The microbiota may increase tumor susceptibility by altering metabolism and immune responses, promoting inflammation, and increasing toxic effects. The microbiota can regulate tumor metastasis by altering multiple cell signaling pathways and participate in tumor angiogenesis through vascular endothelial growth factors (VEGF), endothelial cells (ECs), inflammatory factors and inflammatory cells. Tumor angiogenesis not only maintains tumor growth at the primary site but also promotes tumor metastasis and invasion. Therefore, angiogenesis is an important mediator of the interaction between microorganisms and tumors. The microbiota also plays a part in antitumor therapy. Alteration of the microbiota caused by antibiotics can regulate tumor growth and metastasis. Moreover, the microbiota also influences the efficacy and toxicity of tumor immunotherapy and chemotherapy. Finally, the effects of air pollution, a risk factor for lung cancer, on microorganisms and the possible role of respiratory microorganisms in the effects of air pollution on lung cancer are discussed.
Keywords: Respiratory System, Microbiota, Angiogenesis, Tumor
Introduction
The human microbiome is a complex ecosystem primarily composed of a large number of various bacteria, viruses and fungi. The metagenome is even greater than the human genome by 150 times. These microorganisms mainly settle in human skin and mucous membranes, among other sites, and coevolved with humans [1].
In addition to inflammatory diseases, microorganisms have also been confirmed by many epidemiological studies to be related to many noninflammatory diseases, such as obesity, diabetes, and tumors. The types of microorganisms carried by different people are very diverse, and they are dynamically affected by external factors such as body part, diet, antibiotics, and lifestyle [2]. Meanwhile, the microbiome also affects the host's nutrition, immunity, metabolism and the occurrence or development of diseases including cancer.
At present, it is a good time and a key turning point for human microbiome research. The development of DNA sequencing technology, especially 16S rRNA gene sequencing and the related transcriptome, proteomics, metabolism, and immunological analysis technologies, helps us to determine the changes in the microbiome in health or various diseases more accurately. Based on many studies of different microbiomes in healthy populations or disease states, investigations have gradually transitioned to explore the mechanism.
The carcinogenesis of Helicobacter pylori (H. pylori), which promotes chronic inflammation or host DNA damage through molecules such as CageA, VacA and NapA2, is assured. However, interestingly, H. pylori infection lowers the risk of esophageal adenocarcinoma and is a strong protective factor [3]. However, in addition to the carcinogenesis of H. pylori and chronic inflammation, few studies have convincing evidence of other gastric microbiomes in the pathogenesis of cancer. For colorectal cancer, Fusobacterium nucleatum (F. nucleatum) increased the risk, and in subsequent animal and laboratory experiments, F. nucleatum was verified to increase cancer risk through attachment and invasion mechanisms and tumor immune escape [4, 5]. Tumors are not only related to microbial imbalances in the microenvironment next to the tumor, whose classic example is liver cancer induced by hepatitis B virus and cervical cancer induced by human papilloma virus (HPV), but the microbiota in different parts may also affect each other. The intestinal microbiota and metabolites have been confirmed to promote the development of tumors in regions that are far away from the intestine, such as the liver and prostate [6, 7].
In the lung, which was once considered sterile, the existence of microbiota different from the upper respiratory tract has also been demonstrated [8]. The lung microbiome may partially explain why the lung is an important organ for tumor formation and/or metastasis. The inflammatory and immune responses caused by the lung microbiota affect lung tumorigenesis. Lung bacteria stimulate tissue-resident γδ T cells, promoting the infiltration of neutrophils and the proliferation of tumor cells [9]. Different changes in the immune system caused by the microbiome may lead to different changes in tumor progression. Tumor-associated macrophages (TAMs) are alveolar macrophages (AMs) infiltrated into tumor tissue. The two phenotypes M1 and M2 of TAMs have different effects on lung cancer. The M1 phenotype plays a role in killing tumor cells, while M2 macrophages make the tumor microenvironment conducive to tumor progression [10]. The lung microbiome has become an important object for the study of tumorigenesis and development.
Similar to other solid tumors, the growth and metastasis of lung cancer requires oxygen, nutrient replenishment, and transportation of metabolic wastes, which require angiogenesis. An increase in microvessel density can be observed in early lung cancer, which may confirm this point. Studies in recent years have found that the lung is also a hematopoietic organ, and a large number of megakaryocytes capable of producing platelets can be found in the lungs of mice [11]. At the same time, the role of platelets in promoting angiogenesis has also been demonstrated [12].
The relation between respiratory microorganisms and lung cancer
The respiratory system is an important part of the human body connecting the outside world with the body. Complex and diverse microbial communities in the respiratory tract [13] play unique roles in biological processes. Respiratory microorganisms are of great importance in a variety of diseases. Studying the structure of human respiratory microorganisms and their correlation with disease progression is conducive to clarifying the mechanism of respiratory and other systemic diseases, which is of great importance in prevention and treatment. The pathogenesis of lung cancer is related to changes in bacterial burden and composition in the respiratory system [9]. At present, research on the correlation between respiratory microorganisms and lung cancer is still in its infancy, but it has already been able to explain some aspects of the disease.
Respiratory bacteria and lung cancer
The respiratory system consists of the respiratory tract and lung. There are fixed bacteria in the respiratory system of healthy people [14]. Respiratory microecology will change dramatically under the condition of disease, which is responsible for the occurrence and development of disease [15]. The pathogenesis of lung cancer is a process involving the interaction of many factors, including dynamic changes in respiratory microbial structure. Therefore, the improvement of bacterial structure in the respiratory tract has the potential to regulate the development of cancer, is important for research and thus is valuable to apply.
Bacteria in respiratory tract and lung cancer
In the healthy respiratory tract, Proteobacteria, Firmicutes, Fusobacteria and Bacteroidetes are the main phyla and classes, and Pseudomonas, Streptococcus, Prevotella, Veillonella, Haemophilus and Neisseria are the main genera of flora [14, 16].
There are structural changes in the respiratory tract flora in the case of lung cancer. The correlation between respiratory tract flora and the pathogenesis of lung cancer has previously been revealed. In patients with lung cancer, multiple anaerobic bacteria exist in the lower respiratory tract [17], represented by Actinomyces and Peptostreptococcus. There is also a large quantity of oral bacteria, such as Streptococcus and Wechsler, which are related to the upregulation of the ERK and PI3K signaling pathways [18]. Moreover, both Mycobacterium tuberculosis (M. tuberculosis) [19] and H. pylori [20] infection take part in promoting the development of lung cancer. Compared with adjacent normal tissues, the contents of Modestobacter in lung cancer tissues are higher, but Propionibacterium and Enterobacteriaceae are lower [21]. A lower airway microbiome analysis showed that Bradyrhizobium japonicum could only be detected in lung cancer patients, but the genus Acidovorax could be detected in not only patients with lung cancer but also patients with benign lung disease [22]. Moreover, the numbers of Eubacterium xylanophilum, Eubacterium eligens and Clostridium in the human respiratory tract tend to have a positive correlation with the occurrence of small cell lung cancer (SCLC), and the number of Prevotella and Pseudobutyrivibrio ruminis may have a negative correlation with the occurrence of SCLC [23].
Various microorganisms form a complex regulatory network linked with lung cancer. Different microorganisms in the same type of lung cancer or the same microorganisms in different types of lung cancer may play different roles. Additional elaboration of the detailed regulatory network of microorganisms in lung cancer is expected.
Pulmonary flora and lung cancer
The opinion of the aseptic lung has been repudiated. Many studies have found that the dominant populations in healthy lungs of adults are Firmicutes, Bacteroidetes, Proteobacteria, Actinobacteria, and Fusobacterium [24, 25]. Furthermore, there are also related findings that for healthy adults, the lung microbiome has no significant difference in spatial space [25]. In terms of diversity analysis, there was no significant difference in beta diversity between nonmalignant lung tissue and tumor lung tissues, but the alpha diversity of the microbial group was significantly reduced in tumor lung tissues [26].
Lung microorganisms are closely related to certain lung diseases and health because microbiome dysbiosis may increase tumor susceptibility by changing metabolism, altering the immune response, promoting inflammation, and increasing toxic effects [27]. Therefore, defining the composition and changes of the microbial group in lung cancer patients is conducive to deepening the understanding of the mechanism of lung cancer. At the same time, this knowledge may provide new ideas for preventing lung cancer or treatment intervention. For example, the epidemiological relationship between M. tuberculosis and lung cancer can be explained by the mechanism of chronic inflammation-associated carcinogenesis [28].
The results of an experimental analysis of contralateral bronchoalveolar lavage fluid (BALF) samples via 16S rRNA sequencing showed that the number of operational taxonomic units (OTUs) was significantly different between the lung cancer group and the benign mass-like lesion group, while β-diversity was still no different. The Chao 1 and Shannon indices in the lung cancer group were higher than those in the normal group. Among the five detected phyla and seven detected genera, the phyla with increased relative abundance were Firmicutes and TM7 in the lung cancer group while the most abundant genera were Eillonella, Megasphaera, Atopobium and Selenomonas. The abundance of Atopobium and Selenomonas decreased in both groups [29]. These changes may suggest a way to affect lung cancer by altering the pericancerous microenvironment.
In addition, the proportion of Firmicutes/Bacteroidetes in the lung cancer group with smoking was significantly higher than that in the lung cancer group with nonsmoking, and the relative abundance of some bacteria was different from that in the nonsmoker group. Meanwhile, increased TM7 in both chronic obstructive pulmonary disease (COPD) and lung cancer suggests that TM7 is likely to promote the progression of COPD to lung cancer. Finally, Megasphaera and Veillonella were identified as biomarkers for lung cancer, meaning they could be used to predict lung cancer or as microbial therapeutic targets [29].
Another study collecting saliva and BALF found that the microbial alpha diversity in lung cancer patients was low, and this difference from the control group was not related to the sampling site. However, certain bacterial groups associated with lung cancer are affected by the sampling site and pathological type, and the same is true for the metabolic differences of the microbial population. Via KEGG metabolic pathway enrichment analysis and functional analysis, activated and enriched behaviors usually include antibiotic resistance and sorting, folding and degradation as well as metabolism of amino acids and glycans. However, regardless of the sampling site, the microbiota composition differed between the two groups. Finally, the study also suggests that the good bacterial biomarkers in BALF are Treponema and Filifactor and that Filifactor has good resolution in distinguishing between healthy controls and lung cancer patients [30].
However, symbiotic microbial biomarkers of lung cancer or microbial therapies still require further large-scale studies.
Respiratory system fungi, viruses and lung cancer
A retrospective study suggested that pulmonary lymphoepithelioma-like carcinoma is closely related to Epstein-Barr virus (EBV) infection, and EBV latent infection is the main state of pulmonary lymphoepithelioma-like carcinoma [31]. Among EBV-encoded small nonpolyadenylated RNA (EBER)-positive pulmonary lymphoepithelioma-like carcinoma (LELC) cases, latent membrane protein 1(LMP1) and viral capsid antigen (VCA) are expressed in approximately half and a quarter of these cases, respectively. Many recent related studies confirm that lymphoid-specific helicase (LSH) can affect the occurrence and development of non-small cell lung cancer (NSCLC) in cancer metabolism and epigenetics through different mechanisms or as intermediate products. For example, Go, Ichi, Nii, and San complex subunit 4 (GINS4) is highly expressed in NSCLC and contributes to tumorigenesis. LSH increases GINS4 expression by binding to the 3'-untranslated region [32]. Moreover, LSH promotes lipid metabolism-related genes and becomes a vital inhibitor of ferroptosis in carcinogenesis in vitro or in vivo. The increased expression of LSH is regulated by EGLN1 and c-Myc [33]. The overexpression of LSH in nasopharyngeal carcinoma is linked to EBV infection, and LSH becomes a cancer driver by suppressing the activity of fumarate hydratase [34, 35].
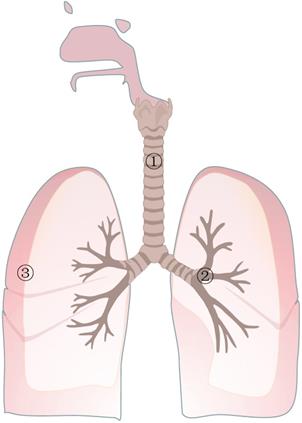
Representative bacteria in different circumstances
| Respiratory Tract ①,② | Lung③ | ||
|---|---|---|---|
| Normal | Phylum and class: Proteobacteria, Firmicutes, Fusobacteria, Bacteroidetes. Genus: Pseudomonas, Streptococcus, Prevotella, Veillonella, Haemophilus, Neisseria. | Firmicutes, Bacteroidetes, Proteobacteria, Actinobacteria, Fusobacterium. | |
| Cancer | Less | Propionibacterium, Enterobacteriaceae, Prevotella, Pseudobutyrivibrio, ruminis. | Atopobium |
| Selenomonas | |||
| More | M.tuberculosis, Helicobacter, Pylori, Actinomyces, Peptostreptococcus, Streptococcus, Wechsler, Modestobacter, Eubacterium xylanophilum, Eubacterium Eligens, Clostridium. | Phylum: | |
| Firmicutes | |||
| TM7 | |||
| Genus: | |||
| Veillonella, Megasphaera Filifactor Selenomonas | |||
A clinical study across Taiwan showed that a high risk of lung cancer involved being exposed to influenza and that there was more risk after cumulative exposure to flu. However, the mechanism is currently unknown [36]. In samples from patients with lung adenocarcinoma, the prevalence of John Cunningham virus (JC virus) and whether virus was present in metastatic lymph nodes were evaluated, suggesting that JC virus may be involved in lung cancer [37].
The data show that the incidence of lung cancer in human immunodeficiency virus (HIV)-infected patients is higher than that in uninfected patients. The cumulative incidence of lung cancer among HIV-infected groups continues to rise. In addition to a reduction in all-cause mortality, immunosuppression, pulmonary inflammation, and systemic inflammation are all specific risk factors for increased risk [38]. There are studies that suggest that direct viral oncogenesis is a mechanism of lung cancer in HIV infection, but there is no exact support for direct HIV oncogenesis in the lungs. Only some early studies have shown some direct oncogenic activity among HIV virus proteins. Studies evaluating antiretroviral treatment did not lead to a higher risk of cancer. In addition, the hypothesis that P450 cytochrome inhibitors may affect the metabolism of lung cancer carcinogens has not been found in The French Hospital Database [39].
In short, the types and quantities of human microorganisms are very large. Although the Human Microbiome Project (HMP) does not use the lung as an organ of research, the traditional view that the lung is sterile has been overthrown, and the study of the microbiome in the lung has developed rapidly as a new field. There are still many areas worth exploring.
Representative bacteria in different circumstances. In the normal respiratory tract, the representative phyla and classes are Proteobacteria and Firmicutes, and the representative genera are Pseudomonas and Streptoccus, respectively. In the respiratory tract with lung cancer, some bacteria tend to be less abundant, such as Propionibacterium and Enterobacteriaceae, and some tend to be more abundant, such as M.tuberculosis and Helicobacter. In the lungs of healthy people, the most common bacterial microorganisms are Firmicutes, Bacteroidetes, Proteobacteria, Actinobacteria, and Fusobacterium. In the lungs of patients with lung cancer, Atopobium and Selenomonas were obviously decreased. In contrast, additional microorganisms were Firmicutes, TM7, Veillonella, Megasphaera, Filifactor and Selenomonas.
Lung cancer-related microorganisms in other tumors
The carcinogenesis of H. pylori, through Helicobacter pylori toxin such as CageA, may also be a way for H. pylori to induce lung cancer [40, 41]. HPV is not only related to cervical cancer and breast cancer but is also a risk factor for lung cancer [41, 42].
Clearly, a healthy commensal microbiome is tightly related to immune cell functioning. The study of Rea Bingula et al. on the gut-lung axis mainly used the gut as an example and tracked the effects of intestinal immune stimulation on the lung to speculate that the situation originating from the lung mucosa and lymph nodes is also similar [43]. The two “cooperate” and influence each other.
The lung cancer-related microorganisms summarized above have also been found in other tumors.
Alterations in these changing microorganisms in the microenvironment of other tumors have also been reported by other papers. For example, Atopobium may not only lead to a high risk of esophageal squamous cell carcinoma [44] but also increase the risk of endometrial cancer in the uterus [45]. Firmicutes was found to be more abundant in the urine of bladder cancer patients, and Veillonella was more abundant in the healthy control group than in the bladder cancer group [48]. Filifactor in the oral cavity is positively related to oral squamous cell carcinoma [126, 127] and is overrepresented in the tongue coating of pancreatic head carcinoma patients [128]. Selenomonas is overrepresented in the stool of colorectal cancer patients [129] and is more abundant in left colorectal cancer patients compared to right colorectal cancer patients [130].
The high risk of esophageal squamous cell carcinoma may be related to Atopobium, a kind of microorganism that is reduced in lung cancer [44]. An in vitro experiment found that Atopobium vaginae (which becomes less abundant in lung cancer) induces proinflammatory cytokine expression in endometrial cells and may lead to significant endometrial cancer if Porphyromonas somerae also exists [45].
In addition, Propionibacterium in the lung is negatively related to lung cancer, while Propionibacterium in the prostate is positively correlated with prostate cancer [46, 47]. For bladder cancer, the most abundant phylum in both groups was Firmicutes, which became more abundant in lungs with lung cancer than in healthy lungs. However, Veillonella, suggested as a lung cancer marker, is more common in healthy urine [48].
The progress and relationship of lung cancer-related microorganisms and other tumors needs further research and discovery.
The influence of pulmonary microorganism on lung cancer
At present, the understanding of the influence of the microbiota on tumors is still insufficient. The influence of microorganisms on tumors involves many aspects, such as canceration, angiogenesis, and therapy.
The possible mechanism of the influence of pulmonary microorganisms on lung cancer canceration
In the process of cancer occurrence, microecology deviates from the normal steady-state structure and gradually forms the microenvironment of promoting cancer to induce a series of chain reactions, finally causing canceration. An imbalance in the lung microecology may affect the canceration of lung cancer through inflammation, immunity, metabolism and genotoxicity.
Inflammation
Chronic lung inflammation is a critical risk factor for lung cancer [49], and proinflammatory cytokines, adhesion molecules, and growth factors can provide a favorable microenvironment to promote the survival and proliferation of tumor cells [50]. Microbial organisms are confirmed to be important causative agents of cancer-inducing inflammation [51].
Membrane receptors play a vital role in cancer caused by microorganisms through the process of inflammation. Membrane receptors such as pattern-recognition receptor (PRR), cluster of designation (CD), and TLR proteins can recognize microorganisms and their production, proinflammatory cytokines and some other signaling molecules, leading to effects on apoptosis and cellular proliferation [52]. In COPD patients, infection with nontypeable Haemophilus influenzae (NTHi) can induce epithelial cells to express IL-17C and neutrophils to promote tumors [53]. Interleukin-6 (IL-6) also exerts a significant function in lung cancer by inducing a COPD-like inflammatory response [54].
Immunity
A small number of cancer cells in the body can be eliminated by the antitumor immune mechanism, which may malfunction with the growth of cancer cells [55]. The regulation of the cancer immune response by microbiota has also received more attention.
Commensal bacteria are involved in maintaining immune homeostasis against cancer through γδT17 responses [56]. Lung immunity after antibiotic treatment tends to be defective, thus creating a more favorable environment for the occurrence of cancer. However, another study suggested that the local microbiota could activate lung-resident γδ T cells to promote inflammation associated with lung adenocarcinoma and tumor cell proliferation [9]. Commensal bacteria also regulate the antitumor response in the lung through alveolar macrophages, maintaining their low level of CCL24 secretion [57]. Although it is unknown and difficult to define whether the immune environment is regulated by specific bacteria with dedicated functions in the lung, the role of some bacteria has been proposed. Morganella morganii and Escherichia fergusonii, two ubiquitous opportunistic pathogens from the Proteobacteria phylum in the lung, can produce different virulence factors to exert an immunostimulatory effect [58]. NSCLC patients react to Streptococcus salivarius and Streptococcus agalactiae with significantly higher frequencies of T helper type 17 (Th17) and Th1 cells than healthy controls [59]. Recently, a study revealed that Pasteurella has a positive correlation with cytotoxic CD8+ TILs and a negative correlation with M2 macrophages. There is a significant positive correlation between Coriobacteriaceae and M2 macrophages and a negative correlation with CD8+ cells [60]. Together, these immune reactions affect the occurrence and development of lung tumors.
These results suggest that the microbiota in the respiratory system has an important influence on the occurrence of lung cancer, which may establish a permissive environment for lung cancer.
Metabolism
Microorganisms have been proven to participate in regulating host metabolism, detoxification, vitamin levels and nutritional status [61].
Changes in microorganisms are related to the production of carcinogens such as acetaldehyde [62] and deoxycholic acid [63]. This metabolic imbalance may result in the formation of carcinogens in the lung or lead to the secondary processing of foreign carcinogens. The lung microbiota is considered to be connected with specific insults related to lung cancer via CD36, which regulates the treatment of cyanobacteria-derived microcystin residues in the lung alveoli, upregulating the expression of poly (ADP-ribose) polymerase 1 (PARP1) [64].
Microorganisms may also play an anticancer role by influencing metabolism. Short-chain fatty acids, as products of intestinal bacteria, exert anti-inflammatory and antiproliferation functions [65]. Moreover, butyrate treatment could lead to an upregulation of miR-3935 and an inhibition of the proliferation and migration of A549 cells [66].
Genotoxicity
Host DNA damage can cause cell death and affect the expression of tumor suppressor genes or oncogenes. Some bacterial molecules have been revealed to be related to genotoxicity. For instance, Bacteroides fragilis toxin and Escherichia coli toxin have been found to be related to inducing double-stranded DNA damage [67, 68]. Chemicals produced by bacteria such as reactive oxygen species (ROS) produced by Porphyromonas, hydrogen sulfide produced by Clostridium cholephilum and superoxide dismutase produced by some bacteria are also responsible for genomic instability [69-71], increasing the probability of cancer. In addition, microorganisms can also play a carcinogenic role through the integration of their DNA and host genome.
The possible mechanism of the influence of pulmonary microorganisms on the metastasis of lung cancer
Microorganisms also play an irreplaceable role in the metastasis of lung cancer. Oral taxa that are rich in the lower airway of lung cancer patients, such as Streptococcus and Veillonella, promote the proliferation, invasion and metastasis of lung cancer cells by activating the ERK and PI3K pathways in airway epithelial cells [18]. Ubiquitin molecules in host cells can activate protein tyrosine phosphatase A (PTPA) secreted by M. tuberculosis, which promotes the proliferation and migration of the human lung adenocarcinoma cell line A549 [72]. Additionally, the metastasis of lung cancer can be affected by microorganisms through inflammatory mechanisms. The stimulation of gram-negative bacteria with TLR4, a receptor that mediates inflammation, increases adhesion, migration, and metastatic spreading of NSCLC cells [73]. Similarly, gram-positive pneumonia augments NSCLC metastasis via TLR2 activation [74]. Inflammation also causes crosstalk between bacteria and tumor cells through quorum sensing peptides to promote tumor metastasis [75]. In terms of immunity, a decrease in bacterial load is linked with reduced regulatory T cells and enhanced T cell and natural killer (NK) cell activation, which contributes to a resultful reduction in lung metastases [58].
Angiogenesis is an intermediary between microorganisms and tumors
Human microbiota can affect angiogenesis [75, 76]. An important possible reason is that inflammatory factors related to microbiota-induced inflammation affect angiogenesis. Angiogenesis has an effect on infection, tumors and other pathological processes. In recent decades, many scholars have studied how angiogenesis affects tumor growth and metastasis. There are many clinical data to prove that antiangiogenic drugs are effective in cancer patients. Therefore, the effect of the microbiome on angiogenesis makes it possible to manipulate the human microbiome to change angiogenesis in the disease process.
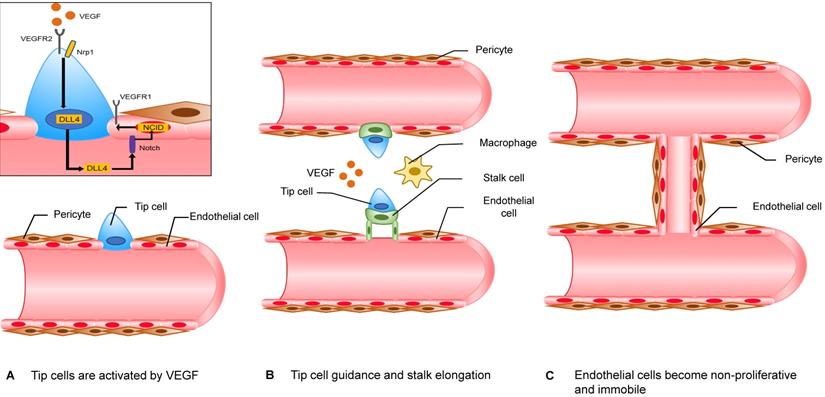
Accepted mechanisms of neovascularization include sprouting angiogenesis, vasculogenesis, vasculogenic mimicry and vascular intussusception, and the above four mechanisms are thought to act on pathological neovascularization at the same time. Among them, sprouting angiogenesis models have been extensively studied (Figure 2). To avoid necrosis that is due to hypoxia, cells in hypoxic regions secrete vascular endothelial growth factors (VEGF). VEGF promotes endothelial cells (ECs) to secrete matrix metalloproteases (MMPs), which can degrade their basement membrane (BM) and extracellular matrix (ECM) [77]. BM is located on the basal surface of ECs and has a supporting, connecting and fixing effect on endothelial cells. When the stability of BM is disrupted, endothelial cells gain the ability to proliferate and to migrate. In this model, there are two important types of cells, namely, “tip cells” and “stalk cells.” “Tip cells” migrate under the stimulation of angiogenic signals. They are the foremost cells. The cells after “tip cells” are called “stalk cells” and have a strong ability to proliferate. Similar to their name, these cells are comparable to the stalk of an angiogenic sprout. When adjacent “tip cells” meet, two angiogenic sprouts fuse into a single branch. Once the branch is perfused, ECs lose their ability to proliferate and to migrate. After recruiting enough pericytes, a mature blood vessel forms [78].
The mechanism of sprouting angiogenesis. The “tip cells” at the forefront of the angiogenic sprout migrate along the angiogenic signal. “Stalk cells” continue to proliferate behind “tip cells”. However, the classification of “tip cells” and “stalk cells” is not fixed and depends on the ratio of VEGFR1/VEGFR2 on the cell membrane. ECs that express more VEGFR1 have lower reactivity to VEGF, while ECs that express more VEGFR2 have higher reactivity to VEGF. The specific mechanism is as follows. EC with a higher VEGFR2 ratio binds to the receptor Notch of the adjacent EC through its expressed DLL4, which can promote the expression of NICD by the adjacent EC. NCID can increase the expression of VEGFR1 and decrease the expression of VEGFR2. When adjacent "tip cells" meet, the two angiogenic sprouts fuse into a branch with the help of macrophages. Once the branch is perfused, ECs lose their ability to proliferate and to migrate. After recruiting enough pericytes, a mature blood vessel is formed.
Microorganisms participate in the abovementioned angiogenesis mechanism through VEGF, ECs, inflammatory factors and inflammatory cells. Irradiated mice treated by fecal microbiota transplantation show increased expression of VEGF, promoting angiogenesis [79]. The immune response to bacteria in vitro and in a sepsis mouse model alters angiogenesis [80]. Alteration of intestinal microbiota was observed in mice fed stool from colorectal cancer patients, with enhanced expression of genes associated with angiogenesis and accelerated tumorigenesis [81]. Positive effects of specific microorganisms on tumor angiogenesis have been previously identified. In the CCSPcre/K-rasG12D mouse lung cancer model exposed to NTHi lysate weekly, the infiltration of Th17 cells in the lung tissue increased, with tumor growth accelerating [82]. The Th17 cell population is capable of producing IL-17. IL-17 can increase angiogenesis by inducing the generation of multiple angiogenic mediators, angiogenic chemokines, and cytokines that promote angiogenesis [83-85], which suggests that NTHi-induced Th17 cells may promote lung cancer development by promoting angiogenesis. Another link to tumor vascularization of carcinoma of the lungs is H. pylori through the assumption of direct damage and chronic inflammation, and H. pylori multiplication induces a systemic immune response coupled with the host's susceptibility and may induce lung cancer [86]. Helicobacter pylori VacA (exotoxin) exists in the lung and can induce lung cells to generate IL-8 and IL-6. Vac has cytotoxic effects and can specifically act on airway epithelial cells to induce interleukins [87]. IL-8 is a promoting factor for tumor angiogenesis; therefore, H. pylori in the lung is closely connected with tumor vascularization of lung carcinoma. HPV can also significantly enhance the angiogenesis of lung cancer cells in vitro [88], which may correlate with the PI3K/Akt signaling pathway and c-Jun [89].
Microorganisms that inhibit tumor vascularization have also been confirmed. The fungi zj-14, zj-17 and zj-36 can significantly inhibit HPV-16E7-stimulated microtubule formation [90]. Therefore, they may ultimately suppress the angiogenesis of lung carcinoma by suppressing the angiogenic effect of HPV on lung cancer. Moreover, A. muciniphila and E. hirae induce dendritic cells to secrete interleukin-12 (IL-12) [91], which can inhibit tumor angiogenesis [92]. This action may be one of the mechanisms by which the microbiota exerts tumor suppressive effects by affecting angiogenesis. Therefore, microorganisms are indeed closely related to tumor angiogenesis, which will also be an important direction for future research. It is worth noting that different types of tumors in various sites have different microenvironments. The mechanisms by which different microorganisms influence angiogenesis in cancer are also worth comparing.
The effect of vascularization on tumors
Tumor angiogenesis plays a significant role in tumor initiation, dormancy, patterned advancement and diversion.
In addition to maintaining tumor growth in the primary site, tumor angiogenesis can also promote tumor metastasis and invasion, thereby affecting the formation of secondary tumor foci. VEGF has a vital impact on neoplasm endotheliocytes including upregulating the production of proteases, promoting degradation of the basilar membrane, and prompting the expression of molecules that connect neoplasms with endotheliocytes [93].
Matrical cells, except endotheliocytes, are involved in angiogenesis-driven diversion processes, such as abnormal pericytes in structure or number. This process causes the cell gap to become larger, the tumor cells passively to leak out, and then to metastasize [94].
Therefore, an increased level of VEGF, a vital angiogenic factor produced in angiogenesis, is related to an unfavorable outcome in NSCLC patients [95, 96].
The uniqueness of blood supply to lung cancer
There are four known models of blood supply to lung cancer: basal, nipple, diffuse, and alveolar blood vessel dependent. The fourth blood supply mode of lung cancer is exactly the difference between lung cancer and other tumors. There is no substantial damage in this model. The lack of tumor-related stroma and new blood vessels indicates that tumors do not undergo angiogenesis. We found that the tumor was full of alveoli, so it can be seen that lung cancer depends on the original alveolar blood vessels for blood supply [97].
The role of the microbiota in tumor therapy
The microbiota has also played a part in antitumor therapy. Aerosolized antibiotics can reduce the lung microbiota, which is associated with a decrease in regulatory T cells and a reduction in the growth of melanoma B16 lung metastases [58]. Possible therapy targeting the respiratory microbiota has also been found in the therapy of primary lung cancer. In a genetically engineered lung adenocarcinoma mouse model, mice treated with antibiotics showed decreased cancer progression. Reduced lung bacterial load is associated with diminished tumor burden [9]. In NSCLC patients, clarithromycin treatment is related to prolonged survival and increased benefit, which is associated with decreased IL-6 levels. Its antitumor effect may be related to regulating the microbiome and reducing the release of cytokines that promote inflammation [98].
In recent years, immune checkpoint blockers (ICBs) have shown great development potential in tumor therapy. Two targets, cytotoxic lymphocyte antigen-4 (CTLA-4) and programmed cell death protein 1 ligand 1 (PD-L1), have drawn great attention. As a critical regulator of the immune system, the microbiota influences the effect of immunotherapy. Bacteroidales plays a role in the antitumor effects of CTLA-4 blockade [99]. The relationship between Bifidobacterium and the efficacy of PD-L1 antibody therapy has also been reported in which enhanced function of dendritic cells is involved [100]. However, an abnormal or unfavorable microbiome composition can lead to primary resistance and unsatisfying clinical outcomes [101, 102]. Moreover, carcinoma-related microbiota has inspired specific targets of immunotherapy. The continual presence of EBV in nasopharyngeal carcinoma promotes vaccine trials of specific epitopes (such as LMP1 and LMP2) and the transfer of certain cytotoxic T lymphocytes [103].
The microbiota has also become nonnegligible to maximize the efficacy and to minimize the toxicity of tumor chemotherapy. The microbiota can cause an increase in the level of ROS produced by myeloid cells, which can promote cancer cell death induced by oxaliplatin [104]. The anticancer efficacy of cyclophosphamide is also closely related to the alteration of the microbiota and the resultant immune response [105]. Despite the capacity, the microbiota also affects drug resistance and chemotherapy toxicity. Gammaproteobacteria can mediate tumor tolerance to gemcitabine through the long isoform of cytidine deaminase [106], while the modulated autophagy pathway is related to chemoresistance mediated by Fusobacterium (F.) nucleatum [107]. Diarrhea, a common side effect caused by irinotecan, is correlated with bacterial β-glucuronidase, which can regulate the bioactive form of irinotecan [108].
However, current studies mainly focus on the intestinal microbiota, but the intestinal and respiratory microbiota differs in many ways, such as composition and function. The difference between the role of the respiratory microbiota and intestinal microbiota in tumor therapy deserves more research, especially on the influence of local microenvironment changes in lung cancer during therapy.
The microbiota in the effect of air pollution on lung cancer
The relationship between the microbiota and lung cancer may explain some lung cancer risk factors that affect the respiratory microbiota, such as air pollution.
Air pollution, the microbiota and lung cancer
Air pollution induces alterations in the respiratory microbiota. Exposure to biomass smoke led to elevated bacterial abundance as well as diversity in rat lungs [109], but bacterial diversity was reduced after exposure to particulate matter 2.5 (PM2.5) in the mouse respiratory tract [110], which suggests that the specific components of the pollutants and the response of different species to pollutants need to be considered to draw a conclusion. There were differences in the oropharyngeal microbiota at different levels of air pollution among healthy young subjects from northeastern China [111]. The relative abundance of Actinobacteria and Firmicutes Proteonacteria was higher in samples from polluted areas, with Bacteroidetes and Fusobacteria being significantly lower. Domestic biomass fuel combustion can cause household air pollution (HAP). The abundance of Streptococcus and Neisseria was higher in the high HAP-related particulate exposure group than in the low-exposure group [112]. Regarding microbial activity, benzo[a]pyrene (B[a]P) altered the transcriptome of the human gut microbiome but had no significant impact on the microbiome composition [113].
Epidemiological studies clearly demonstrate that air pollution is related to the morbidity and mortality of lung cancer [114-117]. Exposure to PM2.5 induced an increase in tumor nodules in tumor-bearing mice, with elevated levels of 12 angiogenesis factors and a perturbed microecosystem of the upper respiratory tract [118]. However, there are difficulties in confirming the causal relationship between changes in the microbiome and changes in tumor development after air pollution exposure.
The microbiota and the effects of air pollution on lung cancer
Air pollution has an impact on the occurrence of lung cancer from multiple aspects related to various biochemical pathways. Genotoxicity plays a vital role in the impacts of air pollution on lung cancer. DNA oxidative damage is related to the initial stage of carcinogenesis. The premutagenic lesion 8-hydroxy-2'-deoxyguanosine (8-OHdG) is a biomarker of DNA oxidative damage. PM can promote DNA damage in vitro via 8-OHdG formation and strand breaks [119, 120]. Interestingly, genotoxicity is also an important mechanism by which the lung microbiome affects lung cancer (as mentioned before), which may be the key to linking the three. Otherwise, DNA oxidative damage can also be caused by ROS and reactive nitrogen species (RNS) that are excessively formed by inflammatory cells during the particle-elicited inflammation process [121]. Microorganisms are associated with cancer-inducing inflammation [51]. However, the exact inflammatory cytokines and cells need to be identified to confirm this possible relationship.
Abnormal promoter methylation of tumor suppressor genes is another possible link between air pollution-related microbiome changes and lung cancer. Continuous PM2.5 exposure induced methylation of the tumor suppressor gene p53 promoter by promoting methylation of DNA (cytosine-5-)-methyltransferase 3β, thereby leading to inactivation of p53 in human alveolar epithelial cells [122]. Particulate carcinogens cause lung cancer in rats, half of which show p16 methylation [123]. Microorganisms are also related to aberrant DNA methylation [124], but the role of the microbiome in DNA methylation changes associated with air pollution needs to be verified by further research.
The specific status of respiratory microorganisms in the effect of air pollution on lung cancer needs to be additionally explored. Increasing research in this field has found novel directions for tumor prevention and treatment. For example, vitamins B6 and C can partially attenuate the gene-specific hypermethylation and hypomethylation caused by B[a]P [125], which may have preventive effects on air pollution-related lung cancer.
Prospective
It is expected that related research will further clarify the relationship between the respiratory microbiome and respiratory tumors, provide clues for exploring new tumor therapies and developing antitumor drugs targeting microbes, and promote the rational use of angiogenesis inhibitors in antitumor therapy from the perspective of the human microbiota. At the same time, the abnormal structure of the tumor blood vessels mentioned above will reduce the various treatment methods to reach the tumor tissue, thereby reducing the treatment efficiency of the tumor. Therefore, perhaps we can propose a different idea and perspective for tumor treatment: improving abnormal angiogenesis of the tumor will limit the hypoxia of the tumor and prevent the selective effect of hypoxia on the highly invasive tumor.
Abbreviations
IL-17: interleukin-17; H. Pylori: Helicobacter pylori; F. nucleatum: Fusobacterium nucleatum; HPV: human papilloma virus; AMs: alveolar macrophages; TAMs: tumor-associated macrophages; M. tuberculosis: Mycobacterium tuberculosis; SCLC: small cell lung cancer; BALF: bronchoalveolar lavage fluid; OTUs: operational taxonomic units; EBV: Epstein-Barr virus; EBER: EBV-encoded small nonpolyadenylated RNA; LELC: lymphoepithelioma-like carcinoma; LMP1: latent membrane protein 1; VCA: viral capsid antigen; LSH: lymphoid-specific helicase; NSCLC: non-small cell lung cancer; GINS4: Go, Ichi, Nii, and San complex subunit 4; HIV: human immunodeficiency virus; JC virus: John Cunningham virus; HMP: the Human Microbiome Project; PRR: pattern-recognition receptor; CD: cluster of designation; NTHi: nontypeable Haemophilus influenza; IL-6: interleukin-6; NK cell: natural killer cell; PARP1: poly (ADP-ribose) polymerase 1; ROS: reactive oxygen species; PTPA: protein tyrosine phosphatase A; VEGF: vascular endothelial growth factor; Th17: T helper type 17; ECs: endothelial cells; TLR: toll-like receptor; DLL4: delta-like 4; IL-8: interleukin-8; GPCRs: G protein-coupled receptors; MetAP2: methionine aminopeptidase 2; BM: basement membrane; ECM: extracellular matrix; MMP: matrix metalloprotease; EPC: endothelial progenitor cell; PDGF: platelet derived growth factor; HGF: hepatocyte growth factor; FGF: fibroblast growth factor; VEGFR: vascular endothelial growth factor receptor; CAF: cancer-associated fibroblasts; NO: nitric oxide; TGF-β: transforming growth factor-β; COPD: chronic obstructive pulmonary disease; HIF-1α: hypoxia-inducible factor-1α; ICBs: immune checkpoint blockers; CTLA-4: cytotoxic lymphocyte antigen-4; PD-L1: programmed cell death protein 1 ligand 1; IL-12: interleukin-12; PM: particulate matter; HAP: household air pollution; B[a]P: benzo[a]pyrene; 8-OHdG: 8-hydroxy-2'-deoxyguanosine; RNS: reactive nitrogen species.
Acknowledgements
We would like to thank all laboratory members for their critical discussion of this manuscript, and apologize to those publications not mentioned due to space limitations.
Financial support
This work was supported by the National Natural Science Foundation of China [82072594(Y.Tao), 81872285(Y.Shi), 81874139(S.Liu)], and the Overseas Expertise Introduction Project for Discipline Innovation (111 Project, No. 111-2-12).
Ethics approval and consent to participate
The ethics committee of the Cancer Research Institute of Central South University has approved this study.
Author contributions
Dan Wang, Jingyi Cheng, Jia Zhang, Fangyu Zhou and Xiao He made equal contributions to this review. They collected the related papers, designed and drafted the manuscript. Yongguang Tao and Ying Shi critically revised and finalized the manuscript. All authors read and approved the final manuscript.
Competing Interests
The authors have declared that no competing interest exists.
References
1. Kovacs T, Miko E, Ujlaki G, Sari Z, Bai P. The Microbiome as a Component of the Tumor Microenvironment. Adv Exp Med Biol. 2020;1225:137-53
2. Gilbert JA, Blaser MJ, Caporaso JG, Jansson JK, Lynch SV, Knight R. Current understanding of the human microbiome. Nat Med. 2018;24:392-400
3. Xie FJ, Zhang YP, Zheng QQ, Jin HC, Wang FL, Chen M. et al. Helicobacter pylori infection and esophageal cancer risk: an updated meta-analysis. World J Gastroenterol. 2013;19:6098-107
4. Abed J, Emgard JE, Zamir G, Faroja M, Almogy G, Grenov A. et al. Fap2 Mediates Fusobacterium nucleatum Colorectal Adenocarcinoma Enrichment by Binding to Tumor-Expressed Gal-GalNAc. Cell Host Microbe. 2016;20:215-25
5. Gur C, Ibrahim Y, Isaacson B, Yamin R, Abed J, Gamliel M. et al. Binding of the Fap2 protein of Fusobacterium nucleatum to human inhibitory receptor TIGIT protects tumors from immune cell attack. Immunity. 2015;42:344-55
6. Sha S, Ni L, Stefil M, Dixon M, Mouraviev V. The human gastrointestinal microbiota and prostate cancer development and treatment. Investig Clin Urol. 2020;61:S43-s50
7. Rajagopala SV, Vashee S, Oldfield LM, Suzuki Y, Venter JC, Telenti A. et al. The Human Microbiome and Cancer. Cancer Prev Res (Phila). 2017;10:226-34
8. Morris A, Beck JM, Schloss PD, Campbell TB, Crothers K, Curtis JL. et al. Comparison of the respiratory microbiome in healthy nonsmokers and smokers. Am J Respir Crit Care Med. 2013;187:1067-75
9. Jin C, Lagoudas GK, Zhao C, Bullman S, Bhutkar A, Hu B. et al. Commensal Microbiota Promote Lung Cancer Development via gammadelta T Cells. Cell. 2019;176:998-1013.e16
10. Aras S, Zaidi MR. TAMeless traitors: macrophages in cancer progression and metastasis. Br J Cancer. 2017;117:1583-91
11. Lefrancais E, Ortiz-Munoz G, Caudrillier A, Mallavia B, Liu F, Sayah DM. et al. The lung is a site of platelet biogenesis and a reservoir for haematopoietic progenitors. Nature. 2017;544:105-9
12. Wojtukiewicz MZ, Sierko E, Hempel D, Tucker SC, Honn KV. Platelets and cancer angiogenesis nexus. Cancer Metastasis Rev. 2017;36:249-62
13. Dickson RP, Erb-Downward JR, Martinez FJ, Huffnagle GB. The Microbiome and the Respiratory Tract. Annu Rev Physiol. 2016;78:481-504
14. Filkins LM, Hampton TH, Gifford AH, Gross MJ, Hogan DA, Sogin ML. et al. Prevalence of streptococci and increased polymicrobial diversity associated with cystic fibrosis patient stability. J Bacteriol. 2012;194:4709-17
15. D'Journo XB, Bittar F, Trousse D, Gaillat F, Doddoli C, Dutau H. et al. Molecular detection of microorganisms in distal airways of patients undergoing lung cancer surgery. Ann Thorac Surg. 2012;93:413-22
16. Dickson RP, Martinez FJ, Huffnagle GB. The role of the microbiome in exacerbations of chronic lung diseases. Lancet. 2014;384:691-702
17. Rybojad P, Los R, Sawicki M, Tabarkiewicz J, Malm A. Anaerobic bacteria colonizing the lower airways in lung cancer patients. Folia Histochem Cytobiol. 2011;49:263-6
18. Tsay JJ, Wu BG, Badri MH, Clemente JC, Shen N, Meyn P. et al. Airway Microbiota Is Associated with Upregulation of the PI3K Pathway in Lung Cancer. Am J Respir Crit Care Med. 2018;198:1188-98
19. Park SK, Cho LY, Yang JJ, Park B, Chang SH, Lee KS. et al. Lung cancer risk and cigarette smoking, lung tuberculosis according to histologic type and gender in a population based case-control study. Lung Cancer. 2010;68:20-6
20. Deng B, Li Y, Zhang Y, Bai L, Yang P. Helicobacter pylori infection and lung cancer: a review of an emerging hypothesis. Carcinogenesis. 2013;34:1189-95
21. Mao Q, Hu J. P3.03-03 Differential Microbiota Features in Lung Tumor and Adjacent Normal Tissues in Lung Cancer Patients. Journal of Thoracic Oncology. 2018; 13
22. Jin J, Gan Y, Liu H, Wang Z, Yuan J, Deng T. et al. Diminishing microbiome richness and distinction in the lower respiratory tract of lung cancer patients: A multiple comparative study design with independent validation. Lung Cancer. 2019;136:129-35
23. Zhang L, Chen J. 90P Study of intestinal flora differences between small cell lung cancer (SCLC) patients and healthy people. Journal of Thoracic Oncology. 2018 13
24. Blainey PC, Milla CE, Cornfield DN, Quake SR. Quantitative analysis of the human airway microbial ecology reveals a pervasive signature for cystic fibrosis. Sci Transl Med. 2012;4:153ra30
25. Dickson RP, Erb-Downward JR, Freeman CM, McCloskey L, Beck JM, Huffnagle GB. et al. Spatial Variation in the Healthy Human Lung Microbiome and the Adapted Island Model of Lung Biogeography. Ann Am Thorac Soc. 2015;12:821-30
26. Morgan XC, Huttenhower C. Chapter 12: Human microbiome analysis. PLoS Comput Biol. 2012;8:e1002808
27. Mao Q, Jiang F, Yin R, Wang J, Xia W, Dong G. et al. Interplay between the lung microbiome and lung cancer. Cancer Lett. 2018;415:40-8
28. Pilaniya V, Gera K, Kunal S, Shah A. Pulmonary tuberculosis masquerading as metastatic lung disease. Eur Respir Rev. 2016;25:97-8
29. Lee SH, Sung JY, Yong D, Chun J, Kim SY, Song JH. et al. Characterization of microbiome in bronchoalveolar lavage fluid of patients with lung cancer comparing with benign mass like lesions. Lung Cancer. 2016;102:89-95
30. Wang K, Huang Y, Zhang Z, Liao J, Ding Y, Fang X. et al. A Preliminary Study of Microbiota Diversity in Saliva and Bronchoalveolar Lavage Fluid from Patients with Primary Bronchogenic Carcinoma. Med Sci Monit. 2019;25:2819-34
31. Han AJ, Xiong M, Zong YS. Association of Epstein-Barr virus with lymphoepithelioma-like carcinoma of the lung in southern China. Am J Clin Pathol. 2000;114:220-6
32. Yang R, Liu N, Chen L, Jiang Y, Shi Y, Mao C. et al. LSH interacts with and stabilizes GINS4 transcript that promotes tumourigenesis in non-small cell lung cancer. J Exp Clin Cancer Res. 2019;38:280
33. Jiang Y, Mao C, Yang R, Yan B, Shi Y, Liu X. et al. EGLN1/c-Myc Induced Lymphoid-Specific Helicase Inhibits Ferroptosis through Lipid Metabolic Gene Expression Changes. Theranostics. 2017;7:3293-305
34. He X, Yan B, Liu S, Jia J, Lai W, Xin X. et al. Chromatin Remodeling Factor LSH Drives Cancer Progression by Suppressing the Activity of Fumarate Hydratase. Cancer Res. 2016;76:5743-55
35. Liu S, Tao YG. Chromatin remodeling factor LSH affects fumarate hydratase as a cancer driver. Chin J Cancer. 2016;35:72
36. Weng CF, Chen LJ, Lin CW, Chen HM, Lee HH, Ling TY. et al. Association between the risk of lung cancer and influenza: A population-based nested case-control study. Int J Infect Dis. 2019;88:8-13
37. Sinagra E, Raimondo D, Gallo E, Stella M, Cottone M, Rossi F. et al. JC Virus and Lung Adenocarcinoma: Fact or Myth? Anticancer Res. 2017;37:3311-4
38. Sigel K, Makinson A, Thaler J. Lung cancer in persons with HIV. Curr Opin HIV AIDS. 2017;12:31-8
39. Bruyand M, Le Marec F, Lavole A, Leffondre K, Spano JP, Le Moing V. et al. Protease inhibitors exposure is not related to lung cancer risk in HIV smoker patients: a nested case-control study. Aids. 2015;29:1105-9
40. Koshiol J, Flores R, Lam TK, Taylor PR, Weinstein SJ, Virtamo J. et al. Helicobacter pylori seropositivity and risk of lung cancer. PLoS One. 2012;7:e32106
41. Corrales L, Rosell R, Cardona AF, Martin C, Zatarain-Barron ZL, Arrieta O. Lung cancer in never smokers: The role of different risk factors other than tobacco smoking. Crit Rev Oncol Hematol. 2020;148:102895
42. de Oliveira THA, do Amaral CM, de Franca Sao Marcos B, Nascimento KCG, de Miranda Rios AC, Quixabeira DCA. et al. Presence and activity of HPV in primary lung cancer. J Cancer Res Clin Oncol. 2018;144:2367-76
43. Bingula R, Filaire M, Radosevic-Robin N, Bey M, Berthon JY, Bernalier-Donadille A. et al. Desired Turbulence? Gut-Lung Axis, Immunity, and Lung Cancer. J Oncol. 2017;2017:5035371
44. Wang Q, Rao Y, Guo X, Liu N, Liu S, Wen P. et al. Oral Microbiome in Patients with Oesophageal Squamous Cell Carcinoma. Sci Rep. 2019;9:19055
45. Caselli E, Soffritti I, D'Accolti M, Piva I, Greco P, Bonaccorsi G. Atopobium vaginae And Porphyromonas somerae Induce Proinflammatory Cytokines Expression In Endometrial Cells: A Possible Implication For Endometrial Cancer? Cancer Manag Res. 2019;11:8571-5
46. Capoor MN, Birkenmaier C, Wang JC, McDowell A, Ahmed FS, Bruggemann H. et al. A review of microscopy-based evidence for the association of Propionibacterium acnes biofilms in degenerative disc disease and other diseased human tissue. Eur Spine J. 2019;28:2951-71
47. Feng Y, Ramnarine VR, Bell R, Volik S, Davicioni E, Hayes VM. et al. Metagenomic and metatranscriptomic analysis of human prostate microbiota from patients with prostate cancer. BMC Genomics. 2019;20:146
48. Bucevic Popovic V, Situm M, Chow CT, Chan LS, Roje B, Terzic J. The urinary microbiome associated with bladder cancer. Sci Rep. 2018;8:12157
49. Garcia-Castillo V, Sanhueza E, McNerney E, Onate SA, Garcia A. Microbiota dysbiosis: a new piece in the understanding of the carcinogenesis puzzle. J Med Microbiol. 2016;65:1347-62
50. Kipanyula MJ, Seke Etet PF, Vecchio L, Farahna M, Nukenine EN, Nwabo Kamdje AH. Signaling pathways bridging microbial-triggered inflammation and cancer. Cell Signal. 2013;25:403-16
51. Elinav E, Nowarski R, Thaiss CA, Hu B, Jin C, Flavell RA. Inflammation-induced cancer: crosstalk between tumours, immune cells and microorganisms. Nat Rev Cancer. 2013;13:759-71
52. Sommariva M, Le Noci V, Bianchi F, Camelliti S, Balsari A, Tagliabue E. et al. The lung microbiota: role in maintaining pulmonary immune homeostasis and its implications in cancer development and therapy. Cell Mol Life Sci. 2020;77:2739-49
53. Jungnickel C, Schmidt LH, Bittigkoffer L, Wolf L, Wolf A, Ritzmann F. et al. IL-17C mediates the recruitment of tumor-associated neutrophils and lung tumor growth. Oncogene. 2017;36:4182-90
54. Ochoa CE, Mirabolfathinejad SG, Ruiz VA, Evans SE, Gagea M, Evans CM. et al. Interleukin 6, but not T helper 2 cytokines, promotes lung carcinogenesis. Cancer Prev Res (Phila). 2011;4:51-64
55. Kunimasa K, Goto T. Immunosurveillance and Immunoediting of Lung Cancer: Current Perspectives and Challenges. Int J Mol Sci. 2020 21
56. Cheng M, Qian L, Shen G, Bian G, Xu T, Xu W. et al. Microbiota modulate tumoral immune surveillance in lung through a gammadeltaT17 immune cell-dependent mechanism. Cancer Res. 2014;74:4030-41
57. Cheng M, Chen Y, Wang L, Chen W, Yang L, Shen G. et al. Commensal microbiota maintains alveolar macrophages with a low level of CCL24 production to generate anti-metastatic tumor activity. Sci Rep. 2017;7:7471
58. Le Noci V, Guglielmetti S, Arioli S, Camisaschi C, Bianchi F, Sommariva M. et al. Modulation of Pulmonary Microbiota by Antibiotic or Probiotic Aerosol Therapy: A Strategy to Promote Immunosurveillance against Lung Metastases. Cell Rep. 2018;24:3528-38
59. Ma QY, Huang DY, Zhang HJ, Wang S, Chen XF. Upregulation of bacterial-specific Th1 and Th17 responses that are enriched in CXCR5(+)CD4(+) T cells in non-small cell lung cancer. Int Immunopharmacol. 2017;52:305-9
60. Zheng L, Xu J, Sai B, Zhu Y, Wang L, Yin N. et al. Microbiome Related Cytotoxically Active CD8+ TIL Are Inversely Associated With Lung Cancer Development. Front Oncol. 2020;10:531131
61. O'Keefe SJ. Diet, microorganisms and their metabolites, and colon cancer. Nat Rev Gastroenterol Hepatol. 2016;13:691-706
62. Vanhaecke L, Knize MG, Noppe H, De Brabander H, Verstraete W, Van de Wiele T. Intestinal bacteria metabolize the dietary carcinogen 2-amino-1-methyl-6-phenylimidazo[4,5-b]pyridine following consumption of a single cooked chicken meal in humans. Food Chem Toxicol. 2008;46:140-8
63. Keren N, Konikoff FM, Paitan Y, Gabay G, Reshef L, Naftali T. et al. Interactions between the intestinal microbiota and bile acids in gallstones patients. Environ Microbiol Rep. 2015;7:874-80
64. Apopa PL, Alley L, Penney RB, Arnaoutakis K, Steliga MA, Jeffus S. et al. PARP1 Is Up-Regulated in Non-small Cell Lung Cancer Tissues in the Presence of the Cyanobacterial Toxin Microcystin. Front Microbiol. 2018;9:1757
65. Singh N, Gurav A, Sivaprakasam S, Brady E, Padia R, Shi H. et al. Activation of Gpr109a, receptor for niacin and the commensal metabolite butyrate, suppresses colonic inflammation and carcinogenesis. Immunity. 2014;40:128-39
66. Xiao X, Cao Y, Chen H. Profiling and characterization of microRNAs responding to sodium butyrate treatment in A549 cells. J Cell Biochem. 2018;119:3563-73
67. Cuevas-Ramos G, Petit CR, Marcq I, Boury M, Oswald E, Nougayrede JP. Escherichia coli induces DNA damage in vivo and triggers genomic instability in mammalian cells. Proc Natl Acad Sci U S A. 2010;107:11537-42
68. He Z, Gharaibeh RZ, Newsome RC, Pope JL, Dougherty MW, Tomkovich S. et al. Campylobacter jejuni promotes colorectal tumorigenesis through the action of cytolethal distending toxin. Gut. 2019;68:289-300
69. Attene-Ramos MS, Wagner ED, Plewa MJ, Gaskins HR. Evidence that hydrogen sulfide is a genotoxic agent. Mol Cancer Res. 2006;4:9-14
70. Huycke MM, Gaskins HR. Commensal bacteria, redox stress, and colorectal cancer: mechanisms and models. Exp Biol Med (Maywood). 2004;229:586-97
71. Carbonero F, Benefiel AC, Alizadeh-Ghamsari AH, Gaskins HR. Microbial pathways in colonic sulfur metabolism and links with health and disease. Front Physiol. 2012;3:448
72. Wang J, Ge P, Qiang L, Tian F, Zhao D, Chai Q. et al. The mycobacterial phosphatase PtpA regulates the expression of host genes and promotes cell proliferation. Nat Commun. 2017;8:244
73. Chow SC, Gowing SD, Cools-Lartigue JJ, Chen CB, Berube J, Yoon HW. et al. Gram negative bacteria increase non-small cell lung cancer metastasis via Toll-like receptor 4 activation and mitogen-activated protein kinase phosphorylation. Int J Cancer. 2015;136:1341-50
74. Gowing SD, Chow SC, Cools-Lartigue JJ, Chen CB, Najmeh S, Jiang HY. et al. Gram-positive pneumonia augments non-small cell lung cancer metastasis via host toll-like receptor 2 activation. Int J Cancer. 2017;141:561-71
75. Wynendaele E, Verbeke F, D'Hondt M, Hendrix A, Van De Wiele C, Burvenich C. et al. Crosstalk between the microbiome and cancer cells by quorum sensing peptides. Peptides. 2015;64:40-8
76. Zhang YJ, Li S, Gan RY, Zhou T, Xu DP, Li HB. Impacts of gut bacteria on human health and diseases. Int J Mol Sci. 2015;16:7493-519
77. Ausprunk DH, Folkman J. Migration and proliferation of endothelial cells in preformed and newly formed blood vessels during tumor angiogenesis. Microvasc Res. 1977;14:53-65
78. Welti J, Loges S, Dimmeler S, Carmeliet P. Recent molecular discoveries in angiogenesis and antiangiogenic therapies in cancer. J Clin Invest. 2013;123:3190-200
79. Cui M, Xiao H, Li Y, Zhou L, Zhao S, Luo D. et al. Faecal microbiota transplantation protects against radiation-induced toxicity. EMBO Mol Med. 2017;9:448-61
80. Xia S, Menden HL, Korfhagen TR, Kume T, Sampath V. Endothelial immune activation programmes cell-fate decisions and angiogenesis by inducing angiogenesis regulator DLL4 through TLR4-ERK-FOXC2 signalling. J Physiol. 2018;596:1397-417
81. Wong SH, Zhao L, Zhang X, Nakatsu G, Han J, Xu W. et al. Gavage of Fecal Samples From Patients With Colorectal Cancer Promotes Intestinal Carcinogenesis in Germ-Free and Conventional Mice. Gastroenterology. 2017;153:1621-33.e6
82. Chang SH, Mirabolfathinejad SG, Katta H, Cumpian AM, Gong L, Caetano MS. et al. T helper 17 cells play a critical pathogenic role in lung cancer. Proc Natl Acad Sci U S A. 2014;111:5664-9
83. Chauhan SK, Jin Y, Goyal S, Lee HS, Fuchsluger TA, Lee HK. et al. A novel pro-lymphangiogenic function for Th17/IL-17. Blood. 2011;118:4630-4
84. Numasaki M, Watanabe M, Suzuki T, Takahashi H, Nakamura A, McAllister F. et al. IL-17 enhances the net angiogenic activity and in vivo growth of human non-small cell lung cancer in SCID mice through promoting CXCR-2-dependent angiogenesis. J Immunol. 2005;175:6177-89
85. Lee JW, Wang P, Kattah MG, Youssef S, Steinman L, DeFea K. et al. Differential regulation of chemokines by IL-17 in colonic epithelial cells. J Immunol. 2008;181:6536-45
86. Samareh-Fekri M, Hashemi Bajgani SM, Shafahi A, Asadi-Zarandi M, Mollaie H, Jamali Paghalhe A. Detection of Helicobacter pylori in the Bronchoalveolar Lavage of Patients with Lung Cancer Using Real-Time PCR. Jundishapur J Microbiol. 2016;9:e32144
87. Nakashima S, Kakugawa T, Yura H, Tomonaga M, Harada T, Hara A. et al. Identification of Helicobacter pylori VacA in human lung and its effects on lung cells. Biochem Biophys Res Commun. 2015;460:721-6
88. Deng C, Huang C, Wu Q, Pang J, Lin Y. A new sesquiterpene from the mangrove endophytic fungus Aspergillus terreus (No. GX7-3B). Nat Prod Res. 2013;27:1882-7
89. Zhang E, Feng X, Liu F, Zhang P, Liang J, Tang X. Roles of PI3K/Akt and c-Jun signaling pathways in human papillomavirus type 16 oncoprotein-induced HIF-1alpha, VEGF, and IL-8 expression and in vitro angiogenesis in non-small cell lung cancer cells. PLoS One. 2014;9:e103440
90. Liu X, Wu X, Ma Y, Zhang W, Hu L, Feng X. et al. Endophytic fungi from mangrove inhibit lung cancer cell growth and angiogenesis in vitro. Oncol Rep. 2017;37:1793-803
91. Daillere R, Vetizou M, Waldschmitt N, Yamazaki T, Isnard C, Poirier-Colame V. et al. Enterococcus hirae and Barnesiella intestinihominis Facilitate Cyclophosphamide-Induced Therapeutic Immunomodulatory Effects. Immunity. 2016;45:931-43
92. Li T, Kang G, Wang T, Huang H. Tumor angiogenesis and anti-angiogenic gene therapy for cancer. Oncol Lett. 2018;16:687-702
93. Carmeliet P. VEGF as a key mediator of angiogenesis in cancer. Oncology. 2005;69(Suppl 3):4-10
94. Jain RK. Normalization of tumor vasculature: an emerging concept in antiangiogenic therapy. Science. 2005;307:58-62
95. Fleitas T, Martinez-Sales V, Vila V, Reganon E, Mesado D, Martin M. et al. VEGF and TSP1 levels correlate with prognosis in advanced non-small cell lung cancer. Clin Transl Oncol. 2013;15:897-902
96. Chen P, Zhu J, Liu DY, Li HY, Xu N, Hou M. Over-expression of survivin and VEGF in small-cell lung cancer may predict the poorer prognosis. Med Oncol. 2014;31:775
97. Offersen BV, Pfeiffer P, Hamilton-Dutoit S, Overgaard J. Patterns of angiogenesis in nonsmall-cell lung carcinoma. Cancer. 2001;91:1500-9
98. Van Nuffel AM, Sukhatme V, Pantziarka P, Meheus L, Sukhatme VP, Bouche G. Repurposing Drugs in Oncology (ReDO)-clarithromycin as an anti-cancer agent. Ecancermedicalscience. 2015;9:513
99. Vétizou M, Pitt JM, Daillère R, Lepage P, Waldschmitt N, Flament C. et al. Anticancer immunotherapy by CTLA-4 blockade relies on the gut microbiota. Science. 2015;350:1079-84
100. Sivan A, Corrales L, Hubert N, Williams JB, Aquino-Michaels K, Earley ZM. et al. Commensal Bifidobacterium promotes antitumor immunity and facilitates anti-PD-L1 efficacy. Science. 2015;350:1084-9
101. Routy B, Le Chatelier E, Derosa L, Duong CPM, Alou MT, Daillère R. et al. Gut microbiome influences efficacy of PD-1-based immunotherapy against epithelial tumors. Science. 2018;359:91-7
102. Gopalakrishnan V, Spencer CN, Nezi L, Reuben A, Andrews MC, Karpinets TV. et al. Gut microbiome modulates response to anti-PD-1 immunotherapy in melanoma patients. Science. 2018;359:97-103
103. Chua MLK, Wee JTS, Hui EP, Chan ATC. Nasopharyngeal carcinoma. Lancet. 2016;387:1012-24
104. Iida N, Dzutsev A, Stewart CA, Smith L, Bouladoux N, Weingarten RA. et al. Commensal bacteria control cancer response to therapy by modulating the tumor microenvironment. Science. 2013;342:967-70
105. Viaud S, Saccheri F, Mignot G, Yamazaki T, Daillère R, Hannani D. et al. The intestinal microbiota modulates the anticancer immune effects of cyclophosphamide. Science. 2013;342:971-6
106. Geller LT, Barzily-Rokni M, Danino T, Jonas OH, Shental N, Nejman D. et al. Potential role of intratumor bacteria in mediating tumor resistance to the chemotherapeutic drug gemcitabine. Science. 2017;357:1156-60
107. Yu T, Guo F, Yu Y, Sun T, Ma D, Han J. et al. Fusobacterium nucleatum Promotes Chemoresistance to Colorectal Cancer by Modulating Autophagy. Cell. 2017;170:548-63.e16
108. Wallace BD, Wang H, Lane KT, Scott JE, Orans J, Koo JS. et al. Alleviating cancer drug toxicity by inhibiting a bacterial enzyme. Science. 2010;330:831-5
109. Li N, He F, Liao B, Zhou Y, Li B, Ran P. Exposure to ambient particulate matter alters the microbial composition and induces immune changes in rat lung. Respir Res. 2017;18:143
110. Yang B, Zhang Y, Li B, Zou Y, Xiao C. Fine particulate matter alters the microecology of the murine respiratory tract. Environ Sci Pollut Res Int. 2019;26:8623-32
111. Li X, Sun Y, An Y, Wang R, Lin H, Liu M. et al. Air pollution during the winter period and respiratory tract microbial imbalance in a healthy young population in Northeastern China. Environ Pollut. 2019;246:972-9
112. Rylance J, Kankwatira A, Nelson DE, Toh E, Day RB, Lin H. et al. Household air pollution and the lung microbiome of healthy adults in Malawi: a cross-sectional study. BMC Microbiol. 2016;16:182
113. Defois C, Ratel J, Denis S, Batut B, Beugnot R, Peyretaillade E. et al. Environmental Pollutant Benzo[a]Pyrene Impacts the Volatile Metabolome and Transcriptome of the Human Gut Microbiota. Front Microbiol. 2017;8:1562
114. Jerrett M, Burnett RT, Ma R, Pope CA 3rd, Krewski D, Newbold KB. et al. Spatial analysis of air pollution and mortality in Los Angeles. Epidemiology. 2005;16:727-36
115. Katanoda K, Sobue T, Satoh H, Tajima K, Suzuki T, Nakatsuka H. et al. An association between long-term exposure to ambient air pollution and mortality from lung cancer and respiratory diseases in Japan. J Epidemiol. 2011;21:132-43
116. Cao J, Yang C, Li J, Chen R, Chen B, Gu D. et al. Association between long-term exposure to outdoor air pollution and mortality in China: a cohort study. J Hazard Mater. 2011;186:1594-600
117. Cohen AJ, Brauer M, Burnett R, Anderson HR, Frostad J, Estep K. et al. Estimates and 25-year trends of the global burden of disease attributable to ambient air pollution: an analysis of data from the Global Burden of Diseases Study 2015. Lancet. 2017;389:1907-18
118. Yang B, Xiao C. PM2.5 exposure significantly improves the exacerbation of A549 tumor-bearing CB17-SCID mice. Environ Toxicol Pharmacol. 2018;60:169-75
119. Storr SJ, Woolston CM, Zhang Y, Martin SG. Redox environment, free radical, and oxidative DNA damage. Antioxid Redox Signal. 2013;18:2399-408
120. Valavanidis A, Vlachogianni T, Fiotakis C. 8-hydroxy-2' -deoxyguanosine (8-OHdG): A critical biomarker of oxidative stress and carcinogenesis. J Environ Sci Health C Environ Carcinog Ecotoxicol Rev. 2009;27:120-39
121. Vineis P, Husgafvel-Pursiainen K. Air pollution and cancer: biomarker studies in human populations. Carcinogenesis. 2005;26:1846-55
122. Zhou W, Tian D, He J, Wang Y, Zhang L, Cui L. et al. Repeated PM2.5 exposure inhibits BEAS-2B cell P53 expression through ROS-Akt-DNMT3B pathway-mediated promoter hypermethylation. Oncotarget. 2016;7:20691-703
123. Belinsky SA, Snow SS, Nikula KJ, Finch GL, Tellez CS, Palmisano WA. Aberrant CpG island methylation of the p16(INK4a) and estrogen receptor genes in rat lung tumors induced by particulate carcinogens. Carcinogenesis. 2002;23:335-9
124. Qin Y, Wade PA. Crosstalk between the microbiome and epigenome: messages from bugs. J Biochem. 2018;163:105-12
125. Jiang CL, He SW, Zhang YD, Duan HX, Huang T, Huang YC. et al. Air pollution and DNA methylation alterations in lung cancer: A systematic and comparative study. Oncotarget. 2017;8:1369-91
126. Yang CY, Yeh YM, Yu HY, Chin CY, Hsu CW, Liu H. et al. Oral Microbiota Community Dynamics Associated With Oral Squamous Cell Carcinoma Staging. Front Microbiol. 2018;9:862
127. Zhao H, Chu M, Huang Z, Yang X, Ran S, Hu B. et al. Variations in oral microbiota associated with oral cancer. Sci Rep. 2017;7:11773
128. Lu H, Ren Z, Li A, Li J, Xu S, Zhang H. et al. Tongue coating microbiome data distinguish patients with pancreatic head cancer from healthy controls. J Oral Microbiol. 2019;11:1563409
129. Allali I, Boukhatem N, Bouguenouch L, Hardi H, Boudouaya HA, Cadenas MB. et al. Gut microbiome of Moroccan colorectal cancer patients. Med Microbiol Immunol. 2018;207:211-25
130. Kim K, Castro EJT, Shim H, Advincula JVG, Kim YW. Differences Regarding the Molecular Features and Gut Microbiota Between Right and Left Colon Cancer. Ann Coloproctol. 2018;34:280-5
Author contact
Corresponding authors: Y.T. E-mail: taoyongedu.cn; Y.S. E-mail: yingshiedu.cn; Department of Pathology, Xiangya Hospital, Central South University, Hunan, 410078 China. Tel.: +(86) 731-84805448; Fax: +(86) 731-84470589.

 Global reach, higher impact
Global reach, higher impact