Impact Factor
ISSN: 1449-2288
Int J Biol Sci 2022; 18(8):3298-3312. doi:10.7150/ijbs.71431 This issue Cite
Research Paper
Quantitative proteomics of HFD-induced fatty liver uncovers novel transcription factors of lipid metabolism
1. Central Laboratory, Department of Liver Diseases, ShuGuang Hospital Affiliated to Shanghai University of Chinese Traditional Medicine, Shanghai, China
2. Institute of Infectious Disease, ShuGuang Hospital Affiliated to Shanghai University of Chinese Traditional Medicine, Shanghai, China
3. Department of Ultrosound, Shuguang Hospital Affiliated to Shanghai University of Chinese Traditional Medicine, Shanghai, China
4. Department of General Surgery, Shanghai Ninth People' Hospital, Shanghai Jiao Tong University School of medicine, Shanghai, China
* These authors contributed equally to this study.
Abstract
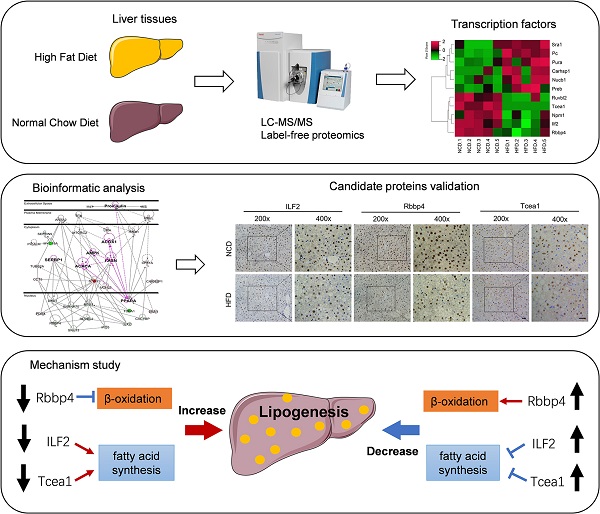
Nonalcoholic fatty liver disease (NAFLD) has become the most common chronic liver disease, which progression is tightly regulated by transcription factors (TFs), nuclear receptors, and cellular enzymes. In this study, a label-free quantitative proteomic approach was used to determine the effect of the high-fat diet on the proteomics profile of liver tissue and to identify novel NAFLD related TFs. Mice were fed with HFD for 16 weeks to establish a NAFLD mouse model. Mice fed with normal chow diet were taken as controls. Liver samples were collected from each group for proteomics analysis. A total of 2298 proteins were quantified, among which 106 proteins were downregulated, while 256 proteins were upregulated in HFD-fed mice compared with the controls with fold change more than 1.5 and p value less than 0.05. Bioinformatic analysis revealed that metabolic-related functions and pathways were most significantly enriched. A subgroup of 11 TFs were observed to share interactions with metabolic-related enzymes and kinases by protein-protein interaction analysis. Among them, 7 TFs were selected for verification, and 3 TFs were finally validated, including Rbbp4, Tcea1, and ILF2. Downregulating each of the 3 TFs could significantly promote lipid accumulation in AML12 hepatocytes, by regulating the expression of fatty acid synthesis- or β-oxidation-related genes. In contrast, overexpression of Tcea1, Rbbp4, and ILF2, respectively, could ameliorate hepatocyte steatosis. These findings propose novel lipid metabolism related TFs, which might have potential roles in preventing NAFLD.
Keywords: quantitative proteomics, NAFLD, transcription factor, lipid metabolism

 Global reach, higher impact
Global reach, higher impact