Impact Factor ISSN: 1449-2288
Int J Biol Sci 2022; 18(13):4837-4852. doi:10.7150/ijbs.71411 This issue Cite
Research Paper
MiR-138-5p Targets MACF1 to Aggravate Aging-related Bone Loss
1. Lab for Bone Metabolism, Xi'an Key Laboratory of Special Medicine and Health Engineering; Key Lab for Space Biosciences and Biotechnology, Research Center for Special Medicine and Health Systems Engineering, NPU-UAB Joint Laboratory for Bone Metabolism, School of Life Sciences, Northwestern Polytechnical University, Xi'an, Shaanxi, 710072, China.
2. Department of Spine Surgery, Shenzhen People's Hospital, Southern University of Science and Technology, Jinan University School of Medicine, Shenzhen, 518020, China.
3. Key Laboratory of Biomedical Information Engineering of Ministry of Education, and Institute of Molecular Genetics, School of Life Science and Technology, Xi'an Jiaotong University, Xi'an, Shaanxi, 710049, China.
4. Law Sau Fai Institute for Advancing Translational Medicine in Bone and Joint Diseases, School of Chinese Medicine, Hong Kong Baptist University, Hong Kong, SAR, 999077, China.
5. Department of Gynaecology and Obstetrics, Xijing Hospital, Fourth Military Medical University, Xi'an, Shaanxi, 710032, China.
#Co-first authors.
Received 2022-1-25; Accepted 2022-6-26; Published 2022-7-18
Abstract
Senile osteoporosis is one of the major health problems in an aging society. Decreased bone formation due to osteoblast dysfunction may be one of the causes of aging-related bone loss. With increasing evidence suggesting that multiple microRNAs (miRNAs) play important roles in osteoblast function, the relationship between miRNAs and senile osteoporosis has become a popular research topic. Previously, we confirmed that mechanoresponsive miR-138-5p negatively regulated bone anabolic action. In this study, the miR-138-5p level was found to be negatively correlated with BMD and osteogenic markers in bone specimens of senile osteoporotic patients by bioinformatic analysis and experimental verification. Furthermore, high miR-138-5p levels aggravated the decrease of aged osteoblast differentiation in vitro and led to worse bone loss in aged osteoblastic miR-138-5p transgenic mice in vivo. We also previously identified that the target of miR-138-5p, microtubule actin cross-linking factor 1 (MACF1), could attenuate senile osteoporosis. Here, miR-138-5p was demonstrated to regulate aged osteoblast differentiation by targeting MACF1. Finally, the therapeutic inhibition of miR-138-5p counteracted the decrease in bone formation and aging-related bone loss in aged mice. Overall, our results highlight the crucial roles and the molecular mechanism of miR-138-5p in aging-related bone loss and may provide a powerful therapeutic target for ameliorating senile osteoporosis.
Keywords: senile osteoporosis, miR-138-5p, MACF1, osteoblast differentiation, aging-related bone loss
Introduction
Senile osteoporosis is a degenerative disease and has become one of the major health problems worldwide with the demographic shifts caused by the dramatic increase in human life expectancy [1, 2]. Only 31-36% of people above the age of 70 have normal bone density, while the remainder suffer from some form of osteopenia or osteoporosis, which is a major contributor to the loss of independence due to bone fractures in the elderly [3]. Substantial changes in bone structure occur with aging, including decreases in the thickness and mass of trabecular bone, loss rather than sparing of cortical bone, and an increase in bone marrow fat [4]. With aging, bone loss occurs due to a process in which the bone remodeling balance is biased toward bone resorption rather than bone formation due to the decrease in the number and activity of osteoblasts [5]. The changes in aging bone that lead to osteoporosis are mediated at multiple levels, including hormonal changes, bone unloading, and the accumulation of senescent cells [4].
MicroRNAs (miRNAs) are small noncoding RNAs that play important roles in posttranscriptional gene regulation [6]. Accumulating evidence has suggested that multiple miRNAs play essential regulatory roles in aged osteoblast differentiation and aging-related bone loss [7-10]. Recently, miR-138-5p was reported to play vital roles in osteogenic differentiation and bone formation [11-13]. In our previous studies, we have confirmed that miR-138-5p, as a mechanoresponsive miRNA, accounts for the mechanosensitivity of the bone anabolic response [14]. In addition, miR-138-5p has been demonstrated to negatively regulate osteoblast differentiation by inhibiting β-catenin under simulated microgravity [15]. However, the relationship between miR-138-5p and senile bone loss has not been investigated, and the underlying mechanisms need to be studied.
In this study, we identified miR-138-5p as a novel aging-related miRNA in humans, mice and cultured cells for the first time. Moreover, the miR-138-5p level was negatively correlated with osteoblast differentiation and bone formation. We subsequently validated the relationship of miR-138-5p with the target gene microtubule actin cross-linking factor 1 (MACF1) in aged osteoblast differentiation and senile osteoporosis by loss- and gain-of-function of miR-138-5p in vitro. Moreover, we verified the involvement of miR-138-5p in senile osteoporosis in vivo by comparing young and aged mice with osteoblastic miR-138-5p transgenic (TG) mice. We observed that aged TG mice showed more severe osteoporosis symptoms. Finally, the increased bone formation and reduced bone loss induced by administering miR-138-5p inhibitor to aged mice also suggested that the inhibition of miR-138-5p may be a potential therapeutic strategy for senile osteoporosis.
Materials and Methods
MicroRNA expression profile analysis of differentially expressed miRNAs in plasma of osteoporotic patients
The microRNA expression profile was tested in plasma of healthy patients (T-score ≥ -1, n = 6) or osteoporotic patients (T-score ≤ -2.5, n = 6) [16, 17]. In this study, we further used this expression profile to analyze the relationship of miR-138-5p expression and bone mineral density (BMD) in osteoporotic patients. The microRNA expression profile has been uploaded to GEO database (GSE93883).
Human bone specimen collection
The bone specimens of osteoporotic patients (11 female between 60 and 79 years of age were recruited) were collected from the Shenzhen People's Hospital (Table S1 and Table S2). The patients with malignancy, osteoarthritis or systemic inflammatory diseases, liver and kidney diseases, chronic corticosteroid use, or other severe diseases in the previous five years were excluded from the present study (exclusion criteria). All the clinical procedures were approved by the Committees of Clinical Ethics of the Shenzhen People's Hospital and conformed to the principles of the Helsinki Declaration. We obtained informed consent from all the participants.
Aged Mice
The C57BL/6 mice used in the present study were purchased from the Laboratory Animal Center of Air Force Medical University (Xi'an, Shaanxi, China). The mice were maintained in normal humidity (55-60%), temperature (25 °C), and light-dark (12 h/12 h) SPF conditions, given free access to water and feed. Naturally aged male mice used in this study were monitored continuously until the mice were more than 20-month-old, and the unhealthy mice were excluded. All mouse experiments were in accordance with the Guide for the Care and Use of Laboratory Animals and approved by the Laboratory Animal Ethics & Welfare Committee of Northwestern Polytechnical University.
The osteoblastic miR-138-5p transgenic mouse model
The osteoblastic miR-138-5p transgenic (TG) mouse model was constructed as described previously [14]. In brief, the recombinant plasmid including the miR-138-5p sequence with osteoblastic marker Runx2 promoter was microinjected into a zygote to obtain stable TG mouse model. In this study, by mating the 2-6 month-old male TG mice with 2-4 month-old wild-type female mice, the genotype of transgenic mice were +/- mice, and the genotype of littermate WT mice were -/- mice.
RNA extraction and Real-time PCR
High-quality total RNA was extracted from tissue or cell samples by the Omega miRNA kit (Omega, USA) according to the manufacturer's instruction. The quality of RNA was then tested by ultraviolet spectrophotometry. For miRNA and mRNA, both cDNAs were synthesized with the PrimeScript™ RT reagent Kit (Takara, Japan). Real-time PCR was performed by the using of SYBR® Premix Ex Taq™ II kit (Takara, Japan) following the manufacturer's instruction. The mRNAs and miRNAs relative expression was analyzed by the 2-ΔΔCT method. What's more, Gapdh was used as an internal control for mRNAs, and U6 was used as an internal control for miRNAs.
Primary osteoblast isolation and cell culture
Primary osteoblasts in Figure 1 were isolated from the femora and tibias of 6-month-old or 21-month-old male mice according to bone research protocol [18]. In brief, femora and tibias were excised with removing all muscles and placed in a petri dish with the α-MEM medium after euthanize adult mice. Then, the epiphyses of both ends were cut off and the bone marrow was flushed out by 27-gauge needle. The cleaned diaphyses were cut into little pieces of approximately 1-2 mm2 using scissors and washed with α-MEM medium several times. The bone pieces were incubated in collagenase II (Gibco, USA) solution at 37 °C in a shaking watch bath (150-200 rpm) for 2 hours (4 times, 30 min every time). From the fifth to the seventh incubation (30 min every time), primary long bone osteoblasts were collected for culture.
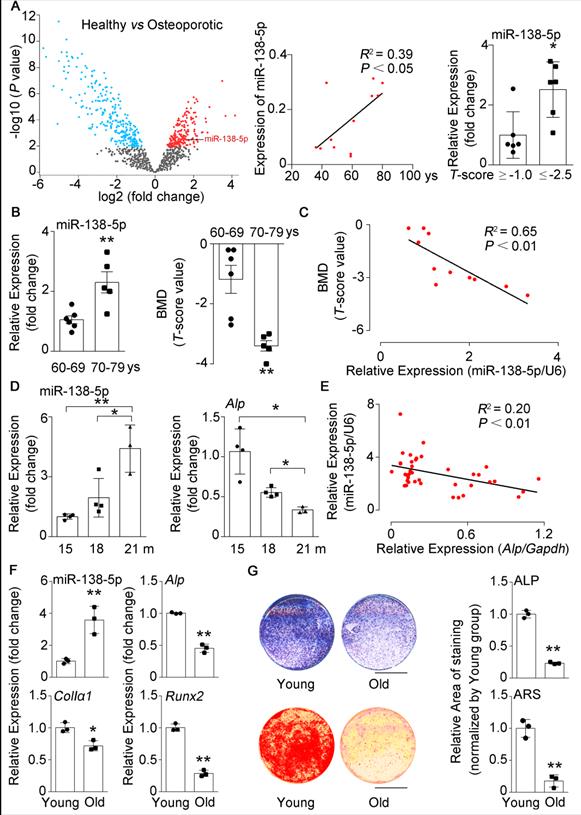
miR-138-5p levels are elevated with age in humans, mice and cultured cells. (A) The volcano plot analysis of differentially expressed miRNAs (left), linear regression analysis of miR-138-5p expression and age association (middle), relative miR-138-5p expression (right) in plasma of healthy patients (T-score ≥ -1, n = 6) or osteoporotic patients (T-score ≤ -2.5, n = 6). (B) Real-time PCR analysis of miR-138-5p (left) in bone specimens, and T-score (right) of spine from aged female fracture patients (60-69 years, n = 6; 70-79 years, n = 5). (C) Linear regression analysis of miR-138-5p and T-score association in aged female fracture patients. (D) Real-time PCR analysis of miR-138-5p (left) and Alp (right) levels in tibias from 15, 18 and 21-month-old male C57BL/6 mice, respectively. (15 m, n = 4; 18 m, n = 4; 21 m, n = 3). (E) Linear regression analysis of miR-138-5p and Alp association in wild-type C57BL/6 mice with different age. n = 39. (F) Real-time PCR analysis of miR-138-5p and osteogenic marker gene (Alp, ColIa1 and Runx2) mRNA levels in primary osteoblasts from 6-month-old (young) and 21-month-old (old) mice, respectively. n = 3 for each group. (G) Representative images of ALP staining (upper, left) and quantification of ALP staining areas (upper, right), and Alizarin Red S (ARS) staining (lower, left, 10 d) and quantification of ARS staining areas (lower, right) in primary osteoblasts from 6-month-old (young) and 21-month-old (old) mice, respectively. Scale bar, 5 mm. n = 3 for each group. U6 small nuclear RNA was used as the internal control for miR-138-5p. Gapdh was used as the internal control for mRNAs. For human experiments, Data are represented as mean ± sem. For mice and cells experiments, Data are represented as mean ± s.d. Statistical differences among three groups were analyzed via one-way ANOVA and significances were determined using student's t-test between two groups. P value less than 0.05 was considered significant in all cases (*P < 0.05, **P < 0.01).
Mouse primary calvarial osteoblasts in Supplement Figure 4 were isolated from the calvaria of 1-2 d neonatal mice pups according to bone research protocol [18]. Briefly, the calvaria of neonatal mice pups was excised with removing the edges and sutures and placed in a petri dish with the α-MEM medium. The calvaria was washed with α-MEM medium for several times. Then cleaned calvaria was incubated in collagenase I (Gibco, USA) solution at 37 °C in a shaking watch bath (150-200 rpm) for 2 hours and removed into a 6-well plate for culture.
The primary osteoblasts were maintained in α-MEM (Gibco, USA) with 10% FBS (Corning, USA), 1% penicillin and streptomycin (Amresco, USA). The primary osteoblasts were cultured under 5% CO2, 95% humidity and 37 °C incubator, and confluent cells were passaged with 0.25% trypsin. The primary osteoblasts were not used beyond passage 4.
Construction of stable osteoblastic cell line and cell culture
For the construction of the stable miR-138-5p overexpression osteoblastic cell line (Hi-138) in murine osteoblastic MC3T3-E1 cells, recombinant lentivirus vectors carrying miR-138-5p precursor or one scrambled negative sequence were designed and synthesized by Genepharma (Shanghai, China) as described previously [15]. For the construction of the stable knockdown MACF1 osteoblastic cell line (KD-Macf1) in MC3T3-E1 cells, recombinant lentivirus vectors carrying shRNA targeting mice Macf1 (NM_001199136.1) or its scramble control were designed and synthesized by Genepharma (Shanghai, China) as described previously [19]. The cells were maintained in α-MEM supplemented with 10% FBS (Biological Industries, Israel), 1% penicillin and streptomycin (Amresco, USA). The cells were cultured under 5% CO2, 95% humidity and 37 °C incubator, and confluent cells were passaged with 0.25% trypsin.
Cell transfection
The osteoblasts were seeded at plates with a density of 105 cells/well until the confluence reached 80% ~ 90%, cells were transfected with 50 nM antagomir-138-5p or agomir-138-5p, or 1 μg/ml MACF1 siRNA by lipofectamine 2000 (invitrogen, USA) following the manufacturer's instructions. The serum-free medium was changed as the standard α-MEM medium after 6-8 h. The osteoblasts were incubated at 37 °C for 48 h and then harvested for real-time PCR, western blot analysis and cell staining. Antagomir-138-5p (Gene Pharma, China) sequences were 5'-CGGCCUGAUUCACAACACCAGCU-3'. Agomir-138-5p (Ribobio, China) sequences were 5'- AGCUGGUGUUGUGAAUCAGGCCG-3' (positive-sense strand), 5'-UCGACCACAACACUUAAUCCGGC-3' (antisense strand), MACF1 siRNA (Xi'an RQC, China) effective sequences were 5'-GCUAGUGAAUAUCCGCAAUGA-3'.
Alkaline phosphatase (ALP) staining
Alkaline phosphatase staining was examined by BCIP/NBT alkaline phosphatase color development kit (Beyotime, China) as reported previously [14, 15, 20]. Briefly, cells were fixed and rinsed with PBS. Then fixed cells were added BCIP/NBT liquid substrate until the color of cells turned blue/purple. Finally, cells were rinsed in deionized water for several times. The plate with stained cells was imaged with a scanner (Canon, Japan).
Alizarin Red S (ARS) staining
For osteoblast differentiation, the osteoblasts were cultured in medium with 10% FBS, 1% streptomycin and penicillin, 50 μg/ml ascorbic acid (Sigma, USA) and 10 mM β-glycerophosphate (Sigma, USA) for about 2-3 weeks. Then, the osteoblasts were fixed and rinsed with PBS. The fixed cells were subsequently stained with 0.5% alizarin red s (Sigma, USA) solution (pH 4.2) for 15-30 min. Finally, cells were rinsed in deionized water for several times. The plate with stained cells was imaged with a scanner (Canon, Japan).
Aged osteoblast induced by etoposide
The osteoblasts were seeded at plates at a density of 105 cells/well. Cells were added with 2 μM etoposide (Solarbio, China) subsequently upon reaching 70% ~ 80% confluency. 48 h later, the medium was changed as the standard α-MEM medium. The osteoblasts were for cell transfection or incubated at 37 °C for 48 h and then harvested for real-time PCR, western blot analysis and cell staining.
β-galactosidase staining
β-galactosidase activity was examined by β-galactosidase staining kit according to manufacturer's instructions (Beyotime, China). Briefly, cells were fixed and rinsed with PBS. Then fixed cells were added β-galactosidase liquid substrate for 24-48 h. Finally, cells were rinsed in deionized water for several times.
Micro-computed tomography (micro-CT) analysis
The femur samples were fixed in 4% paraformaldehyde and scanned by micro-CT devices (μCT40, Switzerland) to assess and image bone microarchitecture in Figure 3 and Supplement Figure 5 as reported previously [7, 14]. Briefly, the scanning resolution was set as 10 μm and 423 slices were scanned from the growth plate of the distal femur beginning. For micro-CT analysis, 80 continuous slices were selected as the region of interest (ROI) from beginning the distal growth plate disappearing to calculate the following parameters, bone mineral density (BMD), bone volume to tissue volume (BV/TV), trabecular spacing (Tb.Sp), trabecular number (Tb.N), trabecular thickness (Tb.Th) and structure model index (SMI). For 3D reconstruction of trabecular bone, the related parameters were set as sigma = 1.2, supports = 2 and threshold = 200.
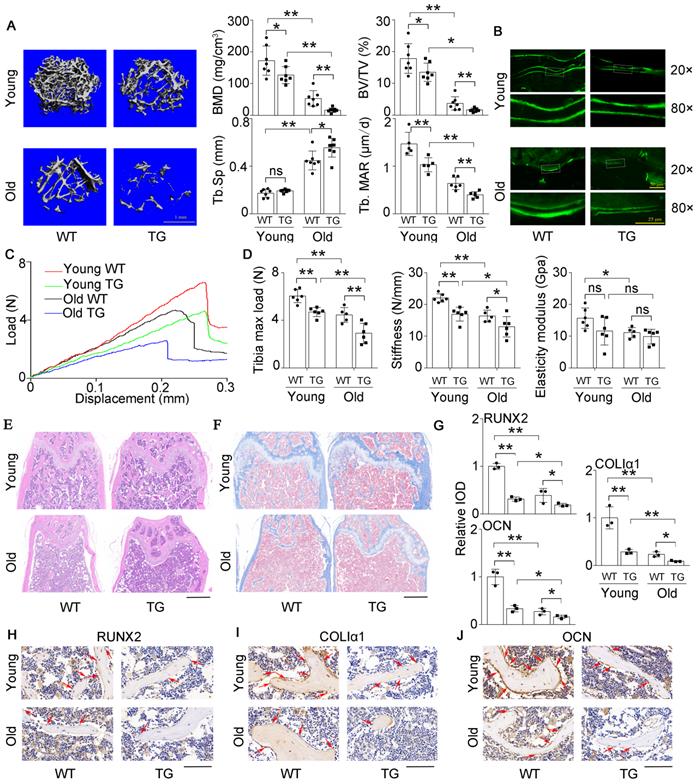
High miR-138-5p levels aggravate aging-related bone loss. (A) Representative 3D reconstruction images showing microarchitecture, and microCT analysis of BMD, BV/TV and Tb.Sp in distal femur of young or old osteoblastic miR-138-5p transgenic (TG) mice and wildtype (WT) mice, respectively. BMD, bone mineral density; BV/TV, Bone volume per tissue volume; Tb.Sp, trabecular spacing, young, 3-month-old, old, 19-month-old. Scale bar, 1 mm. Young-WT, n = 7; Young-TG, n = 7; Old-WT, n = 7; Old-TG, n = 8. Dynamic histomorphometric analysis of MAR showing bone formation capacity in femur trabecular (lower, right) bone of young or old TG mice and WT mice, respectively. MAR, mineral apposition rate. Young-WT, n = 5; Young-TG, n = 5; Old-WT, n = 6; Old-TG, n = 5. (B) Representative calcein double labeling images showing bone formation capacity in femur trabecular bone of young or old TG mice and WT mice. Upper scale bar, 50 µm. Lower scale bar, 25 µm. young, 3-month-old, old, 19-month-old. (C) Representative load-deflection curves for the respective groups in tibias of young or old TG mice and WT mice. young, 3-month-old, old, 19-month-old. (D) Three-point bending measurement of tibia max load, stiffness and elasticity modulus in young or old TG mice and WT mice. young, 3-month-old, old, 19-month-old. Young-WT, n = 6; Young-TG, n = 6; Old-WT, n = 5; Old-TG, n = 6. (E) Representative images of H&E staining analysis in distal femur of young or old TG mice and WT mice. Scale bar, 500 µm. young, 3-month-old, old, 19-month-old. (F) Representative images of Masson's trichrome staining analysis in distal femur of young or old TG mice and WT mice. Scale bar, 500 µm. young, 3-month-old, old, 19-month-old. (G, H) Representative images of immunohistochemical (IHC) staining and quantification analysis of RUNX2 (left), COLIα1 (middle) and OCN (right) in distal femur of young or old TG mice and WT mice. Scale bar, 100 µm. young, 3-month-old, old, 19-month-old. n = 3 mice for each group. Data are represented as mean ± s.d. Two-way ANOVA was performed to study the interaction between two independent variables. Significances were determined using student's t-test between two groups. P value less than 0.05 was considered significant in all cases (*P < 0.05, **P < 0.01).
The femur samples were fixed in 4% paraformaldehyde and scanned by micro-CT devices (General Electric, WI) to assess and image bone microarchitecture in Figure 6 and Supplement Figure 1 as reported previously [14, 21]. Briefly, the scanning parameters were set as following, Energy: 80 kVp/80 μA; Angle of Increment: 0.5°; Exposure Time: 3000 ms/frame; Scanning Time: 120 min and the scanning resolution were set as 8 μm. For micro-CT analysis, 1 mm was selected as the region of interest (ROI) from beginning the distal growth plate disappearing to calculate the following parameters, bone mineral content (BMC), BMD, BV/TV, Tb.N, Tb.Th and Tb.Sp. The 3D reconstruction of trabecular bone was used by the microview software.
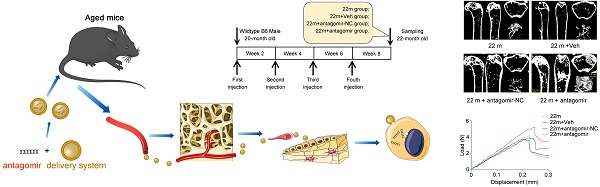
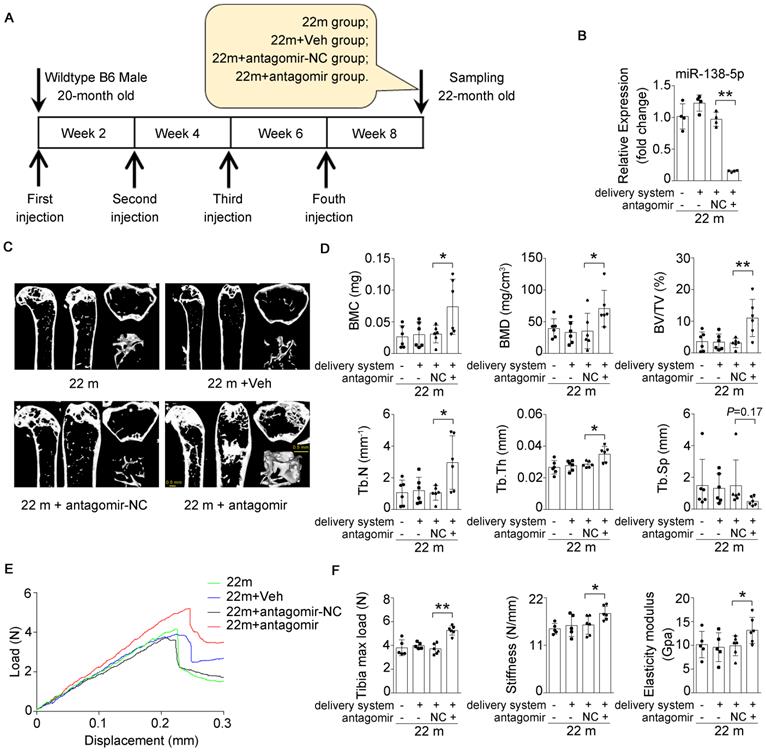
Bone-targeted delivery of miR-138-5p inhibitor alleviates aging-related bone loss. (A) Schematic graph showing experiment grouping, antagomir-138-5p treatment in aged mice. Mice were injected with antagomir-138-5p four times every two weeks during two-month therapy. 22 m (22-month-old mice, without any injection), 22 m + Veh (22-month-old mice injected with osteoblast-targeted delivery system), 22 m + antagomir-NC (22-month-old mice injected with osteoblast-targeted delivery system and antagomir-NC), 22 m + antagomir (22-month-old mice injected with osteoblast-targeted delivery system and antagomir-138-5p). (B) Real-time PCR analysis of miR-138-5p relative level in bone tissue of aged mice after antagomir-138-5p treatment. n = 4 for each group. (C) Representative 3D reconstruction images showing microarchitecture in distal femur of aged mice after antagomir-138-5p treatment. Scale bar, 0.5 mm. (D) The microCT statistical analysis of BMC, BMD, BV/TV, Tb.Th, Tb.N and Tb.Sp in distal femur of aged mice after AMO treatment. BMC, bone mineral content; BMD, bone mineral density; BV/TV, bone volume to tissue volume; Tb.N, trabecular number; Tb.Th, trabecular thickness; Tb.Sp, trabecular spacing. n = 6 mice for each group. (E) Representative load-deflection curves for the respective groups in tibias of aged mice after antagomir-138-5p treatment. (F) Three-point bending measurement of tibia max load, stiffness and elasticity modulus in tibias of aged mice after antagomir-138-5p treatment. 22 m, n = 5; 22 m + Veh, n = 5; 22 m + NC, n = 6; 22 m + antagomir, n = 6. U6 small nuclear RNA was used as the internal control for miR-138-5p. Data are represented as mean ± s.d. Two-way ANOVA was performed to study the interaction between two independent variables. Then, statistical differences among three or more groups were analyzed via one-way ANOVA and significances were determined using student's t-test between two groups. P value less than 0.05 was considered significant in all cases (*P < 0.05, **P < 0.01).
Bone mechanical property
The three-point bending mechanical test system (UniVert, Canada) was used to examine the bone mechanical properties of tibias. The anterior surface on the two lower support points spaced 10 mm apart. The tibias were fixed on a bracket with a loading speed of 0.6 mm/min until fracture occurred at the anterior surface on the two lower support points spaced 10 mm apart. Load-displacement curves were acquired and used to calculate the maximum load at failure, stiffness and elasticity modulus.
Double calcein labeling
All mice were intraperitoneally injected with 10 mg/kg calcein green (Sigma, USA) at 10 d and 2 d before euthanasia. After dehydration and embedding into gradient methyl methacrylate (MMA), the fixed femur samples were then sliced into 20-40 μm sections by the EXAKT 300CP tissue cutting system and the EXAKT 400CS micro grinding system (EXAKT Apparatebau, Germany). Sections were examined with a fluorescence microscope (Nikon, Japan), bone dynamic parameter (MAR) was calculated with ImageJ software (NIH, USA) [14, 20].
Bone histological decalcified sections staining
The femurs were fixed with 4% PFA for 2 days, decalcified with EDTA for two weeks, and then embedded in paraffin. Serial 4 μm thick tissue sections for hematoxylin and eosin (HE) staining, Masson's trichrome staining or immunohistochemical (IHC) staining following the manufacturer's instructions as we have reported (Servicebio, China) [14, 22]. The stained sections were scanned by a Digital Whole Slide Scanner (Leica, Germany).
Western blot
The osteoblasts were harvested and then lysed with Cell Lysis Buffer (Beyotime, China) containing a protease inhibitor cocktail (Merck, Germany) on ice. The protein extracts were collected after 4 °C centrifugation (15,000 g for 5 min) and then boiled with SDS-PAGE protein loading buffer (Biosharp, China). The protein was transferred to nitrocellulose membranes (PALL, USA) after separation via 8% SDS-PAGE (MACF1) or 10% SDS-PAGE (COLIα1 and GAPDH). The separated protein fractions on the membranes were blocked with 5% skim milk and incubated overnight at 4 °C with specific primary antibodies on a shaking device. HRP-labeled secondary antibody (Beijing CoWin Bioscience, China) was added after primary antibodies rinsed off, and then protein bands were visualized using ECL chemiluminescence reagents (Rockford, United States) by X-ray film (Kodak, NY) or T5200 Multi Chemiluminescence Detection System (Tanon, China). The primary antibodies included MACF1 Rabbit pAb (1:1000, Abcam, USA), COLIα1 Rabbit pAb (1:1000, Cell Signaling Technology, USA) and GAPDH Rabbit pAb (1:1000, Calbiochem, Germany) [23].
Dual-luciferase reporter assay
MACF1 3′UTR vector and its mutant vector which we previously reported [14] were used to further detect direct interaction between MACF1 and miR-138-5p. Briefly, the reporter plasmids were co-transfected with pRL-TK Renilla plasmid into MC3T3-E1 cells (8×106 cells per well), and antagomir-138-5p or antagomir-NC transfection was performed as reported [14]. The osteoblasts were harvested for the luciferase assay 48 h after transfection, and then performed with the dual-luciferase reporter assay system (Promega, USA) according to the manufacturer's instructions. Each value of quantified luminescent signals from the firefly luciferase was normalized by Renilla luciferase.
Antagomir-138-5p therapy in aged mice
20-month-old male C57BL/6 mice received antagomir-138-5p (antagomir) or antagomir-NC (10 mg/kg body weight) with the osteoblast-targeted delivery system [7] by tail vein injection for four consecutive injections every two weeks. The groups were as followed: 22 m group (no treatment), 22 m + Veh group (injection of the osteoblast-targeted delivery system alone), 22 m + antagomir-NC group (injection of antagomir-NC with the osteoblast-targeted delivery system) and 22 m + antagomir group (injection of antagomir-138-5p with the osteoblast-targeted delivery system). Before euthanasia, all mice were intraperitoneally injected with 10 mg/kg calcein green at 10 d and 2 d before euthanasia.
Statistical analysis
For data consisting of two independent variables, two-way analysis of variance (ANOVA) was performed to study the interaction between fixed factors. Then, statistical differences among three or more groups were analyzed via one-way ANOVA and significance between two groups was determined using Student's t-test. For human experiments, all numerical data were presented as the mean ± sem. For the mouse and cell experiments, all numerical data were presented as the mean ± sd. All statistical analyses were performed with GraphPad Prism 6 software and P < 0.05 was considered statistically significant for all comparisons (*P < 0.05, **P < 0.01).
Results
miR-138-5p levels are elevated with age in humans, mice and cells
In our previous study, we found that silencing of mechanoresponsive miR-138-5p could partially neutralize disuse osteoporosis [14]. Interestingly, our miRNA expression profile (GSE93883) analysis of plasma of healthy or osteoporotic patients showed that the miR-138-5p expression was increased with age and elevated in osteoporotic patients (T-score ≤ -2.5) (Figure 1A). Hence, the role of miR-138-5p in senile bone loss attracted our attention. Furthermore, we have previously confirmed that the miR-138-5p level was increased with age in bone specimens of aged male osteoporotic patients [14]. Here, we further detected and verified that the expression of miR-138-5p was also elevated and spine bone mineral density (T-score) was significantly decreased in bone specimens of female osteoporotic patients (Figure 1B). Moreover, the miR-138-5p level was negatively correlated to T-score (Figure 1C). Furthermore, the relationship between miR-138-5p level and age was validated in bone tissue of mice and primary osteoblasts, and the results showed that the expression of miR-138-5p was increased with age at mice or cellular levels, respectively (Figure 1D, F). In contrast, the bone architecture, trabecular bone mass-related parameters (bone mineral density (BMD), bone volume to tissue volume (BV/TV), trabecular thickness (Tb.Th.), trabecular t (Tb.N.)) and markers of osteoblast differentiation or bone formation (osteogenic marker genes expression, alkaline phosphatase (ALP) activity and matrix mineralization) were inversely correlated with miR-138-5p level in bone tissue and primary osteoblasts of aged mice (Figure 1D-G, Figure S1A, B). These results suggested that miR-138-5p is an aging-related miRNA and that its level was negatively correlated to osteoblast differentiation and bone formation in human tissues, mice and cultured cells.
High miR-138-5p levels aggravate the decrease in osteoblast differentiation induced by aging
To further validate the roles of miR-138-5p in regulating osteoblast differentiation, we first detected the expression of miR-138-5p at different time points (1 day, 7 days, 14 days) in the osteogenic differentiation process. We found that the expression of osteogenic marker factors (Alp, ColIα1, Runx2) increased while miR-138-5p expression was gradually decreased during primary calvaria osteoblast differentiation (Figure S2A, B). Furthermore, primary calvaria osteoblasts were transfected with agomir-138-5p (miR-138-5p agonist, agomir) or antagomir-138-5p (miR-138-5p inhibitor, antagomir). Real-time PCR analysis confirmed that the expression level of intracellular miR-138-5p was significantly upregulated by agomir-138-5p, and substantially downregulated by antagomir-138-5p compared to their corresponding controls (Figure S2C, G). Functionally, the expression levels of Alp, ColIα1, and Runx2, ALP activity and matrix mineralization nodules formation were weakened by agomir-138-5p, whereas enhanced by antagomir-138-5p (S-Figure 2C-J). Previously, we have constructed a stable miR-138-5p overexpression osteoblastic cell line (Hi-138) in murine osteoblastic MC3T3-E1 cells with lentivirus [15]. Compared to the negative control cell line (Hi-NC), Hi-138 cells showed higher miR-138-5p level and lower levels of osteoblast differentiation markers (marker gene expression (Alp, ColIα1, and Runx2), ALP activity and matrix mineralization nodule formation)). However, silencing of miR-138-5p could partly rescue the reduction of osteoblast differentiation markers induced by high miR-138-5p level in Hi-138 cells treating with antagomir-138-5p (Figure 2A-D).
High miR-138-5p levels aggravate the decrease in osteoblast differentiation induced by aging. (A) Real-time PCR analysis of miR-138-5p and osteogenic marker gene (Alp, ColIa1 and Runx2) mRNA levels in stable miR-138-5p overexpression osteoblastic cell line (Hi-138) treating with either antagomir-138-5p (antagomir) or antagomir-NC (NC), or in negative control cell line (Hi-NC) for 48 h, respectively. n = 3 for each group. (B) Representative images of ALP staining in Hi-138 cells treating with either antagomir-138-5p or antagomir-NC, or in Hi-NC cells for 48 h, respectively. Scale bar, 5 mm. (C) Representative images of ARS staining in Hi-138 cells treating with either antagomir-138-5p or antagomir-NC, or in Hi-NC cells for 14 d respectively. Scale bar, 5 mm. (D) Quantification of ALP staining areas (upper) and ARS staining areas (lower, 14 d) in Hi-138 cells treating with either antagomir-138-5p or antagomir-NC, or in Hi-NC cells, respectively. n = 3 for each group. (E) Real-time PCR analysis of miR-138-5p and osteogenic marker gene (Alp, ColIa1 and Runx2) mRNA levels in Hi-138 cells or Hi-NC cells treating with 2-day etoposide-induced aging, respectively. n = 3 for each group. (F) Representative images of ALP staining in Hi-138 cells or Hi-NC cells treating with 2-day etoposide inducing aging, respectively. Scale bar, 5 mm. (G) Representative images of ARS (18 d) staining in Hi-138 cells or Hi-NC cells treating with 2-day etoposide-induced aging, respectively. Scale bar, 5 mm. (H) Quantification of ALP staining areas (upper) and ARS staining areas (lower, 18 d) in Hi-138 cells or Hi-NC cells treating with 2-day etoposide-induced aging, respectively. n = 3 for each group. (I) Real-time PCR analysis of miR-138-5p and osteogenic marker gene (Alp, ColIa1 and Runx2) mRNA levels in MC3T3-E1 cells treating with either antagomir-138-5p or antagomir-NC after 2-day etoposide-induced aging for 48 h, respectively. n = 3 for each group. (J) Representative images of ALP staining in MC3T3-E1 cells treating with either antagomir-138-5p or antagomir-NC after 2-day etoposide-induced aging for 48 h, respectively. Scale bar, 5 mm. (K) Representative images of ARS staining (14 d) in MC3T3-E1 cells treating with either antagomir-138-5p or antagomir-NC after 2-day etoposide-induced aging, respectively. Scale bar, 5 mm. (L) Quantification of ALP staining areas (upper) and ARS staining areas (lower, 14 d) in MC3T3-E1 cells treating with either antagomir-138-5p or antagomir-NC after 2-day etoposide-induced aging, respectively. n = 3 for each group. U6 small nuclear RNA was used as the internal control for miR-138-5p, and Gapdh was used as the internal control for mRNAs. Data are represented as mean ± s.d. Two-way ANOVA was performed to study the interaction between two independent variables. Then, statistical differences among three or more groups were analyzed via one-way ANOVA and significances were determined using student's t-test between two groups. P value less than 0.05 was considered significant in all cases (*P < 0.05, **P < 0.01).
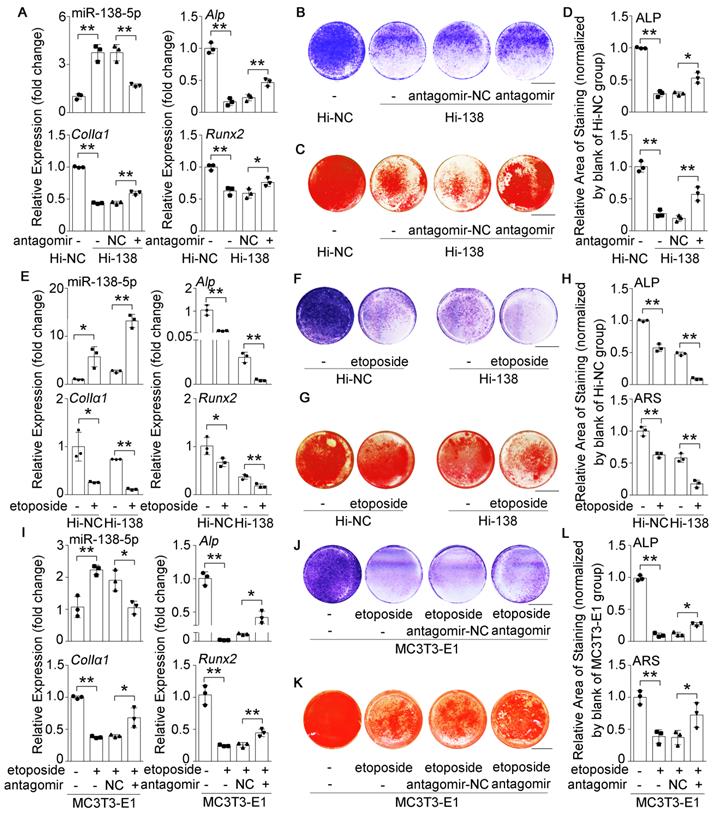
To further identify the effects of miR-138-5p on aging-related decreased osteoblast differentiation, we used etoposide to induce senescence in Hi-138 cells or Hi-NC cells. The results showed that the miR-138-5p level and cell senescence were increased, while osteoblast differentiation markers were decreased after treatment with etoposide. Moreover, the decrease in osteoblast differentiation induced by elevated miR-138-5p level was aggravated by etoposide-induced aging (Figure 2E-H, Figure S3A, C, D). In addition, the decreases in Alp, ColIα1, and Runx2 mRNA levels, COLIα1 protein level, ALP activity and matrix mineralization nodules formation, and the increases in cell senescence induced by etoposide-induced aging were partially neutralized by transfection with antagomir-138-5p in osteoblastic MC3T3-E1 cells (Figure 2I-L, Figure S3B, E, F). These results together demonstrated that high miR-138-5p level could exacerbate aging-related reduced osteoblast differentiation in vitro.
High miR-138-5p levels aggravate aging-related bone loss
Previously, we constructed an osteoblastic miR-138-5p transgenic (TG) mouse model by the Runx2 promoter [14] and found that the bone formation activity of TG mice was decreased compared to that of wild-type (WT) mice. Here, the expression of Alp, ColIα1, and Runx2, ALP activity and matrix mineralization were significantly suppressed in primary calvarial osteoblasts isolated from TG mice, compared to WT mice (Figure S4A-D). To further investigate the effect of miR-138-5p on aging-related bone loss in vivo, TG mice and WT mice were divided into young (3-month) and old (19-month) groups. We scrutinized bone mass and trabecular microarchitecture and found that the bone mineral density and bone mass in the TG mice declined significantly compared to that in the controls in both the young group and old group, and the trabecular microarchitecture appeared to have deteriorated in TG mice. These phenomena worsened with age (Figure 3A, Figure S5A). The calcein-labeled bone formation experiment showed that the bone formation rate and osteoblast number decreased in TG mice and declined more with age (Figure 3A, B and Figure S5B). With increasing age, the skeletal mechanical strength properties (Tibia max load and stiffness) of mice tibias in the TG groups were significantly weaker compared to those in the WT groups (Figure 3C, D). HE staining and Masson's trichrome staining images showed that TG mice had loose bone trabeculae compared to WT mice, this phenotype was more severe in older mice (Figure 3E, F). As expected, the IHC assay also showed that the expression levels of osteogenic genes (Runx2, ColIα1and Ocn) were lower in TG mice than in WT mice. Likewise, the expression levels of these genes further decreased with age (Figure 3G-J). These results indicated that high miR-138-5p level could exacerbate aging-related bone loss and the decreased in bone formation in vivo.
miR-138-5p suppresses osteoblast differentiation by targeting and inhibiting MACF1 in senile bone loss
MiR-138-5p has been demonstrated to directly target MACF1 to regulate osteoblast differentiation in our previous study [14]. However, the relationship between miR-138-5p and MACF1 in senile bone loss needs further validation. We determined that MACF1 expression was inhibited or promoted after transfecting MC3T3-E1 osteoblasts with agomir-138-5p or antagomir-138-5p, respectively (Figure 4A, B and Figure S6A, B). We then cloned the binding region or the mutant sequence into a dual luciferase reporter vector (Figure 4C). We found that luciferase activity was significantly elevated in the cells simultaneously transfected with antagomir-138-5p and Macf1 WT 3' UTR plasmid, while plasmids bearing mutations inmiR-138-5p or the Macf1 3' UTR could not change the luciferase activity significantly (Figure 4D). Additionally, we used Macf1-siRNA (si-Macf1) to knock down Macf1 in miR-138-5p high expression (Hi-138) cells. The results showed that the expressions of Macf1 and osteogenic genes (Alp, ColIα1, and Runx2) mRNA levels, ALP activity and matrix mineralization nodules formation were further attenuated after MACF1 deficiency in Hi-138 cells, compared to Hi-NC cells (Figure 4E-H). These data suggested that miR-138-5p directly targeted MACF1 to regulate osteoblast differentiation.
miR-138-5p suppresses osteoblast differentiationby targeting and inhibiting MACF1. (A) Real-time PCR (left) and Western blot (right) analysis of MACF1 expression level in MC3T3-E1 cells treating with either agomir-138-5p (agomir) or agomir-NC for 48 h, respectively. n = 3 for each group. (B) Real-time PCR (left) and Western blot (right) analysis of MACF1 expression level in MC3T3-E1 cells treating with either antagomir-138-5p (antagomir) or antagomir-NC for 48 h, respectively. n = 3 for each group. (C) Schematic graph showing the design of dual-luciferase reporters with either Macf1 WT 3'UTR or Macf1 mutant 3'UTR. (D) The effects of antagomir-138-5p (antagomir) or antagomir-NC on luciferase activity of Macf1 WT 3'UTR reporter (Luc-WT UTR, left), Macf1 mutant 3'UTR reporter (Luc-Mut UTR, right) in MC3T3-E1 cells, respectively (left). (E) Real-time PCR analysis of Macf1 and osteogenic marker gene (Alp, ColIa1 and Runx2) mRNA levels in stable miR-138-5p overexpression osteoblastic cell line (Hi-138) treating with either MACF1 siRNA (si-Macf1) or negative control siRNA (si-NC), or in negative control cell line (Hi-NC) for 48 h, respectively. n = 3 for each group. (F) Representative images of ALP staining in Hi-138 cells or Hi-NC cells treating with either si-Macf1 or si-NC for 48 h, respectively. Scale bar, 5 mm. (G) Representative images of ARS (18 d) staining in Hi-138 cells or Hi-NC cells treating with either si-Macf1 or si-NC for 48 h, respectively. Scale bar, 5 mm. (H) Quantification of ALP staining areas (upper) and ARS staining areas (lower, 18 d) in Hi-138 cells or Hi-NC cells treating with either si-Macf1 or si-NC for 48 h, respectively. n = 3 for each group. Gapdh was used as the internal control for mRNAs. Data are represented as mean ± s.d. Two-way ANOVA was performed to study the interaction between two independent variables. Then, statistical differences among three or more groups were analyzed via one-way ANOVA and significances were determined using student's t-test between two groups. P value less than 0.05 was considered significant in all cases (*P < 0.05, **P < 0.01).
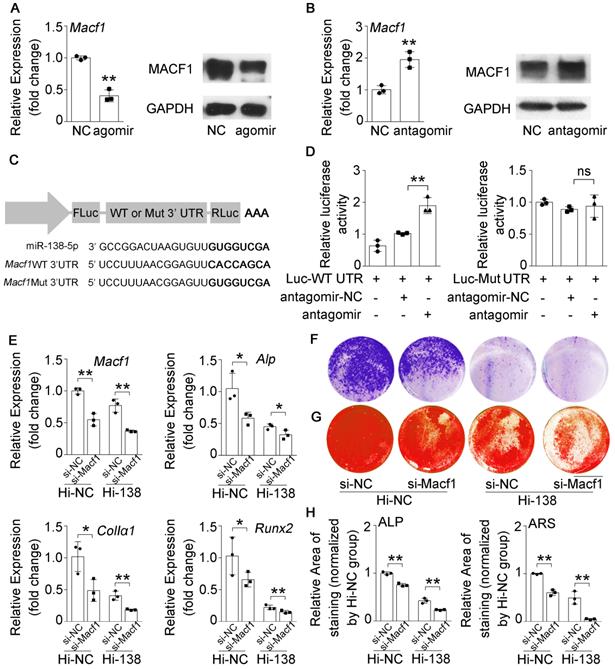
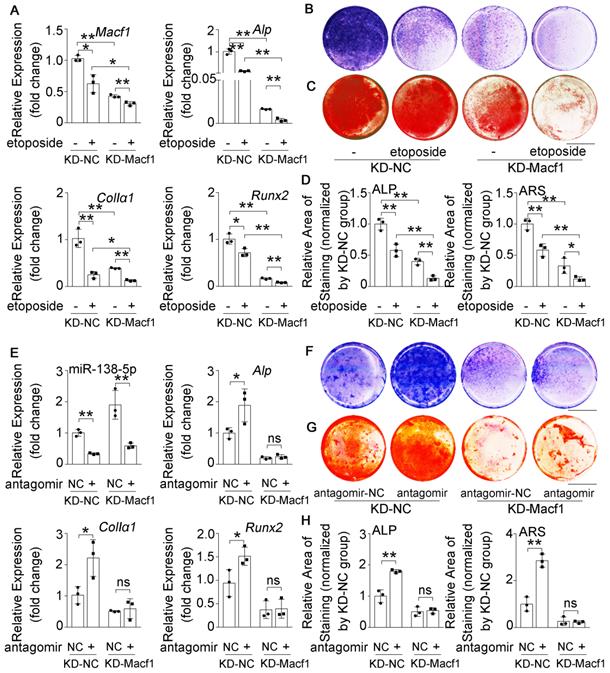
To further validate the relationship between miR-138-5p and MACF1 during aging-related bone loss, we first treated the stable MACF1 low-expressing (KD-Macf1) cells with etoposide to induce osteoblast senescence (Figure S6E, F). The results showed that Macf1 and osteogenic gene (Alp, ColIα1, and Runx2) mRNA levels, COLIα1 protein level, ALP activity and matrix mineralization were further attenuated after etoposide-induced aging in MACF1-deficient osteoblasts (Figure 5A-D, Figure S6C-E), which was similar to the previously observed osteoblast differentiation phenomenon in which miR-138-5p high-expressing osteoblasts were treated with etoposide to induce aging or si-Macf1 to induce Macf1 deficiency. Furthermore, we transfected antagomir-138-5p into either KD-Macf1 cells or negative control (KD-NC) cells. In KD-NC osteoblasts, the miR-138-5p level was significantly decreased and osteogenic marker gene expression, COLIα1 protein level, ALP activity and the formation of mineralized nodules were increased after treating with antagomir-138-5p. However, decreasing the miR-138-5p level in KD-Macf1 osteoblasts led to a no significant difference in the markers of osteoblast differentiation (Figure 5E-H, Figure S6F). These results suggested that in aging-related reduced osteoblast differentiation, miR-138-5p regulated osteogenic differentiation by targeting and inhibiting MACF1.
MACF1 is involved in miR-138-5p regulating aged osteoblast differentiation. (A) Real-time PCR analysis of Macf1 and osteogenic marker gene (Alp, ColIa1 and Runx2) mRNA levels in stable MACF1 low-expressing (KD-Macf1) cells treating with 2-day etoposide-induced aging, or in negative control cell line (KD-NC) for 48 h, respectively. n = 3 for each group. (B) Representative images of ALP staining in KD-Macf1 cells or KD-NC cells either treating with 2-day etoposide-induced aging for 48 h, respectively. Scale bar, 5 mm. (C) Representative images of ARS (16 d) staining in KD-Macf1 cells or KD-NC cells treating with 2-day etoposide-induced aging for 48 h, respectively. Scale bar, 5 mm. (D) Quantification of ALP staining areas (upper) and ARS staining areas (lower, 16 d) in KD-Macf1 cells or KD-NC cells treating with 2-day etoposide-induced aging for 48 h, respectively. n = 3 for each group. (E) Real-time PCR analysis of miR-138-5p and osteogenic marker gene (Alp, ColIa1 and Runx2) mRNA levels in KD-Macf1 cells or KD-NC cells treating with either antagomir-138-5p or antagomir-NC for 48 h, respectively. n = 3 for each group. (F) Representative images of ALP staining in KD-Macf1 cells or KD-NC cells treating with either antagomir-138-5p or antagomir-NC for 8 d, respectively. Scale bar, 5 mm. (G) Representative images of ARS staining (14 d) in KD-Macf1 cells or KD-NC cells treating with either antagomir-138-5p or antagomir-NC for 48 h, respectively. Scale bar, 5 mm. (H) Quantification of ALP staining areas (upper) and ARS staining areas (lower, 14 d) in KD-Macf1 cells or KD-NC cells treating with either antagomir-138-5p or antagomir-NC for 48 h, respectively. n = 3 for each group. U6 small nuclear RNA was used as the internal control for miR-138-5p. Gapdh was used as the internal control for mRNAs. Data are represented as mean ± s.d. Two-way ANOVA was performed to study the interaction between two independent variables. Significances were determined using student's t-test between two groups. P value less than 0.05 was considered significant in all cases (*P < 0.05, **P < 0.01).
Inhibition of miR-138-5p can alleviate aging-related bone loss
Based on our findings that an increased miR-138-5p level was accompanied by bone loss with age and that silencing of miR-138-5p could partially decrease osteoblast differentiation induced by aging in vitro (Figure 1-3) [14], we hypothesized that therapeutic inhibition of miR-138-5p could rescue the aging-related bone loss in vivo as well. To test our hypothesis, we separately treated 20-month-old male wild-type C57/BL6 mice with pharmacological antagomir-138-5p (antagomir) or antagomir-NC (NC) carried by an osteoblast-targeting delivery system [7] for 4 consecutive injections via the tail vein every two weeks (Figure 6A). We first confirmed that antagomir-138-5p could effectively downregulate miR-138-5p level in aged bone tissue (Figure 6B). In addition, we performed microCT analysis and found that antagomir-138-5p relieved the deterioration of trabecular bone mass and trabecular microarchitecture caused by aging (Figure 6C). Furthermore, the trabecular bone parameter analysis showed that BMC, BMD, BV/TV, Tb.N and Tb.Th. were significantly rescued after treatment with pharmacological antagomir-138-5p (Figure 6D). To further investigate the functions of pharmacological antagomir-138-5p on tibial mechanical strength properties in aged mice, we performed a three-point bending mechanical test and found that the tibial max load, stiffness, and elasticity modulus were also significantly increased in the antagomir group (Figure 6E, F).
Inhibition of miR-138-5p can alleviate the decrease in bone formation induced by aging
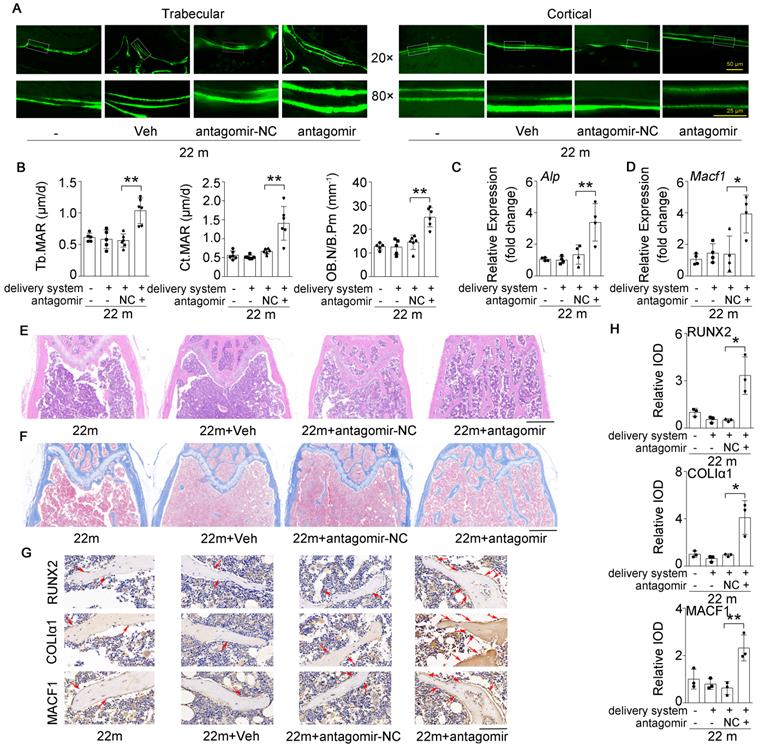
Given that the osteoblastic miR-138-5p level was elevated and negatively correlated with bone formation in aged mice (Figure 1 and Figure 3), we further tested whether therapeutic inhibition of osteoblastic miR-138-5p could prevent the aging-related decrease in bone formation. Calcein double-labeling confirmed that aged mice treated with pharmacological antagomir-138-5p showed a significantly higher bone formation rate and related parameter (MAR) of trabecular and cortical bone than other normal control groups (Figure 7A, B). Moreover, HE staining and Masson's trichrome staining images showed that elevated trabecular bone and increased osteoid were observed in aged mice after pharmacological antagomir-138-5p treatment (Figure 7E, F). To further examine the expression of bone formation marker gene (Alp, Runx2 and ColIα1) and the miR-138-5p target MACF1, we performed the real-time PCR and IHC assay analysis. The results showed that the expression levels of osteogenic genes and MACF1 in the bone tissue of aged mice were significantly increased in the antagomir-138-5p group (Figure 7C, D and Figure 7G, H). These data suggested that the pharmacological miR-138-5p inhibitor could alleviate the decrease in bone formation and improve senile bone loss symptoms in aged mice.
Bone-targeted inhibition of miR-138-5p can alleviate the decrease in bone formation induced by aging. (A) Representative calcein double labeling images showing bone formation capacity in femur trabecular and cortical bone of aged mice after antagomir-138-5p treatment. Upper scale bar, 50 µm. Lower scale bar, 25 µm. 22 m (22-month-old mice, without any injection), 22 m + Veh (22-month-old mice injected with osteoblast-targeted delivery system), 22 m + antagomir-NC (22-month-old mice injected with osteoblast-targeted delivery system and antagomir-NC), 22 m + antagomir (22-month-old mice injected with osteoblast-targeted delivery system and antagomir-138-5p). (B) Dynamic histomorphometric analysis of MAR showing bone formation capacity in femur trabecular (left) and cortical (middle) bone of aged mice after antagomir-138-5p treatment; Static histomorphometric analysis of OB.N/B.Pm (right) showing osteoblast number in femur trabecular bone of aged mice after antagomir-138-5p treatment. OB.N/B.Pm: osteoblast number per bone perimeter. 22 m, n = 5; 22 m + Veh, n = 5; 22 m + antagomir-NC, n = 6; 22 m + antagomir, n = 6. (C, D) Real-time PCR analysis of Alp and Macf1 relative levels in bone tissue of aged mice after antagomir-138-5p treatment. n = 4 for each group. (E) Representative images of H&E staining analysis in distal femur of aged mice after antagomir-138-5p treatment. Scale bar, 500 µm. (F) Representative images of Masson's trichrome staining analysis in distal femur of aged mice after antagomir-138-5p treatment. Scale bar, 500 µm. (G, H) Representative images of immunohistochemical (IHC) staining and quantification analysis of RUNX2 (upper), COLIα1 (middle) and MACF1 (lower) in distal femur of aged mice after antagomir-138-5p treatment. Scale bar, 100 µm. n = 3 for each group. Gapdh was used as the internal control for mRNAs. Data are represented as mean ± s.d. Two-way ANOVA was performed to study the interaction between two independent variables. Then, statistical differences among three or more groups were analyzed via one-way ANOVA and significances were determined using student's t-test between two groups. P value less than 0.05 was considered significant in all cases (*P < 0.05, **P < 0.01).
Discussion
In the present study, we identified miR-138-5p as a novel aging-related miRNA that negatively regulates osteogenesis by targeting and inhibiting MACF1 in aging-related bone loss. Especially for senile osteoporosis, elevated miR-138-5p levels led to greater decrease in osteoblast differentiation and more severe bone loss with aging. In addition, administering a miR-138-5p inhibitor to aged mice alleviated senile bone loss symptoms, which provided a new mechanism and a potential therapeutic strategy for the treatment of senile osteoporosis.
Senile osteoporosis is a growing public health problem worldwide, and it is essential to develop more effective approaches for its treatment [1, 2]. It has been reported that osteoblast dysfunction inducing decreased bone formation may be one of the major causes of senile osteoporosis [9, 14, 24-29]. In the current study, we built upon our previous study [14] and found that BMD and osteogenic marker gene levels gradually decreased with age in bone specimens from aged osteoporotic patients. Furthermore, bone formation and osteoblast differentiation were also reduced with aging in mice and cultured cells. Therefore, it is necessary to systematically clarify the mechanism by which aged osteoblast dysfunction leads to a decrease in osteogenesis during senile osteoporosis.
With the growing evidence of the important regulatory roles of miRNAs in aged osteoblast differentiation and aging-related bone loss [7-9, 25, 30-34], the relationship between miRNAs and senile osteoporosis has become a focus of substantial research. Recently, miR-146a was found to be an essential epigenetic switch controlling osteoblast generation and aging-related bone loss by regulating bone anabolic Wnt signaling [25]. In addition, Sun et al. reported that miR-103-3p was negatively correlated with bone formation in bone specimens from elderly women with fractures [9]. Here, we further confirmed that miR-138-5p was elevated with age and negatively correlated with osteoblast differentiation and osteogenesis in human tissues, mice and cultured cells (Figure 1). In our previous study, we screened and identified several miRNAs, including miR-138-5p, as potential biomarkers or therapeutic targets in human osteoporosis [14, 34, 35]. These data suggested that miR-138-5p might play an important role in senile osteoporosis.
Recently, miR-138-5p was reported to play vital roles in different bone diseases, including osteoarthritis [36, 37], osteosarcoma [38] and osteoporosis [14, 39, 40]. Moreover, the functions of miR-138 in the osteogenetic differentiation process attracted broad attention[11, 12, 39, 41, 42] after Eskildsen et al. verified that miR-138 modulates the osteogenic differentiation of human mesenchymal stem cells [40]. In addition, we previously found that miR-138-5p negatively regulated osteoblast differentiation and bone formation under different mechanical conditions [14] and simulated microgravity [15]. In the present study, we constructed a stable miR-138-5p overexpression osteoblastic cell line (Hi-138) in vitro and an osteoblastic miR-138-5p transgenic (TG) mice model in vivo to deeply investigate the effects of miR-138-5p gain-of-function on aging-related osteogenesis. A high miR-138-5p level enhanced the decrease in osteoblast differentiation induced by aging (Figure 2). Specifically, the bone formation rate, trabecular microarchitecture and skeletal mechanical strength properties became significantly weaker with age in TG mice, compared to the WT mice (Figure 3). These results demonstrated that elevated miR-138-5p aggravated decrease of osteoblast differentiation and bone formation induced by aging. Given that aging-related osteoporosis is a growing public health concern, it is necessary to identify effective approaches for the treatment of this disease. With regard to translational medicine for therapeutic consideration, a pharmacological miR-138-5p inhibitor alleviated the decrease in bone formation and improved senile bone loss symptoms in aged mice (Figures 6 and 7), suggesting a novel potential therapeutic strategy for aging-related osteoporosis.
Previously, multiple miR-138 target genes during the osteogenic differentiation process have been reported, including Fak [40, 43], Zeb2 [42], Eif4ebp1 [44], BMPR2 [45], Trps1 and Sulf2 [46], but these studies mostly focused on the osteogenic differentiation of MSCs. In our previous study, we found that miR-138-5p could directly target microtubule actin crosslinking factor 1 (MACF1) in primary osteoblast differentiation [14], which was further confirmed here (Figure 4D). After mutation of the binding sequence of miR-138-5p and the Macf1 3'UTR, the luciferase activity in osteoblasts was not significantly changed by treatment with agomir-138-5p [14] or antagomir-138-5p in osteoblasts (Figure 4). These data show that MACF1 is a potential target of miR-138-5p in osteoblast differentiation.
MACF1, an important cytoskeletal protein, is crucial in controlling cytoskeleton dynamics through binding to both microtubules and F-actin [47, 48]. In our laboratory, we have long concentrated on the roles and mechanisms of MACF1 in different bone cells and various bone diseases [14, 21-23, 49-55]. In osteoblasts, MACF1 promotes osteoblast proliferation, migration and differentiation by regulating Wnt/β-catenin signaling [23, 49, 50, 52, 55]. Recently, we found that miR-138-5p suppresses osteoblast differentiation by inhibiting β-catenin under simulated microgravity [15]. These results provide a possible underlying mechanism for the involvement of miR-138-5p in senile osteoporosis. Furthermore, we established osteoblast-specific MACF1 conditional knockout mice with Osterix promoter to closely examine the function of MACF1 in bone formation in vivo. The results showed that deficiency of MACF1 in osteoblasts inhibited osteoblast differentiation and bone formation through the Bmp2/Smad/Runx2 pathway [22]. In a recent study, we found that the expression of MACF1 decreased with age in femur tissues of patients with aging-related osteoporosis, and overexpression of MACF1 alleviated the decrease in bone formation and osteoporosis in aged mice [53]. In the present study, we also found that the reduced osteoblast differentiation induced by low MACF1 levels was aggravated after etoposide-induced aging (Figure 5) and that miR-138-5p regulated aged osteoblast differentiation depended on MACF1. In addition, the therapeutic treatment of aged mice with a pharmacological miR-138-5p inhibitor led to the significant elevation of MACF1 expression (Figure 7), which indicated that the inhibition of miR-138-5p alleviated senile osteoporosis by targeting the regulation of MACF1.
In summary, the current findings identify a new aging-related miRNA, miR-138-5p, that negatively modulates osteoblast differentiation and bone formation in senile osteoporosis by targeting MACF1. High miR-138-5p levels aggravate the decrease in osteoblast differentiation and severity of bone loss with age. Moreover, treatment with a pharmacological miR-138-5p inhibitor could alleviate aging-related bone loss, providing a potential therapeutic approach for treating senile osteoporosis.
Abbreviations
ALP/Alp: alkaline phosphatase; agomir: agomir-138-5p; antagomir: antagomiR-138-5p; ARS: alizarin red staining; BMC: bone mineral content; BMD: bone mineral density; BV/TV: bone volume to tissue volume (bone volume fraction); ColIα1/COLIα1: collagen type 1 alpha-1; GAPDH/Gapdh: glyceraldehyde3-phosphatedehydrogenase; Hi-138: stable miR-138-5p overexpression osteoblastic cell line; Hi-NC: Hi-138 corresponding negative control osteoblastic cell line; IHC: immunohistochemical; KD-Macf1: stable MACF1 low-expressing osteoblastic cell line; KD-NC: KD-Macf1 corresponding negative control osteoblastic cell line; MACF1/Macf1: microtubule actin cross-linking factor 1; MAR: mineral apposition rate; microCT: Micro-computed tomography; miRNA: microRNA; NC: antagomir-NC; OCN: osteocalcin; OB.N/B.Pm: osteoblast number per bone perimeter; WT: Wild type mice; Runx2/RUNX2: runt-related transcription factor 2; Si-Macf1: Macf1 siRNA; Si-NC: negative control siRNA; Tb.N: trabecular number; Tb.Th: trabecular thickness; Tb.Sp: trabecular spacing; TG: osteoblastic miR-138-5p transgenic mice; Veh: osteoblast-targeted delivery system.
Supplementary Material
Supplementary figures and tables.
Acknowledgements
The authors would like to thank Dr. Hong Zhou (The University of Sydney, Australia) for generously providing MC3T3-E1 cell line. This work was supported by the Natural Science Foundation of China (32101055, 82072106, 31570940, 81872129, 81700784 and 81772017), the China Postdoctoral Science Foundation (2020M683573 and 2017M613196), the Project Supported by Natural Science Basic Research Plan in Shaanxi Province of China (2021JQ-128 and 2018JM3040), Shaanxi Provincial Key R&D Program (2018KWZ-10 and 2021SF-242), the Innovation Foundation for Doctor Dissertation of Northwestern Polytechnical University (CX201821).
Author contributions
Designing research studies: Qian AR, Zhang G, Chen ZH and Lin X; Conducting experiments: Qian AR, Lin X, Zhang G, Peng SL, Chen ZH and Huai Y; Acquiring data: Chen ZH, Huai Y, Chen GY, Li DJ, Zhang Y, Yin C and Liu SY; Analyzing data: Chen ZH, Huai Y, Chen XF and Zhang Y; bioinformatic analysis: Huai Y, Chen XF and Peng SL; Providing reagents: Chen ZH, Zhang G, Peng SL, Chen GY, Huai Y, Li DJ, Zhang Y, Liu SY, Zhao F, Yin C, Lin X, Yang CF, Mao WJ, Wang XH, Xu X, Ru K, Deng XN, Hu LF and Li Y; and writing the manuscript: Chen ZH, Huai Y, Liu SY and Qian AR. and Revised the manuscript: Chen ZH, Huai Y, Mao WJ and Wang XH. These co-first authors (Chen ZH, and Huai Y) share the first author position.
Competing Interests
The authors have declared that no competing interest exists.
References
1. Farr JN, Xu M, Weivoda MM, Monroe DG, Fraser DG, Onken JL. et al. Targeting cellular senescence prevents age-related bone loss in mice. Nature medicine. 2017;23:1072-9
2. Deng P, Yuan Q, Cheng YD, Li J, Liu ZQ, Liu Y. et al. Loss of KDM4B exacerbates bone-fat imbalance and mesenchymal stromal cell exhaustion in skeletal aging. Cell stem cell. 2021;28:1057-73
3. Chandra A, Rajawat J. Skeletal Aging and Osteoporosis: Mechanisms and Therapeutics. Int J Mol Sci. 2021 22
4. Pignolo RJ, Law SF, Chandra A. Bone Aging, Cellular Senescence, and Osteoporosis. JBMR Plus. 2021;5:e10488
5. Parfitt AM. The coupling of bone formation to bone resorption: A critical analysis of the concept and of its relevance to the pathogenesis of osteoporosis. Metabolic Bone Disease and Related Research. 1982;4:1-6
6. Bushati N, Cohen SM. microRNA functions. Annu Rev Cell Dev Biol. 2007;23:175-205
7. Wang X, Guo B, Li Q, Peng J, Yang Z, Wang A. et al. miR-214 targets ATF4 to inhibit bone formation. Nature medicine. 2013;19:93-100
8. Aquino-Martinez R, Farr JN, Weivoda MM, Negley BA, Onken JL, Thicke BS. et al. miR-219a-5p Regulates Rorbeta During Osteoblast Differentiation and in Age-related Bone Loss. Journal of bone and mineral research: the official journal of the American Society for Bone and Mineral Research. 2019;34:135-44
9. Sun Z, Wang H, Wang Y, Yuan G, Yu X, Jiang H. et al. MiR-103-3p targets the m(6) A methyltransferase METTL14 to inhibit osteoblastic bone formation. Aging cell. 2021;20:e13298
10. Hensley AP, McAlinden A. The role of microRNAs in bone development. Bone. 2021;143:115760
11. Zheng H, Ramnaraign D, Anderson BA, Tycksen E, Nunley R, McAlinden A. MicroRNA-138 Inhibits Osteogenic Differentiation and Mineralization of Human Dedifferentiated Chondrocytes by Regulating RhoC and the Actin Cytoskeleton. JBMR plus. 2019;3:e10071
12. Zhang L, Liu Y, Feng B, Liu LG, Zhou YC, Tang H. MiR-138-5p knockdown promotes osteogenic differentiation through FOXC1 up-regulation in human bone mesenchymal stem cells. Biochemistry and cell biology = Biochimie et biologie cellulaire. 2021;99:296-303
13. Wu K, Liu M, Li N, Zhang L, Meng F, Zhao L. et al. Chitosan-miRNA functionalized microporous titanium oxide surfaces via a layer-by-layer approach with a sustained release profile for enhanced osteogenic activity. Journal of nanobiotechnology. 2020;18:127
14. Chen Z, Zhao F, Liang C, Hu L, Li D, Zhang Y. et al. Silencing of miR-138-5p sensitizes bone anabolic action to mechanical stimuli. Theranostics. 2020;10:12263-78
15. Chen Z, Zhang Y, Zhao F, Yin C, Yang C, Huai Y. et al. miR-138-5p negatively regulates osteoblast differentiation through inhibiting β-catenin under simulated microgravity in MC3T3-E1 cells. Acta Astronaut. 2021;182:240-50
16. Sun M, Hu L, Wang S, Huang T, Zhang M, Yang M. et al. Circulating MicroRNA-19b Identified From Osteoporotic Vertebral Compression Fracture Patients Increases Bone Formation. Journal of bone and mineral research: the official journal of the American Society for Bone and Mineral Research. 2020;35:306-16
17. Sun M, Zhou X, Chen L, Huang S, Leung V, Wu N. et al. The Regulatory Roles of MicroRNAs in Bone Remodeling and Perspectives as Biomarkers in Osteoporosis. BioMed research international. 2016;2016:1652417
18. Bakker AD, Klein-Nulend J. Osteoblast isolation from murine calvaria and long bones. Methods in molecular biology. 2012;816:19-29
19. Hu L, Su P, Li R, Yan K, Chen Z, Shang P. et al. Knockdown of microtubule actin crosslinking factor 1 inhibits cell proliferation in MC3T3-E1 osteoblastic cells. BMB reports. 2015;48:583-8
20. Chen Z, Zhang Y, Zhao F, Yin C, Yang C, Wang X. et al. Recombinant Irisin Prevents the Reduction of Osteoblast Differentiation Induced by Stimulated Microgravity through Increasing beta-Catenin Expression. International journal of molecular sciences. 2020 21
21. Zhao F, Ma X, Qiu W, Wang P, Zhang R, Chen Z. et al. Mesenchymal MACF1 Facilitates SMAD7 Nuclear Translocation to Drive Bone Formation. Cells. 2020 9
22. Qiu WX, Ma XL, Lin X, Zhao F, Li DJ, Chen ZH. et al. Deficiency of Macf1 in osterix expressing cells decreases bone formation by Bmp2/Smad/Runx2 pathway. Journal of cellular and molecular medicine. 2020;24:317-27
23. Zhang Y, Yin C, Hu L, Chen Z, Zhao F, Li D. et al. MACF1 Overexpression by Transfecting the 21 kbp Large Plasmid PEGFP-C1A-ACF7 Promotes Osteoblast Differentiation and Bone Formation. Human gene therapy. 2018;29:259-70
24. Li D, Liu J, Yang C, Tian Y, Yin C, Hu L. et al. Targeting long noncoding RNA PMIF facilitates osteoprogenitor cells migrating to bone formation surface to promote bone formation during aging. Theranostics. 2021;11:5585-604
25. Saferding V, Hofmann M, Brunner JS, Niederreiter B, Timmen M, Magilnick N. et al. microRNA-146a controls age-related bone loss. Aging cell. 2020;19:e13244
26. Lee WC, Guntur AR, Long F, Rosen CJ. Energy Metabolism of the Osteoblast: Implications for Osteoporosis. Endocrine reviews. 2017;38:255-66
27. Zhong D, Xu GZ, Wu JZ, Liu H, Tang JY, Wang CG. Circ-ITCH sponges miR-214 to promote the osteogenic differentiation in osteoporosis via upregulating YAP1. Cell death & disease. 2021;12:340
28. Hu L, Liu J, Xue H, Panayi AC, Xie X, Lin Z. et al. miRNA-92a-3p regulates osteoblast differentiation in patients with concomitant limb fractures and TBI via IBSP/PI3K-AKT inhibition. Molecular therapy Nucleic acids. 2021;23:1345-59
29. Tang Z, Xu T, Li Y, Fei W, Yang G, Hong Y. Inhibition of CRY2 by STAT3/miRNA-7-5p Promotes Osteoblast Differentiation through Upregulation of CLOCK/BMAL1/P300 Expression. Molecular therapy Nucleic acids. 2020;19:865-76
30. Elmansi AM, Hussein KA, Herrero SM, Periyasamy-Thandavan S, Aguilar-Perez A, Kondrikova G. et al. Age-related increase of kynurenine enhances miR29b-1-5p to decrease both CXCL12 signaling and the epigenetic enzyme Hdac3 in bone marrow stromal cells. Bone reports. 2020;12:100270
31. Chen S, Li Y, Zhi S, Ding Z, Huang Y, Wang W. et al. lncRNA Xist Regulates Osteoblast Differentiation by Sponging miR-19a-3p in Aging-induced Osteoporosis. Aging and disease. 2020;11:1058-68
32. Li C, Cheng P, Liang M, Chen Y, Lu Q, Wang J. et al. MicroRNA-188 regulates age-related switch between osteoblast and adipocyte differentiation. The Journal of clinical investigation. 2015;125:1509-22
33. Yin C, Tian Y, Yu Y, Yang C, Su P, Zhao Y. et al. miR-129-5p Inhibits Bone Formation Through TCF4. Frontiers in cell and developmental biology. 2020;8:600641
34. Ma J, Lin X, Chen C, Li S, Zhang S, Chen Z. et al. Circulating miR-181c-5p and miR-497-5p Are Potential Biomarkers for Prognosis and Diagnosis of Osteoporosis. The Journal of clinical endocrinology and metabolism. 2020 105
35. Huai Y, Zhang W, Chen Z, Zhao F, Wang W, Dang K. et al. A Comprehensive Analysis of MicroRNAs in Human Osteoporosis. Frontiers in endocrinology. 2020;11:516213
36. Shi J, Cao F, Chang Y, Xin C, Jiang X, Xu J. et al. Long non-coding RNA MCM3AP-AS1 protects chondrocytes ATDC5 and CHON-001 from IL-1β-induced inflammation via regulating miR-138-5p/SIRT1. Bioengineered. 2021;12:1445-56
37. Chunlei H, Chang Z, Sheng L, Yanchun Z, Lulin L, Daozhang C. Down-regulation of MiR-138-5p Protects Chondrocytes ATDC5 and CHON-001 from IL-1 β-induced Inflammation Via Up-regulating SOX9. Current pharmaceutical design. 2020;25:4613-21
38. Roberto GM, Lira RC, Delsin LE, Vieira GM, Silva MO, Hakime RG. et al. microRNA-138-5p as a Worse Prognosis Biomarker in Pediatric, Adolescent, and Young Adult Osteosarcoma. Pathology oncology research: POR. 2020;26:877-83
39. Yan J, Zhang C, Zhao Y, Cao C, Wu K, Zhao L. et al. Non-viral oligonucleotide antimiR-138 delivery to mesenchymal stem cell sheets and the effect on osteogenesis. Biomaterials. 2014;35:7734-49
40. Eskildsen T, Taipaleenmaki H, Stenvang J, Abdallah BM, Ditzel N, Nossent AY. et al. MicroRNA-138 regulates osteogenic differentiation of human stromal (mesenchymal) stem cells in vivo. Proceedings of the National Academy of Sciences of the United States of America. 2011;108:6139-44
41. Wu J, Zhao J, Sun L, Pan Y, Wang H, Zhang WB. Long non-coding RNA H19 mediates mechanical tension-induced osteogenesis of bone marrow mesenchymal stem cells via FAK by sponging miR-138. Bone. 2018;108:62-70
42. Feng L, Shi L, Lu YF, Wang B, Tang T, Fu WM. et al. Linc-ROR Promotes Osteogenic Differentiation of Mesenchymal Stem Cells by Functioning as a Competing Endogenous RNA for miR-138 and miR-145. Molecular therapy Nucleic acids. 2018;11:345-53
43. Hu J, Liao H, Ma Z, Chen H, Huang Z, Zhang Y. et al. Focal Adhesion Kinase Signaling Mediated the Enhancement of Osteogenesis of Human Mesenchymal Stem Cells Induced by Extracorporeal Shockwave. Sci Rep. 2016;6:20875
44. Sun T, Leung F, Lu WW. MiR-9-5p, miR-675-5p and miR-138-5p Damages the Strontium and LRP5-Mediated Skeletal Cell Proliferation, Differentiation, and Adhesion. International journal of molecular sciences. 2016;17:236
45. Li D, Yu K, Xiao T, Dai Y, Liu L, Li H. et al. LOC103691336/miR-138-5p/BMPR2 axis modulates Mg-mediated osteogenic differentiation in rat femoral fracture model and rat primary bone marrow stromal cells. Journal of cellular physiology. 2019;234:21316-30
46. Tsukamoto S, Lovendorf MB, Park J, Salem KZ, Reagan MR, Manier S. et al. Inhibition of microRNA-138 enhances bone formation in multiple myeloma bone marrow niche. Leukemia. 2018
47. Wu X, Kodama A, Fuchs E. ACF7 regulates cytoskeletal-focal adhesion dynamics and migration and has ATPase activity. Cell. 2008;135:137-48
48. Wu X, Shen QT, Oristian DS, Lu CP, Zheng Q, Wang HW. et al. Skin stem cells orchestrate directional migration by regulating microtubule-ACF7 connections through GSK3beta. Cell. 2011;144:341-52
49. Hu L, Su P, Yin C, Zhang Y, Li R, Yan K. et al. Microtubule actin crosslinking factor 1 promotes osteoblast differentiation by promoting beta-catenin/TCF1/Runx2 signaling axis. Journal of cellular physiology. 2018;233:1574-84
50. Yin C, Zhang Y, Hu L, Tian Y, Chen Z, Li D. et al. Mechanical unloading reduces microtubule actin crosslinking factor 1 expression to inhibit beta-catenin signaling and osteoblast proliferation. Journal of cellular physiology. 2018;233:5405-19
51. Lin X, Xiao Y, Chen Z, Ma J, Qiu W, Zhang K. et al. Microtubule actin crosslinking factor 1 (MACF1) knockdown inhibits RANKL-induced osteoclastogenesis via Akt/GSK3beta/NFATc1 signalling pathway. Molecular and cellular endocrinology. 2019;494:110494
52. Hu L, Yin C, Chen D, Wu Z, Liang S, Zhang Y. et al. MACF1 promotes osteoblast differentiation by sequestering repressors in cytoplasm. Cell death and differentiation. 2021;28:2160-78
53. Yin C, Tian Y, Hu L, Yu Y, Wu Z, Zhang Y. et al. MACF1 alleviates aging-related osteoporosis via HES1. Journal of cellular and molecular medicine. 2021;25:6242-57
54. Lin X, Xiao Y, Zhang K, Yang D, Miao Z, Deng X. et al. Knockdown of MACF1 inhibits the migration and cytoskeletal arrangement of pre-osteoclasts induced by simulated microgravity. Acta Astronaut. 2022;190:149-59
55. Su P, Tian Y, Yin C, Wang X, Li D, Yang C. et al. MACF1 promotes osteoblastic cell migration by regulating MAP1B through the GSK3beta/TCF7 pathway. Bone. 2021;154:116238
Author contact
Corresponding authors: Airong Qian, Tel.: +8602988491840; E-mail: qianairedu.cn; Xiao Lin, E-mail: linxiaoedu.cn; Ge Zhang, E-mail: zhanggeedu.hk; Songlin Peng, E-mail: psling824com.

 Global reach, higher impact
Global reach, higher impact