Impact Factor ISSN: 1449-2288
Int J Biol Sci 2022; 18(13):4984-5000. doi:10.7150/ijbs.75284 This issue Cite
Research Paper
Development and Validation of a DNA Methylation-related Classifier of Circulating Tumour Cells to Predict Prognosis and to provide a therapeutic strategy in Lung Adenocarcinoma
1. School of Medicine, Southeast University, Nanjing 210009, China.
2. Nanjing Medical University, Nanjing, 210011, Jiangsu, China; Department of Oncology, Jiangyin People's Hospital, Jiangyin, 214400, China.
3. Department of oncology, Zhongda Hospital, School of Medicine, Southeast University, Nanjing 210009, China.
Abstract
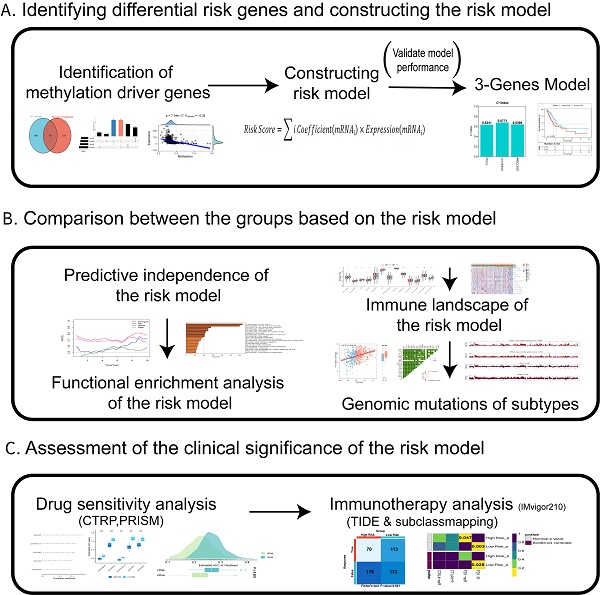
Background: A significant factor influencing the prognosis of lung adenocarcinoma (LUAD) is tumor metastasis. Studies have shown that abnormal DNA methylation in circulating tumor cells (CTCs) is associated with tumour metastasis. Based on the genes expressed in CTCs that play an important role in DNA methylation, we hope to build a risk model to predict prognosis and provide a therapeutic strategy in LUAD.
Methods: The CTC sequencing data for LUAD were obtained from GSE74639, which contains 10 CTC samples and 6 primary tumour samples. To carefully assess the clinical value, functional status, involvement of the tumor microenvironment (TME) based on the risk model, and genetic variants based on based on data from The Cancer Genome Atlas (TCGA) and the Gene Expression Omnibus (GEO), a reliable risk model was successfully built.
Results: Three differentially methylated genes (DMGs) of CTCs for LUAD, including mitochondrial ribosomal protein L51 (MRPL51), STE20-like kinase (SLK), and protein regulator of cytokinesis 1(PRC1), were effectively used to construct a risk model. Both the training and validation cohorts' stability and accuracy of the risk model were evaluated. Each patient in the TCGA-LUAD cohort received a risk score, and based on the median score, they were divided into high- and low-risk groups. The tumors in the high-risk group in this study were classified as "cold" and immunosuppressed, which may be linked to a poor prognosis. The tumors in the low-risk group, however, were deemed "hot" and had immune hyperfunction linked to a positive prognosis. Additionally, patients in the low-risk group showed greater sensitivity to immunotherapy than those in the high-risk group.
Conclusions: Based on DMGs of CTCs from LUAD, we successfully developed a predictive risk model and discovered differences in biological function, TME, genetic variation, and clinical outcomes between those at high and low risk group.
Keywords: Lung adenocarcinoma, Circulating tumour cell, DNA methylation, Risk model, Clinical outcomes

 Global reach, higher impact
Global reach, higher impact