Impact Factor
ISSN: 1449-2288
Int J Biol Sci 2015; 11(12):1413-1423. doi:10.7150/ijbs.13436 This issue Cite
Review
To Know How a Gene Works, We Need to Redefine It First but then, More Importantly, to Let the Cell Itself Decide How to Transcribe and Process Its RNAs
1. Shandong Academy of Pharmaceutical Sciences, Ji'nan, Shandong, 250101, P.R. China
2. Hormel Institute, University of Minnesota, Austin, MN 55912, USA
3. Center for Translational Medicine, Pharmacology and Biomedical Sciences Building, Guangxi Medical University, 22 Shuangyong Road, Nanning, Guangxi 530021, P.R. China.
4. Laboratory of Cell and Molecular Biology, Cancer Institute, Chinese Academy of Medical Science, Beijing 100021, P.R. China
Abstract
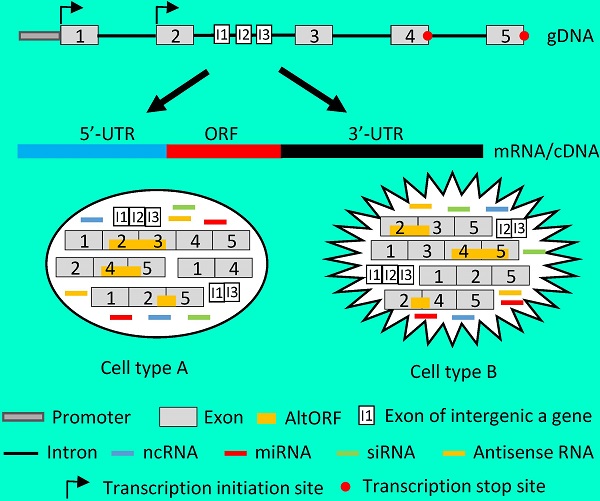
Recent genomic and ribonomic research reveals that our genome produces a stupendous amount of non-coding RNAs (ncRNAs), including antisense RNAs, and that many genes contain other gene(s) in their introns. Since ncRNAs either regulate the transcription, translation or stability of mRNAs or directly exert cellular functions, they should be regarded as the fourth category of RNAs, after ribosomal, messenger and transfer RNAs. These and other research advances challenge the current concept of gene and raise a question as to how we should redefine gene. We can either consider each tiny part of the classically-defined gene, such as each mRNA variant, as a “gene”, or, alternatively and oppositely, regard a whole genomic locus as a “gene” that may contain intron-embedded genes and produce different types of RNAs and proteins. Each of the two ways to redefine gene not only has its strengths and weaknesses but also has its particular concern on the methodology for the determination of the gene's function: Ectopic expression of complementary DNA (cDNA) in cells has in the past decades provided us with great deal of detail about the functions of individual mRNA variants, and will make the data less conflicting with each other if just a small part of a classically-defined gene is considered as a “gene”. On the other hand, genomic DNA (gDNA) will better help us in understanding the collective function of a genomic locus. In our opinion, we need to be more cautious in the use of cDNA and in the explanation of data resulting from cDNA, and, instead, should make delivery of gDNA into cells routine in determination of genes' functions, although this demands some technology renovation.
Keywords: Gene definition, non-coding RNA, Complementary DNA, Gene function, Genomic locus

 Global reach, higher impact
Global reach, higher impact