Impact Factor ISSN: 1449-2288
Int J Biol Sci 2020; 16(12):2205-2219. doi:10.7150/ijbs.42080 This issue Cite
Research Paper
Ligand-receptor interaction atlas within and between tumor cells and T cells in lung adenocarcinoma
1. Department of Thoracic Surgery, Zhongshan Hospital, Fudan University, No. 180, Fenglin Road, Shanghai, 200032, China
2. Shanghai Medical College, Fudan University, Shanghai 200032, China
3. Department of Thoracic Surgery, The Affiliated Suzhou Hospital of Nanjing Medical University, Suzhou 215001, China
#These authors contributed equally: Zhencong Chen, Xiaodong Yang, and Guoshu Bi.
Abstract
Purpose: Lung adenocarcinoma (LUAD) is the leading cause of cancer-related deaths worldwide. Although tumor cell-T cell interactions are known to play a fundamental role in promoting tumor progression, these interactions have not been explored in LUAD.
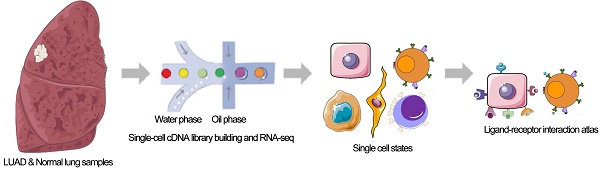
Methods: The 10x genomics single-cell RNA sequencing (scRNA-seq) and gene expression data of LUAD patients were obtained from ArrayExpress, TCGA, and GEO databases. scRNA-seq data were analyzed and infiltrating tumor cells, epithelial cells, and T cells were identified in the tumor microenvironment. Differentially expressed ligand-receptor pairs were identified in tumor cells/normal epithelial cells and tumor T cells/non-tumor T cells based on corresponding scRNA-seq and gene expression data, respectively. These important interactions inside/across cancer cells and T cells in LUAD were systematically analyzed. Furthermore, a valid prognostic machine-learning model based on ligand-receptor interactions was built to predict the prognosis of LUAD patients. Flow cytometry and qRT-PCR were performed to validate the significantly differently expressed ligand-receptor pairs.
Results: Overall, 39,692 cells in scRNA-seq data were included in our study after quality filtering. A total of 65 ligand-receptor pairs (17 upregulated and 48 downregulated), including LAMB1-ITGB1, CD70-CD27, and HLA-B-LILRB2, and 96 ligand-receptor pairs (41 upregulated and 55 downregulated), including CCL5-CCR5, SELPLG-ITGB2, and CXCL13-CXCR5, were identified in LUAD cancer cells and T cells, respectively. To explore the crosstalk between cancer cells and T cells, 114 ligand-receptor pairs, including 11 ligand-receptor pair genes that could significantly affect survival outcomes, were identified in our research. A machine-learning model was established to accurately predict the prognosis of LUAD patients and ITGB4, CXCR5, and MET were found to play an important role in prognosis in our model. Flow cytometry and qRT-PCR analyses indicated the reliability of our study.
Conclusion: Our study revealed functionally significant interactions within and between cancer cells and T cells. We believe these observations will improve our understanding of potential mechanisms of tumor microenvironment contributions to cancer progression and help identify potential targets for immunotherapy in the future.
Keywords: Lung adenocarcinoma, Single-cell RNA-seq, Cell-to-cell interactions, Machine learning, Survival

 Global reach, higher impact
Global reach, higher impact