10
Impact Factor
ISSN: 1449-2288
Int J Biol Sci 2021; 17(1):107-118. doi:10.7150/ijbs.49243 This issue Cite
Research Paper
Deciphering the genomic and lncRNA landscapes of aerobic glycolysis identifies potential therapeutic targets in pancreatic cancer
1. State Key Laboratory of Oncogenes and Related Genes, Ren Ji Hospital, School of Biomedical Engineering, Shanghai Jiao Tong University, Shanghai 200240, P.R. China.
2. State Key Laboratory of Oncogenes and Related Genes, Shanghai Cancer Institute, Ren Ji Hospital, School of Medicine, Shanghai Jiao Tong University, Shanghai 200240, P.R. China.
3. Department of Radiation Oncology, Ren Ji Hospital, School of Medicine, Shanghai Jiao Tong University, Shanghai 200127, PR China.
4. Institute of Oncology, Affiliated Hospital of Jiangsu University, Zhenjiang 212001, P.R. China.
5. The Fengxian Hospital, Southern Medical University, Shanghai 201499, PR China.
6. Department of Biliary-Pancreatic Surgery, Ren Ji Hospital, School of Medicine, Shanghai Jiao Tong University, Shanghai 200217, P.R. China.
*These authors contributed equally to this work.
Abstract
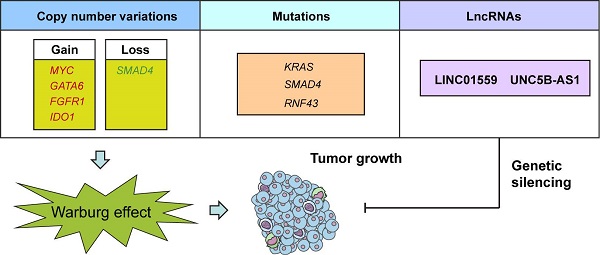
Aerobic glycolysis, also known as the Warburg effect, is emerged as a hallmark of most cancer cells. Increased aerobic glycolysis is closely associated with tumor aggressiveness and predicts a poor prognosis. Pancreatic ductal adenocarcinoma (PDAC) is characterized by prominent genomic aberrations and increased glycolytic phenotype. However, the detailed molecular events implicated in aerobic glycolysis of PDAC are not well understood. In this study, we performed a comprehensive molecular characterization using multidimensional ''omic'' data from The Cancer Genome Atlas (TCGA). Detailed analysis of 89 informative PDAC tumors identified substantial copy number variations (MYC, GATA6, FGFR1, IDO1, and SMAD4) and mutations (KRAS, SMAD4, and RNF43) related to aerobic glycolysis. Moreover, integrated analysis of transcriptional profiles revealed many differentially expressed long non-coding RNAs involved in PDAC aerobic glycolysis. Loss-of-function studies showed that LINC01559 and UNC5B-AS1 knockdown significantly inhibited the glycolytic capacity of PDAC cells as revealed by reduced glucose uptake, lactate production, and extracellular acidification rate. Moreover, genetic silencing of LINC01559 and UNC5B-AS1 suppressed tumor growth and resulted in alterations in several signaling pathways, such as TNF signaling pathway, IL-17 signaling pathway, and transcriptional misregulation in cancer. Notably, high expression of LINC01559 and UNC5B-AS1 predicted poor patient prognosis and correlated with the maximum standard uptakevalue (SUVmax) in PDAC patients who received preoperative 18F-FDG PET/CT. Taken together, our results decipher the glycolysis-associated copy number variations, mutations, and lncRNA landscapes in PDAC. These findings improve our knowledge of the molecular mechanism of PDAC aerobic glycolysis and may have practical implications for precision cancer therapy.
Keywords: Tumor metabolism, Energy metabolism, LncRNA, CNVs, FEZF1-AS1

 Global reach, higher impact
Global reach, higher impact