10
Impact Factor
ISSN: 1449-2288
Int J Biol Sci 2023; 19(6):1764-1777. doi:10.7150/ijbs.81317 This issue Cite
Research Paper
SB Digestor: a tailored driver gene identification tool for dissecting heterogeneous Sleeping Beauty transposon-induced tumors
1. Cancer Center, Faculty of Health Sciences, University of Macau, Macau SAR, China.
2. Centre for Precision Medicine Research and Training, Faculty of Health Sciences, University of Macau, Macau SAR, China.
3. Genomics & Bioinformatics Core, Faculty of Health Sciences, University of Macau, Macau SAR, China.
4. College of Fisheries and Life Science, Shanghai Ocean University, Shanghai, China.
5. MoE Frontiers Science Center for Precision Oncology, University of Macau, Macau SAR, China.
Abstract
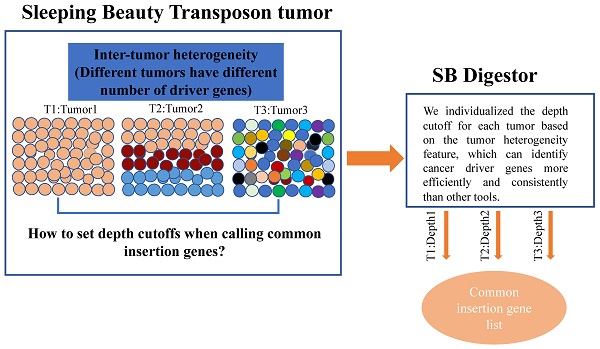
Sleeping Beauty (SB) insertional mutagenesis has been widely used for genome-wide functional screening in mouse models of human cancers, however, intertumor heterogeneity can be a major obstacle in identifying common insertion sites (CISs). Although previous algorithms have been successful in defining some CISs, they also miss CISs in certain situations. A major common characteristic of these previous methods is that they do not take tumor heterogeneity into account. However, intertumoral heterogeneity directly influences the sequence read number for different tumor samples and then affects CIS identification. To precisely detect and define cancer driver genes, we developed SB Digestor, a computational algorithm that overcomes biological heterogeneity to identify more potential driver genes. Specifically, we define the relationship between the sequenced read number and putative gene number to deduce the depth cutoff for each tumor, which can reduce tumor complexity and precisely reflect intertumoral heterogeneity. Using this new tool, we re-analyzed our previously published SB-based screening dataset and identified many additional potent drivers involved in Brca1-related tumorigenesis, including Arhgap42, Tcf12, and Fgfr2. SB Digestor not only greatly enhances our ability to identify and prioritize cancer drivers from SB tumors but also substantially deepens our understanding of the intrinsic genetic basis of cancer.
Keywords: Sleeping Beauty transposon, intertumor heterogeneity, common insertion sites, SB Digestor, Fgfr2

 Global reach, higher impact
Global reach, higher impact