Impact Factor ISSN: 1449-2288
Int J Biol Sci 2023; 19(13):4103-4122. doi:10.7150/ijbs.85724 This issue Cite
Research Paper
Squalene epoxidase/SQLE is a candidate target for treatment of colorectal cancers with p53 mutation and elevated c-MYC expression
1. Experimental and Molecular Pathology, Institute of Pathology, Medical Faculty, Ludwig-Maximilians-Universität München, D-80337 Munich, Germany.
2. German Cancer Consortium (DKTK), Partner site Munich, D-80336 Munich, Germany.
3. German Cancer Research Center (DKFZ), D-69120 Heidelberg, Germany.
*: these authors contributed equally to this work
Abstract
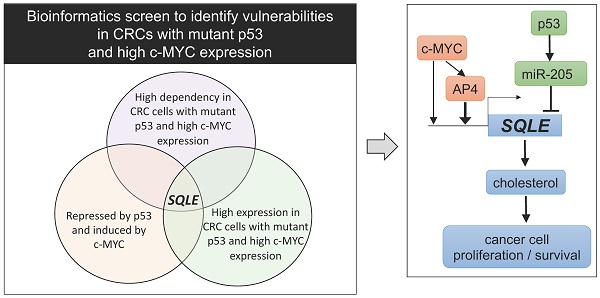
Elevated expression of c-MYC and inactivation of p53 represent two of the most common alterations in colorectal cancer (CRC). However, c-MYC and defective p53 are difficult to target therapeutically. Therefore, effectors downstream of both c-MYC and p53 may represent attractive, alternative targets for cancer treatment. In a bioinformatics screen we identified Squalene epoxidase/SQLE as a candidate therapeutic target that appeared to be especially relevant for cell survival in CRCs, which display elevated c-MYC expression and loss of p53 function. SQLE is a rate-limiting enzyme in the cholesterol synthesis. Here, we show that p53 supresses SQLE expression, cholesterol levels, and cell viability via the induction of miR-205, which directly targets SQLE. Furthermore, c-MYC induced SQLE expression directly and via its target gene AP4. The transcription factor AP4/TFAP4 directly induced SQLE expression and cholesterol levels, whereas inactivation of AP4 resulted in decreased SQLE expression and caused resistance to Terbinafine, an inhibitor of SQLE. Inhibition of SQLE decreased viability of CRC cells. This effect was enhanced in CRCs cells with p53 inactivation and/or enhanced c-MYC/AP4 expression. Altogether, our results demonstrate that SQLE represents a vulnerability for CRCs with p53 inactivation and elevated c-MYC activity.
Keywords: SQLE/Squalene epoxidase, cholesterol synthesis, p53, miR-205, c-MYC, AP4, TFAP4, colorectal cancer, Terbinafine, therapy

 Global reach, higher impact
Global reach, higher impact